Abstract
BACKGROUND:
The option of endocrine therapy in breast cancer remains conventionally promising.
OBJECTIVE
: We aimed to investigate how accurately the pattern of hypermethylation at estrogen receptor (ESR) and progesterone receptor (PgR) genes may associate with relative expression and protein status of ER, PR and the combinative phenotype of ER/PR.
METHODS
: In this consecutive case-series, we enrolled 139 primary diagnosed breast cancer. Methylation specific PCR was used to assess the methylation status (individual test). Tumor mRNA expression levels were evaluated using real-time RT-PCR. Immunohistochemistry data was used to present hormonal receptor status of a tumor (as test reference).
RESULTS
: Methylation at ESR1 was comparably frequent in ER-breast tumors (83.0%,
CONCLUSIONS
: Despite the methylation status of ESR1 showed preponderant contribution to tumoral phenotypes of ER- and ER-/PR-, the hypermethylation of PgRb seem another epigenetic signalling variable actively associate with methylated ESR1 to show lack of ER+/PR+ tumors in breast cancer.
Introduction
Although breast cancer (BC) has been addressed as one of the most common malignancies among females in most Western countries, there is rapid growing incidence rate of BC among Iranian women with increasing burden of poorer survival [1]. Despite the highly demanding research interrogate the pathogenesis of disease, a few insights provided to understand the underlying mechanism to explain the resistance to endocrine therapeutic approaches.
It is clearly established that epigenetic silencing can play a key role in governing the lack of hormonal expression [2, 3]. The hypermethylation status of specific DNA sequence of regulatory regions is mostly studied field in epigenome [4]. Accordingly, it is appealing to unravel aberrant methylation pattern at multiple upstream regulatory regions of different genes, which possibly may imply the inactivation of cis-regulatory sequence of target promoter toward silencing consequences [3, 4, 5]. However, less information has been provided to show a group of promoters of different genes, function as transcriptional factors, to regulate expression of certain putative genes.
Estradiol signaling is a predominant growth promoting pathway for great proportions of hormonal receptor positive tumors during reproductive years [6]. Estradiol mediates as ligand of steroid receptors of nuclear superfamily. Estrogen receptor-alpha (ERalpha) is encoded by ESR1 (locus: chromosome 6q25.1) and estrogen receptor-beta or ERbeta is encoded by ESR2 (locus: chromosome 14q23.2) [7]. Epigenetic inactivation of ESR1 and ESR2 receptors have been linked by numerous studies carried out hypermethylation assays [8, 9]. We previously suggested that the hypermethylation of ESR1 was detected in 51.1% of studied tumors and associated inversely with ER protein expression in breast tumor specimen [9]. It is speculated that active ERbeta functions in antagonizing ERalpha-related signaling to lead anti-proliferative activity in breast cancer cells [8]. The promoter region of ESR2 is also rich in CpG islands [5]. However, studies showed less frequency of methylated ESR2 existed in tumors, where had been detected with downregulated ERbeta in breast cancer [5, 7].
Progesterone receptor gene (PgR) (locus: chromosome 11q22.1) entails two alternative promoters in the same gene to encode two transcript variants of PgRa and PgRb [10]. The PgRb isoform contains a transcriptional activating factor placed in NH2-terminal sequence of 164 amino acids, which is absent in the structure of PgRa isoform [11]. Although progesterone acts through its response element within the active promoters of PgR to transcribe PR as a whole, ligand-dependent ERalpha signaling endorse secondary response element of PgRb to touch the bases of PR regulation [5, 12]. The upstream region of promoter of PgRb is rich in CpG island and gets prone to hypermethylation, where exists in the great deal of PR- tumors [10]. Although some studies believed in that promoters of PgR regulate the transcription of PgRa and PgRb independently [11, 13], Gaudet et al. showed that the upregulation of inactive PgRa needs active PgRb-mediated transcription [10]. Given methylation at either regulatory compartment of PgRs suggested to take part in silencing of PR protein, whereas there is question remain critical in understanding the accuracy of inter-correlations of methylated ones contributed in the regulation of PR expression in breast tumors.
Earlier evidence suggested varied potency of PgRa and PgRb to repress the activity of ER that depends on promoter status and cell type [11]. Horwitz research group indicated that PgRa can repress the activity of ERalpha and PgRb [14, 15]. Therefore it is expected that methylation-dependent silencing of PgRa may contribute to overregulation of ERalpha and PgRb and thereby may cause breast tumor cells to be responsive to endocrine therapy [15]. Moreover, some studies provide additional genetic information such as loss of heterozygosity or copy number loss in the regulatory regions of PgR to figure out a meaningful reason to underlie PgR-dependent ERalpha alterations and consequently become resistant to tamoxifen therapy [12]. However, there is sparse and unaddressed data to support the contribution of corresponding panel of genes at methylation status to be contributed in negativity of hormonal receptor.
Methylation specific PCR (MSP) is a new, low-cost and easy technique with sufficient accuracy and sensitivity to detect the existence of methylation or unmethylation using small amount of DNA [11]. Therefore, there is priority of using MSP rather than quantitative MSP (QMSP) protocols, when QMSP is always quite challenging for researchers to acquire reference cutoffs. Accordingly, it is suggested that MSP is a reasonable accurate tool to provide a new strategy for tumor detection [11, 16, 17].
Hormone receptor status or assessing the coexistence of ER and PR (ER/PR) in tumors addresses the effectiveness of adjuvant endocrine therapy in prognosis of disease in BC patients [5]. Fang et al. substantiated the new term indicated breast CpG island methylator phenotype (B-CIMP) to present hypermethylation status of both ESR and PgR genes [18]. They also discussed the importance of considering B-CIMP in detecting low metastasis-dependent poor prognosis and high rates of breast cancer survival [18, 19].
Although the aberrant methylation at regulatory regions of putative therapeutic genes has already been ascertained [4], studying set of histological grades of breast cancer might be assigned to simultaneous methylation statuses of different genes has less being noticed. Therefore, the aim of present study is to determine how accurately methylation assessment of both and either ESR and/or PgR genes can contribute to negative hormonal receptor status (ER-/PR-) in breast tumors.
Materials and methods
Study subjects
In this consecutive case series study, 139 study subjects who newly diagnosed with breast cancer (age: 35–60 years old) were recruited from Day general Hospital from December 2006 to March 2008. Patients had undergone breast-conserving surgery or radical mastectomy. Dissected breast tumor specimen was histopathological confirmed mostly as invasive ductal carcinoma (IDC) and classified into histological grades of II or III and clinical stages of II or III (tumor-node-metastasis, TNM method). A part of this core biopsy were gotten frozen immediately by liquid nitrogen right after dissection and then stored at
Pedigree analysis was conducted by well-trained questioner in order to verify positive familial history of breast cancer and other malignancies in several lines and degrees of relatives of proband (study subject). The related information detailed in previous article [9, 22].
Laboratory analysis
DNA extraction and bisulfite modification
DNA purification from tissue was carried out using Tris-SDS-based proteinase K digestion on fresh frozen tissue and subsequent phenol-chloroform DNA extraction followed by ethanol precipitation. DNA was treated with sodium bisulfite as described by Herman et al. to convert unmethylated cytosine to uracil, meantime methylated cytosine remained unchanged [9, 23]. Albeit, uracil altered to thymidine through PCR amplification.
Relative distribution of demographic, and pathologic characteristics of breast cancer across dichotomous status of ER and PR (
139)
Relative distribution of demographic, and pathologic characteristics of breast cancer across dichotomous status of ER and PR (
ER, estrogen receptor; PR, progesterone receptor; OCP, oral contraceptive; BC, breast cancer. Invasive lobular carcinoma has not being taken into account in the frequency observed in histopathologic grade stratifications.
The hypermethylation status was determined using MSP as sensitive method [16]. To test the methylation status of promoter or exon-promoter regions of bisulfite-treated DNA, primer sets used to discriminate either unmethylated or methylated sequences of bisulfite-treated DNA of a certain gene [23]. The sequences of primers were listed in supplementary Table 1. These sets of primers of ESR and PgR were selected out of those where had been shown to be prudent in association with methylation-dependent silencing in transcription of genes’ variants [24]. The PCR sets were carried out in the final volume of 25
Typical methylation-specific PCR analysis of ESR1 (U: 120 bp and M: 130 bp), ESR2 (U: 109 bp and M: 109 bp), PgRa (U: 99 bp and M: 99 bp) and PgRb (U: 203 and M: 203 bp) in primary breast carcinoma were shown individually. Visible band of PCR product in column of U and M are presentative of unmethylated and methylated status, respectively. The homogenous methylated or heterogenous methylated/unmethylated alleles were defined as hypermethylated status of gene. A positive control (CP) in every PCR set was used that was yielded from in vitro methylase reaction on peripheral blood lymphocyte DNA (SssI methylase). Bisulfite-treated normal lymphocyte DNA was used in amplification as negative control for methylation (CN). The ladder was shown as a marker size.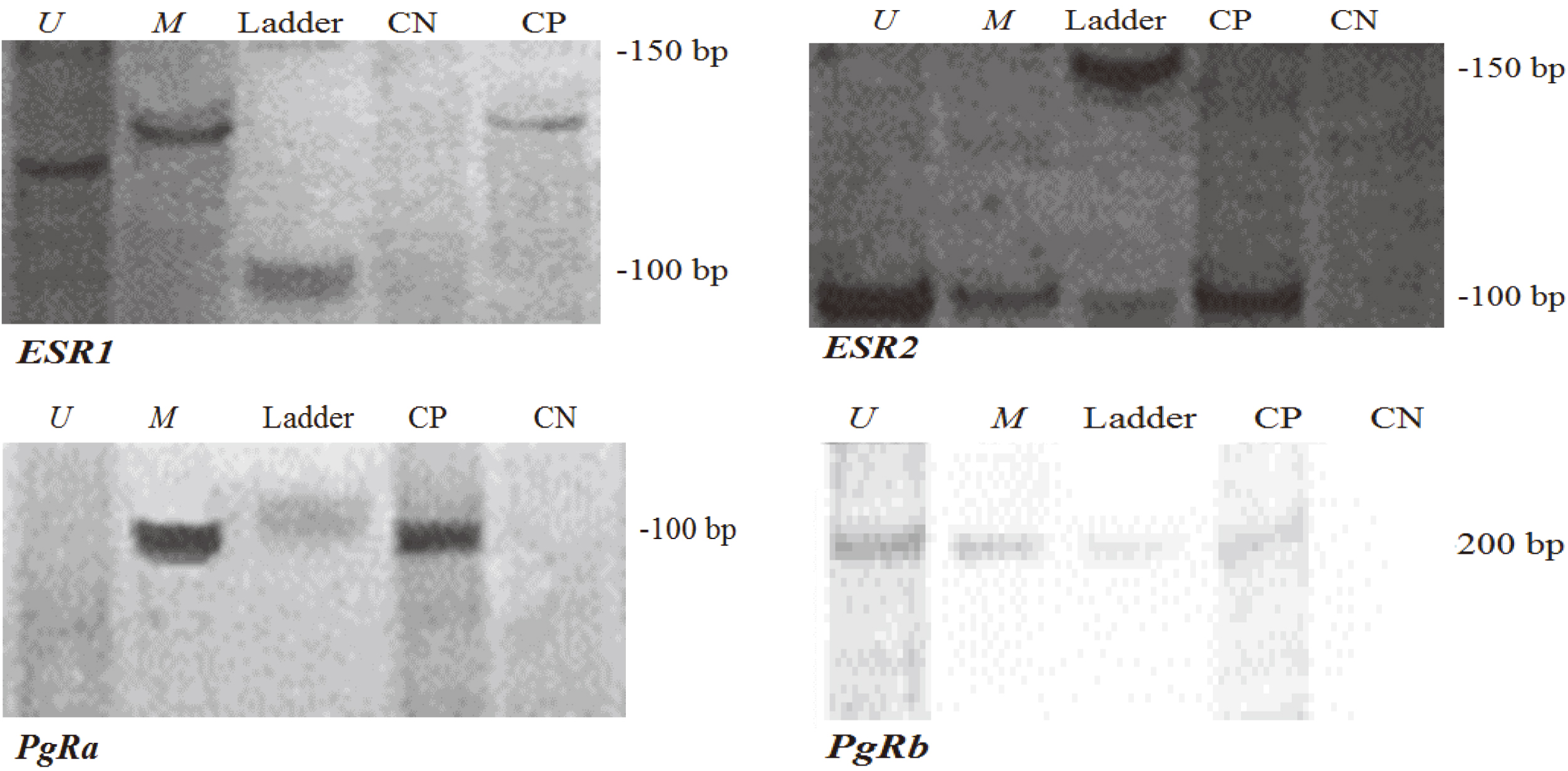
Purified MSP product was cloned into R/T 540 TA cloning vector (Fermentas, Germany), according to manufacturer’s guideline. Plasmid DNA was purified by means of plasmid minikit (Fermentas, Germany). Later MSP assays on plasmid DNA could validate the specificity of plasmids for certain region. Automated sequencer (ABI automated DNA sequencer) was employed to provide DNA methylation pattern out of sequencing results of single amplicon. T7-based sequencing primers were used.
Extraction of total RNA from tissue specimen was performed using QIAzol (Qiagen, Germany; Cat. No. 79306). Extracted RNA were undergone a cleanup procedure to remove DNA by means of RNeasy MiniKit (Qiagene, Germany; Cat. No. 74104). Purified RNA were normalized for concentration and subsequently treated with QuantiTect reverse transcriptase (Fermentase, Germany; Cat. No. 205311, contained reagents to remove genomic DNA contamination) to synthesize complementary DNA (cDNA). Relative expression levels were measured using Corbett real-time PCR (Corbett Research, Canada). Primers used in reaction mixture were RT2qPCR Primer Assay SYBR Green Human ESR1 (SuperArray BioScience Corporation, USA; Cat. No., PPH01001A; RefSeq accession#NM-000125.2), RT2qPCR Primer Assay-SYBR Green Human ESR2 (SuperArray BioScience Corporation, USA; Cat. No., PPH00992B; RefSeq accession#NM-001437.2) and RT2qPCR Primer Assay-SYBR Green Human PGR (SuperArray BioScience Corporation, USA; Cat. No., PPH01007E; RefSeq accession#NM-000926.3). Each expression measurement of each sample was performed in duplicate reactions. The calculation of relative expression level was carried out based on measured cycle threshold (CT) using 2
Typical curves of fluorescence variations detected through PCR amplification (RT-PCR) of different genes were run in duplicate. The relative expression levels of gene of interest (a & b: ERbeta, e & f: ERalpha, g & h: PgR and i: none template controls) relative to reference gene (c & d: ribosomal 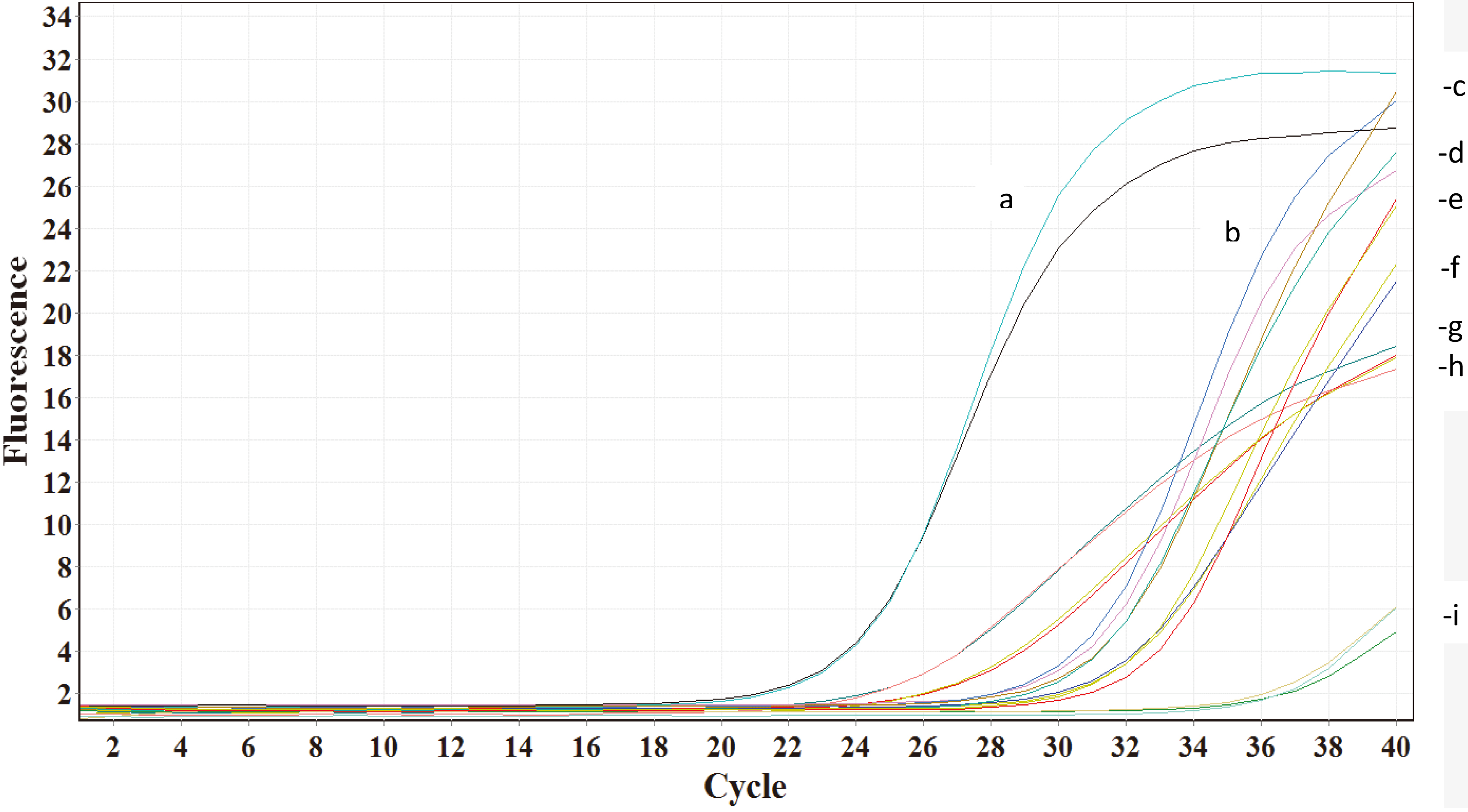
Statistical analysis was conducted using SPSS statistical package for Windows (version 13.0; IL, USA). The association of hypermethylation status with IHC data (as dependent data) was analyzed using chi-square test. When both variables were dichotomous, phi (
The frequency of tumors with methylation at individual gene (ESR1, panel a; ESR2, panel b; PgRa, panel c; PgRb, panel d), where ER/PR features distributed through histopathological grades (tumor grades 1, 2 and 3). Chi-square test was performed to present level of significance (P-value) to compare frequency of tumors between ER+/PR+, ER+/PR- and ER-/PR- tumors within each histological tumor grade. ER, estrogen receptor; PR, progesterone receptor; ESR1, estrogen receptor 1 gene (ERalpha); ESR2, estrogen receptor 2 gene (ERbeta); PgRa, progesterone receptor promoter a; PgRb, progesterone receptor promoter b; 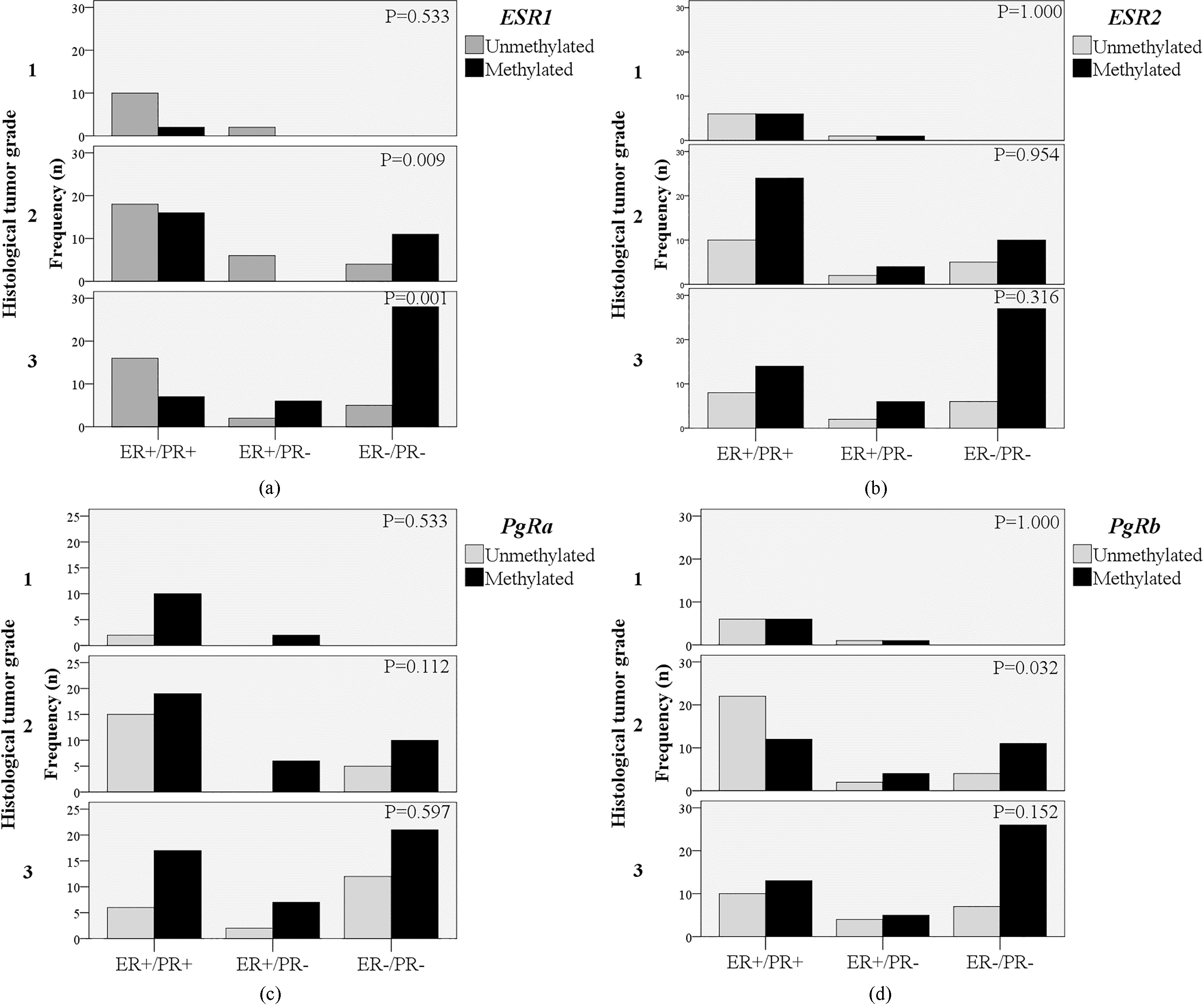
Structural equation modelling (SEM) was carried out using analysis of moment structure (AMOS) by means of IBM SPSS AMOS (ver. 16) to prove the goodness-of-fit (adequacy) of theoretical model based on the hypothesis provided in present study. The path diagram was conceptualized to show the simplest form of theoretical regression model that managed in separate group of menopausal status. Thereafter, a multivariate regression model was identified after revealing potent covariates. In the process of SEM analysis, maximum likelihood estimation (MLE) was performed to estimate the model parameters. Standardized regression weight as beta (
The relative frequency of methylation versus unmethylation statuses of ESR1, ESR2, PgRa and PgRb genes across the dichotomous IHC staining results of ER(-/+) and PR (-/+) of breast tumors (
ER, estrogen receptor; PR, progesterone receptor; ESR1, estrogen receptor 1 gene (ERalpha); ESR2, estrogen receptor 2 gene (ERbeta); PgRa, progesterone receptor promoter a; PgRb, progesterone receptor promoter b;
Pathological data and general characteristics
The hypermethylation status of ESR1 exon 1 was shown in 51.5% of 136 breast tumors. The methylation of ESR2 was detected in 69.6% of 135 tumors. MSP was also able to detect 68.6% of methylations at the promoter of PgRa and 57.7% of PgRb among 137 breast tumors.
Age at diagnosis of participants afflicted with BC was 49.1
The general characteristics of BC participants are summarized in Table 1 (see details in supplementary Table 1). Participants who were characterized with ER- tumors fell into the categories of tumor grade III (68.8%) and tumor size
Figure 3 demonstrates the arrangement of methylated gene through rankings made upon both histopath-ological grades and ER/PR features (details were described in Supplementary Fig. 3). Advanced tumor grades (II) along with ER-/PR- tumors were significantly attributed to ESR1 hypermethylation (
Correlation between methylation at individual gene and IHC data
The relative frequency of methylation status across dichotomous IHC status of ER, PR and also in terms of mutual ER/PR status (ER+/PR+, ER+/PR- and ER-/PR-) in breast tumors were demonstrated in Table 2 (see details in supplementary Table 2). DNA methylation at ESR1 (exon 1) was significantly frequent in ER- (83.0%; 39 out of 47 tumors;
Structural equation modelling (SEM)
SEM was assigned to imply the path diagram entailing the sequential contribution of hypermethylation status and mRNA expression levels toward IHC-based protein expression levels (Fig. 5, details of additional data processing were described in Supplementary Fig. 5). Primarily, a parsimonious model was identified in premenopausal women showing the positive contribution of relative expression of ERalpha gene to support ER protein expression status significantly (coefficient
The relative magnitude of contributions between both ESR and PgR genes (hypermethylated status) in association with certain combinative hormonal receptor status as ER/PR
The relative magnitude of contributions between both ESR and PgR genes (hypermethylated status) in association with certain combinative hormonal receptor status as ER/PR
ER, estrogen receptor; PR, progesterone receptor; ESR1, estrogen receptor 1 gene (ERalpha); ESR2, estrogen receptor 2 gene (ERbeta); PgRa, progesterone receptor promoter a; PgRb, progesterone receptor promoter b;
The average relative expression levels of ERalpha, ERbeta and PgR across ER+/PR+, ER+/PR- and ER-/PR- statuses were compared using analysis of variance (ANOVA) and shown in mean 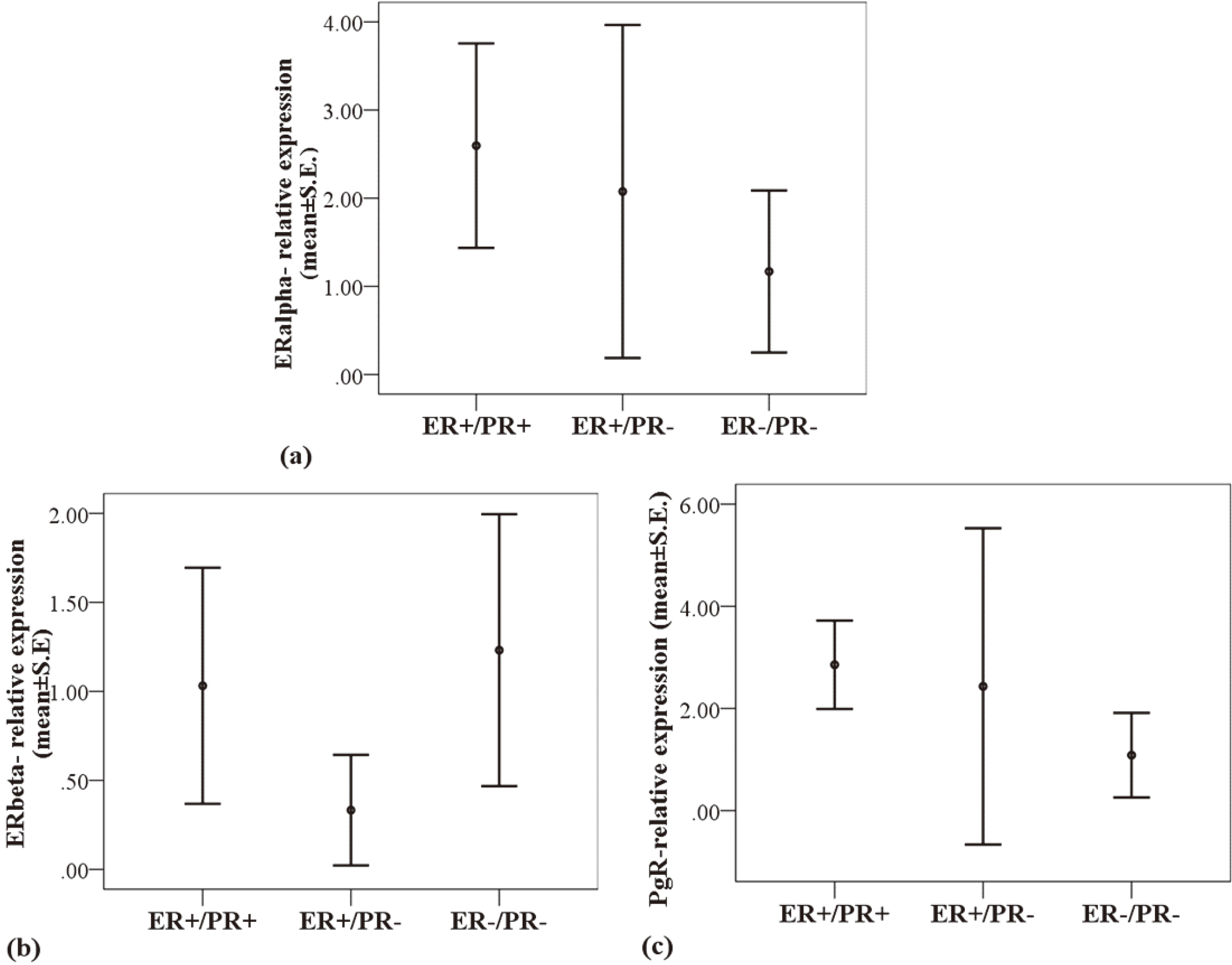
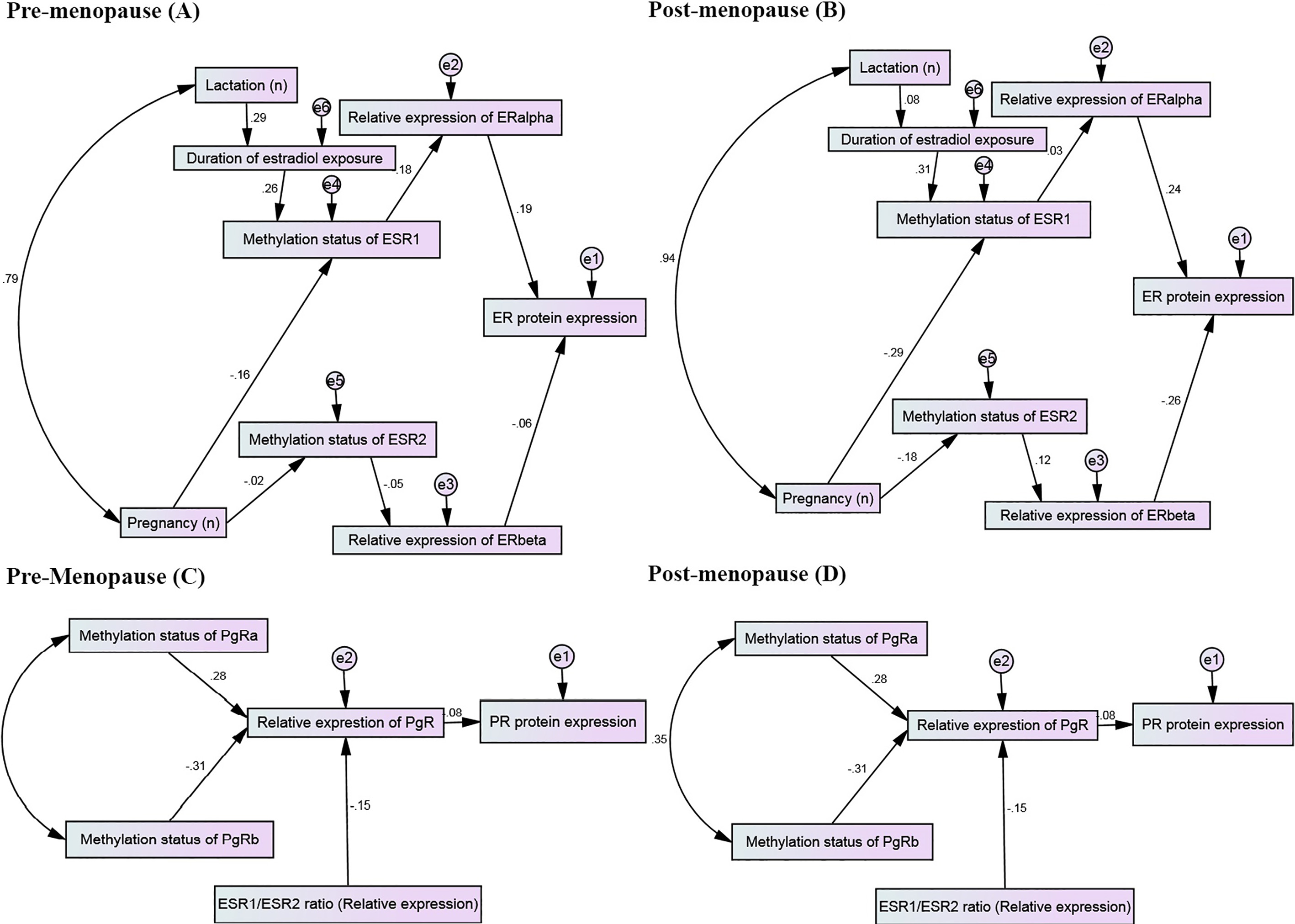
The proposed path diagrams of the implied regression models fitted quite well in parsimonious model to study the regression analyses among hypermethylation status, relative expression levels and protein expression status (immunohistochemistry data). The MSP results of ESR1 and ESR2 genes, relative expression levels of ERalpha and ERbeta and their consequent contribution to tumor ER status where shown in panels A (pre-menopauses) and B (post-menopause). Path diagram identified through confirmatory multi-regression model to assess contributions of MSP results of PgRa and PgRb to variations of relative expressions of PgR and consequently toward determining tumor PR status, where whole diagram depicted for pre-menopause women (panel C) and post-menopause counterparts (panel D) individually. Maximum likelihood estimation was performed to test the fitting function or estimation procedure of parameters in models. Values on arrows were beta (
Table 3 summarizes the results showing the co-contribution of hypermethylations at ESR1 and ESR2 in tumors cross-tabulated with the methylation status of PgRs (promoters of PgRa and PgRb) to figure out how they attribute to the combinative status of hormonal receptors of ER/PR as a dependent categorical variable (see details in supplementary Table 3). Those tumors hypermethylated at ESR1 promoter accompanying with methylated PgRb was often classified into ER-/PR- subclass (29 out of 45 tumors methylated for ESR1; 64.4%,
Sensitivity, specificity and diagnostic accuracy measured to evaluate how accurately methylation of a gene can detect certain hormonal receptor status (
137)
Sensitivity, specificity and diagnostic accuracy measured to evaluate how accurately methylation of a gene can detect certain hormonal receptor status (
ER, estrogen receptor; PR, progesterone receptor; ESR1, estrogen receptor 1 gene (ERalpha); ESR2, estrogen receptor 2 gene (ERbeta); PgRa, progesterone receptor promoter a; PgRb, progesterone receptor promoter b; U, unmethylated and heterogeneously methylated gene; M, methylated gene; DOR, diagnostic odds ratio; LR, likelihood ratio; PPV, positive predictive value; NPV, negative predictive value.
The validity of methylation variables to detect accurately the true ranks of IHC-based ER and PR expressions (consistency of both techniques) were presented in Table 4, using sensitivity, specificity and other diagnostic characteristics (see details in supplementary Table 4). The methylation at ESR1 gene provided 83.0% sensitivity and 65.2% specificity to detect ER- status. DOR resulted by using the hypermethylation status of ESR1 as a test biomarker was 12.0, which showed MSP as a quite potent diagnostic tool to detect tumoral ER status. The accuracy of MSP data to evaluate the concordance of methylated PgRb with PR- status was comparably greater than the performance obtained by using methylation of PgRa that was weakly enabled to detect the PR- status. The related sensitivity (74.1%), specificity (53.0%) and NPV (75.9%) to detect the association of methylated PgRb and PR- were not strong but adequate enough rather than to what extent PgRa took part in diagnostic performance.
The typical characteristics to tests the diagnostic accuracy of hypermethyaltion status to detect the binary status of ER/PR summarized in Table 5 (see details in supplementary Table 5). The concurrent methylations on ESR1 and PgRb was also a state resulted in better diagnostic performance to rank tumors in the category of ER-/PR- stained tumors, by which sensitivity at 78.0% and specificity at 62.5% were attained (DOR
Hypermethylation at specific regions of DNA is a widely studied spectrum of epigenetics implies fundamentally the gene inactivation [2, 20]. Estrogen and progesterone receptors (ER/PR) of primary BC is the single most reliable indicator to detect the responsiveness of BC patients to adjuvant endocrine-therapy [30]. Our main finding provided the importance of methylated ESR1 in association with ER-/PR- status, while the hypermethylation of PgRb was the only ER-dependent downstream co-effector variable to support the possible contribution to PR- and the silent hormone receptor status (ER-/PR-). The validation dataset also supported the impact of hypermethylations at ESR1 and PgRb as reproducible set of indicator genes enabled to accurately diagnose negative hormonal receptors subtype of tumors.
Sensitivity, specificity and other parameters concerning diagnostic accuracy were measured to evaluate how accurately methylation of an ESR gene in connection with another methylated PgR can detect certain hormonal receptor status as ER/PR (
137)
Sensitivity, specificity and other parameters concerning diagnostic accuracy were measured to evaluate how accurately methylation of an ESR gene in connection with another methylated PgR can detect certain hormonal receptor status as ER/PR (
ER, estrogen receptor; PR, progesterone receptor; ESR1, estrogen receptor 1 gene (ERalpha); ESR2, estrogen receptor 2 gene (ERbeta); PgRa, progesterone receptor promoter a; PgRb, progesterone receptor promoter b; U, unmethylated and heterogeneously methylated gene; M, methylated gene; DOR, diagnostic odds ratio; LR, likelihood ratio; PPV, positive predictive value; NPV, negative predictive value; Ref., reference.
The MSP results presented that hypermethylation of exon 1 at ESR1 gene as the prominent factor contributed to the absence of tumoral ER expression. The methylated ESR1 was also notable element of studied profile correlated with tumor status of ER-/PR-. Given that the methylated status of ESR1 can be considered as a sensitive marker in detecting the lack of ER expression with better performance to predict true negative ER protein expression rather than other genes methylated. Although, in MSP assays of frozen tissues, some studies showed the importance of methylation on promoter A of ESR1 in relevant protein expression [10], Widschwendter et al. by performing MethyLight study found out that methylation at ESR1 promoter A can marginally associate with the lack of ER expression [1]. Many epidemiologic and in vitro experimental observations showed that promoter-exon 1 regions of ESR1 gene contain CpG islands, which are highly vulnerable to become methylated in ER- breast tumor or cell lines [9, 13, 24, 31]. Growing evidence showed that ESR1 hypermethylation is a frequent epigenetic change considerably pertinent in terms of becoming resistance to endocrine therapy [10, 32, 33].
SEM analysis gives rise to the concept that ERalpha downregulation can potently attributed to ESR1 hypermethylation, all of which were associated positively with ER protein expression in premenopausal women. Although, ERbeta expression levels correlated negatively with ER status and suggesting the downregulating effects of ERbeta on ER status in postmenopausal women, the related parameters was not statistically significant. Accordingly, Liu et al. showed that exon 1 of ESR1 is rich in clusters of CpGs, where they become methylated cause stable long term repression of ER transcription [33]. When loss of estradiol exposure occurs and persists for instance in the exposure of inhibitors of aromatase, the methylation subsequently could be developed further into the promoter of ESR1 to surrogate original repressors, this is implicated in heritable long term repression attributed to established methylation-dependent heterochromatin structure [33]. Although, ESR2 gene is responsible to encode transcript of ERbeta protein, our findings did not support the correlation between the methylated ESR2 and negative hormone receptor status or ER-.
The hypermethylation at promoter of PgRb gene was correlated significantly with PR- tumors. The present findings also supported the validity of MSP data to test the association of methylated PgRb (rather than PgRa) with PR- tumors with quite acceptable sensitivity. However, when methylated PgRa considered as single gene did not show significant correlation. The expression of PR in tumoral cells provide additional option to the endocrine therapy in favor of PR agonists to inhibit proliferation of breast tumor [34].
Lapidus’s observations, as an earlier evidence, provided a view of tide correlation between hypermethylation at certain CpG islands of PgR promoter and lack of expression [13]. Nevertheless, Pathiraja et al. suggested that the presence of methylation at the promoter sites of PgRa and PgRb could be potentially applied as surrogate markers of specific PR isoforms [15], our results put emphasize on uniquely methylated status at PgRb in PR- tumors. The active PgRb has been addressed to have a stronger effect as transcriptional activator than PgRa on downstream genes, whereas it is highly methylated in tumors of constituent resistance to endocrine-therapy [11, 35, 36].
Present findings showed that the hypermethylation of ESR1 gene was considerable contributor in PR- tumors. Accordingly, observation of Widschwendter et al. indicated that ESR1 methylation is a notable feature in detecting PR status [1]. Given the methylation of PgR is a hallmark epigenetic event against PR expression status [10], our findings showed that ER protein expression or ESR1 methylation was either contributed to altered PgRb expression levels. This significant association between ESR1 methylation and PR status has been addressed to be partially attributed to the region of PR response element induced by upstream ER signaling [30]. Taken together, the contribution of methylation-dependent silent PR may initially depend on epigenetic deactivation of ER [12].
From present study, a sensible correlation between methylated PgRb and ER status was notable evaluated by adequate sensitivity of MSP and true negative result. Some studies supported the role of active PgRb in repressing the estradiol-dependent hyperplasia [11]. Surprisingly, our findings showed that PgRb methylation was associated inversely with ER status, suggesting the possible methylated-dependent PR downregulation may have effects on the transcriptional status of ER+. Consistently, Gaudet et al. showed that DNA methylation at promoters of PgR can associate with less protein expression of ERalpha [10]. Kim et al. have pronounced that BC patients with ER+/PR+ tumors showed better prognosis, when treated with adjuvant ER antagonists rather than those with ER+/PR- tumors [37]. Generally, it is well-known that progesterone can inhibit the ERalpha (protein) related signaling and subsequently estradiol-dependent proliferation of BC cells [12]. In addition, abundant evidence indicates that upregulated PR mutual to ER in BC tumors can warrant better prognosis of disease, when compared to tumors of solo-expression of ER [12, 30, 38].
Although new advances in epigenetics to detect the methylation status at particular region of specific genes is growing rapidly in recent years, there is appealing interests to provide a valid technique farther than existing conventional assays as immunoreactivity of expressed proteins [17]. The MSP is a sensitive method to detect the allele-specific methylation status. There are lots of studies showing that methylation outputs resulted from MSP are in appreciable agreement with data measured by QMSP [15, 39, 40]. Even MSP provides better validity because of greater sensitivity endorsed by MSP [15]. In addition, identifying appropriate cutoff point as a standard reference to rank accurately the correct methylation status with appropriate sensitivity remains as a challenging issue in QMSP [17]. Based on a meta-analysis data, no advantage of QMSP is likely suggested in comparison to MSP data [16]. The possible variation of MSP results on a gene may depends on the MSP assay design such as selected sequence of primers, sample size and population-based bias linked to epidemiologic study [41].
The major limitation of present study was small number of subjects, thereby limited stratification analyses regarding to menopausal status or age categories were undertaken. Therefore, our findings could be cautiously interpreted. Although the isoforms of ESR and PgR genes have been targeted, there could be other potent transcriptional enhancers or repressors act in the regulation and activating hormonal receptors. Another limitations of present study was small amount of cDNA prepared to test expression levels. Therefore, expression levels of each sample was tested in duplicate reactions. Nevertheless background histories of smoking and OCP usage considered as nonsignificant covariates in the present study as potent life-style related factors, which might affect silencing of methylation-related genes, considering the mediating effects of hormonal derivatives are inevitable and remained to be elucidated.
The MSP is a quite sensitive method to detect DNA methylation at ESR1 (exon 1) in association with ER- breast tumors. In addition, methylated ESR1 was a prominent candidate in negativity of hormonal receptor status (ER/PR).The contribution of methylated PgRb was determined in association with PR-, ER- and inactivity of hormonal receptors, suggesting isoform-specific functions of PR. However, the accuracy of MSP to detect both ESR1 and PgRb in the methylation status was sufficiently credited to diagnose tumors with lack of ER/PR expressions. Recent efforts made to assimilate transcriptome in the light of epigenomic markers to provide critical insights into an alternative molecular-based subclassifiaction of breast tumors.
Footnotes
Acknowledgments
We are thankful to all valued and kind participants and supports of our colleagues in Day General Hospital. This study was financially supported by research affair of National Nutrition and Food Technology Research Institute, Shahid Beheshti University of Medical Sciences, Iran. The authors’ responsibilities were as follow: S.P., F.A.T., M.A., and P.M.: designed the research; S.P., M.A., and P.M.: conducted the research; S.P. and S.S.: analyzed the data; and S.P., P.M. and S.S.: wrote the manuscript. All authors have read and approved the final manuscript. This study was funded by National Nutrition and Food Technology Research Institute, Shahid Beheshti University of Medical Sciences (grant number 245).
Conflict of interest
All authors declare that they have no conflict of interest.
Supplementary details describe:
![]()
![]()
![]()
Table 4
![]() ].
].
The probability of generating true results using MSP (test) versus IHC data as golden standard was evaluated by likelihood ratios (LR) as well. The tests with LR+ (sensitivity/false-positive error rate; similar to the ROC curve) larger than one are helpful to screen methylation results with better sensitivity of MSP to be associated with silent hormonal receptor expression, whereas the LR- (false negative error rate/specificity) less than one could reflect a high degree of specificity of MSP to detect methylation in association with hormonal receptor status. Consequently, LRs are independent of the prevalence rate or numbers of observations [1]. LR values near one show no substantial change. The ratio of LR+ to LR- was calculated to estimate the diagnostic odds ratio as another global measure for diagnostic accuracy [2, 3]. Indeed, a test with high specificity and sensitivity with low rates of false positives and false negatives could result in a high odds ratio. For example, a diagnostic odds of 10 means that the true results of a test is 10 times higher than its false results [3]. Required details were expressed in footnote of ![]() .
.
Table 5
![]() ].
].
The probability of generating true results using MSP (test) versus IHC data as golden standard was evaluated by likelihood ratios (LR) and described in details in supplementary of Table 4 [1]. The ratio of LR+ to LR- was calculated to estimate the diagnostic odds ratio [2, 3]. A performance of the candidate diagnostic test (methylation status) was examined against a known reference (Ref.) or golden standard which was IHC data of ER+/PR+. Required details were expressed in footnote of ![]() .
.
![]() ]. In summary, the value of chi-square divided by given degree of freedom (CMIN/df) used as a criterion to show appropriate fitness when the value is
]. In summary, the value of chi-square divided by given degree of freedom (CMIN/df) used as a criterion to show appropriate fitness when the value is
Supplementary data
| Supplementary | |||||||
| List of MSP specified sequences of primers | |||||||
| Gene | U/M | Forward | Reverse | Upstream | Annealing | Product | Reference |
| location | temperature ( |
size (bp) | |||||
| ESR1 |
|
5’-GGATATGGTTTGTATTTTGTTTGT-3’ | 5’-ACAAACAATTCAAAAACTCCAACT-3’ | +225 to +345 | 57.0 | 120 | Lapidus et al. [1, 24] and Pirouzpanah et al. [9] |
|
|
5’-GATACGGTTTGTATTTTGTTCGC-3’ | 5’-CGAACGATTCAAAAACTCCAACT-3’ | +225 to +345 | 57.0 | 130 | Lapidus et al. [1, 24] and Pirouzpanah et al. [9] | |
| ESR2 |
|
5’-TTTGGAAGGTGGGTTTGGTT-3’ | 5’-CACATACAAATATAATAACTAACA-3’ | +191 to +300 | 54.0 | 109 | Hirata et al. [42] and Meyer et al. [43] |
|
|
5’-TTTGGAAGGTGGGTTTGGTC-3’ | 5’-CGCATACAAATATAACTAACG-3’ | +191 to +300 | 56.0 | 109 | Hirata et al. [42] and Meyer et al. [43] | |
| PgRa |
|
5’- ATGGGTTATTTTTTTTTTG -3’ | 5’- TAAAATATACACCCTCCACA -3’ | +894 to +993 | 47.0 | 99 | Sasaki et al. [44] and Kastner et al. [45] |
|
|
5’-ACGGGTTATTTTTTTTTCG-3’ | 5’-TAAAATATACGCCCTCCACG-3’ | +894 to +993 | 47.0 | 99 | Sasaki et al. [44] and Kastner et al. [45] | |
| PgRb |
|
5’-TGATTGTTGTTTGTAGTATG-3’ | 5’-CAACAATTTAATAACACACA-3’ | +156 to +358 | 47.0 | 203 | Sasaki et al. [44] and Chew et al. [46] |
|
|
5’-TGATTGTCGTTCGTAGTACG-3’ | 5’-CGACAATTTAATAACACGCG-3’ | +156 to +358 | 48.5 | 203 | Sasaki et al. [44] and Chew et al. [46] | |
ESR1, estrogen receptor 1gene (ERalpha); ESR2, estrogen receptor 2 gene (ERbeta); MSP, methylation specific PCR; PgRa, progesterone receptor promoter a; PgRb, progesterone receptor promoter b; bp, base pair;
