Abstract
TP53 encodes a major tumor suppressor protein which blocks carcinogenesis process in a variety of tissues including breast tissue. Expression and function of this gene is regulated by a number of long non-coding RNAs (lncRNAs) among them are PANDA, MEG3 and CASC2. We measured expression of TP53 and these transcripts in a cohort of Iranian breast cancer patients. Expression levels of TP53, MEG3, CASC2 and PANDA were significantly lower in tumoral samples compared with non-tumoral samples (Posterior mean differences = −4.26, −1.66, −5.98 and −3.13, respectively; P values < 0.0001). Expression of CASC2 was higher in Her2 1+ cases compared with Her2 negative cases (Beta = 1.85, P value = 0.037). Expression levels of MEG3 and TP53 were lower in grade 2 samples compared with grade 1 (Beta = −1.86, P value = 0.006 and Beta = −2.24, P value = 0.003, respectively). There was no other significant association between expression of genes and clinical variables. CASC2 had the best performance among these genes with area under curve value of 0.78 and sensitivity and specificity values of 56.33% and 88.73%, respectively (P value < 0.0001). The current investigation supports the role of TP53-related lncRNAs in the pathogenesis of breast cancer.
Introduction
Being located on chromosome 17p, TP53 gene encodes a nuclear protein that hinders proliferation of genetically altered cells [1]. This tumor suppressor protein regulates genomic stability by governing cell cycle progression and prompting apoptosis when DNA damage is beyond the repair capacity of the cell [2]. Although TP53 mutations have been detected in minority of breast cancers, changes in levels of interacting molecules or target genes have been shown to decrease the capacity of this protein in management of stress events [2]. Notably, epigenetic alterations in p53 transcriptional targets of p53 are additional mechanisms for modulation of p53 function in breast cancers that do not harbor TP53 mutations [3]. Moreover, mutations in this gene predict poor survival of breast cancer patients [3]. Recent studies have shown interaction between this tumor suppressor gene and a number of long non-coding RNAs (lncRNAs). For instance, P21-associated noncoding RNA DNA damage-activated (PANDA) is an lncRNA whose expression is activated following DNA damage [4]. This lncRNA suppresses apoptosis by inhibiting the function of nuclear transcription factor Y. Moreover, PANDA has been shown to stabilize p53 protein in response to DNA damage [5]. Maternally expressed 3 (MEG3) has been shown to enhance p53 protein levels and induce p53-dependent transcription from a p53-responsive promoter. Moreover, MEG3 down-regulates MDM2 levels to enhance p53 stability [6]. Finally, lncRNA cancer susceptibility candidate 2 (CASC2) is a tumor suppressor lncRNA that down-regulates CDK1 and BCL2 [7,8]. Both CDK1 and BCL2 have suppressive effects on p53 function [9,10]. Therefore, CASC2 indirectly affects p53-related pathways. Forced up-regulation of CASC2 in thyroid cancer tissues has suppressed proliferation of these cells and attenuated their migratory potential and invasiveness. Besides, this lncRNA can suppress epithelial-mesenchymal transition through enhancing E-cadherin levels and decreasing expressions of ZEB1 and N-cadherin [11]. In the current study, we compared transcript levels of TP53, MEG3, CASC2 and PANDA in breast cancer tissues versus non-tumoral tissues to identify their relevance with breast carcinogenesis.
Material and methods
Patients
Expressions of TP53 and related lncRNAs were quantified in 69 pairs of tissues including both tumoral and non-tumoral tissues. Patients recruited for this study were hospitalized in Farmanieh and Sina hospitals, Tehran, Iran during 2017–2020. The study protocol was approved by the ethical committee of Shahid Beheshti University of Medical Science and all methods were performed in accordance with the relevant guidelines and regulations (IR.SBMU.RETECH.REC.1400.087). All enrolled patients signed informed consent forms. Specimens were gathered during surgery and prior to any chemotherapy or radiotherapy. Medical records were collected to obtain pathological and clinical data.
Experiments
All obtained tissues were subjected to RNA extraction using the RiboEx kit (GeneAll, Seoul, South Korea). After that, 75–100 ng of RNA was converted to cDNA using the ExcelRTTM Reverse Transcription Kit II (SMOBIO, Taiwan). Expressions of TP53 and related lncRNAs were measured in all paired samples in the ABI step one plus PCR machine. Expression levels were normalized to transcripts of B2M. Reactions were prepared using RealQ Plus 2x PCR Master Mix (Ampliqon, Odense, Denmark). Forward and reverse primers were designed using Primer3 software (Table 1).
Primer sequences
Primer sequences
Bayesian estimation supersedes the t-test (BEST) was used to test the significance of difference in means of genes expression levels between tumoral and non-tumoral samples. A t student prior family was assumed for parameters with 5000 iterations and 2000 burn-outs. P values were estimated using frequentist methods including median test. The Spearman correlation coefficients were calculated to assess the association between expression levels of genes. These analyses were performed using BEST, RJags, and ggplot2 packages in the R 4.0.3 software.
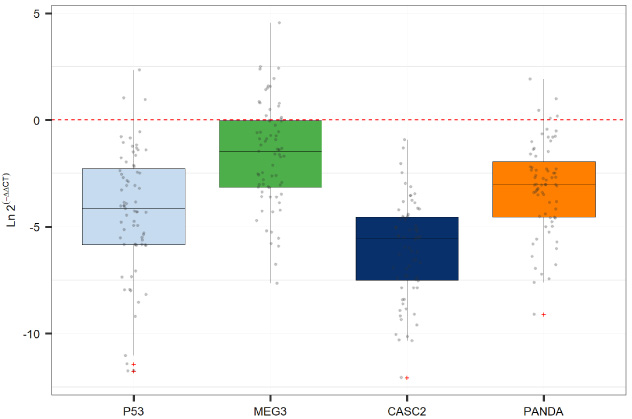
Expression levels TP53, MEG3, CASC2 and PANDA in tumoral tissues versus non-tumoral tissues as calculated by Ln 2(−delta delta Ct). The data is expressed relative to the matched healthy tissue.
The comparison of relative expression of genes between tumoral and non-tumoral samples (Results of Bayesian estimation supersedes t-test)
Abbreviations; SD: Standard deviation. HDI: highest density interval.
Expression assays have been conducted on the same cohort of breast cancer patients as our previous study [12]. Detailed demographic and clinical data of these patients have been reported formerly [12]. Figure 1 shows expression levels TP53 and related lncRNAs in breast cancer tissues versus non-tumoral tissues as calculated by Ln 2(−delta delta Ct).
Expression levels of TP53, MEG3, CASC2 and PANDA were significantly lower in tumoral samples compared with non-tumoral samples (Posterior mean differences = −4.26, −1.66, −5.98 and −3.13, respectively; P values < 0.0001) (Table 2).
Expression of CASC2 was higher in Her2 1+ cases compared with Her2 negative cases (Beta = 1.85, P value = 0.037). Yet, expression of this gene was similar between Her2 2+ or Her2 3+ cases compared with Her2 negative ones. Expression levels of MEG3 and TP53 were lower in grade 2 samples compared with grade 1 (Beta = −1.86, P value = 0.006 and Beta = −2.24, P value = 0.003, respectively). There was no other significant association between expression of genes and clinical variables (Table 3).
Association between expression levels of TP53, PANDA, MEG3 and CASC2 genes and clinical data
Association between expression levels of TP53, PANDA, MEG3 and CASC2 genes and clinical data
While expression of none of mentioned genes was correlated with patients’ age, there were significant correlations between all pairs of genes with correlation coefficients ranging from 0.66 (MEG3/PANDA) to 0.37 (MEG3/CASC2) (Fig. 2).
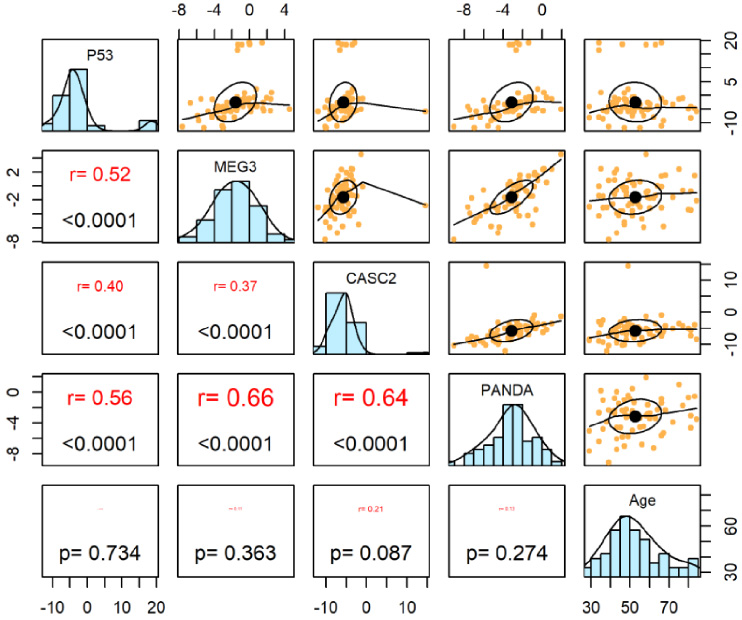
Correlation between expression levels of TP53, PANDA, MEG3 and CASC2. The distribution of expression levels of TP53, PANDA, MEG3 and CASC2 genes is shown on the diagonal. On the bottom of the diagonal, P values and correlation coefficients are shown. The bivariate scatter plots are displayed in the upper part.
Finally, the diagnostic value of TP53, PANDA, MEG3 and CASC2 in differentiation between tumoral and non-tumoral tissues was evaluated through plotting receiver operating characteristic (ROC) curves (Fig. 3). CASC2 had the best performance among these genes with area under curve (AUC) value of 0.78 and sensitivity and specificity values of 56.33% and 88.73%, respectively (P value < 0.0001). TP53 with AUC value of 0.62 ranked second in this regard.
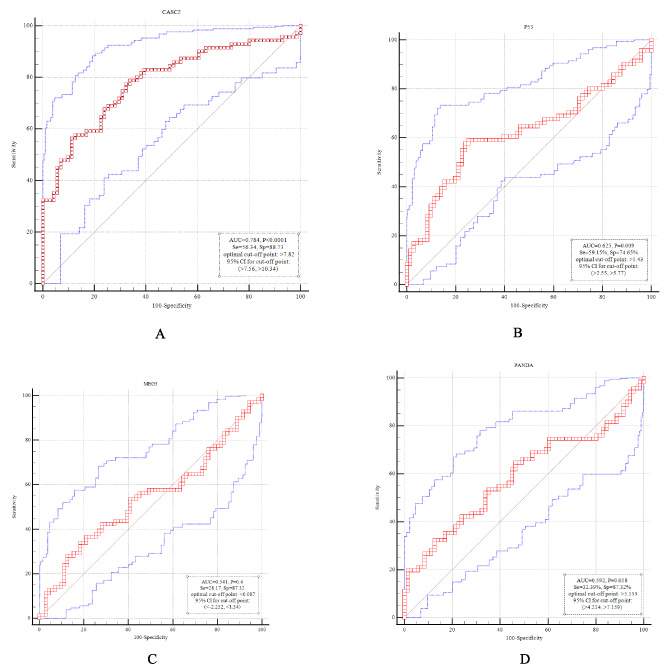
Diagnostic value of CASC2 (A), TP53 (B), MEG3 (C) and PANDA (D) in differentiation between tumoral and non-tumoral tissues. Area under curve (AUC), sensitivity, specificity, optimal cut-off point and P values are shown.
As the first identified tumor suppressor gene, TP53 has been shown to eradicate and suppress the proliferation of genetically abnormal cells, thus averting neoplastic transformation [3]. However, the frequency of mutations in this gene in breast cancer is lower than what has been expected [3], suggesting an alternative route for inactivation of the encoded protein in this kind of cancer. LncRNAs have critical roles in the pathoetiology of breast cancer [13]. Dysregulation of lncRNAs not only affects evolution of breast cancer, but also influences the clinical progression and outcomes of this type of malignancy [14]. Various studies have shown dysregulation of lncRNAs in breast cancer specimens or sera of affected females signifying these transcripts as diagnostic molecules for breast cancer [14].
LncRNAs have been demonstrated to influence breast cancer development through several mechanisms among them is modulation of activity of tumor suppressor genes [15]. In order to find putative mechanism of p53 inactivation in breast cancer, we evaluated expression of three lncRNAs which have been formerly recognized as modulators of p53 stability or function [4,6,7]. Most notably, expression levels of TP53, MEG3, CASC2 and PANDA were significantly lower in tumoral samples compared with non-tumoral samples. PANDA and MEG3 have been shown to directly enhance p53 transcription or stability [5,6]. Meanwhile, CASC2 influences functions of two downstream targets of p53 i.e. CDK1 and BCL2 [7,8]. Therefore, down-regulation of MEG3, CASC2 and PANDA in breast cancer samples not only attenuates p53 stability and function, but also over-activates oncogenes which are normally suppressed by p53. Taken together, the current study suggests abnormality in the function of p53-related molecular cascades in breast cancer which might be an explanation for low frequency of TP53 mutations in this neoplasm. CASC2 has been shown to inhibit breast cancer cell growth and metastasis via modulation of miR-96-5p/SYVN1 cascade [16]. Expression of CASC2 has been remarkably reduced in breast cancer tissues and cell lines compared with neighboring non-affected tissues and non-malignant breast epithelial cells. Most notably, CASC2 up-regulation could inhibit viability, migratory potential and invasive properties of breast cancer cells, while elevating their apoptosis [16]. In addition, MEG3 has been demonstrated to suppress breast carcinogenesis through influencing expression of miR-21 and activity of PI3K/Akt, p53 and NF-κB [17,18]. However, another study in a small-sized cohort of breast cancer patients has demonstrated up-regulation of PANDA in breast cancer samples compared to breast cysts tissues [19]. Low sample size of the mentioned study and the source of control tissues might explain the discrepancy between our results and the results of this study.
Expression of CASC2 was higher in Her2 1+ cases compared with Her2 negative cases. Her 2 has been shown to be up-regulated in 15–30% of invasive breast cancer samples. Moreover, over-expression of this factor has both prognostic and predictive implications [20,21]. Besides, expression levels of MEG3 and TP53 were lower in grade 2 samples compared with grade 1. Although these findings might indicate changes in the expression of mentioned lncRNAs throughout the course of breast carcinogenesis, the exact underlying mechanisms for these observations should be uncovered.
Consistent with the role of these lncRNAs in the regulation of activity of p53, significant pairwise correlations were detected between expression levels of TP53 and these lncRNAs. Moreover, CASC2 had the best performance for differentiation between tumoral and non-tumoral tissues, potentiating this lncRNA as a probable marker for breast cancer.
In brief, we demonstrated abnormal down-regulation of TP53 and its related lncRNAs in breast cancer tissues, which is in line with the critical role of this tumor suppressor gene in prevention of cancer. However, we state lack of functional studies as a limitation of our work.
Footnotes
Conflict of interest
The authors declare they have no conflict of interest.
Role of funding sources
The current study was supported by a grant from Shahid Beheshti University of Medical Sciences.
