Abstract
BACKGROUND:
The long non-coding RNA (lncRNA) DSCAM-AS1 has been demonstrated to participate in the pathogenesis of breast cancer and tamoxifen resistance.
OBJECTIVE:
To evaluate expression profile of DSCAM-AS1 in invasive ductal carcinoma of breast and its suitability as a biomarker for diagnosis of breast cancer.
METHODS:
We evaluated expression of DSCAM-AS1 in 108 breast tissues including tumoral and adjacent non-cancerous tissues (ANCTs) by means of quantitative real time PCR.
RESULTS:
DSCAM-AS1 was up-regulated in tumoral tissues compared with ANCTs (Fold change = 2.86, P = 0.011). Its expression was significantly higher in patients aged less than 55 compared with older patients (P = 0.02). However, its expression levels had not a good performance as a diagnostic biomarker for breast cancer.
CONCLUSIONS:
The significant up-regulation of DSCAM-AS1 in tumoral tissues compared with ANCTs provides further evidences for participation of this lncRNA in the pathogenesis of breast cancer.
Keywords
Introduction
Recent studies have acknowledged the role of long non-coding RNAs (lncRNAs) as regulators of gene expression at different levels of transcription and post-transcription [1]. They participate in many aspects of cellular and molecular processes including but not limited to intracellular trafficking, chromosome remodeling, apoptosis, cell cycle regulation, growth, differentiation, genomic imprinting and alternative splicing [2]. Such diverse modes of function have led to their participation in the pathogenesis of cancer [3]. Several studies have assessed the role of lncRNAs as putative markers in the diagnosis or prognosis of breast cancer. An appropriate marker should have several features including sufficient sensitivity and specificity to facilitate early detection of breast cancer, observation of progression of the disease and prediction of the response to therapeutic options [4]. Although no ideal biomarker has entered clinical practice yet, assessment of expression profile of cancerous cells has led to introducing some potential biomarkers. For instance, elevated expression of the HOX Transcript Antisense RNA (HOTAIR) has been suggested as an independent biomarker for estimation of the possibility of metastasis in patients with estrogen receptor (ER)-positive breast cancer [5]. Moreover, based on the reported association between the expression of lncRNA activated by TGF- 𝛽 (lncRNA-ATB) and trastuzumab resistance, this lncRNA has been proposed as a putative biomarker and therapeutic target for breast cancer invasion and metastasis [6]. In addition, assessment of the Cancer Genome Atlas (TCGA) and the lncRNA profiler has led to identification of a number of lncRNAs with differential expression in breast cancer among them was PVT1 whose expression was inversely correlated with the pathological stage, hormone receptor status and p53 expression in breast cancer [7]. Down Syndrome Cell Adhesion Molecule-antisense RNA 1 (DSCAM-AS1) is among the lncRNAs whose roles in the pathogenesis of certain types of breast cancer have been assessed. The existence of estrogen response elements in this gene along with its up-regulation in human breast cancer samples compared with normal human breast and benign lesions have been noticed previously [8]. In addition, this lncRNA has been recognized as the most ample lncRNAs whose expression is regulated by ER𝛼 in hormone-deprived situations. Moreover, it has a specific expression in ER𝛼+ luminal subtype in association with tumor stage [9]. Through in silico analysis of miTranscriptome database, Miano et al. have reported very low expression of this lncRNA in normal tissues and suggested a cancer-specific signature for it [9] which implies its suitability as a diagnostic biomarker in breast cancer. In the present study, we assessed expression of DSCAM-AS1 in breast cancer samples compared with the paired adjacent non-cancerous tissues (ANCTs) to find it performance in the diagnosis of cancer state in tumoral vs. non-tumoral specimens.
The nucleotide sequences of primers used for expression analysis
The nucleotide sequences of primers used for expression analysis
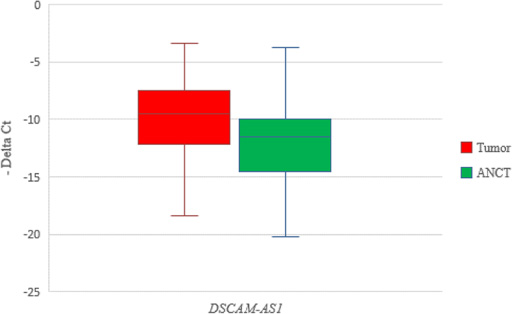
Relative expression of DSCAM-AS1 in tumoral tissues (n = 54) and ANCTs (n = 54) as calculated by –Delta CT values (CT reference gene–CT target gene).
Patients
The current study was performed on tissue samples obtained from 54 patients with invasive ductal carcinoma of breast who underwent mastectomy in Sina and Farmanieh hospitals. Informed consent forms were signed by all patients. The research protocol was approved by the ethical committee of Shahid Beheshti University of Medical Sciences. All methods were performed in accordance with the relevant guidelines and regulations. Tumoral tissues and ANCTs (with a minimum distance of 5 cm from tumor) were excised from all patients before any kind of chemo/radiotherapy. The immediately snap-frozen samples were transferred the medical genetic department and stored in −80 °C. Estrogen receptor (ER) and progesterone receptor (PR) status of samples, tumor grades and other relevant data were retrieved from pathological records. Demographic data were collected through questionnaires.
RNA extraction and quantitative real-time reverse transcription polymerase chain reaction (RT-PCR)
All tumoral and non-tumoral samples were subjected to RNA extraction using the AccuZol TM total RNA extraction solution (Bioneer, Korea) based on the company guidelines except for an additional step of treatment with DNase I. After verification of RNA purity and concentration, one μg of RNA was converted to cDNA using PrimeScript RT reagent kit (Takara Bio, Ohtsu, Japan). Relative expression of DSCAM-AS1 gene was measured in tumoral samples vs. ANCTs in a rotor gene 6000 corbett detection system using SYBR Premix Ex Taq (Takara Bio, Ohtsu, Japan). Thermal cycling program included an activation step at 95 °C for 5 minutes followed by 40 cycles at 95 °C for 15 seconds, 60 °C for 10 seconds and 72 °C for 30 seconds and subsequent melting curve assessment. B2M gene was used as normalizer. Table 1 shows the nucleotide sequences of primers used for expression analysis.
Statistical analysis
Relative expression of DSCAM-AS1 in tumoral tissues compared with ANCTs was calculated using the real-time PCR efficiencies (E) and the threshold cycle (CT) difference in tumoral samples versus ANCTs. The transcript quantities were standardized to the B2M. P values less than 0.05 were regarded as statistically significant. Statistical analyses were performed in SPSSv.18.0.1 (SPSS Inc., Chicago, IL). Chi-square test was used for testing the association between expression of DSCAM-AS1 and categorical variables. The difference between mean values of DSCAM-AS1 transcript levels between distinct categories of patients was assessed using Tukey’s honest significance test. We also plotted the receiver operating characteristic (ROC) curve to assess the diagnostic performance of DSCAM-AS1 transcript levels. The area under the curve (AUC) was used to appraise the accurateness of gene expression levels for discriminating tumoral tissues from ANCTs. The Youden index (j) was applied to get the most difference between sensitivity (true-positive rate) and 1-specificity (false-positive rate).
Results
General statistical information
Demographic and clinical features of study participants have been summarized in Table 2.
General demographic data of study participants
General demographic data of study participants
DSCAM-AS1 was up-regulated in breast cancer samples compared with ANCTs (Fold change = 2.86, P = 0.011). Figure 1 shows relative expression of DSCAM-AS1 in tumoral tissues and ANCTs as calculated by –Delta CT values (CT reference gene–CT target gene).
Associations between DSCAM-AS1 expression and clinical characteristics
We classified patients in down/up-regulation categories according to the relative expression of DSCAM-AS1 gene in tumoral tissue compared with its paired ANCT and evaluated associations between relative expression of DSCAM-AS1 and patients’ demographic/clinicopathological data (Table 3). DSCAM-AS1 relative expression in tumoral tissue vs. ANCT was not associated with any of demographic or clinical characteristics.
Association analysis between relative expressions of DSCAM-AS1 gene in tumoral tissues vs. ANCTs and tumors features (Up/down regulation is defined based on the relative expression of DSCAM-AS1 gene in tumoral tissue compared with its paired ANCT)
Association analysis between relative expressions of DSCAM-AS1 gene in tumoral tissues vs. ANCTs and tumors features (Up/down regulation is defined based on the relative expression of DSCAM-AS1 gene in tumoral tissue compared with its paired ANCT)
Next, we normalized DSCAM-AS1 expression with B2M expression in each tumoral tissue using the (Efficiency ∧CT B2M–Efficiency ∧CT DSCAM-AS1) formula and compared these values across clinicopathological-based categories of breast cancer patients (Table 4). DSCAM-AS1 expression was significantly higher in patients aged less than 55 compared with older patients (P = 0.02). No other significant difference was found in expression of DSCAM-AS1 between other categorizes.
Association between transcript levels of DSCAM-AS1 in tumoral tissues and tumors features (Mean (Standard deviation) values of Efficiency ∧CT B2M–Efficiency ∧CT DSCAM-AS1 are presented)
DSCAM-AS1 expression had 75.9% specificity and 55.6% sensitivity for diagnosis of breast cancer. Table 5 shows the details of ROC curve analysis of DSCAM-AS1 expression for differentiation of disease status between tumoral and non-tumoral tissues.
The results of ROC curve analysis (a: Youden index, b Significance level P (Area = 0.5), Estimate criterion: optimal cut-off point for gene expression)
The results of ROC curve analysis (a: Youden index, b Significance level P (Area = 0.5), Estimate criterion: optimal cut-off point for gene expression)
In the present study, we demonstrated significant up-regulation of DSCAM-AS1 in invasive ductal carcinoma samples compared with the corresponding ANCTs. Although we did not assess the mechanism of DSCAM-AS1 up-regulation in our cohort of patients, gene amplification in the genome seems to be the principle underlying mechanism based on the results of Xu et al. study which analyzed the data from the TCGA, Broad, Sanger and BCCRC databases [10]. Future studies are needed to show whether these amplified regions are presented as double minutes or homogeneously staining regions. We did not find any correlation between its transcript levels and ER/PR status which is consistent with Liu et al. study which reported no significant correlation between its mRNA level and either the existence of ER or the immunocytochemical recognition of the estrogen-responsive PR in a panel of breast cancer samples despite its relation with ER status in breast cancer cell lines which might reflect the intricacy of ER regulation [8]. Miano et al. have reported down-regulation of DSCAM-AS1 following treatment with ER𝛼 siRNA and suggested DSCAM-AS1 as an lncRNA whose expression is regulated by ER𝛼 in an independent manner from estrogen [9]. However, Niknafs et al. demonstrated significant over-expression of DSCAM-AS1 in both MCF7 and T47D cells after estrogen stimulation [11]. Miano et al. also showed significant correlation between DSCAM-AS1 and ER𝛼 expression in both cell lines and tumor samples [9]. We hypothesized that expression of DSCAM-AS1 in clinical samples might be under several regulatory mechanisms which would diminish the observed in vitro effect of ER on DSCAM-AS1 expression in breast cancer cell lines.
We also detected considerable expression of DSCAM-AS1 in ANCTs which contradicts the cancer-specific signature of this lncRNA as suggested previously [9]. We propose expression analysis of this lncRNA in normal breast tissues to assess whether its expression is influenced by the alterations of microenvironment in the histologically normal ANCTs.
Noticeably, we found significant higher expression of DSCAM-AS1 in patients aged less than 55 compared with older patients. This is consistent with Benz’s speculation regarding the less aggressive behavior of late-onset breast cancers compared with early-onset breast cancers even after adjustment of the effects of ER and HER2 status [12]. Besides, Yao et al. have demonstrated the effects of aging on transcriptomes of hormone-dependent breast cancers in a way that specimens with over-expression of a poor outcome-related proliferation profile mainly corresponded to younger patients [13].
Niknafs et al. have previously assessed expression profile of a number of breast cancer samples along with their clinical data and found significant association between DSCAM-AS1 correlated genes and aggressive behavior of tumor, tamoxifen resistance, higher grade, stage and metastasis [14]. Lack of association between expression level of DSCAM-AS1 and clinical features in our study might be explained by relative small sample size and the heterogeneity of samples in our study which consisted of both ER negative and ER positive samples. Based on the ER-regulated nature of DSCAM-AS1, Niknafs et al. have evaluated its significance in breast cancer pathogenesis only in ER-positive breast cancers [11].
Finally, despite the observed significant over-expression of DSCAM-AS1 in tumoral tissues compared with ANCTs, the performance of its transcript levels for diagnosis of breast cancer was poor. Further studies are needed to assess its performance in larger sample sizes of cancer patients in combination with transcript levels of other lncRNAs to find a panel of transcripts with a putative role as biomarkers.
In conclusion, the significant up-regulation of DSCAM-AS1 in tumoral tissues compared with ANCTs provides additional evidences for the participation of this lncRNA in the pathogenesis of breast cancer and warrants further assessment of its role in a larger cohort of patients.
Footnotes
Acknowledgements
The present study has been supported by a grant from Shahid Beheshti University of Medical Sciences.
Conflict of interest
None.
