Abstract
BACKGROUND:
Both QKI and TMEFF2 genes are considered putative tumor suppressor genes (TSGs). In gastric (GC) and colorectal (CRC) cancers, downregulation of their expressions is known to be frequent. However, QKI and TMEFF2 mutations that could potentially inactivate their functions are not reported in cancers.
METHODS:
In a genome database, we observed that both QKI and TMEFF2 harbor mononucleotide repeats, which could be mutated in cancers with high microsatellite instability (MSI-H). For this, we studied 79 GCs and 124 CRCs for the mutations and their intratumoral heterogeneity (ITH).
RESULTS:
Six of 34 GCs (17.6%) and 10 of 79 CRCs (12.7%) with MSI-H exhibited QKI frameshift mutations while five of 79 CRCs (6.3%) with high MSI (MSI-H) exhibited TMEFF2 frameshift mutations. However, we found no such mutation in microsatellite stable/low MSI (MSS/MSI-L) cancers within the mononucleotide repeats. We also studied ITH for the detected frameshift mutations in 16 cases of CRCs and detected that QKI and TMEFF2 frameshift mutations showed regional ITH in 2 (12.5%) and 1 (6.3%) cases, respectively.
CONCLUSIONS:
Our data show that candidate TSG genes QKI and TMEFF2 harbor mutational ITH as well as the frameshift mutations in GC and CRC with MSI-H. From this observation, frameshift mutations of QKI and TMEFF2 may play a role in tumorigenesis through their TSG inactivation in GC and CRC.
Keywords
Introduction
QKI gene encodes an RNA-binding protein that regulates diverse mRNA functions such as splicing, export and stability of mRNA [1]. QKI regulates target mRNAs such as p27, beta-catenin and c-fos that are related to cancer development [2, 3, 4]. Aberrant expression of QKI has been reported in many tumors [5, 6, 7]. For example, QKI gene is silenced in gastric (GC) and colorectal (CRC) cancers [3, 7]. QKI overexpression inhibits proliferation of GC and CRC cells [3, 7]. These reports suggest that QKI has tumor suppressor gene (TSG) activity in multiple cancers. TMEFF2, also known as HPP1, encodes the tomoregulin family of transmembrane proteins [8]. Loss of TMEFF2 expression by promoter hypermethylation has been reported in many tumors, including GC, CRC and breast cancers [8, 9, 10]. TMEFF2 expression promotes apoptosis and induces cell cycle arrest [11, 12], indicating that TMEFF2 gene is a candidate TSG as well.
About one third of GC and CRC are classified as high microsatellite instability (MSI-H) cancers [13]. Many TSGs harbor frameshift mutations at monocleotide repeats in MSI-H cancers [13]. In the human genome database, we observed that both QKI and TMEFF2 genes possess nucleotide repeats in coding sequences that might be mutated in MSI-H cancers. Intratumoral heterogeneity (ITH) is a common phenomenon in cancers, which may result in cancer evolution and influence on clinical outcomes [14, 15, 16]. Thus, identification of genetic ITH is important in understanding biological and clinicopathologic features of the cancers. The current study aimed to find whether QKI and TMEFF2 frameshift mutations are common and harbor ITH in MSI-H GC and CRC.
Summary of pathologic features of gastric and colorectal cancers
Summary of pathologic features of gastric and colorectal cancers
EGC: early gastric cancer, AGC: advanced gastric cancer, TNM: tumor, lymph node, metastasis, MSI-H: high microsatellite instability, MSI-L: low microsatellite instability, MSS: stable microsatellite instability.
Tissue samples and microdissection
In this study, we used 79 GCs and 124 CRCs. The GCs were 34 GCs with MSI-H, 45 GCs with microsatellite stable/low MSI (MSS/MSI-L), 79 CRCs with MSI-H and 45 CRCs with MSS/MSI-L (Table 1). These samples overrepresent MSI-H cases, because we collected MSI-H and MSS/MSI-L cases separately in different times. We adopted an MSI evaluation system using five mononucleotide repeats [17]. In brief, we used multiplex PCR for five quasi-monomorphic mononucleotide repeats (BAT25, BAT26, NR21, NR24, and NR27). Cancers with no allelic size variations in fewer than two of the microsatellites were considered MSS/MSI-L, while those with size variations in more than two microsatellite markers were considered MSI-H. For 16 MSI-H CRCs, we picked 4–7 tumor areas and one normal area per CRC for the ITH analysis. Each ITH area was histologically confirmed under light microscope. These ITH areas were studied for detecting mutational ITH of QKI and TMEFF2. Pathologic features of the cancers were evaluated in all blocks of all cases by a pathologist and are summarized in Table 1. Tumor and normal cells were microdissected as described previously [18, 19]. Approval of this study was obtained from the institutional review board of the Catholic University of Korea.
Representative SSCP and DNA sequencings of repeats in QKI and TMEFF2 in colon and gastric carcinomas. SSCP (A) and DNA sequencing analyses (B) of the A7 repeat in QKI (left) and the A8 repeat in TMEFF2 (right) from tumor (Lane T) and normal tissues (Lane N). A. SSCPs of the PCR products show aberrant bands (arrows in lane T) as compared to SSCP from normal tissues (N).B. Direct DNA sequencing analyses show a heterozygous A deletion (left) and a heterozygous A duplication (right) in the tumor tissues as compared to normal tissues.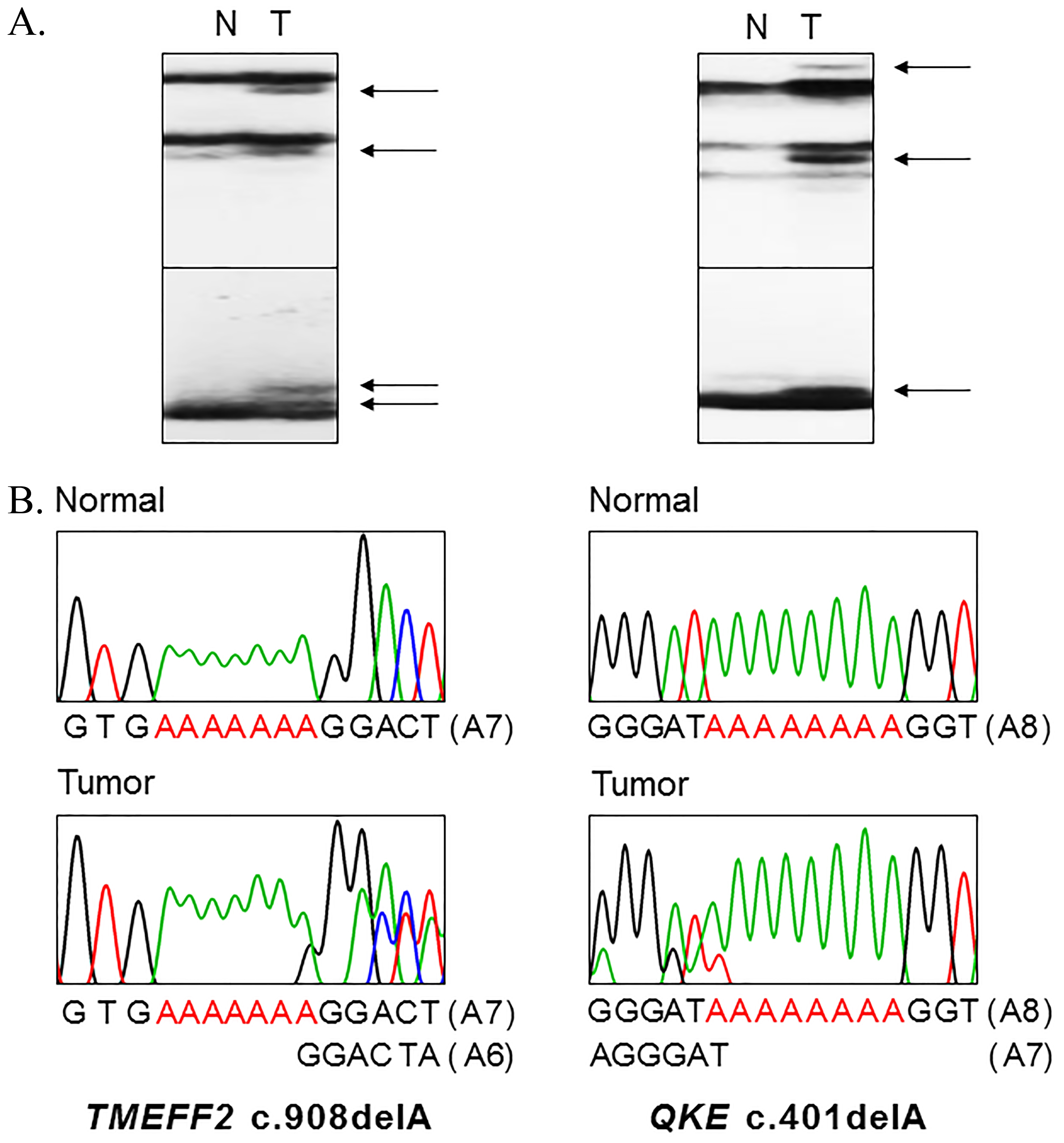
Summary of QKI and TMEFF2 mutations in gastric and colorectal cancers
MSI-H: high microsatellite instability.
Intratumoral heterogeneity of QKI and TMEFF2 frameshift mutations in colorectal cancers with MSI-H
ITH: intratumoral heterogeneity, MSI-H: high microsatellite instability, n.d.: not done.
We analyzed DNA sequences in QKI (A8 in exon 3) and TMEFF2 (A7 in exon 9). Genomic DNA was amplified using polymerase chain reaction (PCR). [
Results
Mutational analysis
Genomic DNAs of the 79 GCs and 124 CRCs were studied to detect frameshift mutations in QKI (A8 in exon 3) and TMEFF2 (A7 in exon 9) mononucleotide repeats. In SSCP, we observed aberrant bands in 20 cases among the 79 GCs and 124 CRCs (Table 2 and Fig. 1). Normal tissues of the corresponding patients revealed no aberrant migration in SSCP, which indicated that the aberrant bands had risen somatically (Fig. 1). Sanger sequencing of the cancer tissues revealed that they were frameshift mutations of QKI and TMEFF2 (Fig. 1). All detected mutations of QKI and TMEFF2 were considered heterozygous mutations by SSCP and direct sequencing patterns (Fig. 1). These mutations were deletion mutations, which would lead to premature termination of QKI or TMEFF2 translation (Table 2). Ten of 79 CRCs (12.7%) and 6 of 34 GCs (17.6%) with MSI-H exhibited QKI mutations while 5 CRCs (6.3%) with MSI-H exhibited TMEFF2 mutations. Any cancers in this study did not harbor both QKI and TMEFF2 frameshift mutations simultaneously.
Intratumoral heterogeneity of QKI and TMEFF2 frameshift mutations in colon cancers. A: Direct DNA sequencing analyses show QKI c.401delA mutation (MT) in 5 regional areas (45-1, -2, -3, -4 and -6) and wild-type (WT) in the other 2 areas (45-5 and -7). B: Direct DNA sequencing analyses show TMEFF2 c.908delA mutation (MT) in one regional area (26-2) and wild-type (WT) in the other 5 areas (26-1, -4, -5, -6 and -7).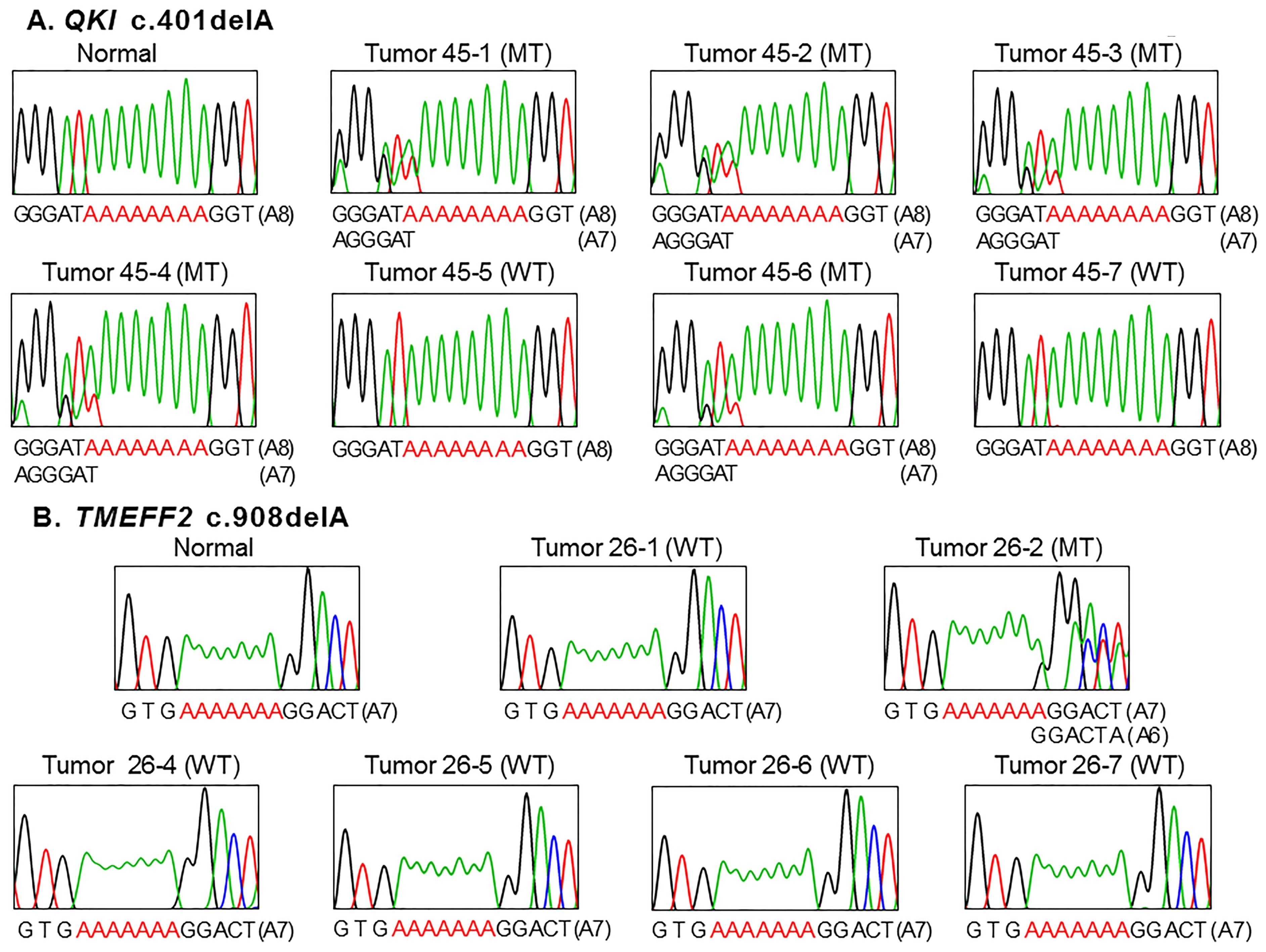
QKI and TMEFF2 frameshift mutations were found in MSI-H cancers, but there was none in MSS/MSI-L cancers (Table 2) (Fisher’s exact test,
Ninety-six areas from 16 CRCs with MSI-H were analyzed to find the ITH in QKI and TMEFF2 frameshift mutations. In two (#45 and 49) of the 16 CRCs (12.5%), the A8 repeat in QKI exhibited one base-pair deletion mutation in one or more regional areas (Table 3 and Fig. 2). In one (#26) of the 16 CRCs (6.3%), the A7 repeat in TMEFF2 exhibited one base-pair deletion (A7 to A6) mutation in one of the 6 regional fragments. There was no significant histological difference between mutation (
Discussion
Inactivation of a TSG by somatic mutations is known to lead the cell to cancer development in combination with other genetic changes. Earlier studies identified that both QKI and TMEFF2 possess TSG activities in cells [3, 7, 11, 12]. Both QKI and TMEFF2 are known to lose their activities by expression losses through promoter hypermethylation in GC and CRC [3, 7, 8, 9]. In the COSMIC database that encompasses overall mutation incidences to date, the QKI recurrently has hotspot p.K134 frameshift mutations in GC (
Two opposite activities (TSG and oncogenic) of TMEFF2 have been identified [8, 9, 21]. In prostate cancers, TMEFF2 promotes cell proliferation in an ADAM17-depencdent manner [22]. By contrast, in GC and CRC, TMEFF2 expression is decreased compared to the normal cells [8, 9]. In addition, loss of TMEFF2 expression is related to the tumor stages and poor clinical outcomes of GC patients [22]. TMEFF2-mediated tumor suppression requires activation of STAT1/2 pathways that induces G0/G1 phase cell cycle arrest [11, 12]. These earlier data may indicate that TMEFF2 function either oncogenic or TSG is context- and cell type-dependent. Together, TMEFF2 may behave as a TSG that is frequently altered by frameshift mutations as well as expression loss in GC and CRC. The QKI and TMEFF2 frameshift mutations may truncate and disable QKI and TMEFF2 proteins and hence be considered loss-of-function mutations. Expression of QKI increases the enterocyte differentiation markers in CRC cells along with p27 protein level and beta-catenin, and inhibits cell proliferation [3]. The truncated mutants of QKI predicted by our data may possibly inactivate their TSG function (maintenance of cell differentiation and inhibition of proliferation), but the cellular consequences as well as clinical implications remain to be further clarified.
In this study, we identified mutational ITH of QKI or TMEFF2 in 3 of 16 CRCs (19%), which is consistent with previous studies that had reported frequent mutational ITH in CRCs with MSI-H [23]. It is well known that cancer ITH of driver mutations could result in poor clinical outcomes in patients. For instance, single or a group of mutations with a metastasis potential might redirect clinical outcomes since such rare clones may accomplish dominance during tumor progression [14, 15, 16]. Based on the TSG properties of QKI and TMEFF2 in CRC, our results suggest that loss of the TSG properties may have regional heterogeneity that could be further selected and influence the clinical outcomes. However, it should be further studied how the ITH change during the progression and the ITH play a causal role in tumor pathogenesis.
TMEFF2 has EGF-like domain that is known to bind with ERBB4 [24]. Given the importance of oncogenic erbB proteins in cancer therapies, further understanding of TMEFF2 alterations in our data (mutational inactivation in MSI-H GC and CRC) here may provide useful information for its potential clinical application. In these points, our data may reveal information on biological mechanisms as well as therapeutic strategies in cancer cells.
Footnotes
Acknowledgments
This study was supported by a grant from National Research Foundation of Korea (2012R1A5A2047939).
Conflict of interest
The authors declare no competing interests.
