Abstract
OBJECTIVE:
Several studies have shown the role of statin added to the patient’s chemotherapy regimen and the role of Hydroxymethylglutaryl-CoA Reductase (HMGCR) expression in predicting breast cancer patient outcomes. In our previous study, adding statins improved clinical and pathological responses in LABC patients. Furthermore, we planned to study statin’s role as a combination to neoadjuvant chemotherapy (NAC) in treating locally advanced breast cancers on the basis of HMGCR expression. Moreover, we aimed to study the association between the patients’ clinicopathological characteristics and HMGCR expression.
METHODS:
This study is a randomized, double-blinded, placebo-controlled trial in two health centers in Indonesia. Each patient enrolled with written informed consent and then randomized to receive either simvastatin 40 mg/day or a placebo, combined with the fluorouracil, adriamycin, and cyclophosphamide (FAC) NAC.
RESULTS:
HMGCR was associated with low staging and normal serum cholesterol in the high Ki67 level group (p = 0.042 and p = 0.021, respectively). The pre-and post-chemotherapy tumor sizes are significantly correlated in two groups (HMGCR negative expression, p = 0.000 and HMGCR moderate expression, p = 0.001) with a more considerable average decrease in tumor size compared to HMGCR strong expression group.
CONCLUSION:
Statin therapy might work better in HMGCR-negative or low-expression tumors, although HGMCR expression is associated with better clinical parameters in our study.
Introduction
Although there is continuing advancement in cancer diagnostics and treatments, breast cancer is still the most common cancer and cause of mortality among women in every country [1]. Besides, the clinical and molecular heterogenicity of breast cancer remains unequivocally acknowledged. Hence, treatment is usually tailored based on the specific subtype. A mandatory objective of oncological treatment is maximizing efficacy besides minimizing overtreatment and toxicity.
Statins are widely used as cholesterol-lowering drugs inhibiting HMG-CoA reductase (HMGCR), the rate-limiting enzyme for the mevalonate metabolic pathway. In breast cancer, statins are recognized as a potent anti-cancer agent since preclinical studies have shown that statins can decrease breast cancer cell’s proliferation growth, induce apoptosis and inhibit breast cancer metastasis [2–5]. Moreover, epidemiological research has affirmed that statins decrease the risk of recurrence after adjuvant treatment [6–9]. However, in clinical studies, the role of statins in oncology remains subject to controversy and shows varied treatment responses [2]. In window-of-opportunity clinical trials, patients treated with two weeks of statin treatment pre-operatively showed anti-tumoral effects through proliferation inhibition and apoptosis promotion [10–12]. Our previous study reported that 40 mg of simvastatin for four weeks as a neo-adjuvant reduced the proliferation index in LABC by 53.3% and inhibited the migration of LABC cells through the Rho/Rock pathway [13]. Furthermore, clinical studies show that giving statin as a monotherapy is not as effective as an anti-cancer because it requires a large dose, significant side effects, and a low response rate. Considering the potential of combining statins with established treatments, some studies have proved that statins enhance the chemotherapy effect. Our previous study showed that adding statins to neoadjuvant chemotherapy improved clinical and pathological responses in patients with locally advanced breast cancer without causing significant adverse effects [13].
Epidemiological and preclinical reports report that an enzyme in the mevalonate pathway is correlated with statin treatment response. However, studies on HMGCR expression as a biomarker of statin sensitivity have shown inconsistent results [14–17]. Based on a study by Bjarnadottir et al., it is suggested that statins’ anti-proliferation effect is more significant in breast cancer with HMG-CoA reductase positive [14]. Another study by Gustbee et al. showed that HMGCR expression was correlated with less aggressive tumors and not associated with disease-free survival [15]. High protein expression of HMGCR correlated with the tumor proliferation decrement by statin response. Based on a trial by Bourquist, tumor proliferation decrement was found only in tumors with HMGCR expression [17]. However, a study by Kimbung et al., shows that low basal HGMCR expression is associated with better clinical outcomes [16].
Despite our previous results study showing that adding statins improved clinical and pathological responses in LABC patients, the role of HMG-CoA expression as a potential predictive marker for simvastatin response in Indonesian breast cancer patients has not been well investigated. Therefore, as further outcomes from the previous study, we aim to evaluate the role of the combination therapy of statin and neoadjuvant chemotherapy in clinicopathological outcomes of patients with LABC based on HMGCR expression and the association between the patients’ characteristics and HMGCR expression, especially in patients receiving FAC and simvastatin combination.
Materials and methods
The trial was conducted from January 2018 to September 2019 as a prospective, multicenter, randomized, double-blind, placebo-controlled study in the treatment-free window period (diagnosis of breast cancer tumor excision surgery). It was used to examine the biological effects of a specific medication. The trial was conducted at Cipto Mangunkusumo General Hospital (CMGH) and Koja District Hospital (KDH) in Jakarta. A power calculation resulted in 60 patients sample size that is adequate to obtain a power of 90% and a 0.5 standard deviation geometric mean Ki67-difference using a two-sided test at 0.05 alpha. A sample size of 50 was chosen to avoid power drop owing to non-eligible patients. This experiment was authorized by the Medical Faculty of Universitas Indonesia Scientific Research Ethics Committee (Jakarta, Indonesia; Number: 10, Date: November 26, 2018). The study has been registered at ClinicalTrials.gov (i.e., ID number: NCT00816244, NIH).
Patients
The inclusion criteria were female patients aged 20–70 years attending the CMGH and KDH breast cancer unit with a confirmed LABC diagnosis (stage IIIA, B, and C). Patients were candidates for radical surgeries with planned FAC neoadjuvant chemotherapy prior. The participants had adequate vital organ functions (cardiac, renal, and hepatic) and a European Cooperative Oncology Group (ECOG) performance status score of ≤2. Participants who were pregnant or previously had statin or doxorubicin therapy within 30 days of enrolling in the trial were excluded. Written informed consent was required from all the patients.
The change in tumor proliferation was used as a marker of statin-induced tumor response (i.e., Ki67 expression). The secondary objective was to investigate the possible predictive function of HMGCR expression prior to statin therapy as measured by changes in tumor size following three cycles of FAC administration. After enrollment, the patients received a study-specific core biopsy before starting statin therapy. The core biopsies were promptly formalin-fixed. After three cycles of FAC and statin combination therapy, breast surgery was done using the usual surgical techniques, and tumor tissue was obtained from the original tumor in the Department of Pathology.
Treatment protocol
Either a statin or a placebo was prescribed in the outpatient clinics. By using block randomization, patients were allocated to one of two therapy groups (FAC + simvastatin or FAC + placebo). Forty mg of simvastatin and placebo were set in identical boxes, then sequentially numbered. The following chemotherapy regimen was used every three weeks based on the hospital protocol: per cycle contains doxorubicin 50 mg/m2, 5-fluorouracil 600 mg/m2, and cyclophosphamide 500 mg/m2. Oral simvastatin 40 mg or placebo was taken once daily for 21 days, since on the first day of per cycle.
Immunohistochemistry
After a core biopsy, estimates of HMGCR expression in the samples were obtained under the microscope before starting chemotherapy. Tissue samples were formalin-fixed and paraffin-embedded, then cut into 3 to 4 m slices and transferred to poly-L-lysine coated slides (Dako IHC Microscope Slides K8020). De-paraffinization was done in xylol I, II, and III, and rehydration was done with graded alcohol (100%, 96%, and 70%) for 10 min. The samples were immersed in 3% hydrogen peroxide in a methanol solution for 10 min to block the endogenous peroxidase activity. The slides were pretreated with Tris-Ethylenediamine Tetraacetic Acid (Tris-EDTA) buffer (pH 9.0) for 10 min in the decloaking chamber at 96 °C before it was cooled down for 25 min and washed with phosphate-buffered saline (PBS) solution (pH 7.4). The slides were incubated overnight with polyclonal anti-HMGCR antibody (Brand). The antibody used was diluted at 1:150. The slides were washed with PBS solution and stained with polymer-histofine for 30 min. Then, it was washed with the PBS solution and incubated with avidin-biotin complex (ABC) for one hour at 37 °C. 3,3 ′ -Diaminobenzidine tetrahydrochloride was applied for 1 to 2 min, and counterstaining was done with hematoxylin solution for 10 s. The slides were washed with distilled water, air-dried, and mounted using a DPX mount. The sections were washed with distilled water, air-dried, and mounted using a DPX mount. One certified senior breast pathologist assessed the samples. HMGCR expression was analyzed based on the cytoplasm staining intensity (i.e., strong = 3, moderate = 2, weak = 1, negative = 0) and the fraction of the HMGCR-positive cells (51–100% = +3, 11–50% = +2, 1–10% = +1, 0% = 0).
HMGCR expression could not be evaluated in 5 out of 60 samples due to damage, loss in processing, or non-representative tissue.
Statistical analysis
Baseline patient and tumor characteristics were described by mean frequency using descriptive statistics. The statistical analyses used IBM SPSS 26.0 The Kolmogorov–Smirnov and Shapiro-Wilk tests were used for the normality test, and the data were considered to have a normal distribution if the p-value was greater than 0.05. If the distribution were expected, the data would be presented as a mean and standard deviation; however, the data would be shown as median and range if the distribution was not expected. The Chi-square test was conducted to evaluate the distribution differences of tumor and clinical characteristics between HMGCR positive and HMGCR negative tumors and the differences adjusted for Ki67 level. The Mann–Whitney U test was used to assess the changes in tumor size between subgroups in the distribution of ordered categorical HMGCR intensity values and the changes in tumor size between the groups receiving statin combination and placebo combination adjusted for HMGCR expression.
Results
From January 2018 to June 2019, there were 70 patients to evaluate. Four patients were excluded. Randomly, 66 LABC patients were allocated to groups (33 to FAC plus simvastatin and 33 to FAC plus placebo), as shown in Fig. 1. Three patients were excluded from the FAC-Simvastatin group because of loss to follow-up, while three patients of the FAC-Placebo group were excluded because a patient had lung metastasis and two patients were lost to follow-up. The TMAs were therefore built using biopsies from 60 individuals, 5 of which were unable to be evaluated for HMGCR expression. This is because of a lack of tumor tissue in the TMA core or poor staining quality. Finally, 55 samples were eligible for analysis of HMGCR expression. Table 1 shows the well-balanced baseline characteristics between the two treatment groups.
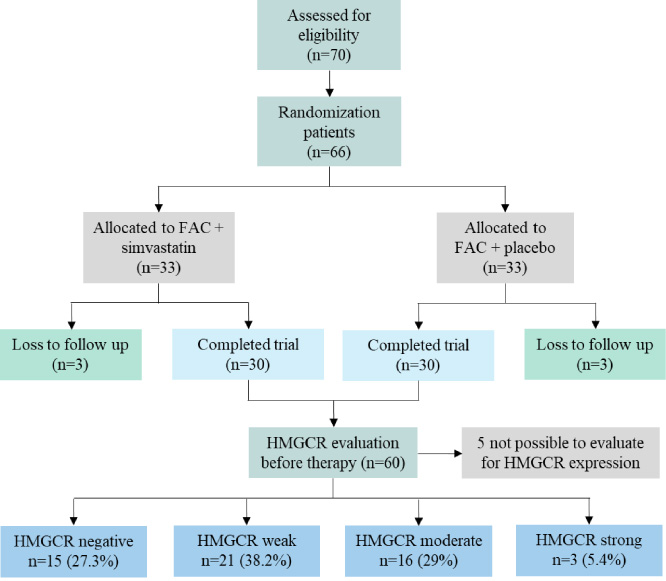
The flow of the study.
Baseline characteristics of the patients
The average age was 48 (range, 28–66 years). Before treatment, most tumor sizes were greater than 300 cm3. Most patients (75%) had invasive nonspecific type cancer and intended to have modified radical mastectomy (70%). Most of the tumors in this study are the Luminal B subtype (61.67%), followed by Luminal A (16.67%), TNBC (15%), and HER-2 (6.66%). Forty-seven out of 60 expressed high Ki67 protein.
The current study assessed HMGCR expression in 55 of 60 patients (91.6%). The remaining cores were lost throughout the immunohistochemistry processing, antibody optimization, and staining. HMGCR was mainly found in the tumor epithelium. It was expressed variedly in the cytoplasm (Fig. 1A). HMGCR expression in the stroma was not seen. Staining intensity was the only factor taken into statistical analysis of HMGCR protein expression. This is due to consistency with prior research. HMGCR was expressed in most tumor cells (>50%). HMGCR nuclear expression was not found; however, membrane expression of HMGCR was seen in a limited number of instances. In a few cases, granular cytoplasmic staining was also seen. Positive HMGCR expression was obtained in 40 patients (72.7%), while 15 out of 55 tumors (27.2%) showed no expression or lacked HMGCR expression (fraction of positive cells <1%), 21 (38.18%), indicating weak expression, 16 (29%) demonstrated moderate expression, and three patients (5.4%) demonstrated strong expression (histological findings were provided in Fig. 2). HGMCR expression in tumor cells was measured at various intensities and fractions.
In order to validate our previous findings, HMGCR expression data were dichotomized into absence versus any staining. As demonstrated in Table 2, HMGCR expression was significantly associated with staging (p = 0.047), histological subtype (p = 0.036), and Ki67 expression (p = 0.011). There was no association found in the rest of the characteristics.
The role of HR status in predicting HMGCR prompted us to investigate the association between HMGCR expression and known clinicopathological parameters stratified by Ki67 status (Table 3).
The data showed differences between the two groups with staging and serum cholesterol levels, significantly associated with HMGCR expression in the high Ki67 level group (p = 0.042 and p = 0.021, respectively).
After administering combination therapy with simvastatin to the group with positive HMGCR expression (n = 40), the average decrease in tumor size was 78.66% (n = 22, p = 0.000). After administering combination therapy with a placebo, the average decrease in tumor size was 60.54% (n = 18, p = 0.003). However, there were no statistically significant differences in tumor decrease between the two groups (p = 0.071). The graph of tumor size change before and after treatment is depicted in Fig. 3.
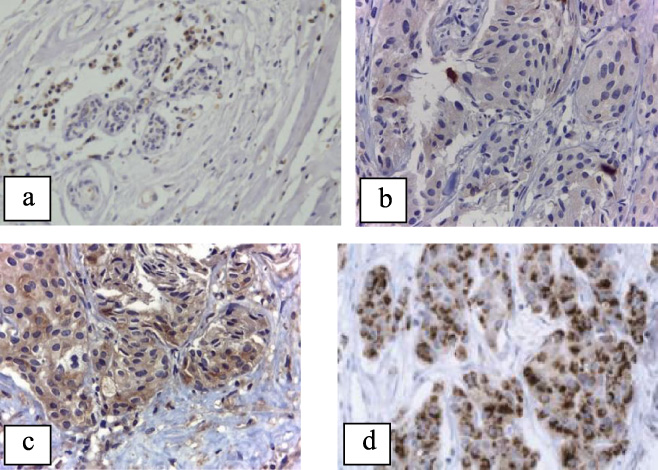
Negative (a), weak (b), moderate (c), and strong (d) HMGCR expression under the microscope (400×).
On the other hand, the average decrease in tumor size was 83.32% (n = 5, p = 0.043) after the administration of combination therapy with simvastatin and 66.08%. (n = 10, p = 0.013) after administration of combination therapy with placebo in the group with negative HMGCR expression (n = 15). There were no statistically significant differences in tumor decrease between the two groups (p = 0.806).
Statistically (as shown in Table 4), there was no difference in tumor size change between the negative or weak with moderate HGMCR expression groups (p = 0.968), the moderate and high HMGCR expression groups (p = 0.737), as well as between the negative or weak and high HMGCR expression groups (p = 0.752). Tumor size changes classified based on HMGCR expression are depicted in Fig. 4. Therefore, HMGCR expression was not associated with tumor size changes.
In the group with negative HMGCR expression (n = 15), there was a statistically significant difference between before and after chemotherapy (p = 0.000), with an average decrease in the tumor size of 70.63%. Likewise, in the group with moderate HMGCR expression (n = 21), there was a significant difference in tumor size (p = 0.001), with an average decrease of 79.49%. However, in the group with high HMGCR expression (n = 3), there was no significant difference in tumor size (p = 0.263), with a 52.99% average decrease in the tumor size.
Tumor characteristics and HMGCR expression associations
Association between HMGCR and relevant clinicopathological parameters stratified by Ki67 expression
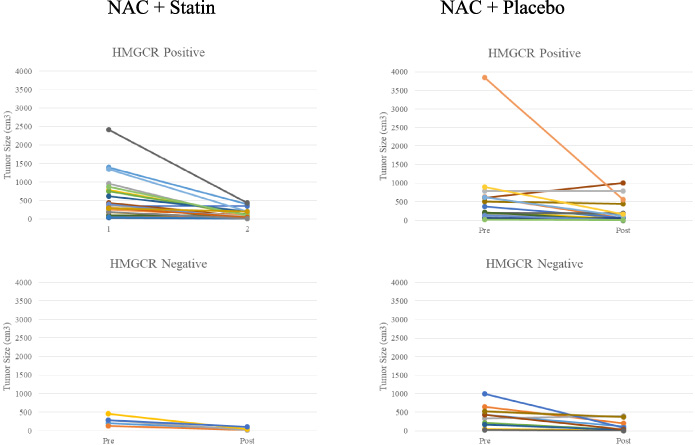
Change in tumor size at baseline (i.e., before combination treatment) and after chemotherapy treatment (i.e., after combination treatment) stratified for HMGCR.
Changes in tumor size stratified by HMGCR expression
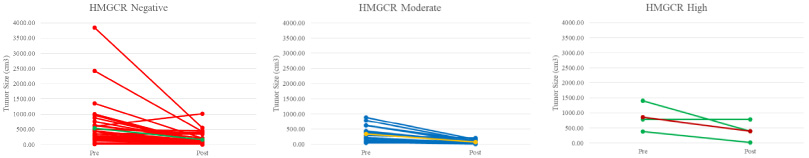
Changes in tumor size stratified by HMGCR expression.
Our study revealed that HMGCR was heterogeneously expressed in terms of intensity and fraction in the breast cancer tumor cell cytoplasm. HMGCR expression was positive in 40 out of 55 cases (72.7%), while other studies show positive results ranging from 75% to 93% [18–20].
Several studies report contradicting results on levels of HMGCR expression and tumor characteristics. Bjarnadottir et al. show that moderate or strong HMGCR expression is associated with prognostically adverse tumor characteristics, such as higher histological grade, high Ki67, and ER negativity [21]. However, other studies report that stronger HMGCR expression is correlated with a less aggressive tumor profile. Borgquist et al. show that strong expression of HMGCR is associated with small size, low histological grade, low proliferation, and ER-positive status [20]. A prospective cohort study on HMGCR expression by Gustbee et al. also shows that HMGCR expression indicates less aggressive tumor characteristics [15].
In this study, we found that positive HMGCR expression is significantly associated with histological subtype, low stage (IIIA), and low proliferation (low Ki67 expression), supporting the study of Borgquist et al. and Gustbee et al. It has been known through several studies that higher grade and higher Ki67 expression; hence, higher proliferation, are associated with poor prognosis. A registry-based study done in Norway shows that immunohistochemistry grade, subtype, size, and nodal status are death-independent predictors in breast cancer. If all prognostic factors were included, the mortality rate would be 20- to 40-fold greater than the group with better clinical characteristics (e.g., the smallest size, the lowest grade, and the status of ER+PR+HER2−). Advanced-stage and high-grade ER+PR-HER2- tumors have a high death rate, similar to the TNBC subtype [22]. TNBC subtype has the worst prognosis out of the four subtypes (luminal A, luminal B, HER-2, and TNBC). One prospective cohort study of 4102 females with breast cancer shows that TNBC has the worst prognosis with LCR of 89.6%, OS of 78.5%, and ROS of 80.1%; while luminal A has the best prognosis with LCR of 99.1%, OS of 95.1%, and ROS of 100% [23].
Association between HMGCR expression and ER/PR positivity has been proved by Borgquist et al. The study shows a significant correlation between HMGCR expression and ER status (p = 0.01). However, HMGCR expression was not associated with PR status (p = 0.79) [20]. Anjum et al. found a significant association between HMGCR expression and ER expression (p = 0.003) and PR expression (p = 0.014) [19]. In our study, there was no significant association between HMGCR expression and either ER or PR expression, with a p-value of 0.192 and 0.354, respectively.
A study by Bjarnadottir et al. examining Ki-67 and HMGCR immunohistochemistry expression in pre-and post-statin matched tumor samples show that statin therapy increased HMGCR expression while decreasing Ki-67 expression. Furthermore, the reduction in Ki-67 status was more pronounced in tumors with HMGCR expression prior to statin regimen therapy [21]. Ki67 expression is associated with proliferation; therefore, HMGCR may be a prognostic marker for statin therapy in breast cancer.
Studies investigating HMGCR expression as a statin sensitivity biomarker have contradictory findings, most likely owing to the lack of specificity in several commercial HMGCR antibodies. These findings further point to a complicated connection between HMGCR expression and statin sensitivity in cancer [15,24]. Another factor that may be taken into account is the variance in HMGCR gene expression, which can increase the response to statin therapy significantly [8].
Kimbung et al. report that ER-negative (basal-like molecular subtype) is strongly associated with statin sensitivity. Statin has been shown to strongly reduce proliferation at low ER positivity levels. A similar result is also observed in genome transcription in which cells with negative ER expression show cell cycle progression regulation and apoptosis-associated genes. They also report that low basal HMGCR expression can predict tumor proliferation reduction following atorvastatin administration. In contrast, previous studies report that high protein expression correlated to tumor proliferation decrement by statin response [16].
When adjusted for Ki67 expression, we found that HMGCR expression is associated with normal serum cholesterol levels. Feldt et al. show that breast cancer tumor cells may be similar to hepatocytes in response to increasing cholesterol levels in the blood. When intracellular cholesterol depletes, HMGCR and LDLR expression are upregulated to increase cholesterol uptake into the cell [25]. In statin-sensitive tumor cells, there is modest or deficient mevalonate feedback response, contributing to dysregulation of the pathway and inability to detect a decrease in HMGCR activity, resulting in tumor cell apoptosis [26].
In addition to promoting carcinogenesis, increased HMGCR expression and dysregulation of the mevalonate pathway can render cancer cells vulnerable to statin treatment. A high level of HMGCR expression in these tumors may indicate tumor dependency, in which a modest reduction in the mevalonate pathway is sufficient to induce tumor-specific cell death. However, when HMGCR expression increases, a higher dose of statin medicine is required to inhibit the mevalonate pathway, decreasing statin sensitivity [24]. As a result, careful attention is necessary when analyzing the usefulness of HMGCR as a statin sensitivity predictive marker in cancer [24].
When a comparison between positive and negative HMGCR expression in groups receiving either simvastatin or placebo in addition to the chemotherapy regimen, a larger average decrease in tumor size is seen in the group with negative HMGCR expression, in the group receiving simvastatin, the average decrease in tumor size is 83.32% in HMGCR negative group versus 78.66% in HMGCR positive group. Similar results are observed in the group receiving a placebo, with an average decrease in tumor size of 66.08% and 60.54%, respectively, favoring the negative expression groups. Irrespective of the treatment given (statin or placebo), a larger average decrease in tumor size is found in the negative and moderate HMGCR expression groups (70.63% and 79.49%, p = 000 and 0.001) compared to the strong HMGCR expression group (52.99%, p = 0.263). These results support the study by Kimbung et al., showing that low basal HGMCR expression is associated with better clinical outcomes. As the average decrease in tumor size is observed larger in HMGCR negative group, it may be implicated that statin therapy, in addition to chemotherapy, brings better clinical outcomes in Asian-Indonesian patients with negative or low HMGCR expression.
Cancer metabolism markers, such as HMGCR, may be affected by dietary choices, lifestyle, medicine, race, etc. IHC was the sole approach employed in the current investigation to measure HMGCR expression. However, combining more advanced methods, such as RNA quantification PCR, with a bigger size cohort study might assist in elucidating the function of HMGCR in breast cancer. Hence, further study on gene expression of HMGCR using RT-PCR is recommended to evaluate quantitative analyses despite our current semi-quantitative study using the immunohistochemistry method. Some similar target gene mutations, rather than HMGCR detection, may also potentially be prognostic indicators for statins [27]. Future research on the potential usage of HMGCR expression as a predictor of statin susceptibility is still needed.
Conclusion
HMGCR expression is related to less aggressive tumor features in this randomized clinical trial investigation of locally advanced breast cancer patients. We also found that statin has a better impact on the HMGCR negative expression group.
Footnotes
Acknowledgements
The authors would like to thank Kevin Varian M., MD, for assisting in the publication of this study.
Conflicts of interest
The authors declare no conflicts of interest.
