Abstract
OBJECTIVE:
Glioma-associated oncogene homolog 1 (Gli1) in Hedgehog signal pathway regulates Cyclin D1 expression, cell cycle or proliferation modulation. Esophageal cancer patients had significantly elevated Gli1 expression, which is related with survival and prognosis. It has been demonstrated that the level of miR-150 was decreased in esophageal cancer patients compared to normal control. As a complementary relationship exists between miR-150 and 3’-UTR of Gli1, this study investigated if miR-150 played a role in regulating Gli1 expression, and proliferation or cell cycle of esophageal cancer cells.
PATIENTS AND METHODS:
Esophageal squamous cell carcinoma (ESCC) patients from our hospital were recruited to collect tumor and adjacent tissues for miR-150 and Gli1 expression. Esophageal carcinoma cell line EC9706 and normal esophageal epithelial cell line HEEC were compared for expression of miR-150, Gli1 and Cyclin D1. Dual luciferase reporter gene assay examined the targeted relationship between miR-150 and 3’-UTR of Gli1. In vitro cultured EC9706 cells were treated with miR-150 mimic, si-Gli1 or the combination of miR-150 mimic and si-Gli1, respectively, to check their gene expression, cell cycle and proliferation.
RESULTS:
ESCC tissues had significantly higher Gli1 expression and lower miR-150 expression. EC9706 cell also had higher Gli1 expression than that in HEEC, whilst miR-150 was down-regulated. Via targeting 3’-UTR of Gli1 gene, miR-150 inhibited its expression. Transfection of miR-150 mimic, si-Gli1 or the combination of miR-150 mimic and si-Gli1, respectively, remarkably decreased expression of Gli1 and Cyclin D1 expression in EC9706 cells, whose cell cycle arresting at G0/G1 phase was enhanced with weakened proliferation.
CONCLUSIONS:
MiR-150 can induce G0/G1 cell cycle arresting and weaken proliferation of esophageal carcinoma cells via targeted inhibition on Gli1 and downstream expression of Cyclin D1.
Introduction
Esophageal cancer (EC) is a type of common malignant tumor that invades digestive system. Esophageal squamous cell carcinoma (ESCC), as a type of EC, was commonly found in China [1]. Glioma-associated oncogene homolog 1 (Gli1) belongs to Gli gene family, which serves as the transcriptional effector of Hedgehog signal pathway [2]. Gli1 contributes to the transduction from extracellular Hedgehog signal pathway into intracellular Gli1 signal, which exerts transcriptional modulation effects on the expression of nuclear target genes [3]. Enhancement of both function and expression of Gli1 can induce the over-activation of Hedgehog signal pathway, and is closely correlated with occurrence, progression and prognosis of various tumors including breast cancer [3], colorectal cancer [2], pancreatic carcinoma [4] and prostate cancer [5]. Previous studies showed significantly elevated expression of Gli1 in tumor tissues of esophageal cancer, and its correlation with patient’s survival and prognosis [6, 7, 8]. MicroRNA (miR) is a group of endogenous non-coding small RNAs, the sequences of which are highly conserved in eukaryotes, and can regulate the expression of target gene via complete or incomplete binding on 3’-untranslated region (3’-UTR) of mRNA, thus mediating cell proliferation, apoptosis and differentiation. Abnormal expression of miR is closely correlated with tumor occurrence, and may exert oncogene or tumor suppressor gene functions depending specific target genes [9, 10]. Previous study showed significantly decreased expression of miR-150 in EC tissues [11], suggesting its possible correlation with EC occurrence. Bioinformatics analysis showed targeted complementary relationship between miR-150 and 3’-UTR of Gli1. This study thus investigated if miR-150 played a role in regulating Gli1 expression and affecting cell proliferation or cell cycle of ES.
Materials and methods
Clinical information
A total of 39 ESCC patients who received treatment in the first affiliated hospital, and college of clinical medicine of Henan university of science and technology from January 2015 to May 2016 were recruited. All patients were diagnosed by pathology examination. There were 22 males and 17 females, aging between 39 and 77 years (average age
Histology of the patients
Pairs of fresh normal esophageal tissues and corresponding esophageal carcinoma tissues from patients with pathologically and clinically confirmed esophageal squamous cell carcinoma were collected by the first affiliated hospital, and college of clinical medicine of Henan University of Science and Technology. Primary tumor regions and the corresponding histological normal esophageal mucosa from the same patients were separated by experienced pathologists, and immediately stored at
Cell culture
EC cell line EC9706 (ScienCell Research Laboratories, US) and normal esophageal epithelial cell line HEEC (Jiniou Bio, China) were seeded into RPMI 1640 medium containing 10% FBS (Gibco, US), 100 U/ml penicillin and 100
Construction of luciferase reporter gene vector and dual luciferase reporter gene assay
Using HEK293T genomic DNA as the template, full length fragment of wild type or mutant forms of 3’-UTR of Gli1 gene was amplified and were cloned into pLUC plasmid (Ambion, US). Recombinant plasmid was then used to transform DH5
Bioinformatics analysis
Bioinformatics analysis was conducted according to the on-line data bases including MiRanda, TargetScan and PITA. Data were downloaded from the MiRanda database MirBase and
EC9706 cell transfection
In vitro cultured EC9706 cells were divided into five groups: miR-NC transfection group; miR-150 mimic transfection group; si-NC transfection group; si-Gli1 group; and miR-150 mimic
qRT-PCR for gene expression assay
Cultured cells were lysed by Trizol method (Invitrogen, US). RNA was obtained by chloroform separation, isopropanol precipitation, washing in 75% ethanol, and was re-suspended in RNAse-free water. PimerScript RT reagent Kit (Takara, Japan) was used to synthesize cDNA from RNA by reverse transcription. cDNA was kept at
Western blot
Cells were collected for proteins extraction using routine method. Proteins were added into three times volumes of 4X loading buffer for 5 min boiling. 40
Flow cytometry for cell cycle
Cells were digested in trypsin and were centrifuged at 1 000 g for 5 min. Cells were re-suspened in pre-cold PBS. After re-centrifugation, the supernatant was discarded. Cells were fixed in 1 ml pre-cold 70% ethanol for 24 h at 4
Lower miR-150 expression and higher Gli1 expression. (A) qRT-PCR for Gli1 mRNA expression; (B) qRT-PCR for miR-150 expression; (C) Western Blot for Gli1 protein expression. *, 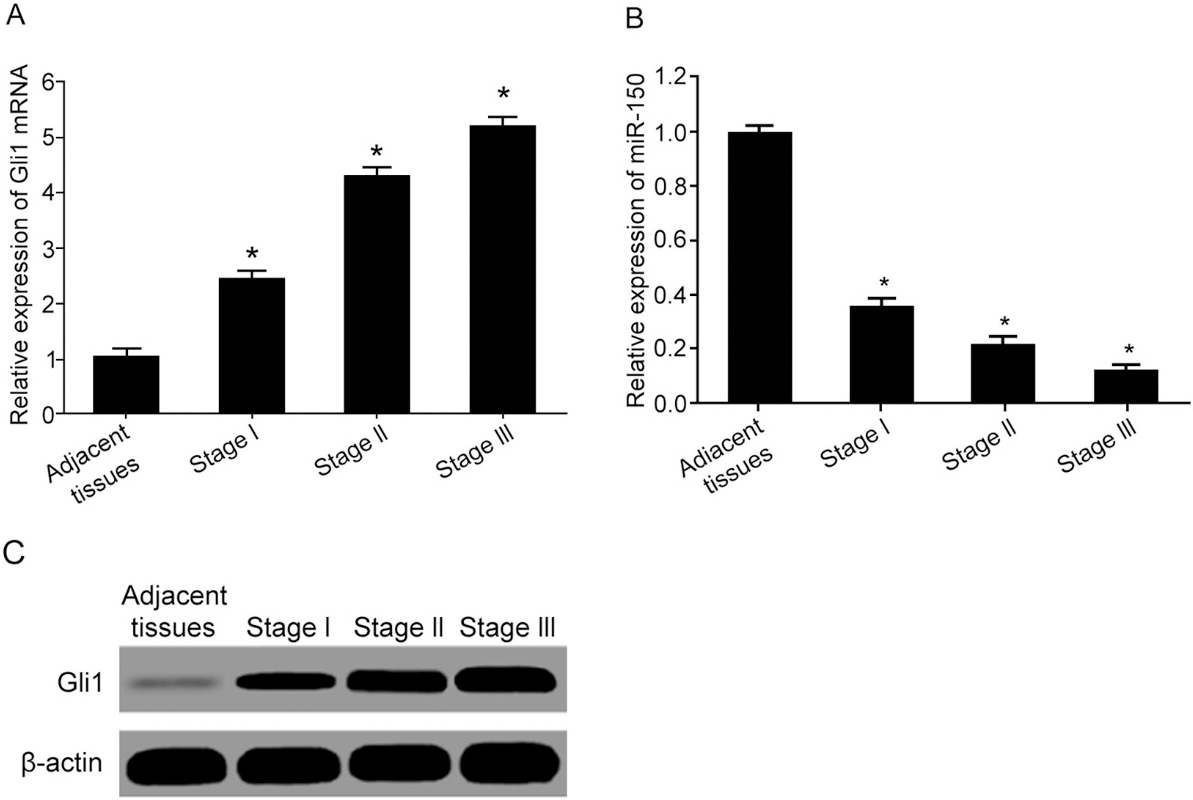
All transfected cells were seeded into 96-well plate at 1
CFSE staining for cell proliferation
1 mL CFDA SE marker buffer was used to re-suspend all transfected cells (Beyotime, China). After mixing with 1 mL 2XCFDA SE stock buffer, cells were incubated at 37
Statistical analysis
SPSS18.0 software was used for data analysis. Measurement data were presented as mean
Lower miR-150 and higher Gli1 expression in EC cells. (A) CFSE staining for cell proliferation; (B) PI staining for cell cycle assay; (C) qRT-PCR for gene expression; (D) Western Blot for protein expression. (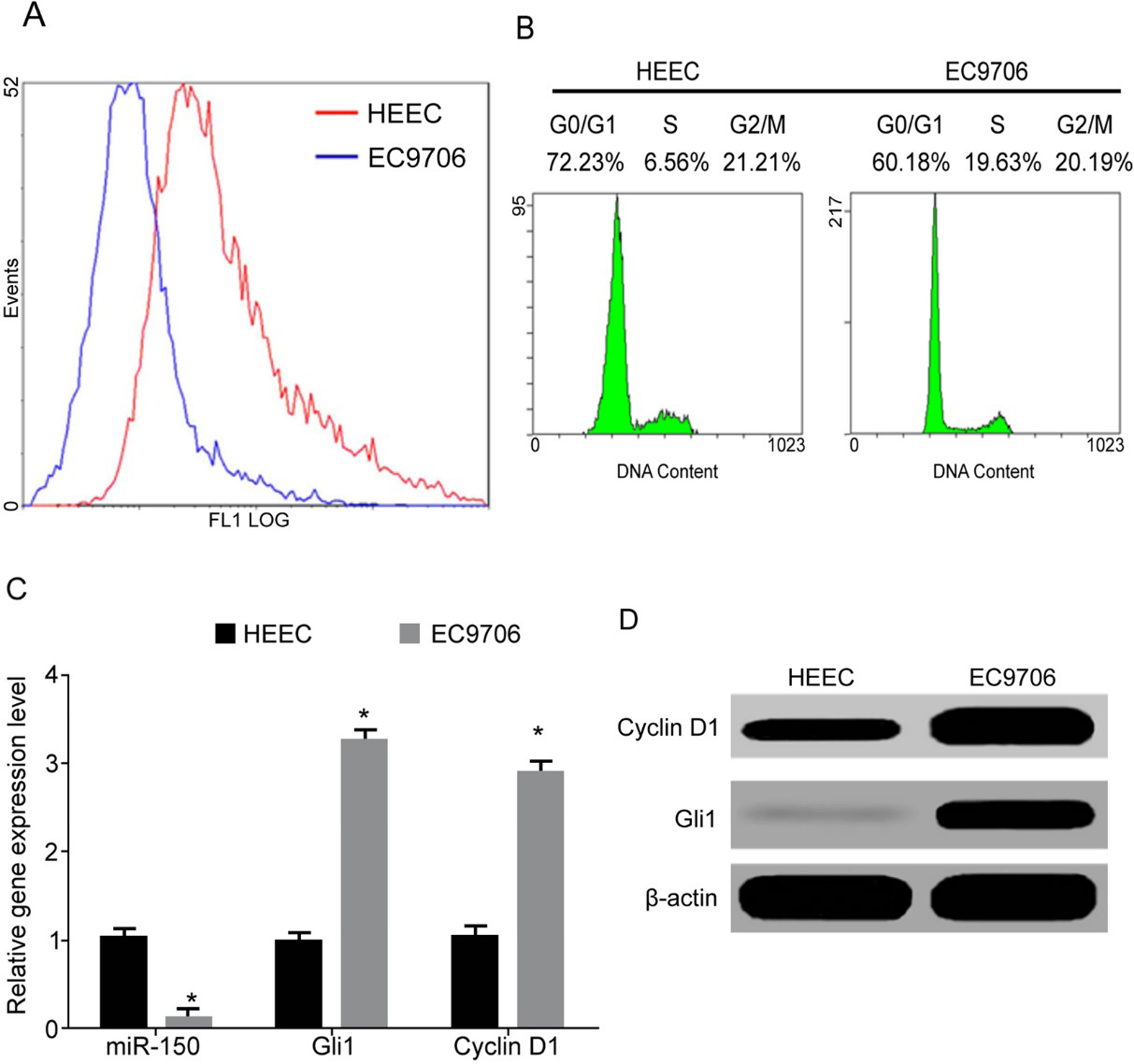
Gli1 as the target gene of miR-150. (A) Function sites between miR-150 and 3’-UTR of Gli1/Gli1 mut mRNA; (B) Dual luciferase reporter gene assay. *, 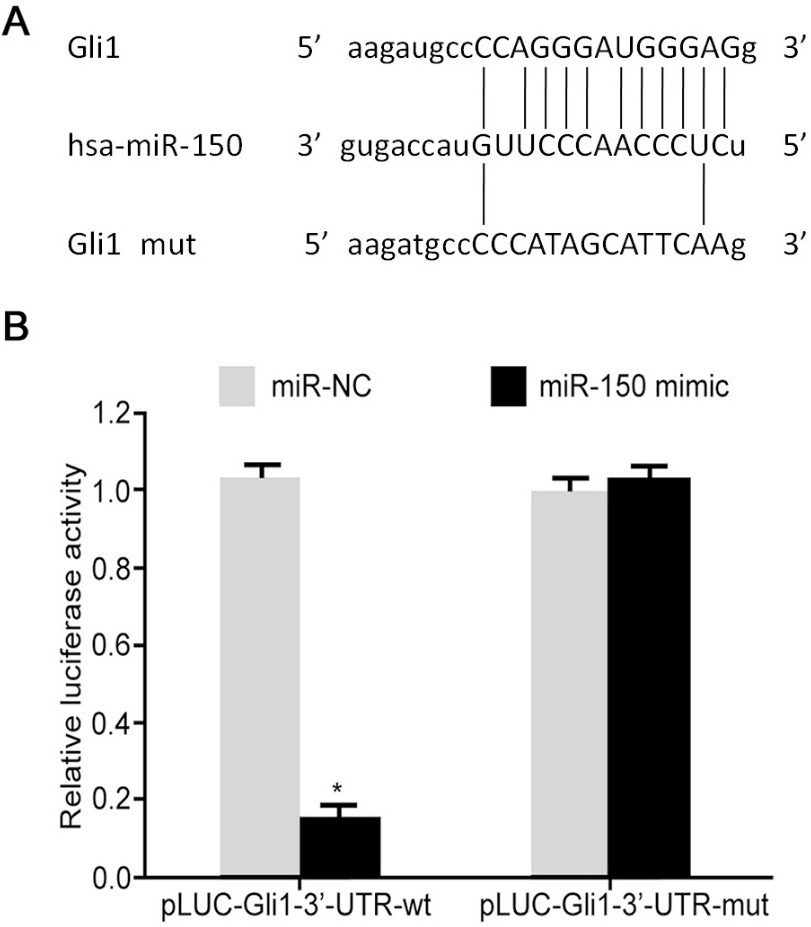
Reduced miR-150 and elevated Gli1 expression in ECSS tissues
Test results showed that, compared to adjacent tissues, ESCC tumor tissues had significantly elevated mRNA (Fig. 1A) and protein of Gli1 (Fig. 1C) expressions, both of which were gradually elevated with advanced TNM stage, indicating possible correlation between Gli1 up-regulation and ESCC occurrence. Compared to adjacent tissues, ESCC tissues had significantly lower miR-150 expression, which was further decreased with advanced TNM stage (Fig. 1B).
Lower miR-150 and higher Gli1 expression in ECSS tumor cell line
CFSE staining and flow cytometry results showed significantly lower fluorescent intensity in EC9706 cells compared to HEEC cells (Fig. 2A), indicating significantly higher proliferation potency of EC cells than normal epithelial cells. DNA quantification showed lower S phase ratio in HEEC cells compared to that in EC9706 cells, whilst G0/G1 ratio was significantly higher (Fig. 2B), suggesting higher DNA synthesis and proliferation potency of EC cells than those in HEEC cells, accompanied with less G0/G1 arresting. Further assay showed remarkably lower miR-150 expression in EC9706 cells compared to HEEC cells (Fig. 2C), whilst Gli1 mRNA or protein level was higher (Fig. 2D). In addition, the level of Cyclin D1, which is important for G1/S transition, was also significantly higher in EC9706 cells (Fig. 2D).
Gli1 is one target gene of miR-150
Bioinformatics analysis showed satisfactory targeted relationship between miR-150 and 3’-UTR of Gli1 mRNA (Fig. 3A). Dual luciferase reporter gene assay showed that the transfection of miR-155 mimic could significantly decreased relative luciferase activity in HEK293T cells transfected by Gli1-3’-UTR-wt plasmid (Fig. 3B), but not on HEK293T cells transfected with mutant form (Gli1-3’-UTR-mut) of plasmids, indicating that miR-150 targeted 3’-UTR of Gli1 mRNA to inhibit its expression.
MiR-150 up-regulation and G0/G1 phase arresting and proliferation inhibition. (A) qRT-PCR for gene, expression; (B) Western Blot for protein expression; (C) PI staining for cell cycle assay; (D) CCK-8 for cell proliferation activity. Note: a, 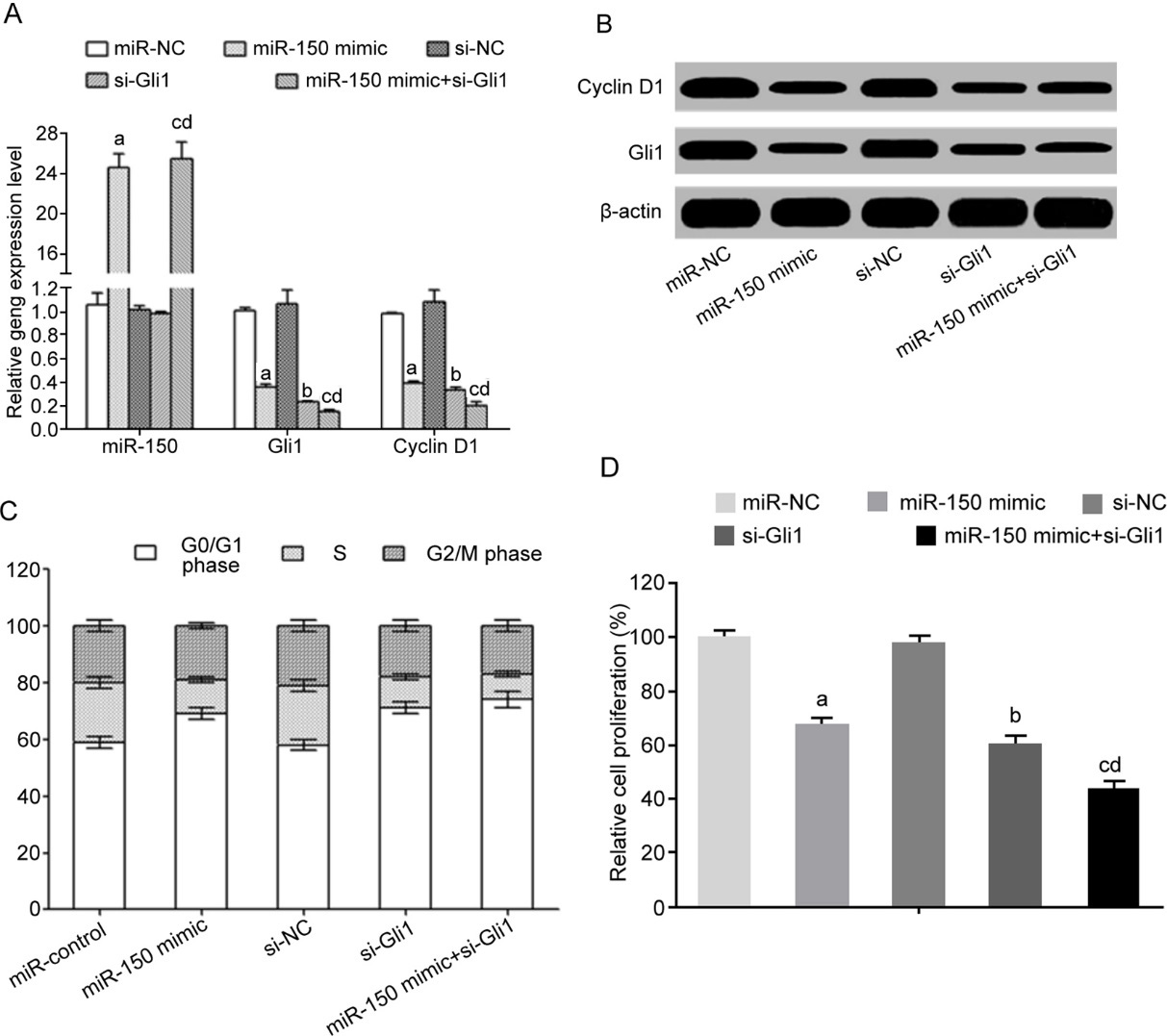
Transfection of miR-150 mimic and/or interference of Gli1 significantly down-regulated mRNA and protein expression of Gli1 (Fig. 4A and B). Meanwhile, cyclin D1 expression was also remarkably inhibited (Fig. 4A and B). Flow cytometry results showed that the transfection of miR-150 mimic and/or interference of Gli1 significantly increased G0/G1 phase arresting (Fig. 4C) and decreased cell proliferation potency (Fig. 4D).
Discussion
EC represents as a type of malignant tumor with relatively higher incidence and mortality. Its rate of occurrence is 8
As the regulator for cell cycle protein dependent kinase (CDKs), cyclin D1 can bind and activate G1-specific CDK4 to phosphorylate G1 phase cycle inhibitory protein Rb, which further dissociates from E2F transcriptional factor for initiating cell cycle regulatory gene, thus pushing cell cycle from G1 into S phase, and facilitating cell proliferation [22]. Previous finding showed that Gli1 played an important role in mediating cyclin D1 expression, cell proliferation and cycle progression [16]. Our data confirmed that, compared to HEEC cells, EC9706 cells presented significantly higher proliferation potency, with the increase of cyclin D1 expression and low G0/G1 ratio. Of note, miR-150 increased the level of cyclin D via the targeting inhibition of Gli1 expression, which validated the previous evidence of positive correlation between Gli1 expression and cyclin D1 level in EC cells [6].
Moreover, it has been demonstrated that the over-expression of Gli1 significantly facilitated epithelial mesenchymal transition (EMT) process of EC cells, and potentiated malignancy including invasion and metastasis [18]. In Gli1-mediated cell cycle regulation, Rizvi et al. found that Gli1 could up-regulate CDK-2 expression via functioning on CDK-2, to facilitate G1/S transition, thus promoting proliferation [19]. Our study further showed that the decrease of Gli1 remarkably caused higher G0/G1 ratio and weakening proliferation potency. Additionally, Yokobori et al. found that miR-150 could weaken EMT process and invasion potency of EC cells via targeting the expression of important EMT regulator, ZEB1 [11]. Wang et al. found that miR-150 could enhance the interaction between rpL11 and c-Myc via targeted inhibition on 5S rRNA expression, thus suppressing the transcriptional regulatory role of oncogene c-Myc and inhibiting EC cell proliferation [23]. This study revealed the role of miR-150 in EC pathogenesis, as consistent with Yokobori et al. [11] and Wang et al. [23]. These results indicated that the down-regulation of miR-150 and consequently upregulated Gli1 and its downstream target gene cyclin D1 might be one important mechanism underlying EC pathogenesis. However, the limitation in our study still exists, and the regulatory effect of miR-150 on other G1/S mediators, such as CDK-2/4/6, cyclin E, along with the validation of other CDK inhibitors via regulating Gli1, requires to be further determined.
Conclusion
Our study demonstrated that miR-150 played a critical role in promoting cyclin D1 expression and inhibiting the proliferation of EC cells via the targeted inhibition of Gli1, which provides academic basis for the further therapy of esophageal cancer.
Footnotes
Conflict of interest
None.
