Abstract
BACKGROUND:
Nitric oxide (NO) may have a dual role in cancer. At low concentrations, endogenous NO promotes tumor growth and proliferation. However, at very high concentrations, it mediates cancer cell apoptosis and inhibits cancer growth. High levels of NO have been observed in blood of breast cancer (BC) patients, which increases tumor blood flow and promotes angiogenesis. To date, the regulation of NO-synthesizing enzyme, eNOS, by miRNAs has not been adequately investigated in BC. Therefore, the main aim of this study is to unravel the possible regulation of eNOS by miRNAs in BC and to examine their influence on NO production and BC progression.
METHODS:
Expression profile of eNOS in Egyptian BC patients and MDA-MB-231 cell lines was investigated using qRT-PCR. In-silico analysis was performed to predict a putative upstream regulator of eNOS. miR-744-5p was selected and its expression was quantified in BC tissues using qRT-PCR. MDA-MB-231 cells were cultured and transfected with miR-744-5p using lipofection method. NO levels were determined using Griess Reagent. Cellular viability and colony-forming ability were assessed using MTT and colony-forming assays; respectively.
RESULTS:
eNOS and miR-744-5p were significantly up-regulated in BC tissues compared to paired normal tissues. In-silico analysis revealed that miR-744-5p putatively binds to eNOS transcript with high binding scores. Transfection of MDA-MB-231 cells with miR-744-5p mimics resulted in a significant up-regulation of eNOS and consequently NO levels. In addition, miR-744-5p transfection led to an increase in cellular viability and colony-forming ability of the MDA-MB-231.
CONCLUSION:
miR-744-5p acts as an upstream positive regulator of the NO synthesizing enzyme, eNOS which in turn elevates NO levels. Furthermore, miR-744-5p is a novel oncogenic miRNA in BC. Thus, targeting miR-744/eNOS/NO axis may act as a therapeutic tool in TNBC.
Introduction
Nitric oxide (NO) acts as a signaling molecule in many different physiological and pathological processes. However, elevated NO synthesis has been involved in many pathophysiological conditions including cancer [1]. NO possesses controversial effects on cellular viability in a dose dependent manner [2,3]. At low concentrations, NO promotes tumor growth by modulating many cancer related events such as angiogenesis, immunosuppression, mutagenicity, cell cycle arrest, invasion, and metastasis [4,5]. In specific, NO plays a variety of roles in various stages of carcinogenesis by damaging the DNA, activating oncogenes, inhibiting apoptosis and metastasis, and inhibiting enzymes for DNA repair and tumor suppressor genes [6]. Whereas, at high concentrations, NO has an anti-neoplastic function by inducing tumor cytostasis, cytotoxicity, reducing tumor blood flow, and apoptosis [7]. The threshold of NO concentration between tumor-promoting and tumor-suppressing effects is still unclear.
eNOS (or NOS3) gene, encoding endothelial nitric oxide synthase, is located on chromosome 7 (7q35-36) in humans. eNOS gene is composed of 26 exons, ranging from 68 to 579 bp and spans 22 kb and encodes a protein of 1203 amino acids [8]. Its core promoter region contains multiple cis-acting elements, including high-affinity binding sites of transcription factors such as specificity protein 1 (Sp1). As an important nuclear transcription factor, Sp1 induces endothelial cell proliferation by enhancing the transcription activity of eNOS gene [3]. The expression level and activity of eNOS enzyme is increased in different malignancies compared with corresponding normal tissues [3]. Thus, eNOS may play an important role in tumor progression via angiogenesis or apoptosis [3].
NO has been strongly implicated in BC. The malignant transformation process occur in the mammary gland has been reported to be correlated with VNTR 4 a/b polymorphism that regulate NOS3 expression [9]. It was reported that both NOS2 and NOS3 expression levels, enzymatic activity and NO levels are significantly high in invasive breast carcinoma tissues when compared to benign and normal tissues [10]. Furthermore, within the IDC BC tissues, it was shown that NO biosynthesis was significantly correlated with tumor grade and aggressiveness of the disease [10,11]. However, up to our knowledge this has not been validated in the Egyptian BC patients. Furthermore, NO biosynthesis was significantly greater in grade III compared with grade II tumors. Tumor-derived eNOS has shown to modulate cancer-related events (inflammation, apoptosis, cell cycle, angiogenesis, invasion, and metastasis) and genetic studies showed that eNOS gene polymorphisms are associated with the development of multiple cancers [2,10,11].
The eNOS expression is regulated by a range of transcriptional and post-translational mechanisms. Among the transcriptional mechanisms is the avid binding of several transcription factors namely Sp1, Sp3, Ets-1, Elf-1, and YY1 to the eNOS promoter region. Post-transcriptionally, eNOS activity is controlled by alterations of the primary transcript, stability of mRNA, its subcellular localization, and intracellular transport. Post-translational modifications of eNOS comprise fatty acid acylation, substrate, coenzymes and cofactors availability, protein-protein interactions, and degree of phosphorylation. Another significant epigenetic mechanism for eNOS gene expression is differential promoter methylation. MicroRNAs are among the key post-transcriptional regulators of eNOS gene expression [12].
MicroRNAs (miRNAs) are defined as a class of approximately 18-22 nucleotide long non-coding, single stranded RNA [13]. They are widely involved in the post-transcriptional regulation of gene expression in both animals and plants by causing translation repression or degradation of mRNA targets [14]. Around sixty percent of all protein coding genes are controlled by miRNAs [14]. The majority of miRNAs encoding genes are located at introns of protein-coding or non-coding genes, although a few miRNAs are encoded by exons of non-coding genes [15,16]. Since, miRNAs are involved in the regulation of almost every cellular process including cell proliferation, apoptosis, cell development and differentiation, it is not surprising that the deregulation of miRNA expression is frequently associated with carcinogenesis [2,13,17–21]. Some miRNAs are overexpressed in cancer and they function as oncogenic miRNA (oncomiRs) that promote cancer development and progression by negatively regulating tumor suppressor genes and/or genes that control cell differentiation or apoptosis. Other miRNAs are described as tumor suppressor. They were found to be under-expressed in several malignancies [13,20,21]. Deregulation of miRNA expression is common in BC [2,17].
Considering the recent interest in the development of novel mi-RNA-based therapies for human pathologies, it is tempting to speculate the role of miRNAs in regulating NO production in cancer. Therefore, the major aims of this study were to investigate the expression profile of eNOS in Egyptian BC patients and MDA-MB-231 cell line (Triple–Negative Breast Cancer cell line, TNBC), to identify potential miRNA targeting eNOS gene using in-silico analysis, to investigate the expression level of miR-744-5p, a potential regulator of eNOS gene, in BC tissues, to investigate the potential regulating role of miR-744-5p on eNOS expression and NO production level, and to uncover the underlying role of miR-744-5p in BC progression.
Subjects, materials & methods
Subjects
Tumor breast cancer samples were obtained from International Medical Centre and Kasr Al-Ainy hospital, Cairo, Egypt. A total of 36 breast tissue samples and their adjacent normal tissues served as control were collected from Egyptian BC female patients undergoing tumor resection surgery. Age ranges for patients were 28 to 68 years with a mean age of 49 years. Tissues were kept fully labeled in 1.8 ml cryo-vials in −80 °C freezer for optimal storage. Pathological examination was performed to assess tumor grade and stage and the tissues were immunohistochemically stained to analyze the expression of the receptors (ER, PR and HER2) to define the molecular subtypes. Characteristic features and clinical parameters of BC patients are shown in Table 1.
Cell culture and transfection
MDA-MB-231 TNBC cell line was purchased from Vacsera - Holding Company for Biological Products and Vaccines (Cairo, Egypt). MDA-MB231 cells were cultured in DMEM (LONZA, Germany) supplemented with 4.5 g/L Glucose, L-Glutamine, 1% penicillin-Streptomycin-Mycozap and 10% FBS. Cells were cultured in culture dishes and incubated under their normal growth conditions (37 °C and 5% CO2) as previously described in [2].
Lipofection technique was employed using HiPerFect Transfection Reagent (Qiagen) to transfect MDA-MB-231 cells with miR-744-5p mimics (Qiagen, Germany), per the manufacturer’s protocol. Mock untransfected cells were exposed to transfection reagent only. This control was used to determine any nonspecific effects that may be caused by the transfection reagent or process, and thus normalize the results of the mimicked well with it as previously described in [2,12].
Characteristic features and clinical parameters of BC patients
Characteristic features and clinical parameters of BC patients
Total RNA, containing both large and small RNAs, was extracted from frozen breast cancer tissues and MDA-MB-231 cells using mirVanaTM microRNA Isolation Kit (Ambion, USA) and Biozol extraction Reagent (Bioer Technology, China), respectively, per the manufacturer’s protocol.
Extracted RNA was then reversed transcribed into complementary DNA (cDNA) using the high capacity cDNA reverse transcription kit (Applied Biosystems, USA). cDNA was amplified and quantified using TaqMan quantitative Real-time polymerase chain reaction (Q-RT PCR) (Applied Biosystems). The housekeeping genes, RNU6B and the endogenous control Beta-actin (β-actin) were used to normalize the relative expression of miR-744 and the target gene, eNOS, in each sample, respectively. StepOne® Real Time PCR instrument was used for detecting the cycle threshold (Ct) values (Applied Biosystems, USA). The 2−ΔΔCt method was used to calculate the relative gene expression as previously described in [2,18].
Bioinformatics analyses
In silico analysis was used to detect the possible miRNAs that may potentially be targeting 3’ UTR of the eNOS gene using microrna.org (www.microrna.org) and Target Scan (http://www.targetscan.org/). The results of these bioinformatics analyses were examined, and based on multiple criteria including miRNA-mRNA complementarity strength, binding score, hybridization energy, and number of predicted binding sites within the transcript 3’UTR, the candidate miRNA was selected. High negative hybridization energy indicates favorable binding.
Nitric oxide (NO) detection assay (griess reagent system)
The Griess Reagent system (Promega, USA) was used for NO measurement in BC cells. Total nitrate/nitrite concentration was detected in a simple two-steps process. Briefly, nitrite standard solution was prepared to plot the standard reference curve. Fifty μl of each experimental sample was added to the respective wells in triplicates. Using a multichannel pipettor, 50 μl of the sulfanilamide solution was added to all experimental samples and the serial dilutions of standard nitrite. The plates were covered with aluminum foil; then it was left to incubate for 5–10 minutes at room temperature. 50 μl of the NED solution was then dispensed to all wells. The plate was covered with aluminum foil; then it was incubated again for 5–10 min at room temperature. The absorbance was measured at 550 nm using a multi–microplate reader Perkin Elmer Wallac 1420 VICTOR2TM (Perkin Elmer, USA) as previously described in [2,12].
MTT assay for cytotoxicity
The MTT (3-(4, 5-Dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide) colorimetric assay is used for assessing cellular viability by measuring the activity of mitochondrial enzymes that reduce yellow MTT dye to a purple formazan compound. Conversion can be directly related to the number of viable cells. Colorimetric measurements and absorbance were performed using Wallac 1420 Victor2 Multilabel Counter (Perkin Elmer, USA) as previously described [22]. MTT was added to MDA-MB-231 cells 48h post-transfection with miR-744-5p oligonucleotides. This was followed by incubation for 4 hours and adding ethanol/DMSO solution for cell lysis. Optical density was measured at 572 nm and relative viability was calculated as previously described in [13,18,21].
Colony-forming (Clonogenicity) assay
Clonogenic assays serve as a useful tool to test the cell proliferation ability of a single cell to form a large colony or a clone. Briefly, MDA-MB231 cells were seeded allowed to achieve 60–80% confluency and were then transfected with miRNA oligonucleotides. A set of untransfected mock cells were also prepared. The culture media was replaced. Then the cells were detached by gentle resuspension by pipetting to form a homogenous cell suspension. Thirty μl cell suspension was transferred into its assigned well in the 6 well plates, and then 2 ml full DMEM was added to each well and incubated for 1–2 weeks under normal growth conditions (37 °C and 5% CO2). The plate was checked frequently during this incubation period. Colony formation was observed after 2 weeks incubation. After 2 weeks, the old media was discarded and 1 ml (6% Glutaldehyde) fixation solution was added to each well and it was kept on the bench for 30–40 minutes to fix the colonies. In the next step, 500 μl crystal violet was added and the plate was left for 1 hour to stain the colonies. After 1 hour, the colonies were washed with normal water two times, then colonies were counted using Zen 2012 software and the number of colonies was compared to those of controls. Colony forming assay experiment were done in triplicate and average of colonies number was taken for mimics and mock cells [13,18,21].
Statistical analysis
Gene expression is expressed in Relative quantitation (RQ = 2−ΔΔCt). For the purpose of comparison between two different studied groups, Unpaired Student’s t test (parametric and two tailed) was employed. Data were expressed as mean ± standard error of the mean (SEM). All experiments were performed in triplicates and repeated at least three times, unless otherwise stated. A p-value less than 0.05 were considered statistically significant **** = P < 0.0001, *** = P < 0.001, ** = P < 0.01, * = P < 0.05 and ns = statistically not significant. All the data were statistically analyzed using GraphPad Prism 7.02 software.
Results
Bioinformatic analysis
Using TargetScan (https://www.targetscan.org) and miRanda software (https://www.microRNA.org), it was found that miR-744-5p potentially targets 3’UTR of eNOS mRNA (Fig. 1).
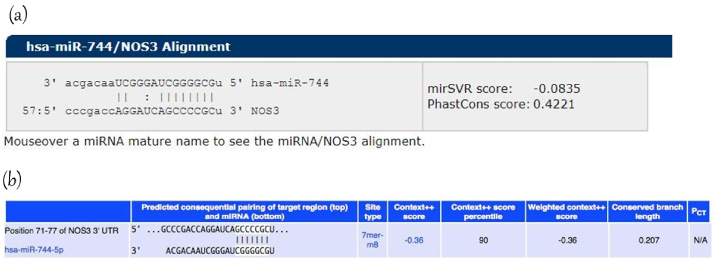
Bioinformatic analysis. Predicted target region-seed sequence binding for miR-744-5p on the 3’UTR of eNOS. (a) Target Scan (b) MiRanda.
The expression profile of the endogenous eNOS mRNA and the expression level of miR-744-5p was analyzed in 36 BC tissues and their paired control tissues. A statistically significant over-expression in eNOS mRNA expression was found in BC tissues up to 16 fold higher than control tissues (p < 0.0001) (Fig. 2). Similarly, miRNA-744-5p expression was significantly upregulated in BC patients (p = 0.0004) up to 10 fold higher than paired control tissues (Fig. 2).
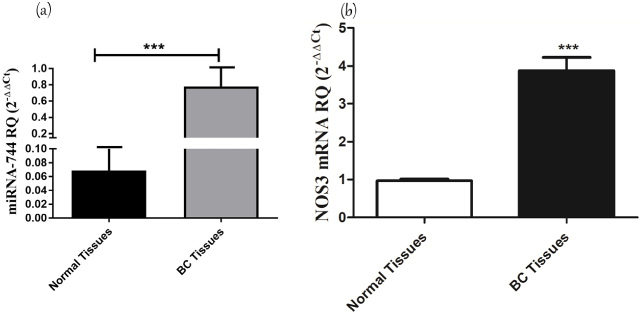
Screening of miR-744 and eNOS in BC patinets. Relative expression level of eNOS (NOS3) mRNA and miR-744 in BC tissues compared to control tissues. The expression of eNOS mRNA was screened in BC tissues and controls and normalized in each sample to β-actin housekeeping endogenous gene as a control. Expression of eNOS was significantly higher in BC compared to adjacent control cells (p = <0.0001). miRNA-744 was screened using qRT-PCR and normalized to RNU6B as an internal control. Screening of miR-744-5p showed a significant upregulation in BC patients compared to control tissues (P = 0.0004).
MDA-MB 231 cells were transfected with oligonucleotide mimics of miR-744-5p. To confirm good transfection efficiency, the change in expression of miR-744-5p, relative to mock cells treated with transfection reagent alone, was determined by qRT-PCR normalized to RNU6B endogenous control. A significant upregulation of miR-744-5p expression up to 1000 folds was observed in miR-744-5p mimicked MDA-MB-231 cells compared to mock cells (p = 0.002) (Fig. 3a). Ectopic expression of miR-744-5p in MDA-MB-231 cells resulted in up-regulation of eNOS mRNA levels up to 2 folds higher than mock cells (p = 0.0068) (Fig. 3b). Next, NO concentration in supernatant of MDA-MB-231 transfected with miR-744-5p mimics was measured relative to mock cells. Results showed an increase in NO production compared to mock cells (p = 0.0143) (Fig. 3c).
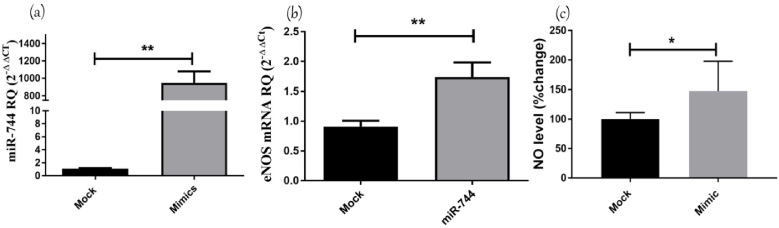
Impact of miR-744 on eNOS/NO axis. The expression of miR-744-5p was determined by TaqMan RT qPCR in MDA-MB-231 cells transfected with miR-744-5p oligonucleotides, relative to their expression in mock MDA-MB231 cells. The expression of miR-744-5p was normalized to RNU6B endogenous control. Transfection of miR-744-5p mimics increased miR-744-5p levels in MDA-MB231 up 1000 folds compared to mock cells (p = 0.002). The relative expression of eNOS in MDA-MB231 transfected with oligonucleotide mimics of miR-744-5p, was determined using TaqMan RT qPCR, relative to mock cells, and gene expression was normalized to endogenous control. eNOS mRNA expression was significantly up-regulated upon mimicking of miR-744-5p compared to mock cells (p = 0.0068). The relative NO level in supernatant of MDA-MB231 transfected with oligonucleotide mimics of miR-744-5p, was increased compared to mock cells (p = 0.0143).
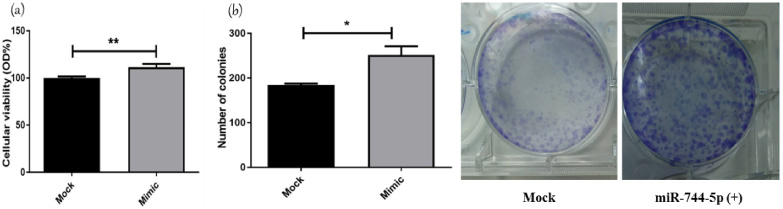
Impact of miR-744-5p on MDA-MB-231 hallmarks. Cellular viability was normalized to mock cells viability of 100%. miR-744-5p mimicking significantly increased cellular viability compared to mock cells (p = 0.0063). Transfection with mimics of miR-744-5p led to a significant increase in colonogenicity compared to mock cells (p = 0.0281).
Several functional analysis experiments were performed to evaluate the overall effect of miR-744-5p on various characteristics of tumor cells. MTT assay was conducted on MDA-MB-231 cells transfected with miR-744 mimics. Transfection with mimics of miR-744-5p led to a substantial increase in cellular viability compared to mock untransfected cells (p = 0.0063) (Fig. 4a). Second functional analysis experiment was conducted to investigate the effect of miR-744-5p on MDA-MB-231 cell line to form a colony that reflects cellular proliferation capacity. Transfection with mimics of miR-744-5p led to a significant increase in clonogenicity compared to mock cells (p = 0.0281) (Fig. 4b).
Discussion
BC is a complex and heterogenic disease associated with genetic and epigenetic alterations. Understanding these alterations will advance the ability to effectively treat BC. Drug therapies directed towards epigenetic changes represent the road for more personalized treatment of BC patients [23]. One of the most powerful epigenetic weapons is the miRNAs, for their high ability to modulate the expression of their target genes. Recently, many studies have focused on the characterization of several miRNAs in different malignancies through their post transcriptional regulatory effect [2,13,17–21].
In Egypt, BC has a high mortality rate due to the lack of awareness, late diagnosis and detection of the disease at an advanced and invasive stage. Thus, the discovery of novel targeted therapeutic approaches directed at epigenetic changes; such as miRNAs has been gaining much attention especially for the treatment of BC [24]. Gasotransmitters and mainly NO play a functional role in several cancers including BC [12]. NO is an endogenously produced radical gas with regulatory roles in many biological events under physiological and pathological conditions [12]. Low concentrations of NO promote tumor growth by genotoxicity, anti-apoptotic effect and angiogenesis. On the other hand, very high concentrations of NO play a tumoricidal effect by stimulation of apoptosis, attenuation of angiogenesis and inactivation of survival [3]. Unfortunately these high tumoricidal concentration levels are barely achieved by the cancer cells [5]. NO levels were found to be elevated in BC patients together with eNOS enzyme [12]. This gives our study an edge of taking the initiative to unravel the mechanisms by which NO levels together with eNOS enzyme were up-regulated in BC, specifically through non-coding RNAs (ncRNAs).
Different algorithms to find miRNA-seeding sites in the 3 ′ -UTR of the eNOS gene were used. miR-744 was among the top ranked miRNAs that modulated eNOS expression. miR-744 was initially identified in 2006 [25]. Its encoding gene is located on chromosome 17p12. It has been reported that miR-744 may act in a cancer specific manner where it can act as tumor suppressor or oncogenic miRNA in a different cellular context [26–28]. Studies have shown that miR-744 functions as oncogenic miRNA in prostate cancer [29] and laryngeal squamous cell carcinoma [30]. However, it functions as tumor suppressor miRNA in colorectal cancer [31], hepatocellular carcinoma [26], and cervical cancer [32,33]. In the literature review, we could not find studies that define the correlation between miRNA-744 and the eNOS expression pattern in BC. Thus, the correlation between miRNA-744-5p and eNOS expression was examined in both BC patients and MDA-MB-231 cell line.
Results showed that miRNA-744 expression was significantly high in BC tissues. Ectopic expression of miRNA-744 in MDA-MB-231 cells enhanced the expression of eNOS and elevated NO levels in the TNBC cells. miRNA-744 was also found to act as oncogenic miRNA by increasing the cellular viability and colony forming ability of the TNBC cells. Therefore, targeting oncogenic signaling pathway “miRNA-744/eNOS/NO” would have very promising results in TNBC. Earlier studies showed controversial results regarding the oncogenic properties of miRNA-744 in cancer. A study by Miyamae et al. showed that miR-744 is highly expressed in patients with pancreatic cancer and cell line when compared to normal tissues, appointing miR-744 oncogenic activity [34]. miR-744 enhances cell proliferation, migration and invasion and inhibits apoptosis, thereby promoting tumor growth through activation of Wnt/B-catenin signaling pathway [35]. On the other hand, Chen and Liu provided evidence that miR-744 functions as tumor suppressor in cervical cancer through targeting Bcl-2 expression. Bcl-2 signaling pathway plays an important role in apoptosis, and proliferation [33]. Another study showed that miR-744 acts as a tumor suppressor miRNA in BC by directly targeting the oncogene eukaryotic translation elongation factor 1A2 (eEF1A2) mRNA and decreasing its corresponding protein eEF1A2, the key component of the translation elongation apparatus. This resulted in the inhibition of MCF7 proliferation [36]. These conflicting findings suggest that the expression pattern and influence of miR-744 has cell and tissue specificity.
Several studies investigated the roles of eNOS in the development and progression of malignancy. A former study showed that many of the cancer-related events in BC, such as inflammation, apoptosis, cell cycle, invasion and metastasis, is modulated by eNOS. The study also assessed the role of the eNOS genetic polymorphism in the BC development [9]. NO was found to be a key regulator for development and progression of cancer through endothelial migration, growth, vascular remodeling and angiogenesis [2]. Only few miRNAs that directly affect eNOS expression have been identified, among those are miR-214 and miR-155. Overexpression of miR-155 and miR-214 decreased eNOS expression and NO production in human umbilical vein endothelial cells [37–39]. Knockdown of miR-155 prevented cytokine induced downregulation of eNOS expression, reduction of NO production, and impairment of endothelium-dependent vascular relaxation [39].
In conclusion, this study revealed that miR-744-5p acts as an upstream positive regulator of eNOS, which in turn elevates NO levels. Furthermore, miR-744-5p is a novel oncomiR in BC. Thus, targeting miR-744/eNOS/NO axis could lead to the development of new approaches and strategies for the effective treatment of BC.
Footnotes
Acknowledgements
Not applicable.
Statement of ethics
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Conflicts of interest
The authors declare that they have no conflict of interest.
Funding
This study was not supported by any grants.
Authors’ contributions
Elkilany F. has performed most of the experimental work and analyzed the data, Youness RA. performed some of the experiments, wrote, edited and revised the manuscript. Assal R. analyzed the data, revised the manuscript and Gad M. wrote, edited and revised the manuscript and supervised the whole work flow. All authors have read and accepted the manuscript in its final submitted form.
