Abstract
BACKGROUND:
Multiple lines of evidence suggest that single nucleotide polymorphisms (SNPs) in genes encoding components of the microRNA processing machinery may underlie susceptibility to various human diseases, including cancer.
OBJECTIVE:
The present study aimed to investigate whether rs6877842, rs642321 and rs10719 SNPs of DROSHA, a key component of the miRNA biogenesis pathway, are associated with increased risk of breast cancer.
METHODS:
A total of 100 patients diagnosed with breast cancer and 100 healthy women were included. Following extraction of DNA, genotyping was performed by tetra primer- amplification refractory mutation system-PCR (T-ARMS-PCR) technique. Under the co-dominant, dominant and recessive inheritance models, the association between DROSHA SNPs and breast cancer risk was determined by logistic regression analysis. The association of DROSHA SNPs with patients’ clinicopathological parameters was assessed. Also, haplotype analysis was performed to evaluate the combined effect of DROSHA SNPs on breast cancer risk.
RESULTS:
We observed a statistically significant association between DROSHA rs642321 polymorphism and breast cancer susceptibility (P < 0.05). Under the dominant inheritance model, DROSHA rs642321 polymorphism was significantly associated with increased risk of breast cancer (OR: 6.091; 95% CI: 3.291–11.26; P = 0.0001). Our findings demonstrated that DROSHA rs642321 T allele can contribute to the development of breast cancer (OR: 3.125; 95% CI: 1.984–4.923; P = 0.0001). We also found that GTC and GTT haplotypes conferred significant risk for breast cancer (OR: 2.367; 95% CI: 1.453–3.856; P = 0.0001 and OR: 7.944; 95% CI: 2.073–30.43; P = 0.0001, respectively).
CONCLUSIONS:
These results provide the first evidence that DROSHA rs642321 polymorphism is associated with increased risk of breast cancer. However, further studies are needed to firmly validate these findings.
Introduction
By surpassing lung cancer for the first time in 2020, breast cancer has become the most commonly diagnosed cancer among women worldwide, with 2.3 million new cases (11.7% of all new cancer cases) [1,2]. Breast cancer ranks as the fifth leading cause of cancer-related deaths globally, accounting for nearly 685,000 deaths (6.9% of all cancer deaths) [2]. According to the American Cancer Society’s annual report, in 2021, it was estimated that 281,550 new cases of breast cancer will be diagnosed in the United States and 43,600 women will die from the disease [3]. Breast cancer is a heterogeneous and multifactorial disease caused by the complex interplay between multiple genetic and lifestyle/environmental factors [4,5]. Despite remarkable advances in elucidating the pathophysiological mechanisms behind development of breast cancer during the last decades, many aspects of the disease have yet to be adequately studied and remains largely unknown [6–8].
Mature microRNAs (miRNAs or miRs), a class of small (∼22 nucleotides), single-stranded non-coding RNAs, regulate gene expression post-transcriptionally through binding to the 3 ′ -untranslated regions (3 ′ UTRs) of target mRNAs and thereby suppressing protein synthesis via promoting mRNA decay and/or translational silencing [9–11]. Based on miRNA target databases, a single miRNA can target several dozen or even hundreds of different mRNAs and one mRNA can be targeted by many miRNAs [12]. Practically, this means that nearly all cellular and molecular processes are subjected to miRNA-mediated regulation of gene expression [13–16]. The majority of miRNA genes are initially transcribed by RNA polymerase II from the genome into primary miRNA (pri-miRNA) and then processed within the nucleus by the microprocessor complex to generate ∼70 nucleotide hairpin-like structure termed precursor miRNA (pre-miRNA) [17]. The core components of the microprocessor complex are Drosha, a nuclear ribonuclease (RNase) III enzyme, and its interacting partner DiGeorge syndrome critical region 8 (DGCR8), a double-stranded RNA binding protein [17–19]. The pre-miRNA is subsequently exported to the cytoplasm by Exportin5 (XPO5) in a RanGTP-dependent manner [20]. In the cytoplasm, the double-stranded pre-microRNA undergo final processing by Dicer, another member of the RNase III family, and trans-activation response (TAR) RNA-binding protein (TRBP) to produce the mature miRNA duplex [21]. Finally, one of the miRNA duplex strands (guide strand) is preferentially loaded onto Argonaute (AGO) proteins to form a ribonucleoprotein machinery called miRNA-induced silencing complex (miRISC), which recognizes and silences cognate mRNA [22].
There is now compelling evidence to show that disrupted miRNA biogenesis can alter the miRNA expression profiles and potentially drive the transformation of normal cells into cancerous cells [23–25]. Dysregulated expression levels of DROSHA, a catalytic core of microprocessor complex, is closely linked to increased risk of developing breast cancer [26–28]. Besides, single nucleotide polymorphisms (SNPs) in DROSHA gene that affect the structure, function and expression level of protein can be involved in the pathogenesis of breast cancer. To date, several studies have examined the association between DROSHA SNPs and breast cancer risk, and attained conflicting results [29–33]. To the best of the authors’ knowledge, no studies thus far have investigated the association of DROSHA polymorphisms with breast cancer susceptibility in Iranian women. Of note, distribution of genotype and allele frequencies for DROSHA rs6877842, rs642321 and rs10719 SNPs in Iranian population can be differ from those seen in other populations. Accordingly, we investigated the possible association between DROSHA rs6877842, rs642321 and rs10719 SNPs and the risk of breast cancer under the co-dominant, dominant and recessive inheritance models. We also addressed the question whether combined genotypes and haplotypes of DROSHA SNPs are associated with breast cancer. Furthermore, the association between DROSHA SNPs and patients’ clinicopathological parameters was assessed.
List of primers used in T-ARMS-PCR procedure for genotyping of DROSHA rs6877842, rs642321 and rs10719 polymorphisms
List of primers used in T-ARMS-PCR procedure for genotyping of DROSHA rs6877842, rs642321 and rs10719 polymorphisms
Abbreviation: T-ARMS-PCR, tetra primer-amplification refractory mutation system- polymerase chain reaction; SNP, single nucleotide polymorphism; Ta, annealing temperature.
Subjects
In this case-control study, one hundred women with histopathologically confirmed breast cancer and one hundred ethnicity-matched healthy women with no personal and/or family history of breast cancer or any other type of cancer were recruited from Anahid Clinic, Isfahan, Iran, between August 2020 and April 2021. Histopathological analysis was conducted by pathologists at department of pathology of the respective hospitals. Patients’ demographic and clinicopathological data, including age at diagnosis, BMI, smoking status, history of taking the oral contraceptive pills, family history of breast cancer, history of chemotherapy and radiotherapy, laterality, histological type, histological grade, tumor size and metastasis were obtained from medical records and review of pathology reports. Patients who underwent neoadjuvant chemotherapy or radiotherapy were excluded from the study. The control subjects were selected from healthy volunteers during the same period, and data were collected on demographic and clinical features using a structured questionnaire. The study protocol was approved by the Institutional Review Board/Independent Ethics Committee (IRB/IEC) of the Shahid Ashrafi Esfahani University, Isfahan, Iran. Written informed consent was obtained from all subjects prior to blood sampling and the study was conducted in accordance with the Helsinki Declaration.
DNA extraction and genotyping of DROSHA SNPs
Peripheral blood samples (∼5 mL) were taken from all subjects into tubes containing ethylene diamine tetraacetic acid (EDTA), and genomic DNA was isolated from leukocytes using Genomic DNA Extraction Kit (ZEAN, Isfahan, Iran). The quality and quantity of the extracted DNA was determined by NanoDrop® ND-1000 UV-Vis Spectrophotometer (ThermoFisher Scientific TM , Waltham, MA, USA). The rs6877842, rs642321 and rs10719 SNPs of the DROSHA were genotyped using tetra primer-amplification refractory mutation system-polymerase chain reaction (T-ARMS-PCR) assays. PCR reactions were carried out in a final volume of 20 μl containing 10 μl 2X Master Mix Red (Ampliqon, Denmark), 10 pmol of each primer and 100 ng genomic DNA. To detect the DROSHA SNPs, the specific primers were designed by Primer3 available at http://primer3.ut.ee (Table 1). The thermal cycling conditions were as follows: initial denaturation at 95 °C for 5 min; 35 cycles of denaturation at 94 °C for 40 s, annealing at temperature that are given in Table 1 for each primer pair for 40 s, and elongation at 72 °C for 2 min, followed by final extension at 72 °C for 5 min. The PCR products were separated by electrophoresis on 2% agarose gel stained with safe DNA gel stain and visualized with ultra-violet (UV) transilluminator.
Linkage disequilibrium and haplotype analysis
To compute Lewontin’s standardized coefficient (D ′ ) and the square of the Pearson’s correlation coefficient (r2), pairwise linkage disequilibrium (LD) between DROSHA SNPs was compared using SHEsis online server available at http://analysis.bio-x.cn/myAnalysis.php. The association between haplotypes of DROSHA SNPs and breast cancer risk was also evaluated using SHEsis platform.
Statistical analysis
Statistical analysis of data was performed by IBM SPSS Statistics for Windows, Version 22.0 (IBM Corp., Armonk, N.Y., USA) and p value less than 0.05 was considered as the threshold of statistical significance. Student’s t-test was used to compare the difference of continuous variables between breast cancer patients and healthy controls. The association between DROSHA SNPs and breast cancer risk under the co-dominant, dominant and recessive inheritance models was evaluated using logistic regression analysis. To this end, the wild-type genotype was used as the reference to calculate the odds ratio (OR) and 95% confidence intervals (CI). Departures of genotype frequencies from Hardy-Weinberg Equilibrium (HWE) among the breast cancer patients and healthy subjects was calculated using Chi-square test. The association of DROSHA SNPs with patients’ clinicopathological characteristics was also estimated by calculating the OR and 95% CI using logistic regression analysis.
Results
Baseline characteristics of participants
The demographic of all subjects and detailed clinicopathological characteristics of breast cancer patients are summarized in Table 2. The mean age of breast cancer patients at the time of diagnosis and healthy controls was 51.56 years (standard deviation (SD) = 11.24 years, range 29–80) and 39.01 years (SD = 10.51 years, range 17–76), respectively. There was statistically significant difference between the two groups with respect to age (P < 0.05). Patients with breast cancer had significantly higher body mass index (BMI) (mean ± SD; 28.09 ± 4.30) compared to the healthy controls (mean ± SD; 25.64 ± 4.59) (P < 0.05). No statistically significant difference was found in terms of smoking status between the two groups (P > 0.05). All breast cancer patients were married (100%) and 49% had history of taking the oral contraceptive pills. Out of 100 breast cancer patients, 30% of them had family history of breast cancer, and 98% had unilateral tumor. A substantial proportion of patients (92%) had invasive ductal carcinoma (IDC), and (8%) had invasive lobular carcinoma (ILC). Histopathologically, 12% of patients had grade I, 62% had grade II, and had 26% grade III. At the time of diagnosis, 89% of patients were diagnosed with stage I, 10% stage II and 1% stage III. Of 100 patients, 91% had a tumor size ≤2 cm, and 9% had tumor size >2 cm. Only 7% of patients had metastasis involving one or more organs.
Demographic and clinicopathological characteristics of breast cancer patients and healthy controls
Demographic and clinicopathological characteristics of breast cancer patients and healthy controls
Abbreviation: SD, standard deviation; BMI, body mass index; IDC, invasive ductal carcinoma; ILC, invasive lobular carcinoma. Values are presented as mean ± SD for age, weight, height and BMI, and as % for other variables. P values were calculated using t-test for continuous variables and Chi-square test for categorical variables. Bold values indicate statistically significant differences (P < 0.05).
Distribution of genotype and allele frequencies for rs6877842, rs642321 and rs10719 SNPs of DROSHA in patients with breast cancer and healthy controls are presented in Table 3. The genotype frequencies of two SNPs, rs6877842 and rs10719, in breast cancer patients and all SNPs in the healthy subjects were in compliance with HWE (P > 0.05) (Table 3). In patients, the distribution of the DROSHA rs642321 C/T genotypes did not conform to HWE (P < 0.05). Using logistic regression analysis, we evaluated all SNPs under co-dominant, dominant and recessive inheritance models to find their association with susceptibility to breast cancer (Table 3). Our results showed a significant association between DROSHA rs642321 and breast cancer (P < 0.05), but not the other two SNPs (P > 0.05) (Table 3). Under the co-dominant inheritance model, both CT and TT genotypes of DROSHA rs642321 polymorphism revealed a significant association with increased risk of breast cancer (OR: 6.191; 95% CI: 3.287–11.66; P = 0.0001) and (OR: 5.361; 95% CI: 1.482–19.37; P = 0.009), respectively (Table 3). The DROSHA rs642321 C/T polymorphism was found significantly associated with breast cancer susceptibility in dominant inheritance model (OR: 6.091; 95% CI: 3.291–11.26; P = 0.0001) (Table 3). Under the recessive inheritance model, DROSHA rs642321 polymorphism showed no association with breast cancer (OR: 2.087; 95% CI: 0.607–7.167; P = 0.373) (Table 3). As regards DROSHA rs642321 polymorphism, we found that T allele significantly increased the risk of breast cancer development (OR: 3.125; 95% CI: 1.984–4.923; P = 0.0001) (Table 3). No significant association was found between DROSHA rs6877842 and rs10719 polymorphisms and breast cancer risk under the co-dominant, dominant and recessive inheritance models (Table 3). However, a trend of increased risk of developing breast cancer can be considered for these polymorphisms (OR > 1) (Table 3). For DROSHA rs6877842 and rs10719 polymorphisms, the frequency of the C allele in breast cancer patients were higher than that in healthy women, but not statistically significant (P > 0.05) (Table 3). Combined genotype analysis of DROSHA rs6877842, rs642321 and rs10719 SNPs was also conducted to assess the combined effects of these polymorphisms on breast cancer risk (Table 4). Among the combined genotypes of DROSHA rs6877842/rs642321 polymorphisms, GG/CT, GG/TT and GC/CT combined genotypes were associated with a higher risk for breast cancer (GG/CT; OR: 7.425; 95% CI: 3.612–15.26; P = 0.0001, GG/TT; OR: 6.444; 95% CI: 1.736–23.92; P = 0.004, GC/CT; OR: 6.444; 95% CI: 2.116–19.62; P = 0.0004) (Table 4). Similarly, carrying the GG/TC combined genotype for DROSHA rs6877842/rs10719 polymorphisms may increase a woman’s risk of developing breast cancer (OR: 2.285; 95% CI: 1.077–4.851; P = 0.039) (Table 4). As demonstrated in Table 4, for DROSHA rs642321/rs10719 polymorphisms, individuals with CT/TT (OR: 4.292; 95% CI: 2.068–8.910; P = 0.0001), CT/TC (OR: 29.94; 95% CI: 6.387–140.3; P = 0.0001), CT/CC (OR: 10.88; 95% CI: 1.139–104.1; P = 0.028) and TT/TT (OR: 5.444; 95% CI: 1.230–24.09; P = 0.024) combined genotypes had higher breast cancer risk.
Association between DROSHA SNPs and patients’ clinicopathological features
We also analyzed the possible association of genotype and allele distribution of DROSHA rs6877842, rs642321 and rs10719 polymorphisms in breast cancer patients with demographic and clinicopathological characteristics. As shown in Tables 5–7, no statistically significant association was found between distribution of genotypes in studied SNPs of DROSHA and age, BMI, family history of breast cancer, histological type, grade, stage, tumor size and metastasis.
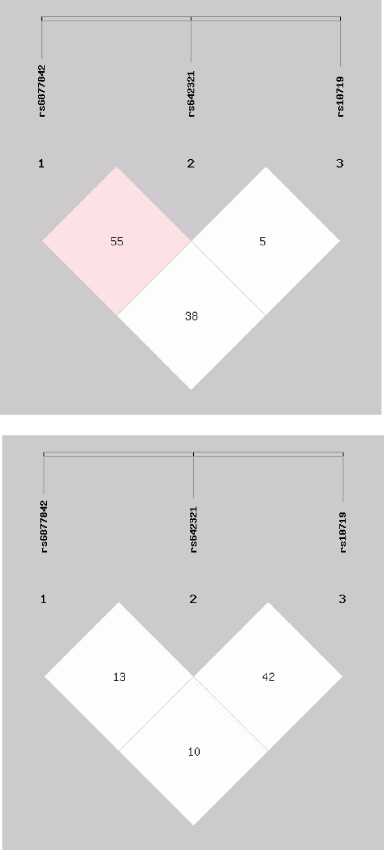
Linkage disequilibrium (LD) plot of DROSHA rs6877842, rs642321 and rs10719 polymorphisms in (a) breast cancer patients and (b) healthy controls.
Genotype and allele distribution of DROSHA rs6877842, rs642321 and rs10719 polymorphisms in breast cancer patients and healthy controls
Abbreviation: OR, odds ratio; CI, confidence interval; HWE, Hardy-Weinberg equilibrium. P value were calculated by Chi-square test. Bold values indicate statistically significant differences (P < 0.05).
Distribution of DROSHA rs6877842, rs642321 and rs10719 polymorphisms combined genotypes in breast cancer patients and healthy controls
Abbreviation: OR, odds ratio; CI, confidence interval; ND, not determined. P value calculated by Chi-square test. Bold values indicate statistically significant differences (P < 0.05).
Association of DROSHA rs6877842 polymorphism with age, BMI, family history of breast cancer, histological type, histological grade, tumor size and other organ metastasis in breast cancer patients
Abbreviation: BMI, body mass index; IDC, invasive ductal carcinoma; ILC, invasive lobular carcinoma; ND, not determined. Values are presented as number (%). P values were calculated using Chi-square test.
Association of DROSHA rs642321 polymorphism with age, BMI, family history of breast cancer, histological type, histological grade, tumor size and other organ metastasis in breast cancer patients
Abbreviation: BMI, body mass index; IDC, invasive ductal carcinoma; ILC, invasive lobular carcinoma; ND, not determined. Values are presented as number (%). P values were calculated using Chi-square test.
Association of DROSHA rs10719 polymorphism with age, BMI, family history of breast cancer, histological type, histological grade, tumor size and other organ metastasis in breast cancer patients
Abbreviation: BMI, body mass index; IDC, invasive ductal carcinoma; ILC, invasive lobular carcinoma; ND, not determined. Values are presented as number (%). P values were calculated using Chi-square test.
On the basis of the genotype frequencies of DROSHA rs6877842, rs642321 and rs10719 SNPs among breast cancer patients and healthy controls, we computed pairwise LD between each pair of the three SNP loci (Table 8), and plotted LD values between all pairs of SNP (Fig. 1). The observed low LD coefficient (D ′ ) values and low r2 in patients with breast cancer and healthy subjects revealed that the studied DROSHA SNPs were not in high linkage disequilibrium with each other (Table 8) (Fig. 1). The association between haplotypes of DROSHA SNPs and breast cancer susceptibility are displayed in Table 9. We found that the GTC and GTT haplotypes (DROSHA rs6877842, rs642321 and rs10719) were significantly associated with increased risk of breast cancer (OR: 2.367; 95% CI: 1.453–3.856; P = 0.0001 and OR: 7.944; 95% CI: 2.073–30.43; P = 0.0001, respectively) (Table 9). There was no significant difference between the two groups regarding the distribution of the CCC, GCC and GCT haplotypes (P > 0.05) (Table 9).
Pairwise LD of DROSHA rs6877842, rs642321 and rs10719 polymorphisms in breast cancer patients and healthy controls
Pairwise LD of DROSHA rs6877842, rs642321 and rs10719 polymorphisms in breast cancer patients and healthy controls
Abbreviation: LD, linkage disequilibrium. r2 reflects statistical power to detect LD.
Haplotype distribution of DROSHA rs6877842, rs642321 and rs10719 polymorphisms in breast cancer patients and healthy controls
Abbreviation: SNPs, single nucleotide polymorphisms; OR, odds ratio; CI, confidence interval. P value were calculated using Pearson’s chi-square test. Bold values indicate statistically significant differences (P < 0.05).
The biogenesis of miRNAs is delicately controlled at multiple steps by enzymes and regulatory proteins involved in the miRNA synthesis pathway [34,35]. Some genetic polymorphisms in the genes encoding miRNA biogenesis components could lead to aberrant expression of miRNAs which are closely related to the development of cancers [36–38]. The SNPs located in the promoter region of gene may affect the expression level of mRNA through altering DNA methylation, histone modifications and chromatin conformational changes, and subsequently transcription factor binding [39]. The SNPs falling in the 3 ′ UTR of gene may modify mRNA secondary structure, mRNA stability and miRNA-mRNA interaction and thereby affect gene expression [39]. Accordingly, we hypothesized that SNPs located in promoter (rs6877842) and 3 ′ UTR (rs642321 and rs10719) of DROSHA gene are contributing factor to breast cancer in Iranian women. To test this hypothesis, the present study was designed to assess the possible association of these SNPs with susceptibility to breast cancer under the co-dominant, dominant and recessive inheritance models. The association between these SNPs and patients’ clinicopathological characteristics was analyzed. We also determined the effects of haplotypes and combined genotypes of DROSHA rs6877842, rs642321 and rs10719 SNPs on breast cancer risk. As far as we know, this is the first report investigating the association between DROSHA SNPs and breast cancer in Iranian women. Overall, a statistically significant association was observed between DROSHA rs642321 SNP and breast cancer risk (P < 0.05). Under the dominant inheritance model, this SNP was found to be associated with increased risk of breast cancer development (OR: 6.091; 95% CI: 3.291–11.26; P = 0.0001). The results revealed that DROSHA rs642321 T allele can affect the susceptibility to breast cancer (OR: 3.125; 95% CI: 1.984–4.923; P = 0.0001). We also found that GTC and GTT haplotypes conferred significant risk for breast cancer (OR: 2.367; 95% CI: 1.453–3.856; P = 0.0001 and OR: 7.944; 95% CI: 2.073–30.43; P = 0.0001, respectively). Based on the literature review, only one study has so far investigated the association between DROSHA rs642321 SNP and breast cancer [30]. In 2013, Jiang et al. conducted the first study on the association between DROSHA rs642321 SNP and risk of breast cancer in the Chinese women [30]. In contrary to our results, they indicated that DROSHA rs642321 SNP was not associated with susceptibility to breast cancer [30]. In another study conducted by Yuan et al. on 685 patients with bladder cancer and 730 healthy controls, no significant association was found between DROSHA rs642321 SNP and susceptibility to bladder cancer in the Chinese population [40]. Although, no details were given on the reasons for the inconsistency in the results of genetic association studies so far, these discrepancies could be attributed in large part to ethnic/racial disparities, limited sample size and multifactorial nature of cancers.
To date, there is no published report regarding the association between DROSHA rs6877842 SNP and breast cancer development. Our results indicate for the first time that this SNP was not significantly associated with susceptibility to breast cancer in Iranian women (P > 0.05). In line with our findings, Kim et al. failed to find any association between DROSHA rs6877842 SNP and lung cancer [41]. Moreover, when Horikawa et al. conducted a case-control study on 279 Caucasian patients with renal cell carcinoma and 278 healthy controls, similar results were obtained [42]. Osuch-Wojcikiewicz et al. demonstrated that there was no significant association between DROSHA rs6877842 SNP and larynx cancer risk [43]. Kim et al. found that DROSHA rs6877842 polymorphism was not associated with pathogenesis of hepatocellular carcinoma in the Korean population [44]. No statistically significant associations between DROSHA rs6877842 SNP and risk of papillary thyroid carcinoma was observed by Mohammadpour-Gharehbagh et al. [45]. In contrast, a meta-analysis by Wen and collaborators demonstrated that DROSHA rs6877842 SNP was associated with an increased cancer risk and may be potential biomarker for cancer-forewarning [46]. These findings lead us to conclude that DROSHA rs6877842 polymorphism is not meaningfully related with development and progression of human cancers. However, these results should be interpreted with caution due to very limited studies.
To our knowledge, only one study has investigated the association between DROSHA rs10719 SNP and risk of breast cancer development [30]. As regards DROSHA rs10719 polymorphism, we did not find any significant association between this SNP and breast cancer risk (P > 0.05). These results reflect those of Jiang et al. who demonstrated that DROSHA rs10719 SNP may not contribute to breast cancer susceptibility [30]. For other types of cancer, some studies have found a significant association between DROSHA rs10719 SNP and cancer risk in diverse populations, while others have not. For instance, Yuan et al. showed that DROSHA rs10719 SNP was significantly associated with bladder cancer susceptibility [40]. Additionally, Cho et al. revealed a significant association between DROSHA rs10719 SNP and colorectal cancer risk in the Korean Population [36]. A meta-analysis by Wen et al. revealed that DROSHA rs10719 SNP is associated with cancer risk [46]. By contrast, Horikawa et al. concluded that there is no significant association between DROSHA rs10719 SNP and risk of renal cell carcinoma [42]. Similarly, Kim et al. failed to find any association between DROSHA rs10719 SNP and hepatocellular carcinoma [44]. No statistically significant associations between DROSHA rs10719 SNP and increased risk of gastric cancer in the Chinese Han population was found by Song et al. [47]. Given these contradictory results, it is far too early to conclude that DROSHA rs10719 SNP contributes to cancer susceptibility.
In conclusion, these results provide the first evidence that DROSHA rs642321 polymorphism is associated with susceptibility to breast cancer in Iranian women.
Footnotes
Acknowledgements
Authors would like to express their sincerest appreciation to all subjects for participating in this study.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee.
Informed consent
The study was approved by the Institutional Review Board /Independent Ethics Committee (IRB/IEC) of the Shahid Ashrafi Esfahani University, Isfahan, Iran and informed written consent was filled in by all participants.
Conflict of interest
The authors declare that they have no conflict of interest.
