Abstract
BACKGROUND/OBJECTIVE:
MED15 is a part of the multiprotein Mediator complex which is involved in the transcription of polymerase (Pol) II-dependent genes. Several studies in this field have reported altered expressions of distinct subunits in human malignancy. However, the role of MED15 in renal cell carcinoma (RCC) has not be investigated yet.
METHODS:
First, we performed an RNA expression and survival analysis of MED15 in RCC by using the database cBioPortal. To confirm these data on the protein level, we executed immunohistochemical (IHC) staining against MED15 on a tissue microarray containing 184 samples of the most common subtypes of the tumour at the various stages. Further, we performed functional analysis including proliferation, migration, and invasion assays on the RCC cell lines A-498 and ACHN following the siRNA-mediated MED15 knockdown.
RESULTS:
On the mRNA level, higher expression of MED15 was associated with worse patient survival rates. IHC staining validated this tendency, unfortunately the results were not significant. However, supporting this tendency, in vitro-assays showed a significant decrease in proliferation, migration, and invasion after knockdown of MED15.
CONCLUSION:
The research concludes that MED15 does seem to play a tumour promoting role in the progression and metastatic spread of renal cell carcinoma.
Introduction
In Germany, renal cell carcinoma (RCC) was the sixth and tenth most common malignancy in men and women respectively, in 2012. Approximately 15,000 people, two-thirds of whom were men and one third women, were diagnosed with RCC that led to approximately 5,200 deaths in 2012. In addition, approximately 90% of all renal tumours in adulthood are RCC, which are classified into several different subtypes based on histomorphological features [1]. The most common subtype is clear cell RCC (ccRCC), followed by papillary RCC (pRCC) and chromophobe RCC (chRCC). In each RCC subtype, distinctive genetic alterations are involved in the development and progression of renal cancer, although the carcinogenic pathways remain to be fully elucidated. Clear cell RCC (ccRCC) was associated with VHL alterations, whereas papillary RCC (pRCC) contained MET mutations. Increased knowledge about the genetic changes led to the development of novel prognostic, diagnostic, and therapeutic targets for this disease [2].
MED15 is a part of the multiprotein Mediator complex, which functions as a bridge between regulatory proteins and RNA polymerase II (Pol II), thereby regulating the Pol II-dependent transcription. It can be divided into four modules – head, middle, tail, and kinase – and is comprised of
However, the tumour promoter role of MED15 in the progression of renal cell carcinoma (RCC) has not been investigated yet, which establishes the importance of conducting this research.
Material and methods
Ethics statement
This investigation was conducted in accordance with ethical standards, the Declaration of Helsinki, and national and international guidelines. Ethical approval for using human material in the present study was obtained from the Internal Review Board of the University Hospital of Bonn (IRB# 036/08). The participants of the study agreed in writing beforehand and were anonymised before inclusion in the study cohort.
RNA expression analysis by cBioPortal
The database cBioPortal for Cancer Genomics (
Tissue microarray construction
The kidney cohort used in this study, provided by the Clinic for Urology of the University Hospital Bonn, contained 24 benign samples, 108 ccRCC samples, 28 pRCC samples, 10 chromophobe samples, and 14 metastases derived by ccRCC; in total,
Clinical pathological data of the kidney cohort
Clinical pathological data of the kidney cohort
RCC
Tissue microarrays (TMA) were performed as described previously [17, 18]. To explain in brief, formal-in-fixed paraffin-embedded (FFPE) tissues were cut into 4-
Prior to performing IHC analyses in the selected TMAs, the specificity of the antibody was confirmed according to the manufacturer’s instructions by using non-seminomatous germ cell tumour (NSGCT) as a positive control. IHC was performed using the Ventana Benchmark automated staining system (Ventana Medical System, Tuscon, AZ, USA). Briefly, slides were incubated with the primary antibody according to the manufacturer’s instructions – anti-MED15 rabbit polyclonal (1:50, clone 11566-1-AP, Proteintech, Chicago, IL) at room temperature; antibody dilution was conducted using a Ventana diluent. For signal detection, the ultraView Universal DAB detection kit (Ventana Medical System, Tuscon, AZ, USA) was utilised. Finally, slides were counterstained with haematoxylin and bluing reagent, dehydrated, and mounted.
After IHC was performed, the staining was evaluated independently by two pathologists (GK, TM). Only cases with at least one assessable TMA core with sufficient tumour tissue were included in the analysis. Nuclear protein expression was evaluated and quantified according a score ranging between 0
Clinical data and statistics
Associations with clinical-pathological parameters were established. Survival analysis was evaluated by the Kaplan-Meier estimator and log rank tests. Statistical evaluation was performed using the Student’s t-test in Microsoft Excel and SPSS. The data were analysed anonymously.
Cell lines and culture conditions
Renal cell carcinoma (RCC) cell lines ACHN and A-498 were purchased from the American Type Culture Collection (ATCC
SiRNA-mediated MED15 knockdown
MED15 siRNA (sc-75767, Santa Cruz, TX, USA) is a pool of three target-specific siRNAs designed to knockdown gene expression. A non-targeting scrambled siRNA was employed as a negative control (Exiqon Life Sciences, Copenhagen, Denmark). Renal cell carcinoma (RCC) cell lines ACHN and A-498 were transfected with 100 nmol/L siRNA using Screenfect A (Genaxxon Bioscience GmbH, Ulm, Germany) for 48 and 72 hours respectively.
Western blot
Post transfected cells were washed with ice-cold phosphate buffered saline (PBS) and lysed in a protein extraction buffer for 30 minutes. In addition, samples were sonicated for 2 minutes and subsequently centrifuged at 16,000
Quantitative reverse transcription PCR (qRT-PCR)
RNA was isolated from cell line pellets using the Total RNA Purification Mini Spin Column Kit (Genaxxon Bioscience GmbH, Ulm, Germany). RNA quantity and quality was analysed using a NanoDrop 2000 spectrophotometer (Thermo Scientific, Wilmington, DE, USA). cDNA was synthesised using 200 ng total RNA and the PrimeScript RT Reagent Kit with gDNA Eraser (Takara Bio, Saint-Germain-en-Laye, France). Quantitative real-time PCR was performed using 5 ng/
EZ4U cell proliferation assay
The EZ4U cell proliferation assay kit was used according to the manufacturer’s protocol (EZ4U; Biom-edica Group, Vienna, Austria). In brief, the siRNA transfections for proliferation assays were performed in 96-well plates. In each well of a flat-bottom 96-well plate, either 1.2
Migration assays
The siRNA transfections for migration assays were performed in 6-well plates. Forty-eight hours post transfection, cells were trypsinised and seeded into migration Boyden chambers. 5
Invasion assays
The siRNA transfections for invasion assays were performed in 6-well plates. Forty-eight hours post transfection, cells were trypsinised and seeded into Matrigel invasion chambers. 1
Results
Transcriptional expression of the MED15 by cBioPortal
Using the cBioPortal database, we analysed the mRNA expression of MED15 in clear cell renal cell carcinoma (ccRCC) samples. In two analyses, 392 and 446 samples of ccRCC were investigated. The results of the expression analyses are shown in Fig. 1A and those for survival and disease-free survival in Fig. 1B–D.
Transcriptional expression analysis of MED15 in ccRCC extended by the survival analysis. To investigate the mRNA expression [RNA Seq V2 RSEM] of MED15 in clear cell renal cell carcinoma (ccRCC), the database cBioPortal for Cancer Genomics was utilized. Two mRNA analyses for ccRCC, named “Kidney Renal Clear Cell Carcinoma TCGA, Nature 2013” and “Kidney Renal Clear Cell Carcinoma TCGA, Provisional”, are available on cBioPortal with 446 and 392 samples respectively. The expression is given in % divided into “upregulation”, “downregulation”, and “no alteration” (A). MED15 overexpression is associated with strongly reduced overall survival (B 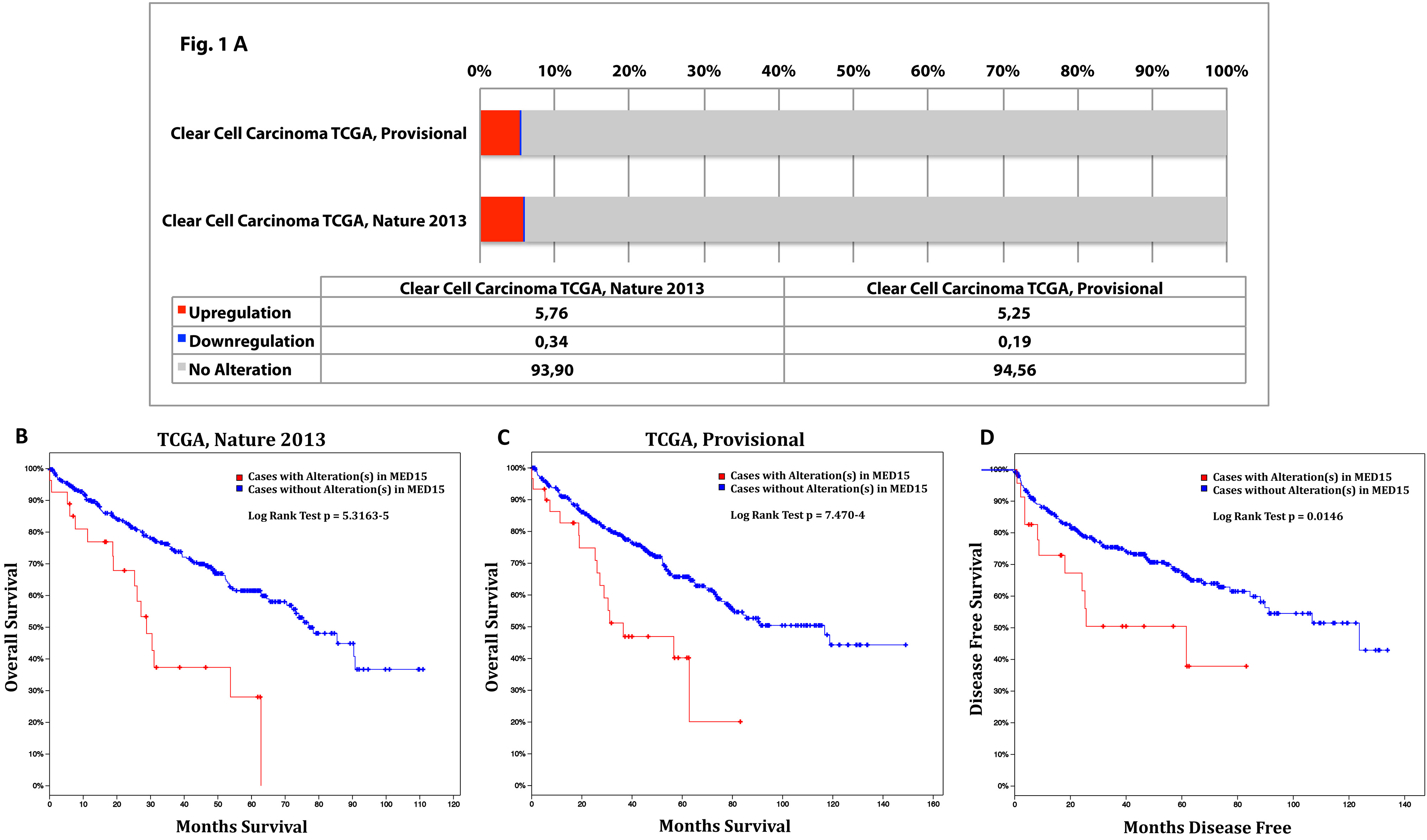
IHC staining of MED15 in RCC. Representative IHC images from tissue of benign kidney, ccRCC, pRCC, and chRCC analysis of the MED15 protein expression. Whole core 10x and inserts 40x objective magnification. Quantification of nuclear protein expression according a score of 0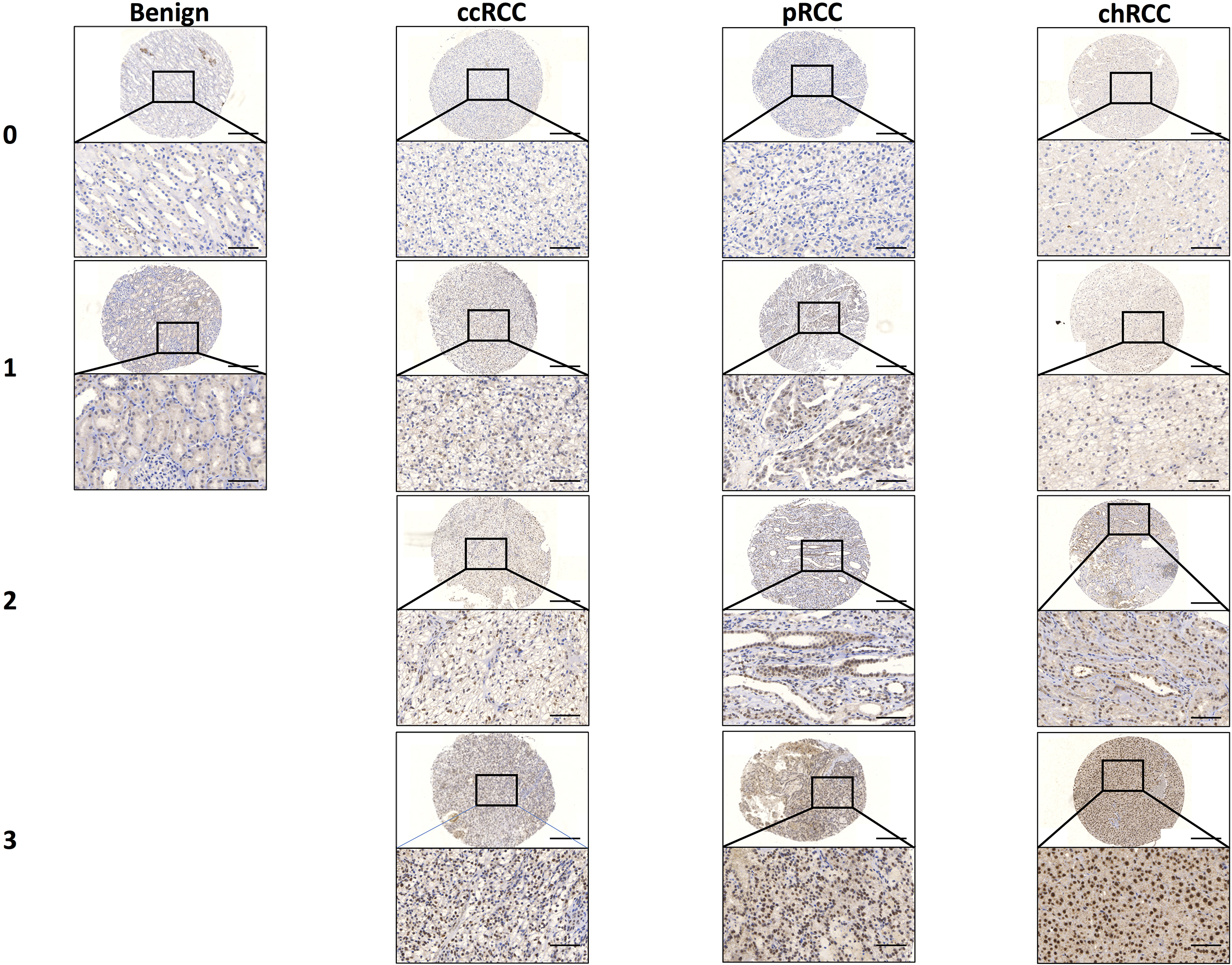 
In the first analysis, named “Kidney Renal Clear Cell Carcinoma TCGA, Nature 2013” an alteration of MED15 is described in 7% (26/391) of the samples. In detail, 23 upregulations and three downregulations of MED15 mRNA expression were detected.
Next, we investigated the datasheet called “Kidney Renal Clear Cell Carcinoma TCGA, Provisional” and found out that MED15 is altered in 7% (29/445) of the cases. In total, 23 upregulations and six downregulations of MED15 mRNA expression were found (Fig. 1A).
Survival analysis regarding transcriptional expression
In the analysis titled “Kidney Renal Clear Cell Carcinoma TCGA, Nature 2013”, cases with alteration in MED15 (predominantly overexpressed) displayed a significantly reduced overall survival rate [
In the investigation titled “Kidney Renal Clear Cell Carcinoma TCGA, Provisional”, cases with alteration in MED15 (predominantly overexpressed) displayed a significantly worse overall survival rate [
In conclusion, the investigation of the database cBioPortal showed that patients overexpressing MED15 suffer from a substantially worse disease outcome in comparison to no overexpression of MED15.
Immunohistochemistry and functional investigations
To validate the transcriptional data obtained from the analysis of the cBioPortal database, we performed protein (IHC) and functional analyses on a tissue microarray (TMA) cohort with the available clinical information and cell lines.
Immunohistochemical (IHC) staining shows MED15 nuclear expression in benign renal tissue, papillary RCC (pRCC), clear cell RCC (ccRCC), chromophobe RCC (chRCC), and metastases derived from ccRCC (Fig. 2). In addition, a MED15 cytoplasmic expression seemed to not be specific. We found MED15 to have a significantly higher expression in pRCC (median nuclear staining intensity
MED15 expression profile and survival analysis based on the MED15 expression in RCC. MED15 protein expression profile of the total kidney cohort including benign tissue, ccRCC, pRCC, and chRCC (A). 
To investigate the functional role of the subunit MED15 in progression and metastatic spread, we performed an in vitro siRNA mediated-knockdown of MED15 in the ccRCC cell lines A-498 and ACHN (Fig. 4A–C). Thereafter, we compared the measured proliferative activity of scrambled control and MED15 knockdown of cells and observed a significant decrease in A-498 (
In conclusion, high MED15 expression showed a nonsignificant tendency towards an unfavourable outcome in ccRCC. Further, the MED15 knockdown led to reduced malignant behaviour in the primary ccRCC cell line A-498 and the metastatic ACHN.
The Mediator complex serves as a hub for pathway activation and specific gene expression, and recent studies identified distinct subunits of the Mediator to be involved in different malignancy types, including RCC. For example, MED8 has been described to be frequently overexpressed in RCC and was associated with parameters of worse outcome and decreased survival rates [8, 9].
MED15 knockdown reduces progression and metastatic spread in ccRCC. Reduced MED15 expression in A-498 and ACHN cells treated with MED15 specific siRNA are shown by qPCR (A) and western blot (B 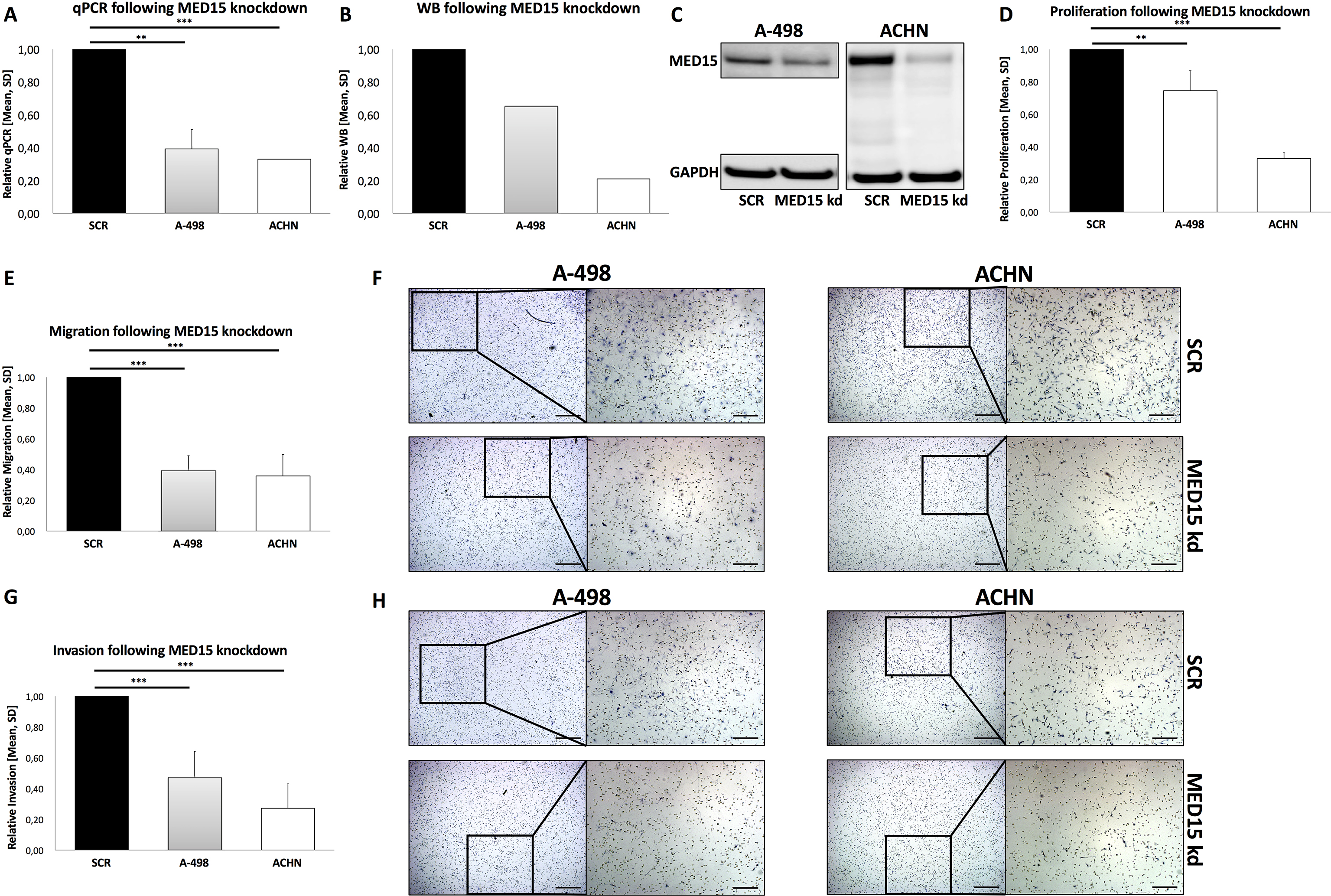
In this study, we investigated whether the Mediator complex subunit MED15, which is essential for TGF-
Supporting our findings, Shaikhibrahim et al. reported that MED15 expression was found to be increased by TGF-
In our investigation, we found an entity-specific MED15 expression profile, especially MED15 that had a significantly higher expression in pRCC compared to ccRCC, although an overexpression of MED15 in pRCC has no influence in the survival analysis. These subtype specific differences might be due to the divergent genetic background including the inactivation of the Von-Hippel-Lindau (VHL) gene, which is specific to ccRCC and is not found in other histological variants, such as papillary RCC [2]. The VHL tumour suppressor gene is located at the short arm of chromosome 3 (3p) and inactivated in VHL disease and in sporadic cases of ccRCC. The pVHL (protein VHL) is involved in the ubiquitination and degradation of hypoxia-inducible-factor (HIF) as well as in other cellular pathways, such as assembly of extracellular matrix (ECM) and stabilisation of p53 and Jade-1 [19]. Papillary RCC (pRCC) has two main subtypes – type 1, which was associated with MET alteration or gain of chromosome 7, whereas type 2 was characterized by increased expression of the NRF2-antioxidant response element (ARE) pathway [20].
Targeted therapy is a growing part of the treatment for many types of cancer. Despite the very promising studies on the role of MED15 in the different tumor entities [10, 11, 12], no inhibitor of MED15 has been developed so far. For the subunit CDK8 and its paralog CDK19, however, this has already been successfully achieved with the development of Senexin B [21]. The use of Senexin B resulted in a suppressed tumour growth or augmented the effects of the estrogen receptor (ER) antagonist fulvestrant in ER-positive breast cancer xenografts. These results identify CDK8 as a novel downstream mediator of ER and suggest the utility of CDK8 inhibitors for ER-positive breast cancer therapy [21]. Due to the comprehensive results it is very likely, an inhibitor for MED15 will be developed promptly and available for investigations. For use in cancer therapy, however, not only the inhibition alone of MED15 must be considered, but possibly also in combination with agents of the participating signaling cascades, e.g. TGF-
In conclusion, we found MED15 to have a frequently higher expression in pRCC compared to ccRCC, and it seems to be a promoter of tumour progression and metastatic spread in renal cell carcinoma.
Footnotes
Acknowledgments
The study was supported by the Ferdinand Eisenberger Fellowship of the German Society of Urology (DGU) (SYI1/FE-13) and the Maria von Linden-Program of the University of Bonn for IS.
Conflict of interest
We declare no conflict of interest.
Supplementary data
Survival analysis based on the MED15 expression in pRCC. Kaplan-Meier analysis for cancer-specific survival (A) and progression free survival (B) according to the MED15 expression in pRCC. No correlation between the expression of MED15 and survival.
