Abstract
OBJECTIVES:
Early diagnosis of triple-negative (TN) and human epidermal growth factor receptor 2 positive (HER2+) breast cancer is important due to its increased risk of micrometastatic spread necessitating early treatment and for guiding targeted therapies. This study aimed to evaluate the diagnostic performance of machine learning (ML) classification of newly diagnosed breast masses into TN versus non-TN (NTN) and HER2+ versus HER2 negative (HER2−) breast cancer, using radiomic features extracted from grayscale ultrasound (US) b-mode images.
MATERIALS AND METHODS:
A retrospective chart review identified 88 female patients who underwent diagnostic breast US imaging, had confirmation of invasive malignancy on pathology and receptor status determined on immunohistochemistry available. The patients were classified as TN, NTN, HER2+ or HER2− for ground-truth labelling. For image analysis, breast masses were manually segmented by a breast radiologist. Radiomic features were extracted per image and used for predictive modelling. Supervised ML classifiers included: logistic regression, k-nearest neighbour, and Naïve Bayes. Classification performance measures were calculated on an independent (unseen) test set. The area under the receiver operating characteristic curve (AUC), sensitivity (%), and specificity (%) were reported for each classifier.
RESULTS:
The logistic regression classifier demonstrated the highest AUC: 0.824 (sensitivity: 81.8%, specificity: 74.2%) for the TN sub-group and 0.778 (sensitivity: 71.4%, specificity: 71.6%) for the HER2 sub-group.
CONCLUSION:
ML classifiers demonstrate high diagnostic accuracy in classifying TN versus NTN and HER2+ versus HER2− breast cancers using US images. Identification of more aggressive breast cancer subtypes early in the diagnostic process could help achieve better prognoses by prioritizing clinical referral and prompting adequate early treatment.
Introduction
Worldwide the total number of new cancer cases and cancer-related deaths continues to rise [1,2]. In women, breast cancer is still the most commonly diagnosed malignancy and a leading cause of cancer-related death, with up to an estimated 2.08 million new cases and 0.63 million deaths globally [1,2]. Breast cancer is a disease comprising of different molecular subtypes based on its receptor protein expression. Triple-negative (TN) tumors lack expression of hormone estrogen receptor (ER), hormone progesterone receptor (PR), and human epidermal growth receptor 2 (HER2). In contrast, HER2+ tumors express the HER2 receptor, with variable ER/PR status. TN and HER2+ subtypes have demonstrated more aggressive biological behaviour resulting in poorer clinical outcomes and prognosis [3,4]. With mammography, conventional breast US forms the cornerstone of identifying abnormal breast lesions and diagnostic workup [5,6]. The standard for describing and classifying breast lesions is the Breast Imaging Reporting and Data System (BI-RADS) lexicon, developed by the American College of Radiology [7]. The BI-RADS system is widely accepted in standard practice and provides a risk assessment of lesions on breast imaging. However, qualitative appearances of benign versus malignant masses on US may overlap [8]. Despite growing evidence that certain imaging features are associated with specific biological phenotypes [9], correlation with pathology currently remains the gold standard in the definite diagnosis of breast cancer [9].
The use of radiomics in combination with Machine Learning (ML) is a hot topic in breast cancer research, aiming to improve the identification of malignant lesions, discriminate tumor grade, and predict prognostic factors such as the response to neoadjuvant therapy and risk of tumor recurrence [9]. Computer-aided diagnostics that predict more aggressive TN and HER2+ molecular subtypes early in the diagnostic process may prove clinically relevant, allowing for prioritizing clinical referral and earlier initiation of treatment [10].
Various techniques are used to capture breast cancer such as Ultrasound Sonography (ULS), Computerized Thermography (CT), Biopsy (Histological images), Magnetic-Resonance-Imaging (MRI), and Digital Mammography breast X-ray images (DMG). Machine learning (ML) is a type of artificial intelligence. Machine learning algorithms (MLs) are suggested as an alternative to human vision and experience for analysing medical images and taking the final decisions with high accuracy. Many ML techniques have been used with mammography, US, and MRI to detect breast cancer and differentiate benign from malignant findings. Yassin et al. reported that 64% of the papers included in their systematic review on the use of ML in the detection of breast cancer reported on digital mammography, whereas only 19% reported on US and 9% on MRI [11]. Support vector machine (52.6%), artificial neural network (26.0%), and K-NN (13.6%) are the most frequently researched ML models.
Using traditional machine learning requires initial pre-processing and feature selection, which take time a computational consumption. While recent studies have used the deep learning approach as it is able to extract suitable features automatically. Different studies employed convolution neural network (CNN) to classify breast tumors, as it is the network architecture of choice for identifying and recognizing objects. A study evaluated three datasets with different sizes and achieved very high accuracy, reaching 97% [12].
High accuracy rates have been found to date ranging from 64.7–100% for mammography, 75.5–98.3% for US and 74.4–98% for MRI [12]. Most of the available studies report on a visual interpretation of US images by the radiologist, where the identification of specific qualitative US features leads to a subtype diagnosis [13,14]. The selected US features used, and outcome parameters vary among studies. Overall, studies by Wu et al., Cho N., Rashmi et al., and Huang et al., reported circumscribed margins and posterior acoustic enhancement to be suggestive of TN breast cancer and microcalcifications, posterior mixed acoustic pattern and high vascularity of HER2+ cancer [15–18]. Machine learning systems based on BI-RADS feature can help in malignant/benign differentiation, but further improvement is needed.
This pilot study aims to evaluate radiomic features associated with TN and HER2+ lesions and test the diagnostic performance of ML classifiers to classify newly diagnosed malignant breast masses as TN or Non-triple-negative (NTN) and HER2+ versus HER2− breast cancer using greyscale US b-mode imaging.
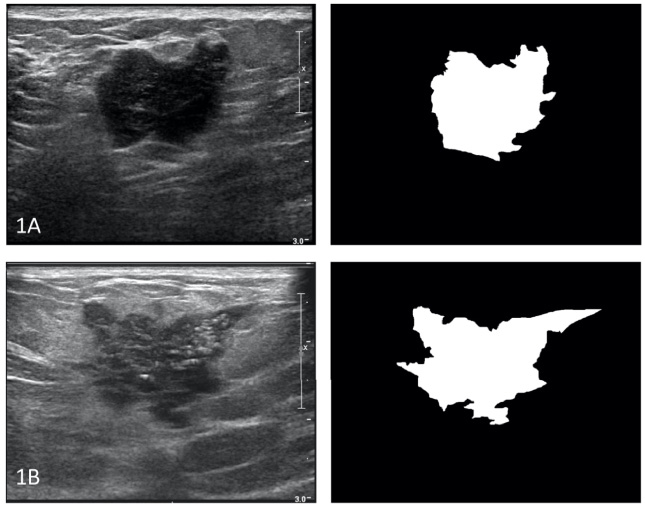
Triple-negative (1A) and HER2+ (1B) breast cancer: representative grayscale US image (left) and corresponding mask (right).
Patient and data acquisition
The institutional research ethics board approved this study, and the need for informed consent was waived. A retrospective chart review of the picture archiving and communications systems (PACS), in addition to the electronic patient records (EPR) database, was conducted for adult female patients who underwent diagnostic workup of breast abnormalities in the Rapid Diagnostic Unit (RDU) of our hospital between June 1, 2011, and July 31, 2019. Additional inclusion criteria included availability of the initial grayscale US images in 2 orthogonal planes, confirmed invasive primary breast malignancy on pathology and molecular subtyping results. Patients with benign pathology results, who solely underwent biopsy of recurrent or metastatic lesions to sites in or outside the breast tissue (i.e. chest wall), and those with breast lesions extending beyond the edge of the US image were excluded. 88 patients were included in our study, 44 TN and 44 NTN as our control group. TN breast cancers comprise only 15% of all breast cancers and are relatively rare.
In concordance with American breast cancer demographics, the initial study cohort was significantly enriched by NTN and HER2− breast cancer cases. Subsequently, PACS and EPR databases were mined for TN and HER2+ cases to balance the patient cohorts. The final study population was randomly selected from the initial NTN and HER2− cases and added to the confirmed TN and HER2+ cases. The two cohorts were classified as TN versus NTN and HER2+ versus HER2−.
Certification in Breast imaging (CBI) or Diagnostic medical sonography (DMS) certified US technologists acquired the US images using a Phillips IU-22 system with 17–6 MHz linear transducers. Our institution’s routine imaging protocol includes grayscale and colour Doppler image acquisition in 2 orthogonal planes.
Summary of patient characteristics
Summary of patient characteristics
Table 1 summarizes the patient characteristics and lesion pathology results. TN = triple negative, NTN = non-triple negative, HER2 + ∕− = human epidermal growth factor receptor 2 positive/negative, IDC = invasive ductal carcinoma, ILC = invasive lobular cancer, IMC = invasive mammary cancer.
A board-certified breast radiologist and a breast imaging fellow outlined the breast masses identified on the US grayscale images’ transverse view (Fig. 1). The manually segmented lesions were then pre-processed for image analysis. An open-source software package was utilized (GNU image manipulation program (GIMP) version 2.10.18 [19]. The outlined breast masses were converted to binary masks, which delineated the breast mass (foreground) from the background (Fig. 1). Using Python programming language version 3.7.6, 249 radiomic features were calculated per US image. Of the 249 features, 213 first and second-order texture features were calculated using Pyradiomics version 3.0 [14]. Additionally, six Fourier shape descriptors [21], eight gradient features [21], twelve intensity features [20], and ten morphological features were calculated.
To avoid overfitting the classifiers, a maximum of 8 features were selected for inclusion into a classification model, based on Harrell’s 1/10 rule. Feature reduction consisted of univariate analyses to compare variables between groups; for this, a two-sided student t-test was used to identify statistically significant differences between groups. Subsequently, a stepwise discriminant function was used to select the most relevant and correlated features (selection criterion; r2 > 0.8, features = 8). The final features were used to train various ML classifiers to label TN versus NTN and HER2+ or HER2−.
Using scikit-learn version 0.21.3, the following supervised ML classifiers were trained: (1) logistic regression (LR), (2) k-nearest neighbour (K-NN), and (3) Naïve Bayes (NB) [21]. As such, six classifiers were trained in total, three per sub-group analysis. Within the ML frameworks, all data were partitioned into a 3:1 ratio. A 10-fold cross-validation strategy was adopted for the training dataset, which optimized the classifiers’ efficacy and generalization. The unseen (test) data were used to report the classification performance of all classifiers. Core biopsy pathology was used as the reference standard for the outcome variables. The area under the receiver operating characteristic curve (AUC), sensitivity (%), and specificity (%) were calculated for each classifier.
Results
There were 88 patient cases identified in the retrospective review. All cases demonstrated a unifocal malignant breast mass, which was confirmed on pathology. Of these, 82 were invasive ductal carcinoma (IDC), 4 were invasive lobular carcinoma (ILC) and 2 were invasive mammary carcinoma (IMC). Molecular subtyping confirmed 22 TN versus 66 NTN cases. Of the 66 NTN cases, 21 were HER2+, and 45 were HER2−. There were 38 ER+ PR+ HER2− cases, 22 TN cases, 13 ER− PR− HER2+ cases, 7 ER+ PR+ HER2+ cases, 5 ER+ PR− HER2− cases, 2 ER− PR+ HER2− cases and 1 ER−, PR+, HER2+ case. A summary of the lesion characteristics can be found in Table 1.
Performance of the diagnostic models
The eight most discriminating, relevant, and correlated features (Table 2) were used to train the ML classifiers per sub-group. Wavelet HH glszm Large Area Emphasisa was the only feature included in both sub-group analyses.
Qualitative features of US images used in differentiating TN and HER2+ breast cancers
Qualitative features of US images used in differentiating TN and HER2+ breast cancers
Table 2 summarizes the 8 radiomic features that most significantly contributed to differentiating TN from NTN and HER2+ from HER2, based on a step-wise discriminant function. TN = triple negative, NTN = non-triple negative, HER2 + ∕− = human epidermal growth factor receptor 2 positive/negative.
In the classification of TN versus NTN breast cancers, the LR classifier demonstrated an AUC of 0.824, sensitivity of 81.8%, specificity of 74.2%, the K-NN classifier an AUC of 0.739, sensitivity of 85.7%, specificity of 65% and the NB classifier an AUC of 0.807, sensitivity of 71.4%, specificity of 90%. In the HER2+ versus HER2− model, the LR classifier demonstrated an AUC of 0.778, sensitivity of 71.4%, specificity of 71.6%, the K-NN classifier an AUC of 0.679, sensitivity of 83.3%, specificity of 52.4% and the NB classifier an AUC of 0.535, sensitivity of 50%, specificity of 57.1%. The LR classifier demonstrated the highest accuracy in both the TN and HER2 model. All ML classifiers performed better in classifying TN versus NTN breast cancers than in classifying HER2+ versus HER2− breast cancers. Table 3 summarizes all classifiers’ performance, and Figs 2 and 3 display the ROC for each classifier per sub-group.
Diagnostic performance TN and HER2 classfication
Table 3 summarizes the diagnostic accuracy results, by means of the area under the operator characteristics curve (AUC), sensitivity and specificity, for each classifier, per model. TN = triple negative, HER2 = human epidermal growth factor receptor 2.
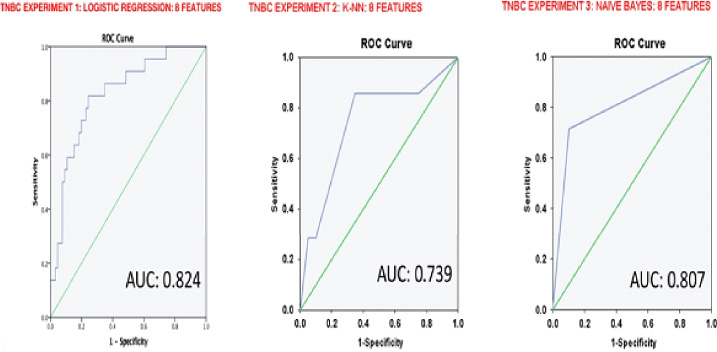
ROC-curve of each classifier for the TN versus NTN model. The logistic regression classifier was the most accurate in classifying the TN subtype with an AUC of 0.824.
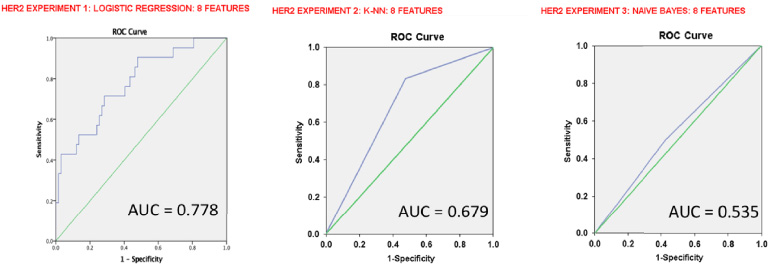
ROC-curve of each classifier for the HER2+ model. The logistic regression classifier was the most accurate in classifying the HER2+ subtype with an AUC of 0.7.
Triple-negative and HER2+ breast cancers have demonstrated more aggressive biological behaviour resulting in poorer clinical outcomes and prognosis [3,4]. TN breast cancer has the worst survival. It arises at an earlier age, is almost exclusively high grade and lacks specific therapeutic targets in the absence of ER/PR/HER2 expression [11]. They have, however, a higher sensitivity to neoadjuvant chemotherapy (NAC) with higher rates of pathological complete response leading to a more favourable outcome [11]. The current studies support that integrating immunotherapy with conventional therapy for better survival outcome in early triple-negative breast cancer-TNBC can be achieved [22].
Unfortunately, patients with an incomplete response to NAC have a poor prognosis due to high relapse rates [11]. HER2+ breast cancers are more likely to metastasize (especially to the central nervous system and viscera) than HER2− breast cancer and HER2-expression and are key risk factors for a de novo diagnosis of metastatic breast cancer (Stage IV disease) [23–25]. Luckily, the HER2-expression also allows targeted immunotherapy with monoclonal antibodies with a new important role for dual HER2 blockade (Trastuzumab + purtuzumab or neratinib) in the neoadjuvant setting leading to an improved rate of pathological complete response and better long-term outcomes [23–26].
This study investigated ML classification of TN or NTN and HER2+ or HER2− breast cancer subtypes using radiomic features derived from grayscale US images. This preliminary study demonstrated that the LR classifier outperformed NB and K-NN in correctly labeling TN and HER2 sub-groups; additionally, all ML classifiers performed better in classifying TN compared to HER2+ cancers. In using basic grayscale US images as the source of radiomic feature extraction, this technique could develop into a widely applicable method to be used in the initial diagnostic workup of a patient with a suspicious breast mass in the day-to-day radiology practice [27–29].
Most of the available studies report on a visual interpretation of US images by the radiologist, where the identification of specific qualitative US features leads to a subtype diagnosis. The selected US features used, and outcome parameters vary among studies. Overall, studies by Wu et al., Cho N., Rashmi et al., and Huang et al., reported circumscribed margins and posterior acoustic enhancement to be suggestive of TN breast cancer and microcalcifications, posterior mixed acoustic pattern and high vascularity of HER2+ cancer [15–18]. Our methodology is similar to an automatic segmentation for tranvsaginal ultrasound images of cervical cancer [30].
More papers are being published on the use of ML analysis of US images to classify the different molecular subtypes. In line with our results, Wu et al. reported an AUC of 0.85 in classifying TN versus NTN breast cancers using the LR ML classifier on greyscale US images [17]. Adding the use of colour Doppler images, Wu and colleagues reported an AUC of 0.88, sensitivity of 86.96%, and specificity of 82.91% [17]. Guo et al. demonstrated that the support vector machine classifier with quantitative US features (shape, margin, boundary, calcification, echo pattern and posterior acoustic pattern) was able to accurately predict the molecular subtype, and achieved an AUC of 0.760, sensitivity of 97.9% and specificity of 60.1%. Guo and colleagues also confirmed that circumscribed margins and posterior enhancement were strongly associated with the TN subtype [31].
To our knowledge, this is the first study reporting the diagnostic performance of a ML HER2 classifier using grayscale US images. This pilot study provides promising results regarding HER2 status prediction. However, lower accuracy was found when compared to the TN classifier. This requires more studies to complete our data.
There are various limitations to this study. First, the major limitation is the lack of any external validation. Second, outlining the breast masses of interest is challenging due to the unclear margins of many tumors [32], making it user-dependent and subject to interobserver variability. Second, while imaging protocols are standardized at our institution, there is variability in the technical sonographic parameters on a case-by-case basis. Third, this study is limited by sampling bias due to its retrospective study design conducted on a subset of patients with known malignancy and a small sample size. Also, the radiomic quality score reaches 10/36, which appears low in 2022. Finally, it is important to acknowledge that the database of patients who underwent breast imaging workup was mined for TN and HER2+ cases because, in ML classification, the underlying training set is assumed to be equally distributed. The class imbalance may result in difficulties accurately predicting the minority classes, in this case, TN and HER2+ [33]. Increasing the number of included patients and automated segmentation in future prospective studies would yield more clinically relevant results. Furthermore, including benign lesions in the study could develop a more comprehensive US-based ML classification model, improving efficacy and non-invasive diagnostic accuracy.
Conclusion
ML classification models of US images have high diagnostic accuracy in classifying either TN and HER2+ breast cancer subtypes depending on the type of the provided algorithm. A radiomic analysis combined with ML showed promising results to differentiate benign from malignant breast lesions on ultrasound images. Identifying these more aggressive breast cancer subtypes early in the diagnostic process could help achieve better prognoses by prioritizing clinical referral and prompting early appropriate aggressive treatment.
Footnotes
Conflicts of interest
None.
