Abstract
BACKGROUND:
tRNA halves (tiRNAs) are produced from mature tRNAs. They have important roles both with in normal cells and cancer cells. However, the diagnostic value of tiRNAs in cancers have not yet been elucidated.
OBJECTIVE:
To explore the diagnostic value of tiRNA-5034-GluTTC-2 in gastric cancer.
PATIENTS AND METHODS:
Quantitative reverse transcription-polymerase chain reaction was used to detect the expression levels of tiRNA-5034-GluTTC-2 in paired gastric cancer tissues and adjacent normal tissues, plasmas from patients with gastric cancer and healthy people, and gastric cancer cell lines. Then, the relationship between its levels and clinicopathological factors of patients with gastric cancer was analyzed. A receiver operating characteristic (ROC) curve was established to predict the diagnostic value.
RESULTS:
tiRNA-5034-GluTTC-2 was first found to be down-regulated in gastric cancer tissues and plasmas. Its levels were significantly associated with tumor size. The area under the ROC curve (AUC) was 0.779 and 0.835 in tissue and plasma, respectively. The sensitivity, specificity and AUC were 84.7%, 92.8%, and 0.915 when tissues and plasmas were used in combination, respectively. The overall survival rate of patients with a lower expression of tiRNA-5034-GluTTC-2 was significantly lower than those with a higher expression.
CONCLUSIONS:
These results indicated that tiRNA-5034-GluTTC-2 may be a novel biomarker for the diagnosis of gastric cancer.
Introduction
Gastric cancer, which has a high morbidity and mortality rate, is one of the most common cancers in the world [1], with approximately one million new cases occurring each year and more than 720,000 deaths. The initial symptoms of gastric cancer are not obvious and there is a lack of specific diagnostic methods for identifying early gastric cancer. The diagnostic value of existing tumor markers, such as carcinoembryonic antigen (CEA) and carbohydrate antigen 19-9 (CA 19-9), are not high enough to aid in early diagnosis [2]. For most patients with gastric cancer diagnosis is made only when they have reached an advanced stage [3]. Therefore, it is essential that new biomarkers be found which assist in the early diagnosis of gastric cancer.
Non-coding small RNAs regulate gene expression at the transcriptional and translational levels. One of the deeply-studied non-coding small RNAs is microRNA (miRNA), which plays an important role in the regulation of organism development, metabolism, cell differentiation, and cell cycle [4]. MiRNAs are closely related to cancers [5] and the elucidation of the relationship between miRNAs and cancers is a promising area of cancer research [6].
With the development of molecular biology, scientists have found that tRNA-derived fragments (tRFs) and tRNA halves (tiRNAs) have important biological roles [7]. As a class of non-coding small RNA molecules, they are produced from mature tRNAs, or precursor tRNAs, by specific cleavage at different sites and widely found in prokaryotes and eukaryotic transcriptomes [8]. There are three major subtypes of tRFs, tRF-5s, tRF-3s, and tRF-1s. tRF-5s are generated from the cleavage between D-ring and anticodon loop to 5’-end; tRF-3 s are generated from the cleavage between T-ring to 3’-end of mature tRNAs; tRF-1s are from the 3’ trailer fragment of precursor tRNAs [7]. tiRNAs, 29–50 nt in length, are induced by stress and produced by specific cleavage at the mature tRNA anticodon loop. Based upon whether a 5’ or 3’ sequence is included, tiRNAs can be divided into two subclasses: 5’tiRNAs and 3’tiRNAs. 5’tiRNAs start from the 5’ end of the mature tRNA to the end of the anticodon loop; 3’tiRNAs start from the anticodon loop to the 3’ end of the mature tRNA [9]. tiRNAs are cleaved by angiogenin (ANG), called RNY1 in yeast, and have a 5’ hydroxyl group instead of a 5’ phosphate [9]. Besides, 5’tiRNAs and 3’tiRNAs can be produced under non-stress conditions [9]. tRFs and tiRNAs have a variety of biological roles including acting as signaling molecules in a stress response and as regulators of gene expression [10]. tRFs and tiRNAs can influence the development of cancers by affecting mRNA stability, regulating reverse transcription, inhibiting translation, and regulating ribosomal biogenesis. They regulate cell proliferation, cell cycle, apoptosis, and migration of cancer cells [11]. tRFs and tiRNAs also bind to Ago and Piwi complexes, suggesting that these tRFs and tiRNAs can function as miRNAs or Piwi-interacting RNAs (piRNAs), respectively [12]. 5’tiRNAs have potential as circulating biomarkers. Dhahbi et al. found that the expression level of specific circulating 5’tiRNAs is related to the clinicopathological features of breast cancer [13]. These features make tRFs and tiRNAs have broad prospects with regards to their application in disease diagnosis and treatments. In this study, we focused upon tiRNA-5034-GluTTC-2 (MINTbase Unique ID tRF-34-86V8WPMN1E8Y2Q;
Materials and methods
Specimens and clinical data collection
Gastric cancer tissues and paired adjacent non-tumorous tissues were obtained from the surgical patients at Yinzhou People’s Hospital Affiliated to Medical School of Ningbo University, China, from September 2012 to April 2016. Healthy peoples’ plasma and plasma from patients with gastric cancer were also obtained on the day before surgery and seven days after surgery, at Yinzhou People’s Hospital Affiliated to Medical School of Ningbo University and Ningbo No. 1 Hospital Affiliated to Medical School of Ningbo University, China. The diagnosis was confirmed by histopathology and the tumor staging was based upon the International Union Against Cancer’s Tumor-Node-Metastasis (TNM) staging system [15]. The histological grades were assessed according to the National Comprehensive Cancer Networks (NCCNs) guidelines for clinical practice of oncology (V.1.2011) [16]. All of the clinical data were collected by experienced physicians. The experimental operators were unaware of the clinical data. Written informed consent was obtained from all patients. The ethics committee of human studies at Ningbo University approved all aspects of this study.
Cell culture
Human gastric epithelial cell line GES-1 and gastric cancer cell lines HGC-27, AGS, BGC-823, SGC-7901, and MGC-803 were purchased from Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences. AGS cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) High Glucose (HyClone, Logan, UT, USA) containing 1% penicillin/streptomycin (Life Technologies, Carlsbad, CA, USA) and 10% fetal bovine serum (FBS) (PAN-Biotech, Aidenbach, Germany) at 37
Total RNA extraction
Total RNA in tissues/cells and plasma samples was first extracted with TRIzol reagent (Invitrogen, Karlsruhe, Germany) and TRIzol LS reagent (Invitrogen), respectively. Next, the purity and concentration were measured by SmartSpec Plus Spectrophotometer (Bio-Rad, Hercules, CA, USA) Finally, RNA was stored at
Assess RNA purity was based upon A260/A280 values, within 1.8–2.1. In addition, the quality of total RNA was further evaluated by 1% agarose gel electrophoresis (Fig. S1).
RNA pretreatment
tRFs and tiRNAs are heavily decorated by RNA modifications. rtStar™ tRF&tiRNA Pretreatment Kit (Arraystar, Rockville, MD, USA) is designed to remove the modifications that interfere with the quantitative polymerase chain reaction (qPCR). These modifications include terminal modifications that block adaptor ligation to the RNA ends and internal methylations that hinder reverse transcription for cDNA synthesis. Pretreatment (de-modification) of RNA was performed following the manufacturer’s protocol (Fig. S2A–E).
Reverse transcription
rtStar™ First-Strand cDNA Synthesis Kit (Arraystar) is designed to prepare cDNA for qPCR detection of tRFs and tiRNAs. The method sequentially ligates 3’ adaptor and 5’ adaptor (Fig. S2F and G). The 3’ adaptor contains a priming site for reverse transcription (RT) primer (Fig. S2H and I). The 3’ adaptor and 5’ adaptor were provided by Arraystar. RT was performed according to the manufacturer’s protocol.
Real-time qPCR
Real-time qPCR was performed with a GoTaq qPCR Master Mix (Promega, Madison, WI, USA) in the Mx3005P real-time PCR system (Stratagene, LaJolla, CA, USA) according to the manufacturer’s protocol (Fig. S2J). The full-length sequence of tiRNA-5034-GluTTC-2 is 5’-UCCCAUAUGGUCUAGCGGUUAG GAUUCCUGGUUU-3’. The sequences of PCR pri- mers for tiRNA-5034-GluTTC-2 were 5’-GTCCGAC GATCTCCCATA-3’ and 5’-TGTGCTCTTCCGATC TAAAC-3’. The primers include part sequences of tiRNA-5034-GluTTC-2 and adaptor specific. This allows for the specific amplification of tRFs and tiRNAs instead of mature tRNA. Therefore, an abnormal E value will occur when performing BLAST search with any of the primers. The primers’ sequences of snRNA (RNU6-2), as a control, were 5’-GCTTCGGCAGCACATATACTAAAAT-3’ and 5’-CGCTTCACGAATTTGCGTGTCAT-3’. These pri- mers were synthesized by Sangon Biotech (Shanghai, China). The
Cloning and sequencing of qRT-PCR products
The qRT-PCR products of tiRNA-5034-GluTTC-2 were first purified using the UNIQ-10 PCR product purification kit and then cloned into the pUCm-T vector (Thermo Fisher Scientific, Hudson, NH, USA), following the manufacturer’s instructions. DNA sequencing was performed by Thermo Fisher Scientific.
Northern blot
After dividing the total RNA molecules by size in a denaturing polyacrylamide gel, they were transferred to a nylon membrane (Amersham Biosciences, Piscataway, NJ, USA) and hybridized with immobilized RNA using a
Statistical analysis
All statistical analyses were performed by GraphPad Prism 6.0 (GraphPad Software, LaJolla, CA, USA) and Statistical Program for Social Sciences (SPSS) 20.0 software (SPSS, Chicago, IL, USA). The comparison of the relative expression levels of tiRNA-5034-GluTTC-2 between gastric cancer tissues and the paired adjacent normal tissues was analyzed by paired
tiRNA-5034-GluTTC-2 (34 nt) and its mature tRNA (75 nt) can be detected and quantified by Northern blotting. Three representative paired gastric cancer tissues (T) and adjacent non-cancerous tissues (N) are shown. 5S rRNA are shown for equal loading.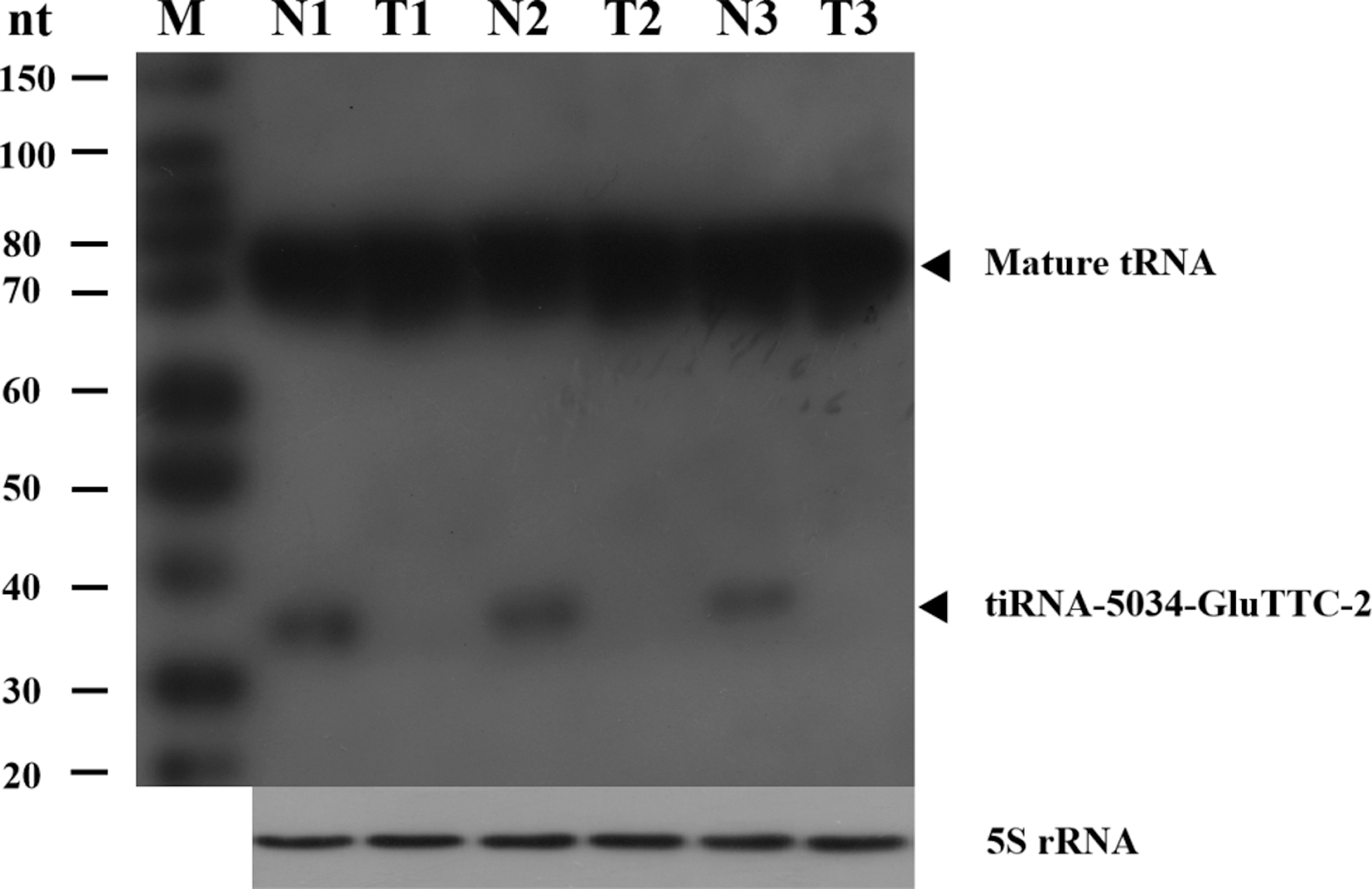
tiRNA-5034-GluTTC-2 expression levels in gastric cancer tissues and plasma samples. (A) The expression levels of tiRNA-5034-GluTTC-2 were significantly lower than those in the corresponding non-tumorous tissues (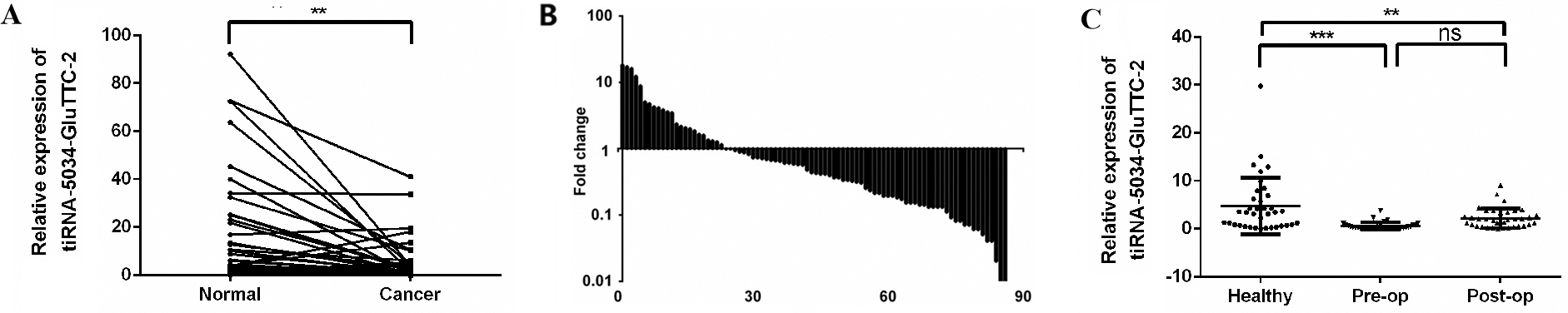
ROC curve of tiRNA-5034-GluTTC-2.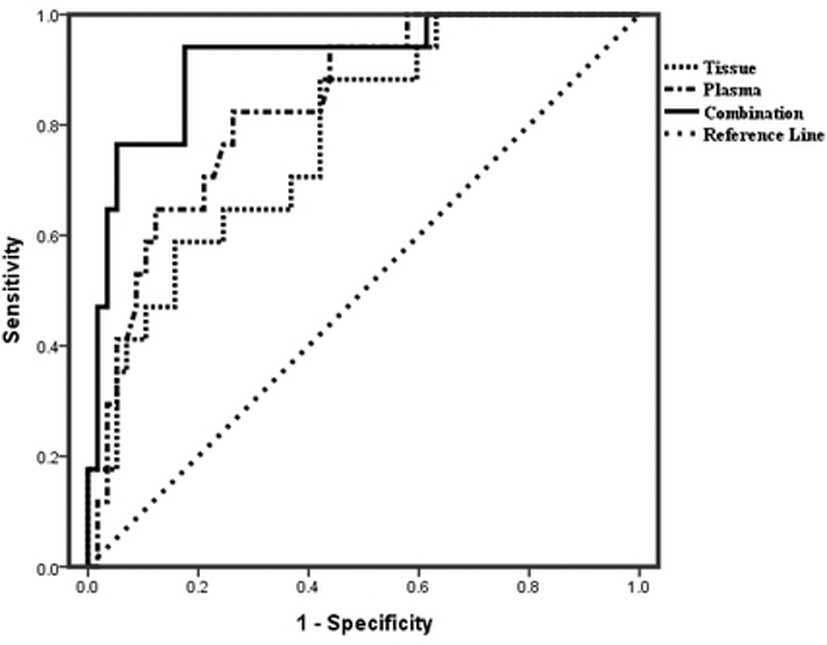
Prognostic significance of tiRNA-5034-GluTTC-2 in gastric cancer patients. Kaplan-Meier analysis of overall survival (OS) based on tiRNA-5034-GluTTC-2 expression level in tissues. 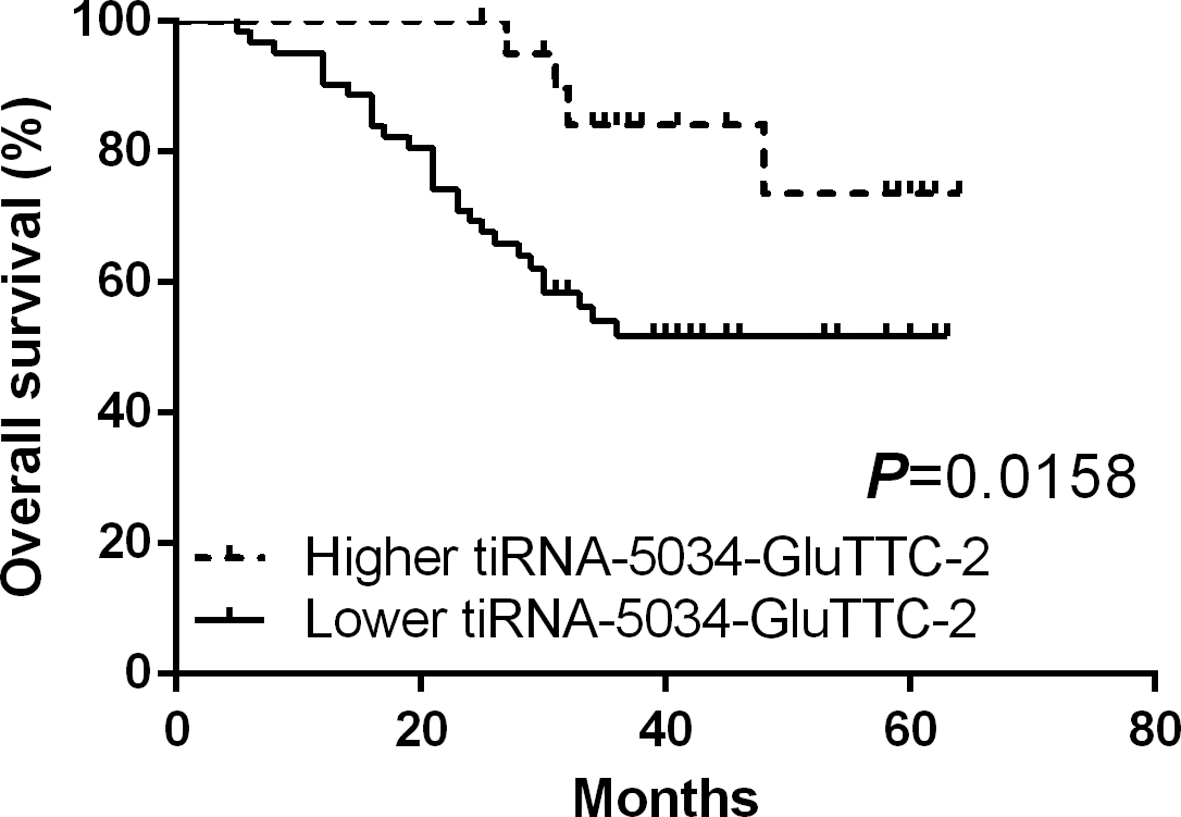
Amplification of tiRNA-5034-GluTTC-2
Since tiRNAs are small non-coding RNAs, its amplification by qRT-PCR needs adaptors (Fig. S2F–J). To analyze the specificity of the amplified tiRNA-5034-GluTTC-2 product, the melting curve analysis was first used, and the result revealed the amplification product as a single peak (Fig. S3), indicating no non-specific amplification or primer-dimers were produced. The Tm value is slightly increased due to the addition of the adaptors. To confirm the correction of qRT-PCR, the PCR products were further analyzed by T-A clone sequencing. The results showed that the sequence of the amplified tiRNA-5034-GluTTC-2 product (Fig. S4) was consistent with that in mintBase (
The expression levels of tiRNA-5034-GluTTC-2 in gastric cancer cell lines and the normal gastric epithelial cell line. Compared with normal gastric epithelial GES-1 cells, the expression levels of tiRNA-5034-GluTTC-2 in five gastric cancer cell lines, AGS, MGC-803, HGC-27, BGC-823, and SGC-7901, were significantly decreased. Relative expression was calculated using the 2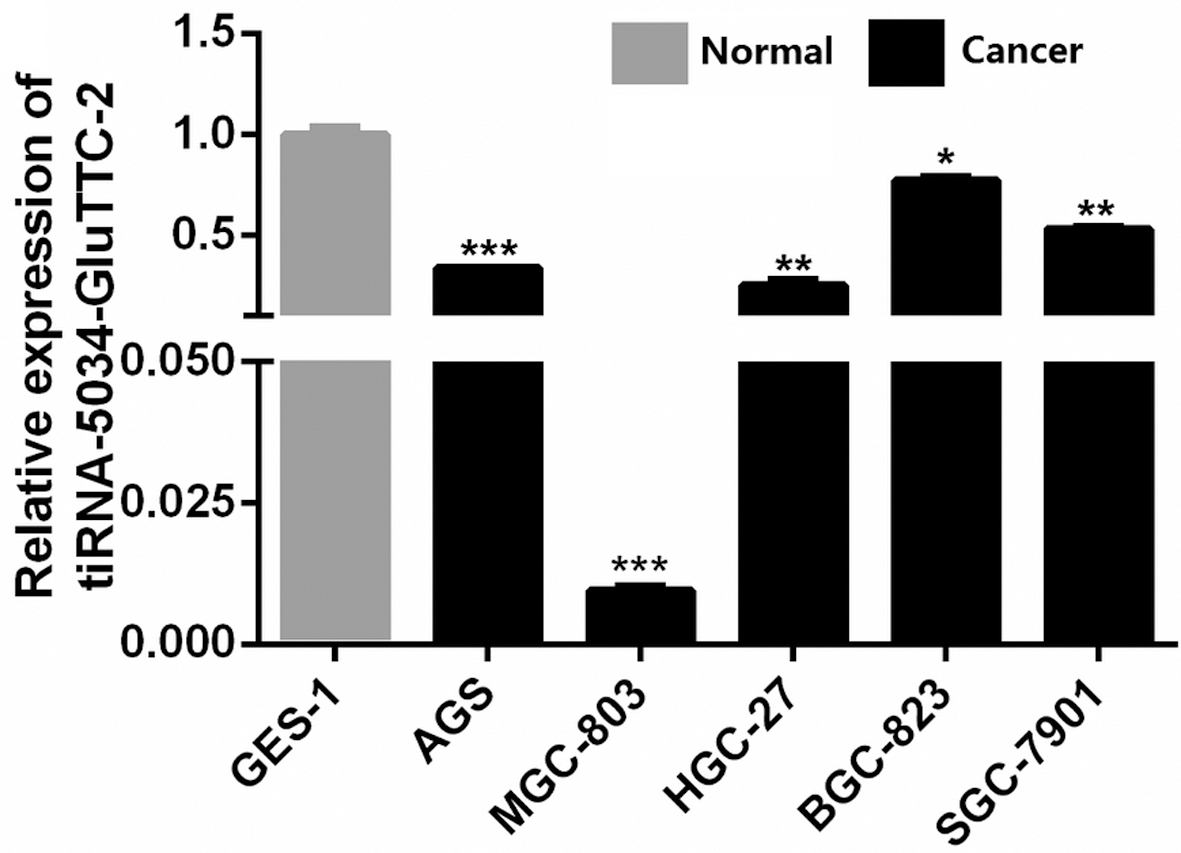
We first detected the expression levels of tiRNA-5034-GluTTC-2 in 86 gastric cancer tissues and their paired adjacent non-cancerous tissues; as shown in Fig. 2A, tiRNA-5034-GluTTC-2 was significantly downregulated in cancer tissues. Among them, the lower expression of samples was accounted for 73.3% (63/86) (Fig. 2B). Since bloods are the most commonly used samples in clinical practice, we next collected paired plasma from patients with gastric cancer on the day before surgery and seven days after surgery, along with the plasma of healthy people. The results indicated that the levels of tiRNA-5034-GluTTC-2 in preoperative and postoperative gastric patients was reduced when compared with the healthy control (Fig. 2C).
The relationship of tiRNA-5034-GluTTC-2 expression levels (
) in gastric cancer tissues with clinicopathological factors of patients with gastric cancer
The relationship of tiRNA-5034-GluTTC-2 expression levels (
Next, we performed an analysis to identify whether tiRNA-5034-GluTTC-2 expression levels were associated with clinicopathological features of patients with gastric cancer. We divided the expression levels of tiRNA-5034-GluTTC-2 into high and low groups and then analyzed the relationship between the expression levels of tiRNA-5034-GluTTC-2 and the clinicopathological factors. The results showed that tiRNA-5034-GluTTC-2 expression levels were associated with tumor size (
The area under the ROC curve (AUC) of tiRNA-5034-GluTTC-2 was 0.779 and 0.835 in tissues and plasmas, respectively. In combined use, the AUC was up to 0.915, with 84.7% sensitivity and 92.8% specificity (Fig. 3). The cut-off was 9.7 and 8.9 in tissues and plasmas, respectively.
Further tiRNA-5034-GluTTC-2 levels in gastric cancer tissues from patients before the operation were categorized into two groups; the higher or lower group, according to the expression levels. The results of survival curves showed that patients with lower tiRNA-5034-GluTTC-2 expression group had a much shorter overall survival than those in the higher expression group (
Univariate and multivariate Cox regression analysis of the overall survival (OS) of tiRNA-5034-GluTTC-2 expression in tissues of patients with gastric cancer
Univariate and multivariate Cox regression analysis of the overall survival (OS) of tiRNA-5034-GluTTC-2 expression in tissues of patients with gastric cancer
To obtain more information about tiRNA-5034-GluTTC-2 expression in gastric cancers, we further analyzed tiRNA-5034-GluTTC-2 expression in human gastric cancer cell lines. As expected, the results showed that tiRNA-5034-GluTTC-2 was significantly downregulated in five gastric cancer cell lines, AGS, MGC-803, HGC-2, BGC-823, and SGC-7901, compared with the human normal gastric epithelial cell line GES-1 (Fig. 5).
Discussion
tRNA fragments were first found in stress and starvation-induced cells, and then found in cells under non-stress conditions [19, 20, 21]. With the development and widespread application of RNA sequencing, more and more tiRNAs and tRFs have been found [14, 22].
Many bases in tRNAs are modified during tRNA maturation and these modifications are affected by angiogenin (ANG) processing [23, 24]. Therefore, understanding the complete information about the tRNAs’ modifications will help our understanding of the tRFs and tiRNAs’ biogenesis. tRFs may also form double-stranded molecules with complementary RNAs and act as a primer binding site, or base pairing, with targeted RNAs [9, 25]. In-depth study of the formation of tiRNAs and tRFs and their binding proteins will help us to understand their molecular mechanisms.
Several tiRNAs and tRFs have been found to be differentially regulated under varied pathological conditions. Olvedy et al. first discovered that the ratio of tRNA
Although the molecular mechanisms and potential roles of tiRNAs and tRFs have received more attention [14, 24, 25], their clinical diagnostic values are still largely unknown. In our study, we found that tiRNA-5034-GluTTC-2 was down-regulated in gastric cancer tissues, plasmas, and gastric cancer cell lines (Figs 2 and 5). More importantly, we first identified its potential diagnostic value.
The differentiation, tumor size, TNM stage, invasion, lymphatic metastasis, and distal metastasis are all vital factors affecting a cancer patients’ outcome [16]. In most cases, the increasing numbers for tumor diameter, invasion, lymphatic metastasis, and distal metastasis result in a decreasing five-year survival rate [29]. Our study found that low levels of tiRNA-5034-GluTTC-2 in gastric cancer tissues was associated with tumor size (Table 1). The larger the tumor diameter was, the lower the expression of tiRNA-5034-GluTTC-2. In gastric cancer tissues, the AUC was 0.779 (Fig. 3). The sensitivity and specificity were 73.3% and 46.5%, respectively; the false negative rate and false positive rate were 26.7% and 53.5%, respectively. The negative predictive value (NPV) and positive predictive value (PPV) were 73.4% and 65.2%, respectively. In plasma samples, the AUC was 0.835, the sensitivity and specificity were 97.3% and 59.5%, respectively (Fig. 3). As a biomarker, the AUC for the combined use of tissue and plasma tiRNA-5034-GluTTC-2 was up to 0.915 (Fig. 3). The sensitivity and specificity were 84.7% and 92.8%, respectively. Besides, the OS of the lower tiRNA-5034-GluTTC-2 expression group was significantly lower than that of the higher expression group (Fig. 4). Univariate analysis revealed that TNM stage, lymphatic metastasis, and expression of tiRNA-5034-GluTTC-2 in tissues was associated with OS (HR
Since tiRNAs and tRFs are produced from tRNAs which have rich modification [7, 9], they are more stable in tissues, plasmas, and cells than other non-coding RNA, such as long non-coding RNAs (lncRNAs) and miRNAs [30, 31, 32, 33, 34, 35, 36]. tiRNAs and tRFs are highly enriched in plasma [9]. Besides, removing the modifications and adding adaptors produced higher yields and lower adaptor-dimer formation in qRT-PCR [37]. These properties make tiRNAs and tRFs become potent novel biomarkers for the non-invasive diagnosis of gastric cancer.
Conclusion
In conclusion, this study demonstrates for the first time that tiRNA-5034-GluTTC-2 is lowly expressed in gastric cancer tissues, plasma samples, and gastric cancer cells relative to the normal control group. As a biomarker, the expression of tiRNA-5034-GluTTC-2 can be detected by qRT-PCR and Northern blot. These findings indicate that tRFs and tiRNAs, such as tiRNA-5034-GluTTC-2, may be novel potential biomarkers for gastric cancer diagnosis.
Footnotes
Acknowledgments
This study was supported by grants from National Natural Science Foundation of China (no. 81772279), the Scientific Innovation Team Project of Ningbo (no. 2017C110019), and the K. C. Wong Magna Fund in Ningbo University.
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary data
The supplementary files are available to download from
