Abstract
BACKGROUND:
Colorectal cancer (CRC) is one of the most prevalent cancers and microRNAs are involved in colorectal carcinogenesis and progression. The role of our candidate microRNAs (miR-143-3p, -424-5p, -212-3p and -34a-3p) have been investigated in various cancers.
OBJECTIVE:
The aim of the current study was to evaluate expression levels of microRNAs (miR-143-3p, -424-5p, -212-3p and -34a-3p) in the sera of patients with CRC in order to identify potential non-invasive biomarker for CRC and investigate the relationship between their expression and clinicopathological features of CRC.
METHODS:
The serological expression of candidate microRNAs were measured in 124 participants, including 62 CRC patients and 62 healthy controls and the serum expression levels of candidate miRNAs were quantified by stemloop reverse transcriptionquantitative polymerase chain reaction.
RESULTS:
In the present study, results showed a significant upregulation expression level of miR-424-5p (
CONCLUSIONS:
The present investigation suggested that low expression of miR-143-3p and increasing level of miR-424-5p in CRC patients may play an important role in development of CRC and they could function as potential non-invasive biomarkers for CRC.
Introduction
Colorectal cancer (CRC) is the third most prevalent malignancy and the fourth leading cause of cancer-related death worldwide [1] and a total of 1 million new cases of CRC, 660,000 deaths were expected worldwide each year [2]. The development and progression of CRC is a multi-factorial, multi-step and multi-stage process, which is resulted from the incorporated actions of environmental factors and genetic factors [3]. Identification of novel biomarkers is a need in order to choose appropriate therapy for various patients [4].
MicroRNAs (miRNAs/miRs) are small non-coding single-stranded RNA molecules [5], which can be paired to the coding region [6] or 3’ untranslated region (UTR), the 5’ UTR [7] of target messenger RNAs (mRNAs) which can induce either mRNA degradation or translational repression [8]. MicroRNAs are found in serum, plasma and other body fluids [9, 10], and exist in an apparently stable extracellular form [11]. Exosomes which contain miRNA produced by tumor cells may have a substantial role in metastasis by promoting angiogenesis, cell proliferation and/or tumor cell invasion [11, 12, 13].
Growing evidence demonstrates that miRNAs play an important role in cancer development, differentiation and progression, functioning as tumor oncogenes or suppressors [14, 15, 16, 17].
Studies have represented the different expression of miRNAs in the sera of patients with tumor compared with the ones for healthy people. In addition, recent studies are indicative of the fact that miRNAs, released by cancer cells, circulating in the plasma are potential candidates for non-invasive detection of cancer [18, 19].
The role of candidate microRNAs (miR-143-3p, -424-5p, -212-3p and -34a-3p) were investigated in progression of cancer [20, 21, 22, 23] and some of them were studied as prognostic biomarkers in other malignant neoplasms [24, 25].
Therefore, the present study was performed to evaluate the expression of miR-143-3p, -424-5p, -212-3p and -34a-3p in the sera of patients with CRC and to define the relationships between candidate miRNAs expression and clinicopathological features of CRC.
Materials and methods
Patients and samples
A total of 62 patients pathologically diagnosed with CRC, who had been admitted to Shahid Sadoughi Hospital (Yazd, Iran) between November 2016 and December 2017, were enrolled in this study. Sixty two people undergoing health check-up for the same period made up the control group. The blood samples were collected and utilized after receiving informed consent from all participants. Exclusion criteria included genetic CRC related to genetic syndrome, previous chemotherapy and radiotherapy. Some information included age, gender, tumor size, TNM staging, lymph node status, distant metastasis and perineural invasion were extracted from patient’s medical record. The present study was approved by the ethics committee of Shahid Sadoughi University of Medical Sciences (ID: IR.SSU.MEDICINE.REC.1395.200).
Serum preparation, RNA extraction and reverse transcription reaction
Venous blood (5 ml) of CRC patients and control subjects were withdrawn before surgery and all blood samples were centrifuged at 3500 g for 5 min within 1 hour after collection. Supernatant was removed and stored in RNase/DNase free tubes at
Serum samples were lysed with TRIzol LS reagent (Invitrogen Inc, San Diego, CA, USA). The aqueous phase was isolated after the addition of chloroform and centrifugation. Isopropanol was added to precipitate the RNA from the aqueous phase and ethanol was used to wash RNA pellet and The resulting RNA was dissolved in DEPC treated water. DNase treatment was accomplished in order to eliminate any contaminating DNA (DNase I, Invitrogen, Carlsbad, CA, USA) The concentration and quality of RNA were analyzed by the NanoDrop Spectrophotometer (A260/A280
In order to detect miRNA, total RNA was reverse transcribed subsequently to cDNA with the stem-loop reverse transcription primer. cDNA synthesis was carried out using the NG dART RT kit (Eurex). The manufacturer’s instructions were observed during all experiments. The relation mixture leading to RT reactions included 5X NG cDNA Buffer, primers, NG dART RT mix, RNase-free Water and RNA (The concentration of RNA used for conversion to cDNA was 700 ng/
RT-qPCR for miRNAs
RT-qPCR was conducted using the ABI StepOne system (Qiagen GmbH) and amplifications were performed using the HyperScript TM First strand synthesis kit (Geneall Biotechnology Co., Korea). Real-time PCR conditions were 95
Relative expression levels of (A) miR-143-3p, (B) miR-424-5p, (C) miR-212-3p and (D) miR-34a-3p in the sera of control subjects and colorectal cancer patients. The line was drawn to demonstrate the median value and the Mann-Whitney U test was used to calculate statistical significance. miR, microRNA.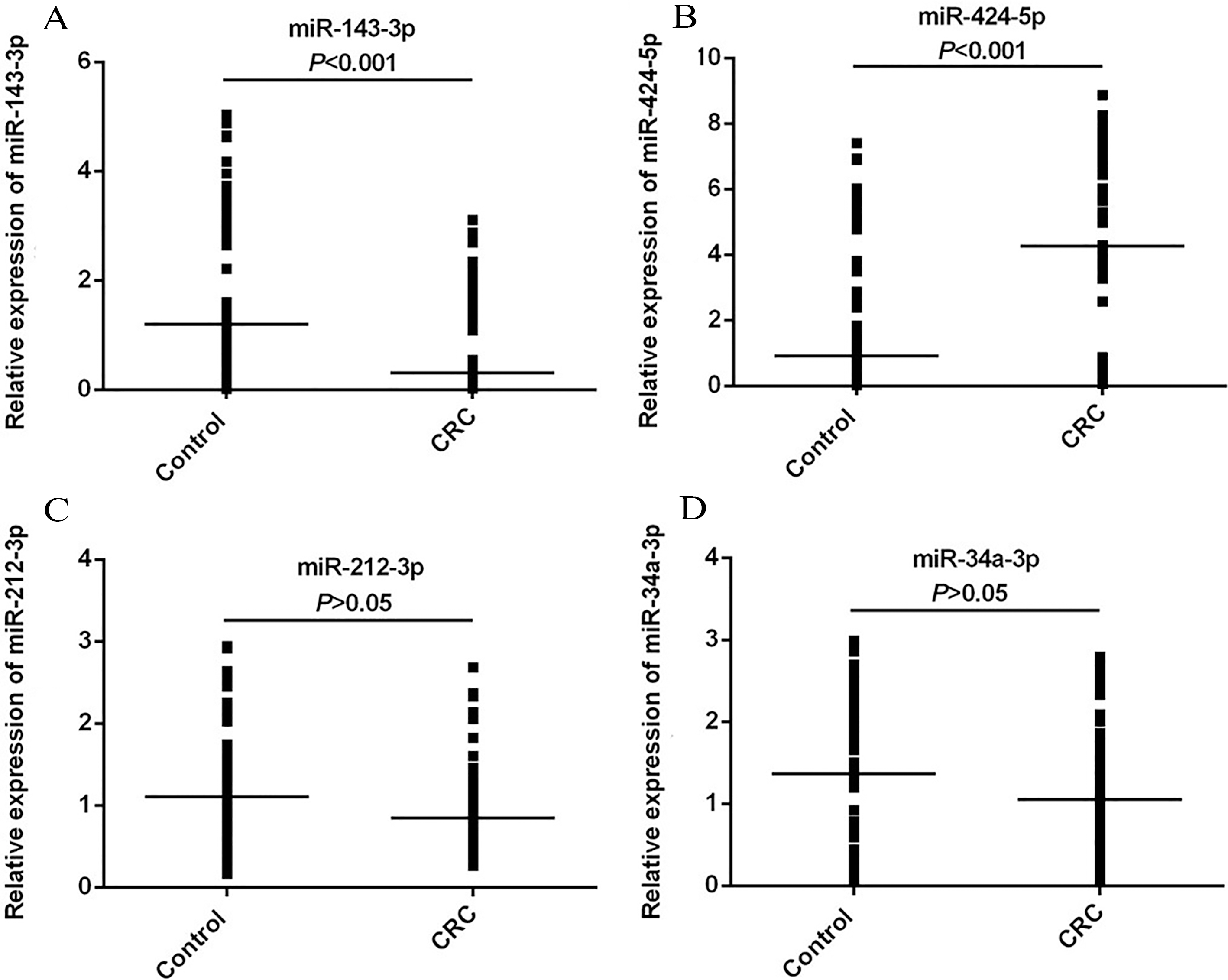
Mann-Whitney U test was employed to analyze the difference in the candidate miRNAs expression levels between the CRC patients and healthy control group. The chi-square test was performed to evaluate the association between candidate miRNAs expression and the clinicopathological parameters of CRC. In order to interpret the ability of miRNA in discriminating patients from healthy controls, a receiver operating characteristic (ROC) curve was plotted based on the DCt (Ct[miRNA]-Ct[internal control]) of miRNA expression in samples. The area under the curve (AUC), specificity and sensitivity at the optimal cut-off were calculated in order to evaluate the diagnostic application of these miRNAs as cancer biomarkers. Statistical analyses were performed using GraphPad Prism 6 (GraphPad Software Inc., CA, USA). All statistical tests were two-sided and a value of less than 0.05 was considered as a statistically significant difference.
Results
Patient characteristics
A total of 62 patients pathologically diagnosed with CRC were enrolled in the present study. Of these patients, 32 were males and 30 females, with the age range of 48–75 years and mean age of (60.7
The relative expression of miR-424-3p, -143-3p, -212-3p and -34a-3p in sera of colorectal cancer patients in comparison with healthy controls
The relative expression of miR-424-3p, -143-3p, -212-3p and -34a-3p in sera of colorectal cancer patients in comparison with healthy controls
CRC: Colorectal cancer.
Receiver operating characteristic curve analysis of (A) miR-424-5p and (B) miR-143-3p. miR, microRNA.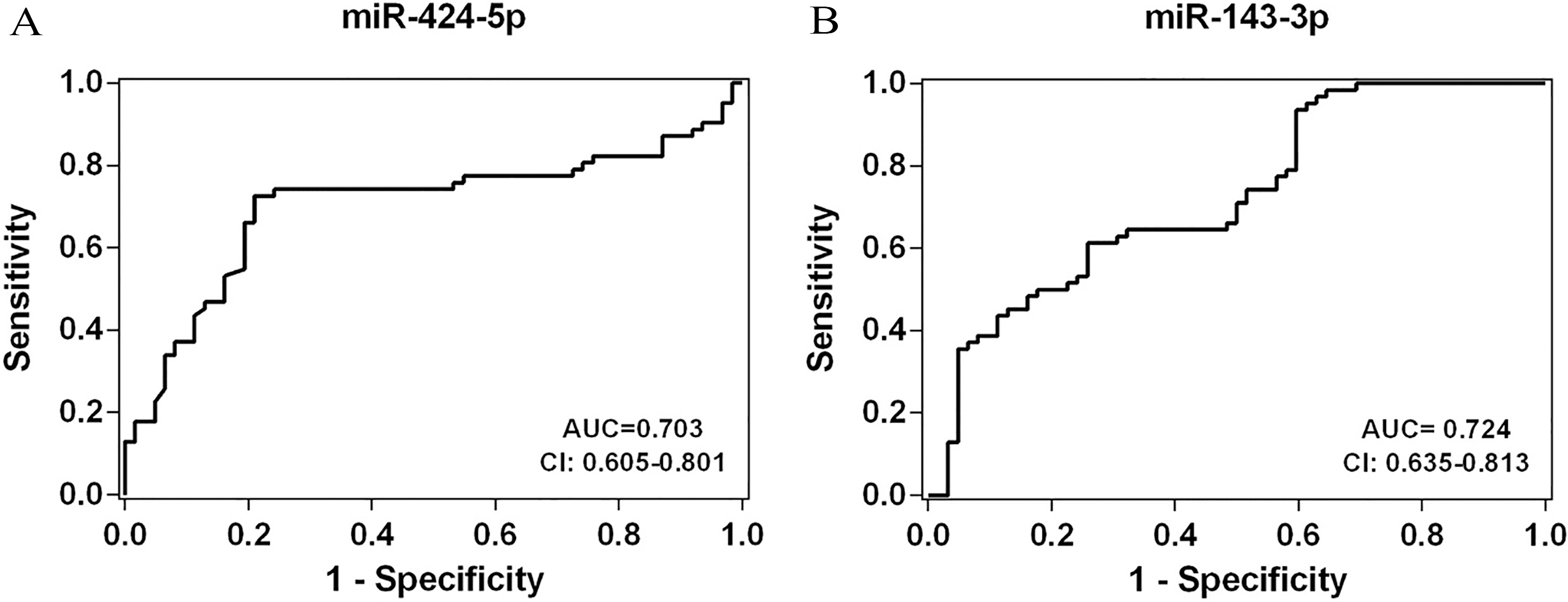
Receiver operating characteristic curve analysis
AUC: Area under curve.
The expression of miR-143-3p, -424-5p, -212-3p and -34a-3p in the serum were analyzed using RT-qPCR. The relative expression levels of candidate miRNAs were illustrated in Fig. 1. Compared with the sera of the control group, the expression of miRNA-143-3p in the serum was significantly decreased in the CRC patients (
In addition, the serum level of miR-212-3p and miR-34a-3p were lower in patients with CRC than healthy control but the results were not statistically significant (
The relative expression of miR-143-3p, -424-5p, -212-3p and -34a-3p in sera of CRC patients in comparison with healthy controls were listed in Table 1.
Correlations of miR-143-3p expression with clinicopathological parameters of patients with CRC
To evaluate the correlation of serum miR-143-3p and miR-424-5p expression with clinicopathological parameters, the 62 CRC patients were classified into high and low expression groups using the optimal cut-off calculated by the ROC curve for miR-143-3p and miR-424-5p. The association of miR-143-3p expression with clinicopathological features of patients with CRC was represented in Table 3. Low expression of miR-143-3p was significantly correlated with tumor size and lymph node metastasis in CRC (
Correlation of miR-143-3p and -424-5p expression with clinicopathological features in colorectal cancer
Correlation of miR-143-3p and -424-5p expression with clinicopathological features in colorectal cancer
TNM, tumor-node-metastasis.
Colorectal tumorigenesis is a complicated and multi-stage process comprising a set of genetic alterations [27]. Considering dysregulation expression of genes seems vital for having an extensive investigation of initiation and progression of colorectal carcinogenesis [28]. Some tumor-specific expression alterations cause non-invasive biomarkers which is detectable in patients’ serum [29, 30]. MicroRNAs are biomarkers having a prominent role in various cellular processes such as proliferation, invasion, and apoptosis [11, 12, 13]. The function of miRNAs might be different in various tissues or it could be influenced by environmental and population features [31], a thorough understanding of their role in the process of carcinogenesis is a difficult task. Furthermore, different dysregulation expression level of miRNAs may be reported in different cancers. Some miRNAs are promising biomarkers for cancer detection [32]. For instance, it has been illustrated that miR-181a and miR-372 can be utilized as prognostic biomarker for CRC [33, 34]. In the current study expressions alterations of the following four miRNAs, miR-424, -143, -212, miR-34a, and their association with clinicopathological parameters has been examined. The present study represents that the circulating miR-143 downregulated significantly in patients with CRC. In addition, serum miR-143 low expression level was associated with tumor size and lymph node metastasis. The role of miR-143 has been investigated in proliferation, cell migration and invasion of cancer cells and its association with clinicopathological characteristics in various carcinomas [35, 36]. Yang et al. reported that miR-143 regulates proliferation and apoptosis of CRC cells and suggested it could be used as a new target for diagnosis [37]. These results were in agreement with the corresponding findings of Slaby et al. stating that expression level of miR-143 decreased in CRC [38]. Also, the results of the current study showed that miR-424 upregulated significantly in the sera of CRC patients. MiR-424 as the circulating miR has been investigated in age-related macular degeneration [39]. Under hypoxia, mir-424 expression in endothelial cells regulates hypoxia inducible factor 1 (HIF-1
Several studies investigated dysregulation of miR-212 and miR-34 expression levels in various cancers. Dong et al. reported low expression of miR-212-3p can be used as a prognostic biomarker in colorectal cancer [45] and Nugent et al. found circulating miR-34a level was significantly downregulated in CRC [46]. In the current study, Serum expression downregulation of miR-212 and -34a were observed in patients with CRC but the results were not statistically significant (
Our study has some limitations. Kirschner et al. represented the impact of hemolysis on alteration of circulating miRNAs content in plasma/serum [47]. Evaluating the level of hemolysis is suggested for future studies, to eliminate this confounding factor in miRNAs analysis in blood derived fluids by considering miR-424 expressed in red blood cells [48]. Also, the selection of internal control is an important step for accurate quantification by RT-PCR. However, there is still no well-recognized internal control to quantify miRNA in serum. In current study, we selected miR-93-5p as the candidate internal control based on a study that evaluated appropriate reference genes in sera of CRC patients [26]. In addition, the present study includes small sample size and the AUC for miR-424-5p and -143-3p were 0.703 and 0.724 (
In conclusion, we have demonstrated that low expression level of miR-143-3p is associated with tumor size and lymph node metastasis in CRC patients. The current investigation indicated that decreased expression of miR-143-3p and increasing level of miR-424-5p in CRC patients may play an important role in progression of CRC and they could act as potential non-invasive biomarkers for CRC.
Footnotes
Acknowledgments
We would like to extend our gratitude to Dr. Saeed Kargar and Mrs. Zahra Khazaei for their contribution in collecting patients’ samples and information as well as Dr. Mojgan Afifezade for her valuable and constructive suggestions during article writing.
Conflict of interest
There are no conflicts of interest.
