Abstract
BACKGROUND:
Many studies have demonstrated that the long non-coding RNA (lncRNA), NEAT1_1, plays critical roles in various human tumor entities and is related to the survival of patients with malignancies. However, the role of NEAT1_1 in diffuse large B cell lymphoma (DLBCL) remains unclear. The aim of this study was to investigate the role of NEAT1_1 in DLBCL.
METHODS:
The expression of NEAT1_1 was evaluated in paraffin-embedded tissues from 64 DLBCL patients and 15 lymphnoditis patients using the ISH method. The correlations between the expression levels of NEAT1_1 and clinical-pathological features and patients’ survival were also analyzed. After knocking down NEAT1_1 using shRNA in the DLBCL cell lines OCI-Ly1 and SUDHL-4, cell viability, apoptosis and migration were assessed by performing CCK8 assays, annexin V-FITC/PI double staining assays and migration filter assays, respectively.
RESULTS:
NEAT1_1 expression was increased in DLBCL tissue compared to lymphnoditis tissue samples (
CONCLUSIONS:
Our results showed that NEAT1_1 plays an oncogenic role in DLBCL. NEAT1_1 expression may serve as a predictive marker for DLBCL patients.
Introduction
Diffuse proliferation of large neoplastic B lymphoid cells with a nuclear size equal to or exceeding that of normal macrophage nuclei is defined as diffuse large B-cell lymphoma (DLBCL), the most common subtype of non-Hodgkin lymphoma (NHL), accounting for 30%–40% of adult NHLs. In Europe and the USA, the current annual incidence of NHL is estimated at 15–20 cases/100,000, comprising approximately 5% of all malignancies in western countries [1]. The incidence of DLBCL in Asia is approximately 3.26/100,000 [2]. The pathological morphology of DLBCL is complex and volatile. During the last decade, the International Prognostic Index (IPI) has become the most frequently used prognostic factor in DLBCL [3]. The IPI scoring system includes age, performance status, serum lactate dehydrogenase (LDH) level, stage and extranodal involvement. Although IPI scoring systems have been useful for risk stratification for several decades, it is unreliable for predicting outcomes in individual patients because of the biological heterogeneity of DLBCL. New prognostic tools and molecular methods have led to the identification of a novel basis for subtype/subset-specific index divisions, demonstrating molecular heterogeneity within morphologically similar tumors and linking gene expression profiles (GEP) to prognosis. DLBCL is usually aggressive, and 80% of aggressive lymphomas are DLBCL. Approximately 60% of DLBCL patients’ disease progresses into advanced stage lymphoma (usually stage III or IV) [4]. Different treatment paradigms should be performed at an early date to improve the survival of these patients. Exploring a novel specific and sensitive biomarker is indispensable for predicting the clinical course of DLBCL.
Non-coding RNAs recognized as a large group of endogenous RNA molecules without protein coding capacity that have specialized cellular and molecular functions. Molecules of 200 nt in size are standard, and those that are greater than or equal to 200 nucleotides are named long non-coding RNAs (lncRNAs), serving as a molecular signal, decoy, guide and scaffold [5]. Verma performed a systematic analysis of novel lncRNAs from the poly-adenylated transcriptomes of 116 primary DLBCL samples, and they identified 2,642 novel, multiexonic lncRNAs expressed in more than one tumor, two-thirds of which are not expressed in normal B cells [6]. Increasing evidence has shown that lncRNAs work as a biomarkers in various human cancers, such as lung cancer, breast cancer, gastric cancer, to reflect disease progression [7, 8, 9]. Aberrantly expressed lncRNAs exhibit unique profiles in different tumors. The specific expression of lncRNAs may correlate with DLBCL biological characteristics, such as treatment response and prognosis. The NEAT1 gene is transcribed as two major isoforms that overlap completely at their 5’ ends: the shorter, polyadenylated NEAT1_1 and the longer NEAT1_2 isoform (also called MEN
Materials and methods
Patients and tissue samples
Paraffin sections were obtained from 64 patients who were diagnosed with DLBCL enrolled at the department of Hematology at the time of diagnosis between January 2007 and June 2010. The diagnosis of DLBCL was confirmed based on the 2008 WHO classification criteria. All of these patients underwent molecular and phenotypic classification with the available clinical data. Paraffin sections from 15 lymphnoditis patients were also from these two hospitals. Data were obtained by outpatient chart review and contacting the patients by telephone or mail. The study was approved by the Ethics Committee these two Hospitals and was performed in accordance with the ethical standards of the 1964 Declaration of Helsinki and its later amendments. The ethical permission and informed consent were obtained from all participants. All study subjects provided written informed consent for the use of their tissue samples in the present study. We retrospectively collected the patients’ clinicopathological characteristics, including sex, age, stage, pathological type, IPI score, LDH level, and survival information. The follow-up was 60 months.
In situ hybridization
Hematoxylin and eosin-stained slides from each tumor block were reviewed, and representative areas with the highest percentage of tumor cells were selected for TMA construction. ISH was performed on 3-
Cell culture
OCI-Ly1 and SUDHL-4 cells were maintained in RPMI-1640 medium (Gibco, Carlsbad, CA, USA) containing 10% heat-inactivated fetal bovine serum albumin (Logan, USA) in a 95% humidified incubator with 5% CO
shRNA plasmid construction and transduction
The shRNAs of the human NEAT1_1 construct lentiviral vector were constructed by Jierui BioCom, with a sequences of 5’-GUGAGAAGUUGCUUAGAA ACUUUCC-3’. The empty vector pLKO.1 is control. First, OCI-Ly1 and SUDHL-4 cells were starved in RPMI-1640 medium without fetal bovine serum for 24 h, then transduced with the shRNA-NEAT1_1 using Lipofectamine3000. After 8 hours, we replaced the medium with fresh RPMI-1640 medium with 10%FBS and G418.
Quantitative real-time PCR analysis
Total RNAs from untreated cells and those transduced with NC or shRNA-NEAT1_1 cells were extracted using TRIzol (Invitrogen, USA) according to the manufacturer’s instructions. The cDNA was synthesized using the MLV transcriptase Kit (Invitrogen, USA). The quantitative analysis of the NEAT1_1 mRNA level was evaluated using qRT-PCR, and
Cell viability assay
The cells of three groups were seeded into 96-well culture plates at a density of 1.5
Cell apoptosis assay
Apoptotic and dead cells from the three groups were determined using FITC-labelled Annexin V and propidium iodide (PI) staining (BD Bioscience, USA), followed by flow cytometry. The cells were collected and resuspended in binding buffer at a concentration of 3
NEAT1_1 was highly expressed in DLBCL. Representative images of in situ hybridization for NEAT1_1 in DLBCL tissues and lymphnoditis tissues. For positive NEAT1_1 staining, purple-blue circumnuclear grains were observed. For the strong positive group, navy blue grains were widely distributed in the cell, particularly in the nucleus. A) and D) are negative in lymphnoditis tissue; B) and E) are weak positive of NEAT1_1 in DLBCL tissue; C) and F) are extra strong positive of NEAT1_1 in DLBCL tissue. A, B and C are at 200 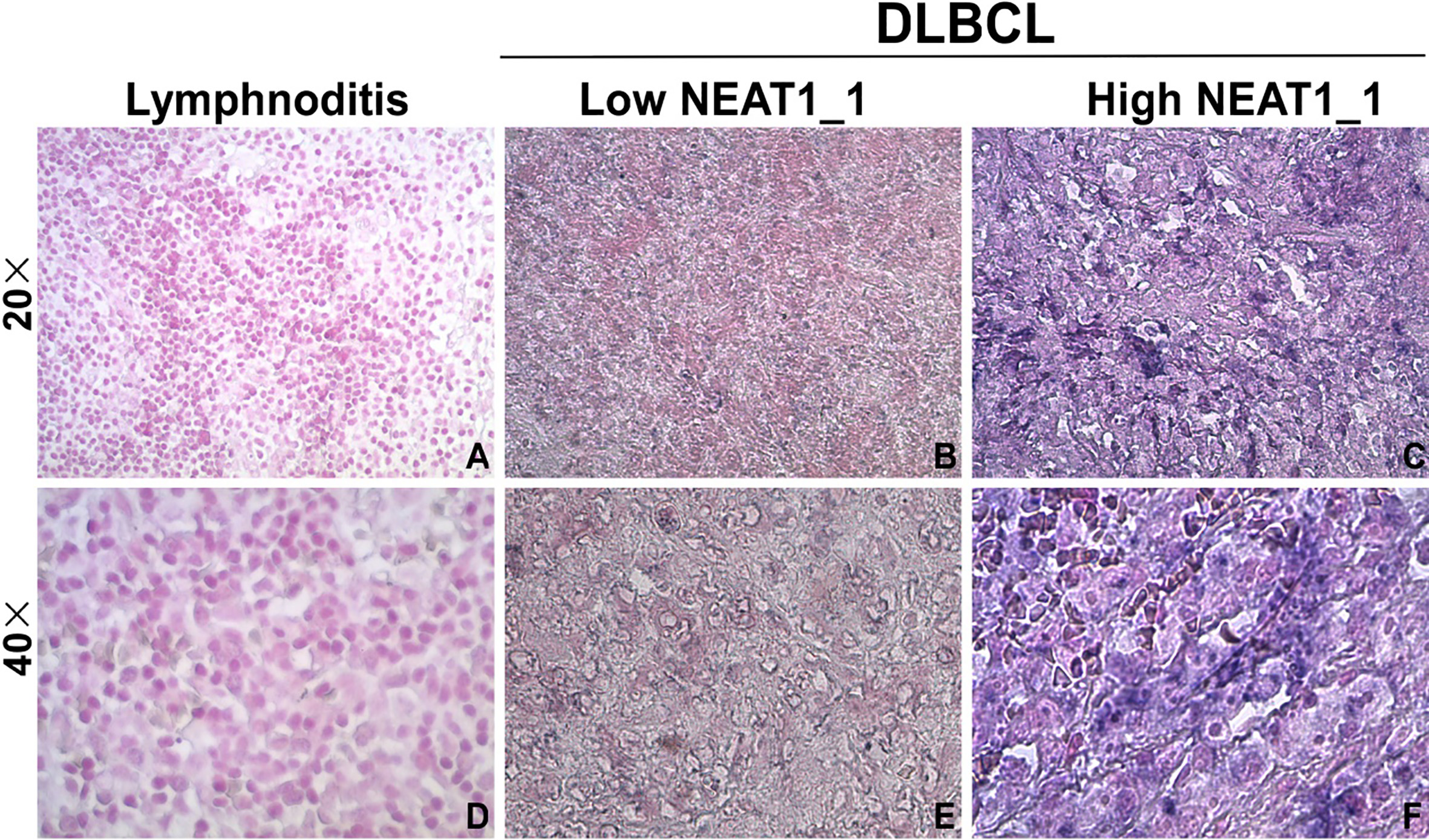
The migration of OCI-Ly1 and SUDHL-4 cells was assayed using a Boyden chamber (Haimen, Jiangsu, China). Cells from the three groups were plated at 1
Statistical analysis
Comparisons of NEAT1_1 expression between DLBCL and lymphnoditis tissues were assessed using an unpaired t-test. One-way ANOVA was used to identify the impact of clinicopathological factors and NEAT1_1 expression in patients with DLBCL. Patient survival rates were calculated using the Kaplan-Meier method, and statistically significant differences in survival were identified using the log-rank test. All
Results
NEAT1_1 upregulation within DLBCL tissue
We compared the NEAT1_1 expression levels between the DLBCL and lymphnoditis tissues. NEAT1_1 expression in the DLBCL tissues was significantly increased compared to that in the lymphnoditis tissues (10.20
Correlation between NEAT1_1 expression and clinicopathological characteristics
We correlated NEAT1_1 expression levels with the clinicopathological characteristics of the patients with DLBCL (Table 1). A total of 64 DLBCL patients were enrolled in this study. Among them, 36 (56.25%) were male, 33 (51.56%) were
Clinicopathological characteristics of the 64 DLBCL patients
Clinicopathological characteristics of the 64 DLBCL patients
GCB is germinal center B cell-like DLBCL; ABC is activated B-cell-like DLBCL.
We evaluated the NEAT1_1 level using the ISH method. A total score equal to or greater than 10 was used as a cutoff value to divide all 64 patients into two groups: the low expression group (
NEAT1_1 is a factor associated with a poor prognosis of DLBCL. A) The expression of NEAT1_1 was detected in two groups using the ISH method. B) Kaplan-Meier survival analysis of 64 patients with DLBCL comparing high and low NEAT1_1 expression (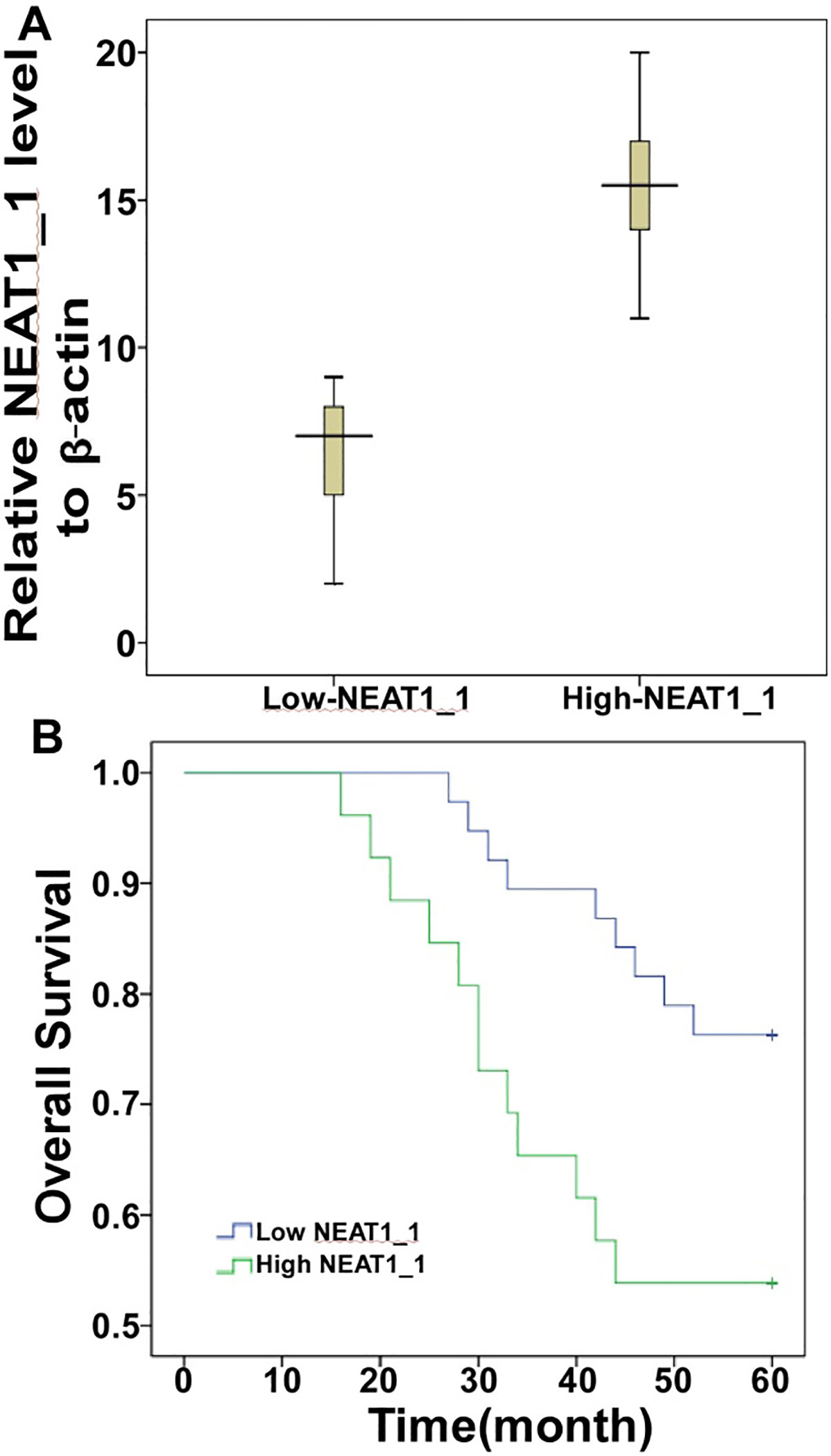
NEAT1_1 plays an oncogenic role in various cancers. Here, to determine the function of NEAT1_1 in DLBCL, we showed that shRNA-NEAT1_1 significantly suppressed NEAT1_1 expression in OCI-Ly1 and SUDHL-4 cells (Fig. 3A). For the cell viability of both cell lines, there was a significant reduction in cell proliferation in the shRNA-NEAT1_1 group (Fig. 3B). To determine the function of NEAT1_1 in migration, we used the Boyden chamber assay to assess OCI-Ly1 and SUDHL-4 cells. As shown in Fig. 3C, we found that there were significantly less migration through the basement membrane in the shRNA-NEAT1_1 group. Apoptosis experiments were also performed (Fig. 3D). The death and apoptosis rates were much higher in the shRNA-NEAT1_1 group than in the other two control groups.
Knocking down NEAT1_1 inhibited DLBCL proliferation and migration in vivo. (A) After transducing the siRNA-NEAT1_1 into OCI-Ly1 and SUDHL-4 cells for 24 h, we detected NEAT1_1 using the QRT-PCR method. The siRNA effectively inhibited NEAT1_1expression in DLBCL cells (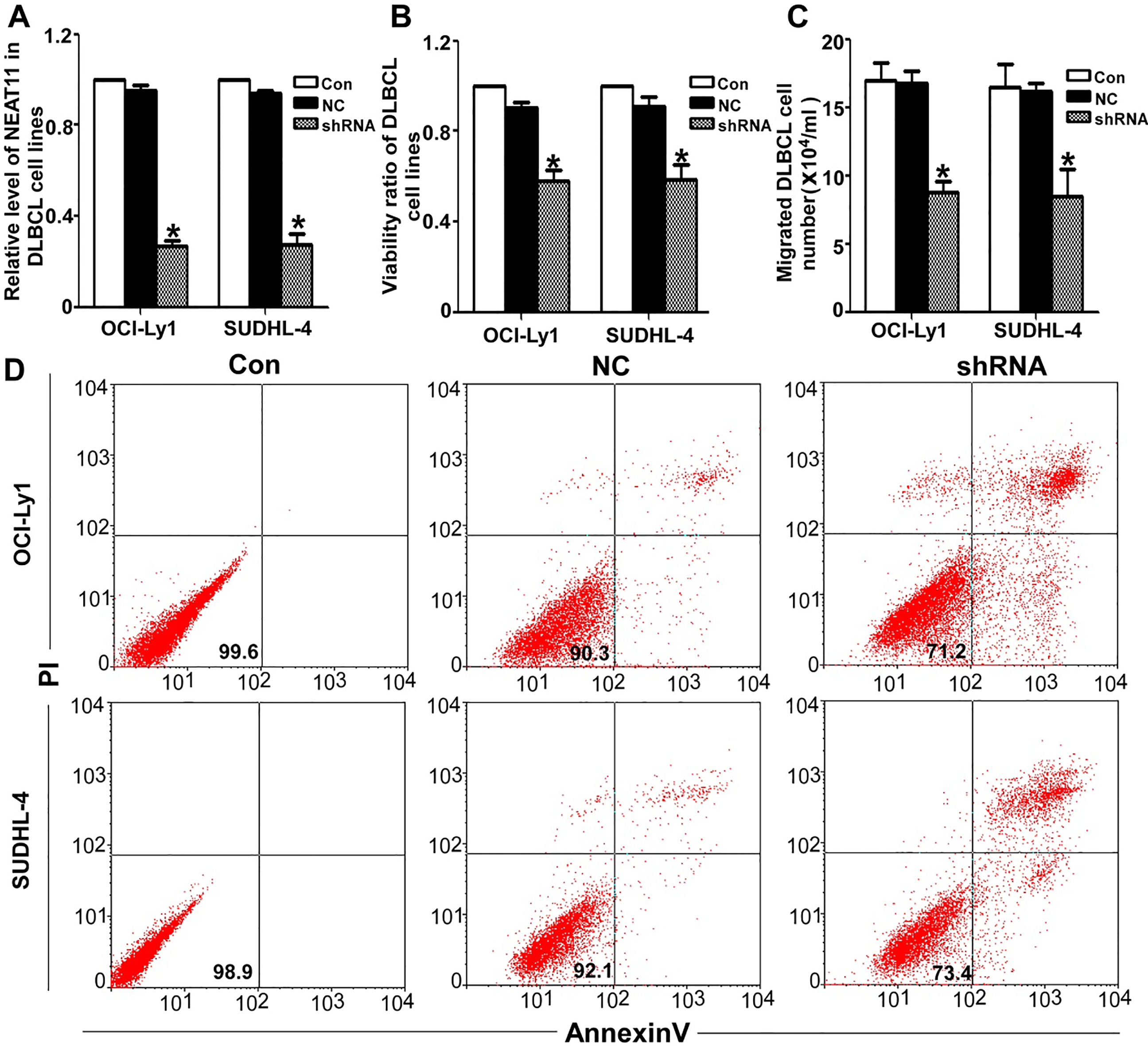
Nuclear enriched abundant transcripts are located less than 70 kb apart on human chromosome 11. There are no more than three transcripts, including XIST, and two unique noncoding nuclear enriched abundant transcripts (NEAT) RNAs. Although NEAT1 and NEAT2 (also known as MALAT-1) share no significant homology with each other, both of them are conserved within the mammalian lineage. MALAT-1, as a metastasis driver, is widely credited, but NEAT1 has not been studied thoroughly as MALAT-1. NEAT1 comprises of two isoforms produced by alternative 3’-end processing (NEAT1_1 and NEAT1_2). NEAT1_1 is a 4-kb, unspliced, polyadenylated, nuclear restricted, noncoding transcript with two small regions of high conservation across several species, indicating its potentially important function. Clemson found that the large, tightly packaged structural aggregates of NEAT1_1 RNA, containing many self-complementary sequences and complex intra-molecular structures, forms quickly at the transcription site, which suggests that NEAT1_1 RNA may self-associate. Except for mitosis, NEAT1_1 RNA is tightly localized in paraspeckles, consistently overlapping with these structured concentrations of both PSP1 and p54. Overexpressed NEAT1_1 can increase the number of paraspeckles, which suggests that it is the functional isoform for paraspeckle formation. p54nrb, PSF, and PSP1 are characteristically enriched in paraspeckles, and NEAT1_1 is required for their localization. NEAT1_1 affects the formation and maintenance of these discrete nuclear domains [11]. In 2007, Hutchinson [12] first identified NEAT1_1 and reported that it was enriched by 25.6-fold in the nuclei of lymphoblasts, compared to 2.1-fold in fibroblasts. Blume [13] showed that the landscape of p53-dependent microRNA/non-coding RNA was induced in response to DNA damage in chronic lymphocytic leukemia (CLL). NEAT1_1 and long intergenic non-coding RNA p21 (lincRNA-p21) are induced in response to DNA damage in the presence of functional p53 but not in the presence of p53 mutations. This p53 dependence of NEAT1_1 and the lincRNA-p21 model were also verified in a lymphoma cell line. NEAT1_1 is highly expressed in lung cancer [14], colorectal cancer [15] and glioma [16]. NEAT1_1 expression patterns and the clinicopathological features of lymphoma are not well described thus far. Approximately 31% of all NHLs and 80% of aggressive lymphomas are DLBCLs. The WHO criteria classify DLBCL into several subtypes and a few distinct disease entities. Many molecular and immunophenotypic subgroups were described as prognostic indicators and provide insights into therapy. However, a large number of DLBCLs remain biologically heterogeneous without clear and acceptable criteria. A number of investigators have attempted to identify several highly specific and sensitive indices to be used for diagnosis or predicted prognosis. Here, our results showed that the NEAT1_1 expression level was increased in DLBCL tissue compared with acute lymphadenitis tissue (
In conclusion, our analysis showed that NEAT1_1 expression was increased in DLBCL tissues and positively related to a poor prognosis and shorter OS than patients with weak NEAT1_1 expression. Accordingly, NEAT1_1 expression may serve as a risk marker for patients with DLBCL. Furthermore, in vitro assay results also indicate that NEAT1_1 plays an oncogenic role in DLCBL. However, future studies are warranted to investigate the mechanism of NEAT1_1 in DLBCL.
Footnotes
Acknowledgments
This work was supported by grants from the Natural Science Foundation of Guangdong Pr (No. S20120100 08748 and S2013010014715) and National Natural Science Foundatio of China (No. 81400103).
