Abstract
Schizophrenia research on postmortem human tissue has matured in the past decade, largely due to two factors: (i) the widespread availability of larger cohorts that are more extensively screened; and (ii) the readily available quantitative molecular tools especially for gene expression studies. In the past, lack of replication of basic biological findings in brains of people with schizophrenia across laboratories hampered progress. The renewed research effort on postmortem brains is finally yielding numerous replicable findings across independent laboratories and independent cohorts, which is a critical advance for postmortem brain research on schizophrenia. Studies of postmortem human brain provide a valuable avenue for elucidating the underlying cellular and molecular mechanisms of schizophrenia that are not accessible through live imaging of people or through animal models. Thus, there is a need for direct studying of postmortem human brains from people with schizophrenia, although collecting and characterizing human brains is methodologically challenging [1–3], and standards regarding inclusion criteria necessary for rigorous postmortem research are not universally agreed upon and continue to change [4–11]. Numerous premortem, antemortem and postmortem factors can influence the quality of human brain tissue and need to be considered, and the relative importance of confounding factors for specific molecules are often not known.
Despite these limitations, postmortem human studies of psychiatric illnesses such as schizophrenia remain an extremely valuable tool because they have already begun to shed light on a number of molecular mechanisms that contribute to the aetiology and development of schizophrenia [12–17]. The establishment of multiple large cohorts of well-characterized brains of people with schizophrenia and controls from around the world contribute to our understanding of the biological basis for schizophrenia and will be critical to advancing the field. Therefore, given our experience, and with access to a fairly large collection of postmortem tissue samples, we set out to establish a new cohort of schizophrenia and control brains from Sydney, Australia.
The assembly and characterization of this cohort has been driven by the Schizophrenia Research Institute (SRI), a virtual institute supporting the collaboration of Australian scientists and clinicians who bring a variety of knowledge, skills and techniques together to find ways to prevent and cure schizophrenia [18]. In 2007, through the Institute's Developmental Neurobiology Panel, SRI commenced a process of actively facilitating the development of a large-scale collaborative research programme based on the use of a common collection of postmortem brains for schizophrenia research from the Tissue Resource Centre at the University of Sydney. The results from this multifaceted investigation will be collected and collated, thereby allowing for testing of relationships among a variety of measures. This report represents the first step in this collaborative process whereby the collection and quality control processes for this new SRI tissue cohort are described and the selection of reference genes is considered.
Methods
Subjects
All research was approved and conducted under the guidelines of the Human Research Ethics Committee at the University of New South Wales (HREC 07261). The study cohort included 37 schizophrenia/ schizoaffective disorder patients and 37 matched controls (Table 1). Controls were selected prior to RNA extraction based on the following parameters (in descending order of importance for matching), brain pH (±1), age at death (within 10 years), and postmortem interval (PMI; within 10 h). Other factors such as hemisphere, gender, time in freezer and agonal state were matched on a stratum (groupwise) basis. Details of brain collection and determination of clinical and tissue factors can be found in Supplementary Materials and Methods, and in Tables 1S, 2S. Supplementary material to be found online, at http//www.informaworld.com/10.1080/00048670903393662.
Subject characteristics
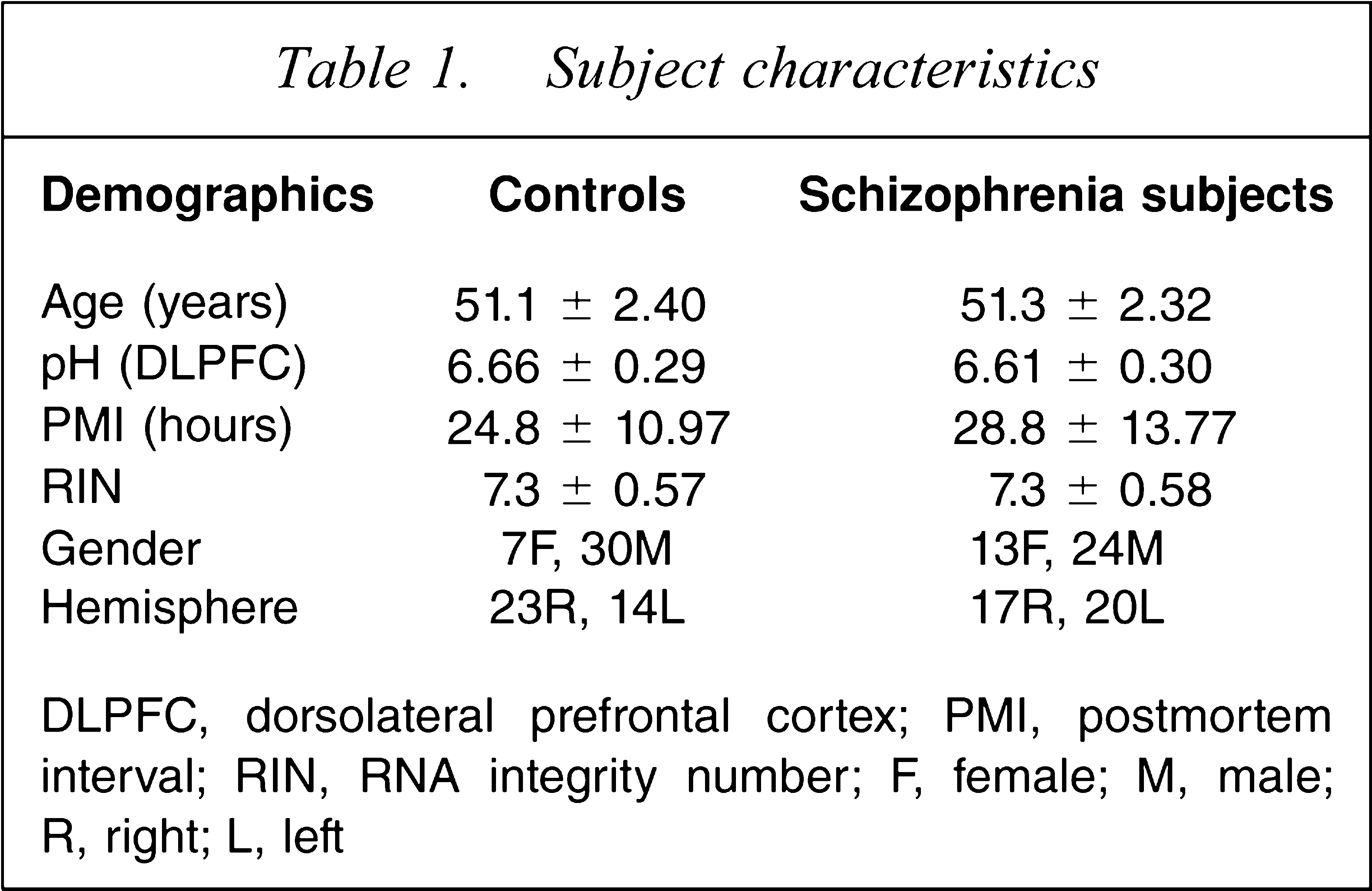
DLPFC, dorsolateral prefrontal cortex; PMI, postmortem interval; RIN, RNA integrity number; F, female; M, male; R, right; L, left
Detailed cohort demographic information.
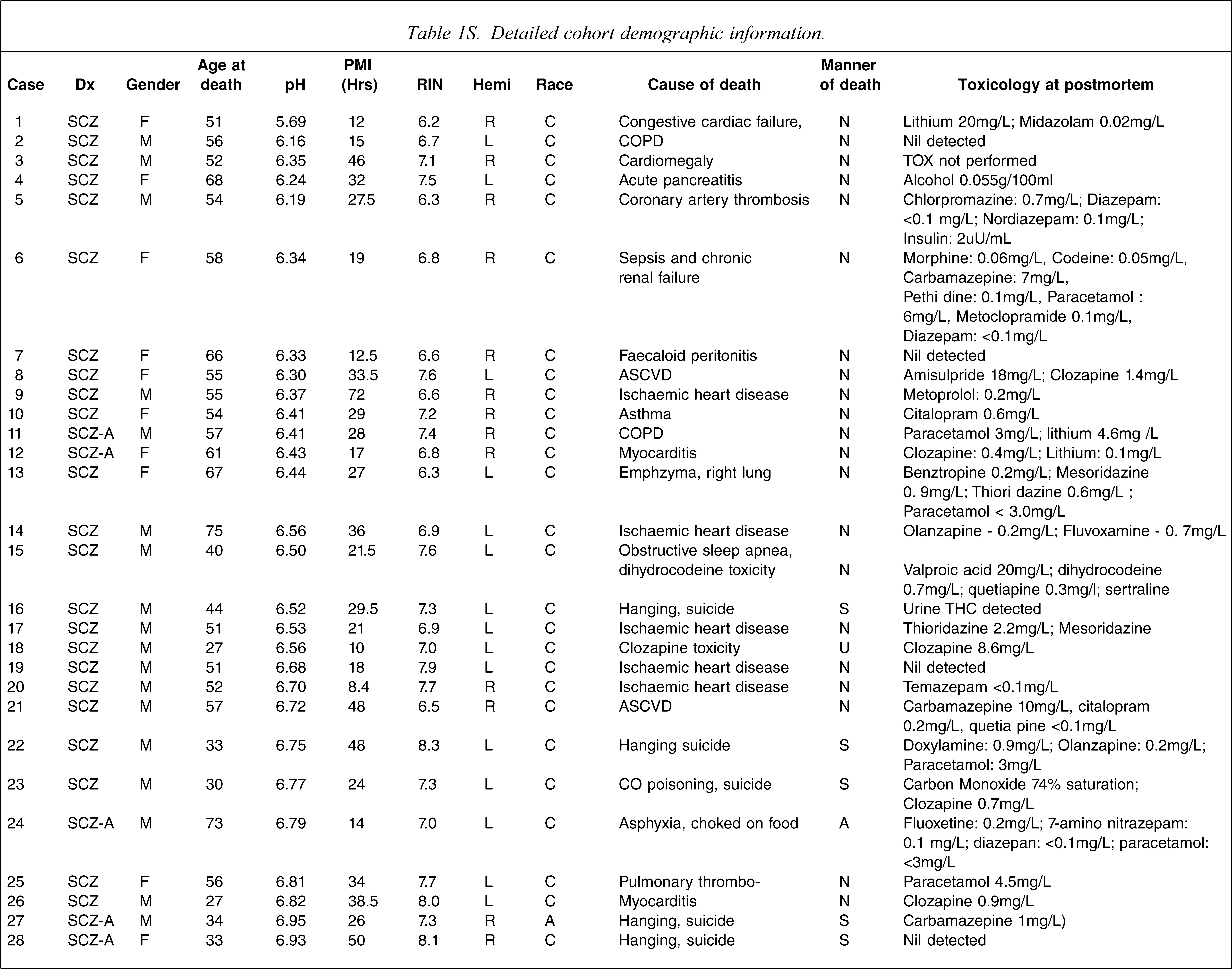
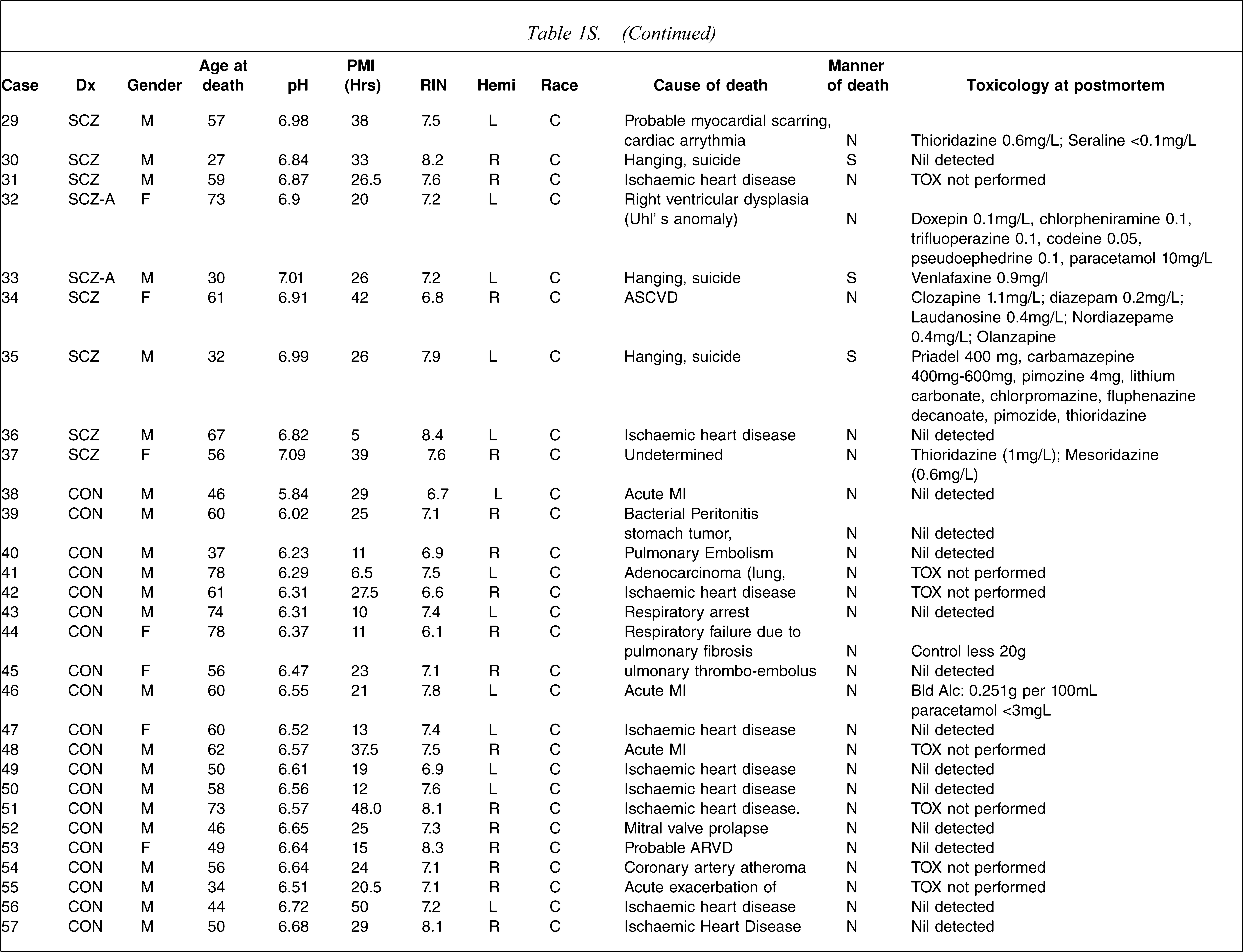
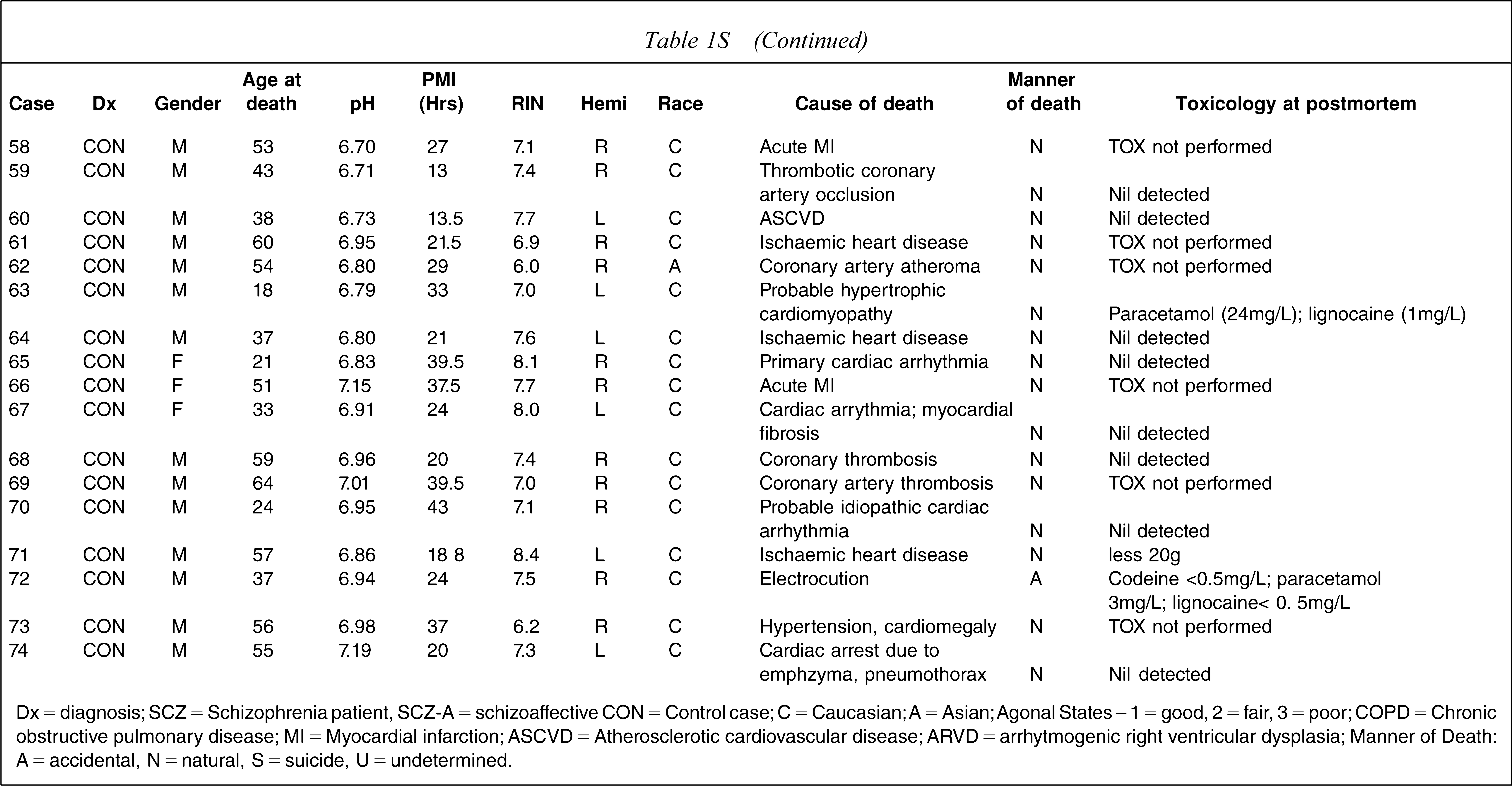
Dx = diagnosis; SCZ = Schizophrenia patient, SCZ-A = schizoaffective CON = Control case; C = Caucasian; A = Asian; Agonal States-1 = good, 2 = fair, 3 = poor; COPD = Chronic obstructive pulmonary disease; Ml = Myocardial infarction; ASCVD = Atherosclerotic cardiovascular disease; ARVD = arrhytmogenic right ventricular dysplasia; Manner of Death: A = accidental, N = natural, S = suicide, U = undetermined.
Clinical characteristics of schizophrenia cohort
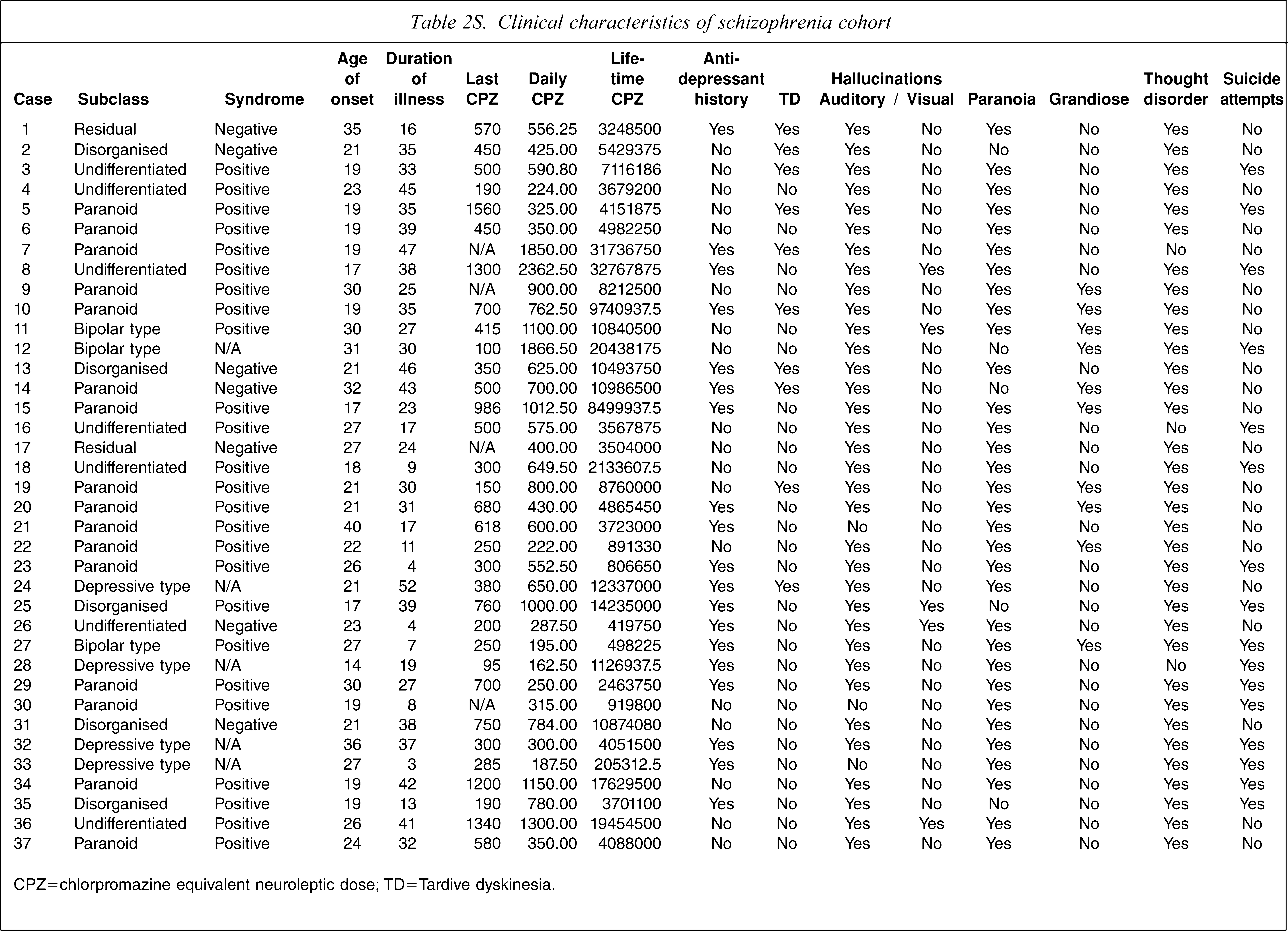
CPZ=chlorpromazine equivalent neuroleptic dose; TD=Tardive dyskinesia.
RNA extraction and quality assessment
Grey matter tissue from dorsolateral prefrontal cortex (DLPFC) was pulverized over dry ice. Approximately 300 mg was weighed while frozen and stored at −80°C until extraction. Total RNA was extracted with TRIzol Reagent (Cat. No. 15596-018; Invitrogen Life Science, Melbourne, Vic., Australia), using a polytron as previously described [19]. Total RNA pellets were resuspended in DEPC-treated water (Cat. No. 95284-100ML; Sigma-Aldrich, Castle Hill, NSW, Australia). The yield of total RNA was then analysed using a spectrophotometer (Nanodrop ND-1000; Thermo Scientific, Wilmington, DE, USA). Total RNA was not further purified through a column. The quality of extracted total RNA was determined on high-resolution capillary electrophoresis using an Agilent Bioanalyzer 2100 (Agilent Technologies, Palo Alto, CA, USA). Approximately 100–200 ng RNA was applied to an RNA 6000 Nano LabChip (Ajilent Technologies, Waldbronn, Germany), without heating prior to loading, and subsequent steps performed according to the manufacturer's protocol. Two internal controls (samples with established high-quality RNA integrity number (RIN) = 10) were loaded on each chip, and all samples analysed on the same day. The RIN was calculated using an algorithm incorporating information from the entire electrophoretic trace and used as an indicator of RNA quality, ranging from 1 (lowest quality) to 10 (highest quality) [20]. Three separate aliquots of total RNA were used to synthesize cDNA from 3 μg of total RNA in a 26.25 μL reaction using the SuperScript First-Strand Synthesis kit (Invitrogen, Carlsbad, CA, USA) and were pooled to reduce any potential variability in cDNA reactions prior to diluting for reverse transcription–polymerase chain reaction (RT-PCR).
Quantitative reverse transcription-polymerase chain reaction
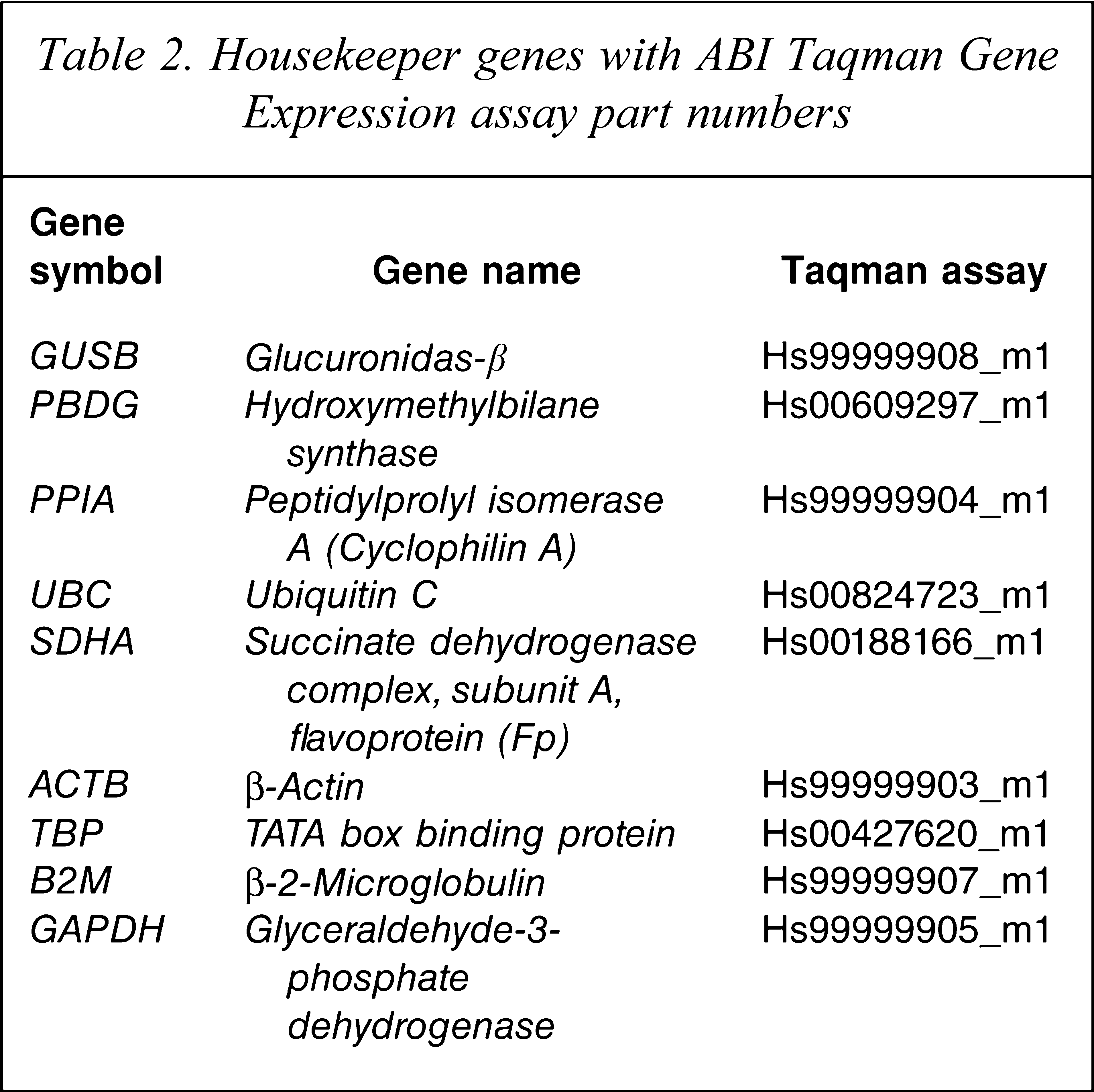
Transcript levels for nine housekeeper genes – glucuronidase-β (GUSB), cyclophilin A (PPIA), ubiquitin C (UBC), porphobilinogen deaminase (PBGD), succinate dehydrogenase complex subunit A (SDHA), β-actin (ACTB), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), TATA box binding protein (TBP) and β-2-microglobulin (B2M) – were measured on quantitative real-time PCR (qPCR) in each cohort sample using an ABI Prism 7900HT Fast Real Time PCR system with a 384-well format (Applied Biosystems, Foster City, CA, USA) and TaqMan Gene Expression Assays (Applied Biosystems, Foster City, CA, USA; Table 2). Samples were run with a seven-point standard curve using serial dilutions of pooled cDNA derived from a representative sample of subjects (three controls and three patients). The no-template control did not produce a signal in any assay. All amplifications from each subject were performed in triplicate and relative quantities were determined from the standard curve. Outliers due to measurement errors were omitted if the percent variance of the triplicates was >30% of the quantity mean value as previously described [21–23] and the mean re-calculated based on two values (this occurred in <5% of the samples). Stability of mRNA expression was measured using the program geNorm VBA applet for Microsoft Excel version 3.5, developed by Vandesompele et al., in which more stable genes will have a lower M [24].
Housekeeper genes with ABI Taqman Gene Expression assay part numbers
Statistical analysis
Statistical analyses were conducted using Statistica version 7 (StatSoft, Tulsa, OK, USA). After measurement errors were removed, population outliers were determined and samples >2SD from the mean for each group were removed (on average 1–2 subjects per group). We also defined population outliers using the Grubb's test, which yielded similar results. Both paired and non-paired Student's t-tests were used to determine if the diagnostic groups differed on individual housekeepers or geomean. To maximize power, the analyses were carried out with the schizophrenia patients combined with the schizoaffective patients as a single diagnostic category, but it is recognized that other analyses focusing only on the schizophrenia group are also planned. Correlations were run on continuous variables to determine if any of the demographic factors (age, pH, PMI or RIN) related to housekeeper mRNA levels or their geometric mean. If this was the case, analysis of covariance (ANCOVA) was used. Independent t-tests were conducted to examine whether gender or hemisphere were related to housekeeper gene expression. Factorial ANOVA was used to determine the effect of gender or hemisphere on gene expression in the diagnostic groups. Statistical significance was set at p ≤ 0.05. Significance level is reported as p rounded to three significant figures.
Results
Matching of diagnostic groups
The patients with schizophrenia and schizoaffective disorder (n = 37) were on average 51 years old, with a brain pH of 6.61 and a PMI of 28.8 hours (Table 1). The normal control subjects selected to match these patients (n = 37) had an average age of 51 years, a brain pH of 6.66 and a PMI of 24.8 hours. The mean RINs were 7.3 for both groups and had a low coefficient of variation (between 7.8 and 8.0%). As anticipated age, pH, PMI, freezer storage time, and RIN did not differ significantly between the groups (all t < 1.3, all p > 0.210). Freezer storage time of brain tissue for patients averaged 81 months and did not differ statistically as compared to normal controls at 70 months. The number of men was higher than women in both groups, although the number of men was slightly lower in the schizophrenia group (n = 24) as compared to the normal control group (n = 30). Women with schizophrenia (n =13) outnumbered their control group counterparts (n = 7). The number of right and left hemispheres studied was comparable for both the control (L = 14, R = 23) and patient (L = 20, R = 17) groups. One patient with a comorbid diagnosis for alcohol dependency and another schizophrenia patient with comorbid drug abuse at the time of death were included in the cohort.
Relationship of demographic variables to each other and RIN
The subjects spanned a considerable age range (18–78 years) and we found that age at death negatively correlated with the overall brain-weight (r = −0.32, p = 0.005, n = 74), overall brain volume (r = −0.26, p = 0.026, n = 74), DLPFC pH (r = −0.27, p = 0.018, n = 74) and RIN (r = −0.27, p = 0.020, n = 74; Figure 1). As expected, women (n = 20, 1300 mL) had smaller brains as compared to men (n = 54, 1468 mL, t = 5.13, df = 72, p < 0.0001), and women (1288 g) had lighter brains as compared to men (1469 g; (t = 5.55, df = 72, p < 0.0001). Patients with schizophrenia (1394 g) had a lighter brain-weight compared to non-affected controls (1446 g) although this difference was not statistically significant (t = −1.53, df = 72, p = 0.131). People with schizophrenia tended to smoke (n = 23) rather than not smoke (n = 7) and were more likely to be confirmed smokers than controls (n = 9). Smokers tended to have lower brain pH in both frontal cortex (6.63 vs 6.66) and cerebellum (6.45 vs 6.57), and frontal cortex RIN was marginally lower in the smokers (7.2 vs 7.3). The pH and RINs in the smokers and non-smokers, however, did not differ statistically.
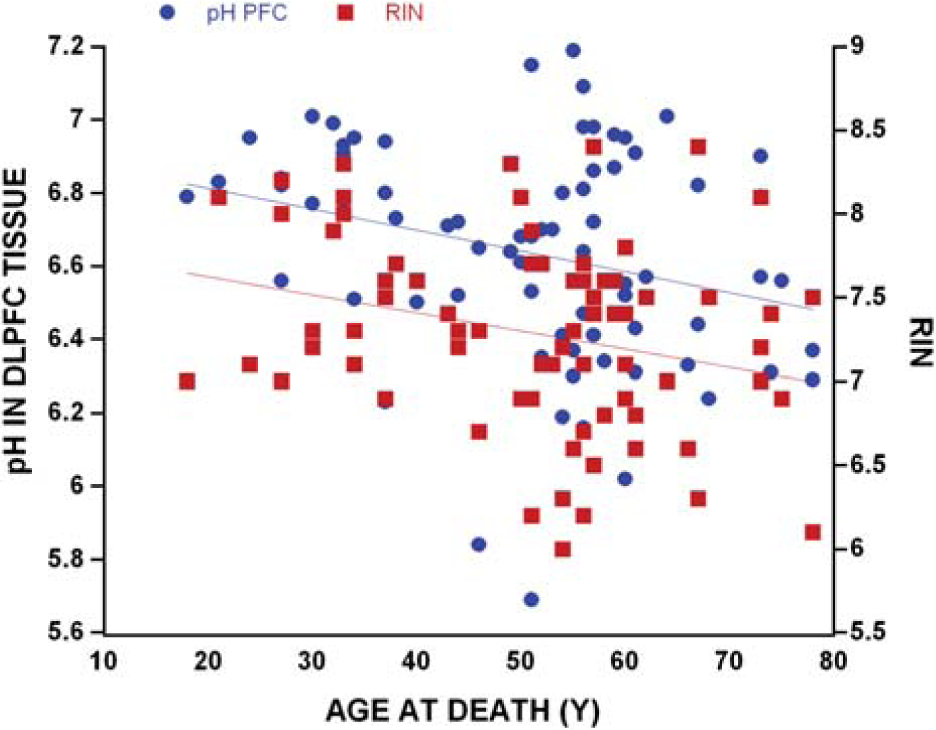
Overall pH and RNA integrity number (RIN) similarly decreased with increasing age (both r = −0.27, p < 0.020, n = 74).
As expected, the RIN measurement for DLPFC total RNA positively correlated with brain pH taken from the same tissue (r = 0.39, p < 0.001) and also correlated with cerebellar pH (r = 0.31, p = 0.007; Figure 6a). Supplementary material to be found online, at http://www.informaworld.com/10.1080/00048670903393662. We found that the pH measurements from the cerebellum and middle frontal gyrus correlated significantly with each other (r = 0.48, p < 0.001, n = 74). PMI did not correlate with RIN (r = 0.02, p = 0.873; Figure 6b-Supp) or cerebellar pH (r = 0.17, p = 0.216). Additionally, we extracted the same amount of RNA per gram of frozen tissue in people with schizophrenia and unaffected controls (both, 0.66 μg mg−1, t = −0.06, df = 72, p = 0.958).
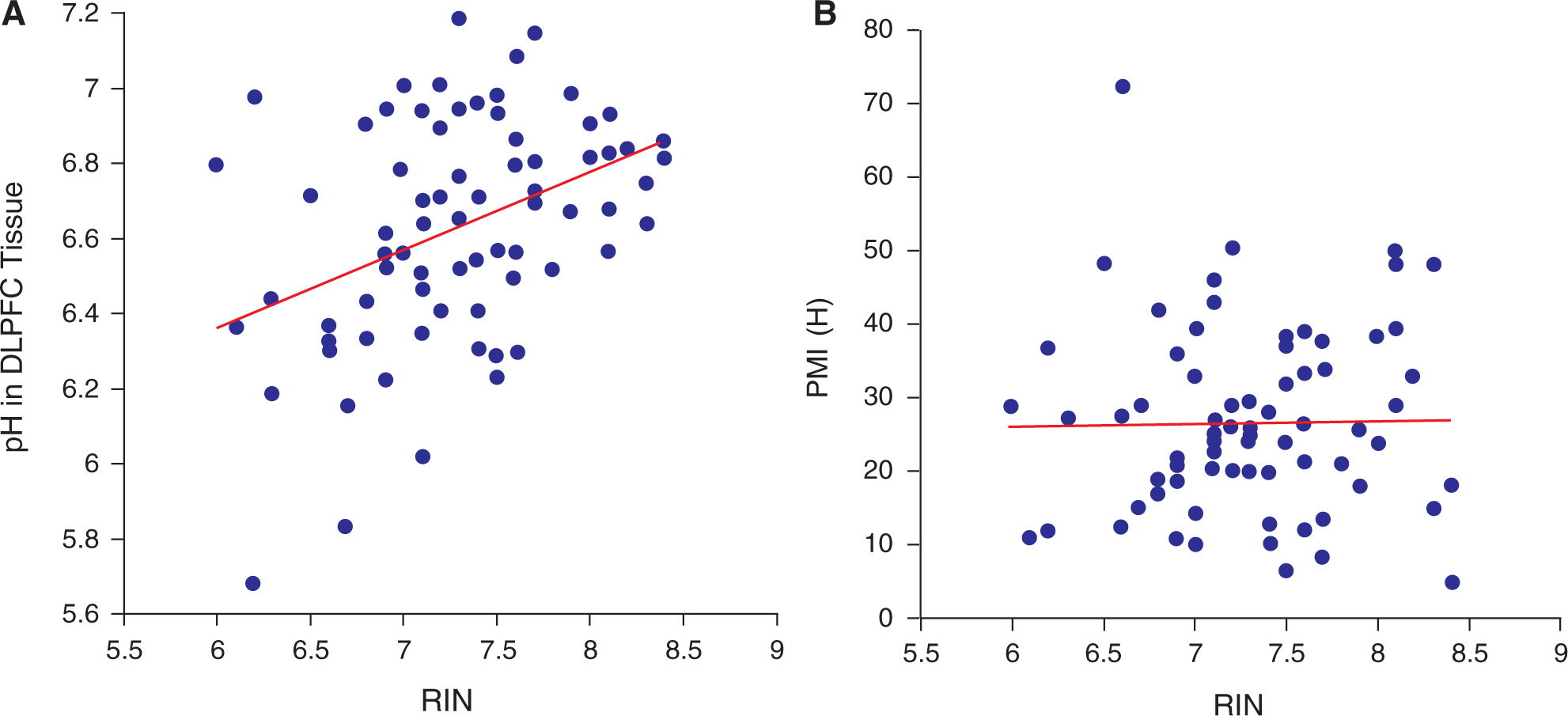
(A) RIN values are positively correlated with pH (p=0.001) (B) and not with PMI (p=0.87).
Transcript stability and calculation of geometric mean
We determined that ACTB and GADPH were the two most stable genes generating the lowest M, and that B2M and GUSB were the least stable genes with the highest M (Figure 2A). Pairwise variation analysis suggests that using the two most stable housekeeping genes (ACTB and GADPH) is sufficient to determine the normalization factor because V2/3 is below the suggested threshold of 0.15 (Figure 2B) [24]. For the purposes of normalizing the data to housekeeper control genes, however, we chose a further two genes because it has also been recommended that genes used for normalization are not significantly altered in schizophrenia compared to controls, and represent transcripts of different levels of expression. We chose four genes, two (ACTB and GAPDH) with higher expression levels, one (UBC) being in the midrange of expression, and one (TBP) with lower expression levels [25]. When the geomean was recalculated using these four housekeeping genes (herein geomean(4)) and outliers removed (one case removed) we did not detect a significant difference between the normalizing factor geomean in patients with schizophrenia compared to controls (unpaired t-test, t = 0.82, df = 71, p = 0.415; paired t-test, t = −0.79, df = 35, p = 0.420; Figure 3A).
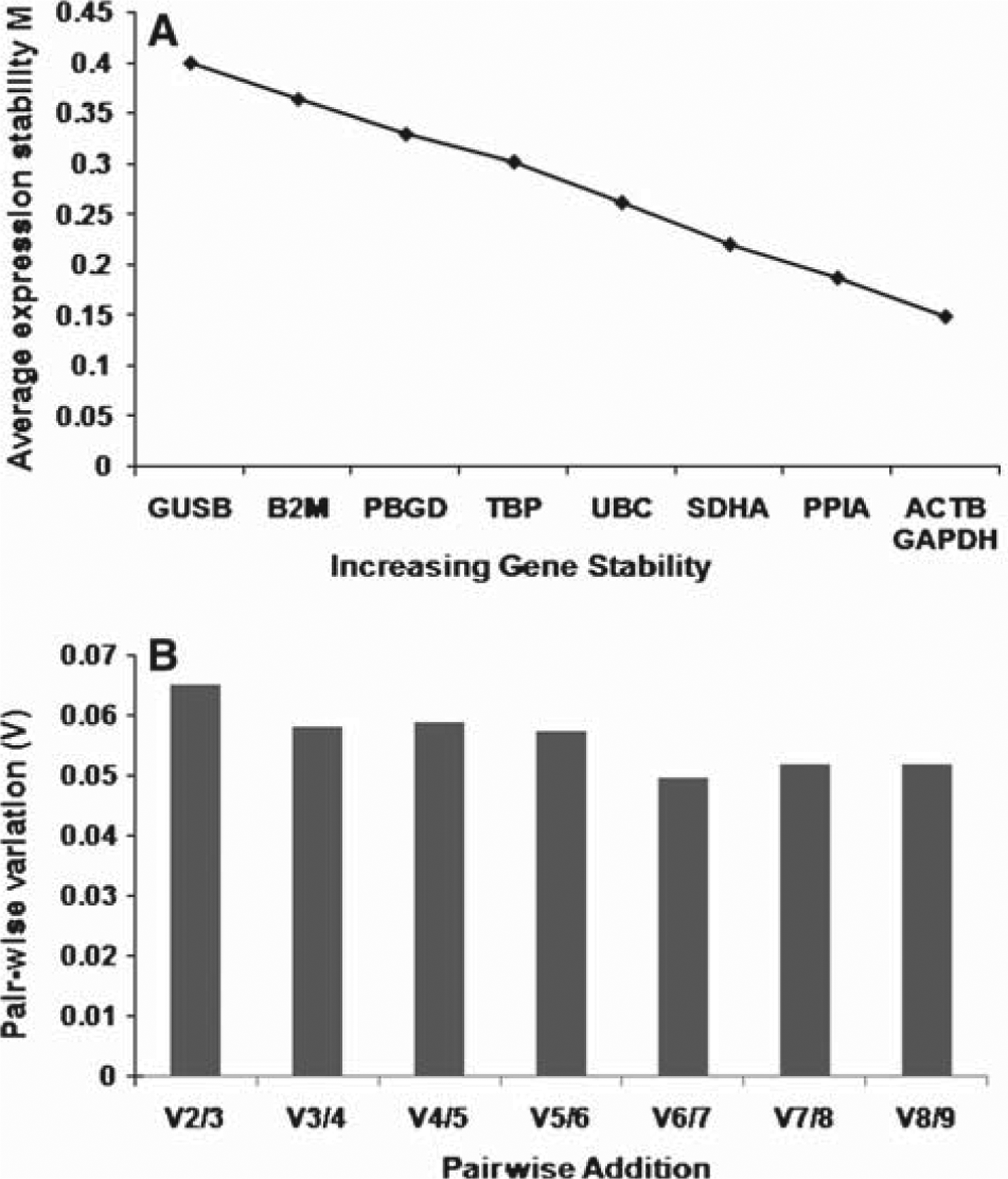
Average expression stability and optimal number of housekeeper genes for normalization determined using the program geNorm. (A) Genes were ranked according to average expression stability. ACTB and GAPDH were the most stable genes (lowest M), while GUSB was the least stable gene (highest M) (B) Pairwise variation with the stepwise addition of more housekeeping genes shows that the use of the two most stable housekeeping genes will be sufficient for accurate normalization because this satisfies the cut-off (0.15 M) recommended by Vandesompele et al. [24].
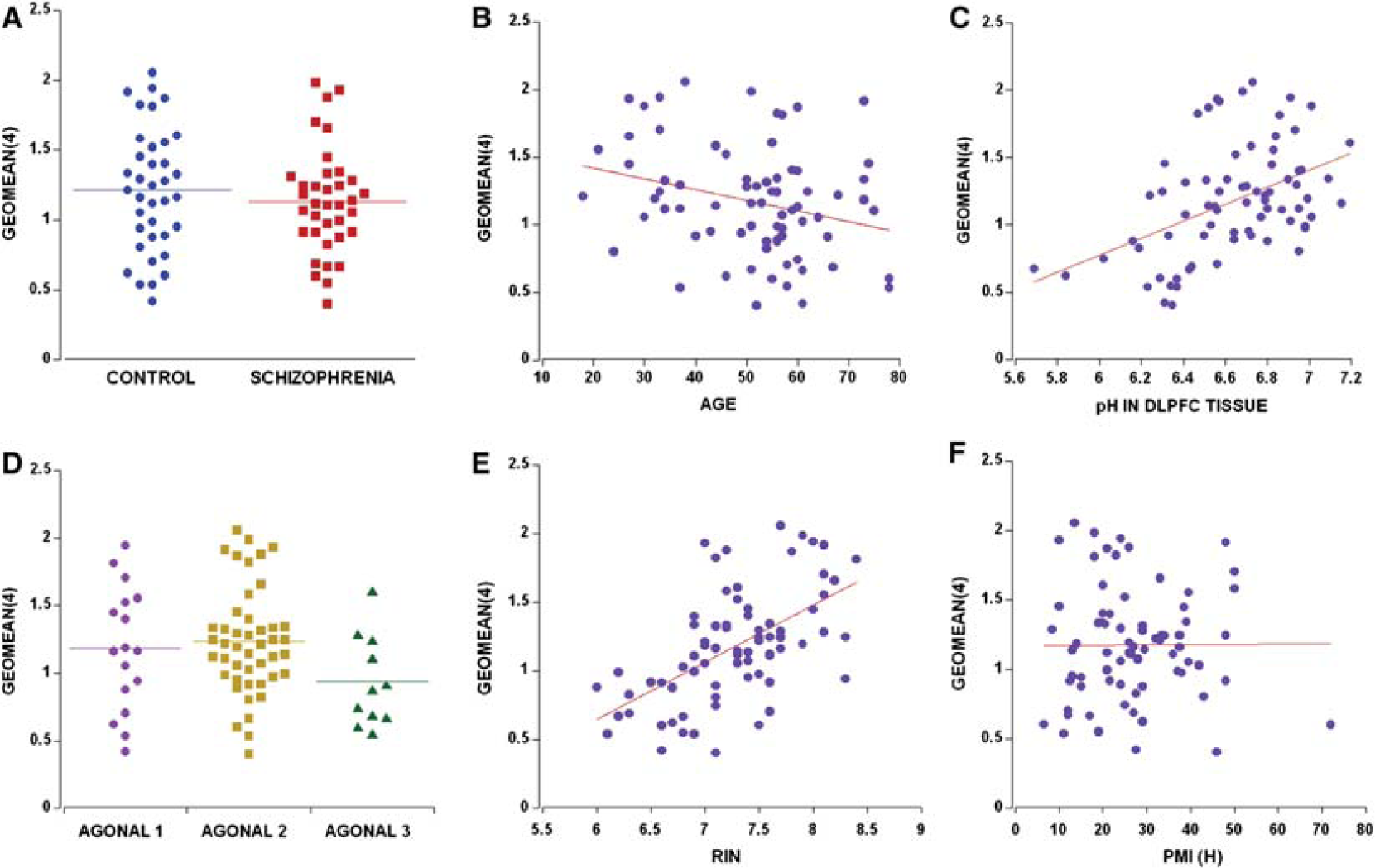
(A) The calculated geomean(4) was comparatively similar in both diagnostic groups whether paired (t = −0.79, df = 35, p = 0.420) or unpaired (t = 0.82 df = 71, p = 0.415); (B) geomean(4) decreased with an increase in age (r = −0.28, p = 0.018); (C) overall pH in dorsolateral prefrontal cortex (DLPFC) tissue was positively correlated with geomean(4) for controls (r = 0.41, p = 0.012) and patients (r = 0.52, p = 0.001); (D) agonal state 1 vs 3 showed a significant decrease in the geomean(4) (p = 0.031 by planned post-hoc LSD); (E) RNA integrity number (RIN) strongly correlated with the geomean(4) (r = 0.57, p < 0.0001); (F) postmortem interval (PMI) had no effect on the geomean(4) (r = 0.006, p = 0.96).
Correlation of demographic variables and housekeeper mRNAs
Age
When considering the whole cohort, the expression of six of nine housekeeping genes correlated negatively with age at death (SDHA: r = −0.26, p = 0.026; ACTB: r = −0.28, p = 0.018; GAPDH: r = −0.28, p = 0.016; PBGD: r = −0.26, p = 0.030; UBC: r = −0.26, p = 0.03; PPIA: r = −0.26, p = 0.025) as did the geomean(4) (r = −0.28, p = 0.018; Figure 3B). When we explored which diagnostic group contributed more to these overall correlations, we found that the normal group did not show any significant correlations with any housekeeper control genes or geomean(4) and age at death (all p > 0.08). In contrast, the expression of five of nine housekeeping genes and geomean(4) correlated negatively with age in schizophrenia patients only (SDHA: r = −0.45, p = 0.006; ACTB: r = −0.47, p = 0.004; GAPDH: r = −0.37, p = 0.024; UBC: r = −0.37, p = 0.028, PPIA: r = −0.40, p = 0.016; geomean(4): r = −0.45, p = 0.006).
pH
Most (seven of nine) reference genes showed a significant positive correlation with DLPFC tissue pH (range: r = 0.26–0.58, p = 0.02 to <0.001). The only two potential reference genes that did not show a significant relationship with tissue pH were those deemed least stable, B2M (r = 0.14, p = 0.237) and GUSB (r = 0.11, p = 0.364). In general, most reference mRNAs were positively correlated to tissue pH in both control and schizophrenia subjects. Additionally, geomean(4) showed a significant correlation to pH in both diagnostic groups (controls: r = 0.41, p = 0.012; patients: r = 0.52, p = 0.001; Figure 3C).
Agonal state
Agonal state had an influence on DLPFC pH (F = 3.07. df = 2,71, p = 0.051) but this did not reach statistical significance with cerebellar pH (F = 3.01, df = 2,71, p = 0.057). In both the prefrontal cortex and the cerebellum, the individuals with the highest agonal state rating had the lowest pH (6.44 and 6.32, respectively). Each agonal state rating had significantly different DLPFC pH on post-hoc least significant difference testing (LSD; all p ≤ 0.05). Although individuals with the highest agonal state scores had the lowest RNA quality as assessed on RIN, this did not reach statistical significance. Those individuals with the longest agonal state in the present study did show a 27% decrease (F = 2.68, df = 2,33, p = 0.086) in expression of the geomean(4) of the housekeeper normalizer genes (agonal state: 1 vs 3, p = 0.031 on planned post-hoc LSD; Figure 3D).
RNA integrity number
All nine potential reference genes had a significant positive correlation with DLPFC RNA quality as determined on RIN. Most of the genes (eight of nine) and the geomean(4) (Figure 3E) showed a strong and statistically significant correlation (range r = 0.43–0.63, all p < 0.001, except GUSB mRNA (r = 0.24, p = 0.042)). All nine housekeeper mRNAs in the control group displayed a correlation with RIN (all r > 0.46, all p < 0.006). Similarly, eight of nine housekeeping genes showed a correlation with RIN in the schizophrenia group (excluding GUSB; all r > 0.36, all p < 0.029).
Postmortem interval
Of the housekeeper control mRNAs, most of the housekeeper genes and the geomean(4) were not correlated with PMI (Figure 3F; all r < 0.067, all p > 0.572 except B2M r = −0.20, p = 0.088). The expression of GUSB in schizophrenia patients only (r = −0.35, p = 0.041) was significantly negatively correlated with PMI.
Diagnostic differences in housekeeper control mRNA
T-tests
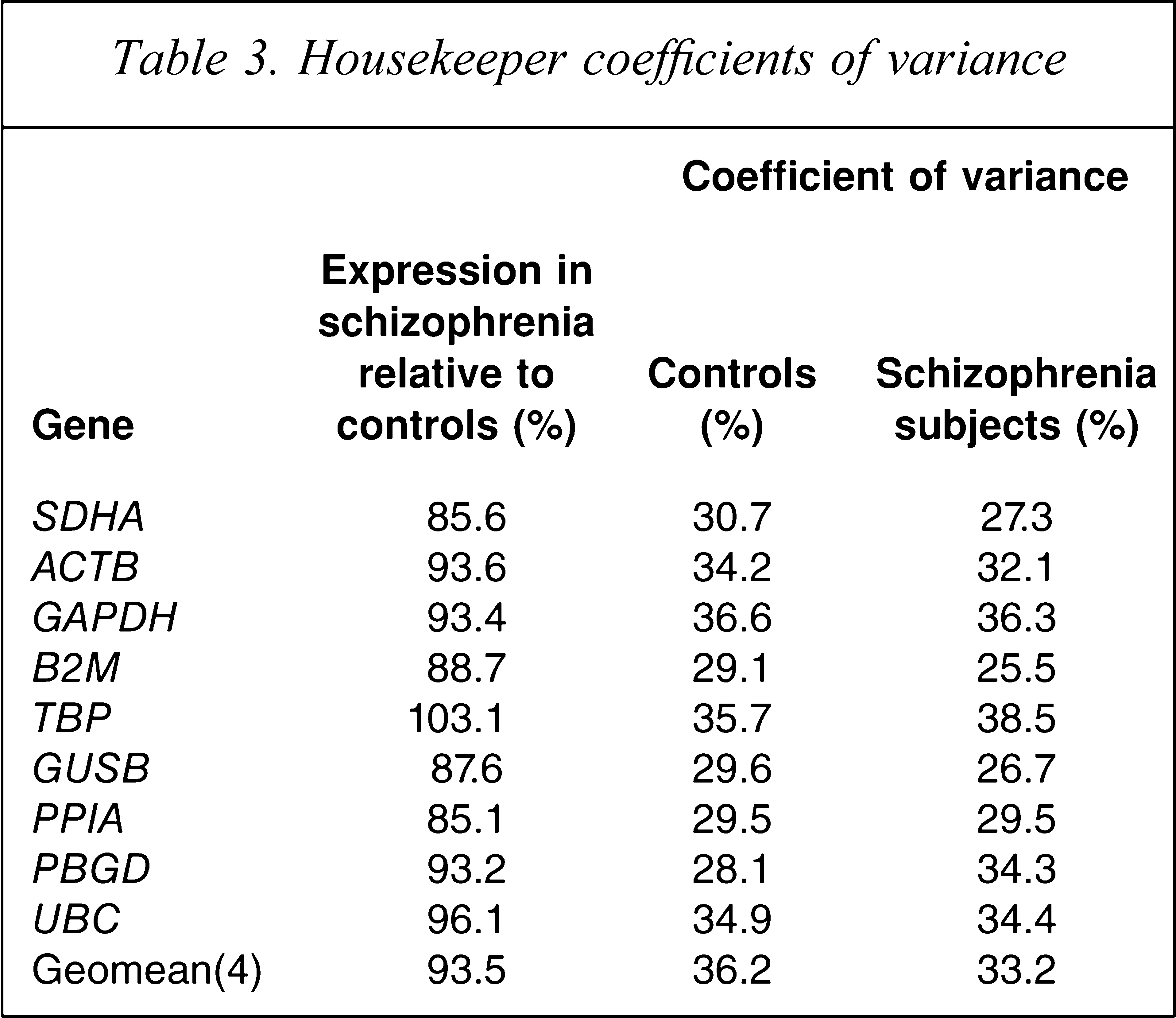
In the present study we found approximately 30% coefficient of variation in housekeeper mRNA in both groups (Table 3), which is less variable than previously found [6]. Surprisingly, we found that some housekeeping control genes were reduced in the DLPFC of patients with schizophrenia by as much as 10–15% (Figure 4A). We found that SDHA (t = −2.21, df = 69, p = 0.030 unpaired; t = −2.55, df = 33, p = 0.016 paired), PPIA mRNA (t = −2.32, df = 71, p = 0.023 unpaired; t = −2.55, df = 35, p = 0.015 paired) and GUSB (t = 1.93, df = 68, p = 0.057 unpaired; t = 2.02, df = 32, p = 0.052 paired) were significantly lower in the schizophrenia and schizoaffective disorder patient group by 14.4%, 14.9%, and 12.4%, respectively. The five other housekeeper genes examined (UBC, ACTB, GAPDH, TBP, PBGD) and the geomean(4) did not significantly differ in patients compared to controls (all p > 0.343; Figure 4B). After ANCOVA analysis, the significant reduction in SDHA mRNA (F = 4.54, df = 69, p = 0.036) and PPIA mRNA (F = 5.78, df = 68, p = 0.018) was maintained. No significant differences were found in any of the other mRNAs or the geomean(4) on ANCOVA (p > 0.05).
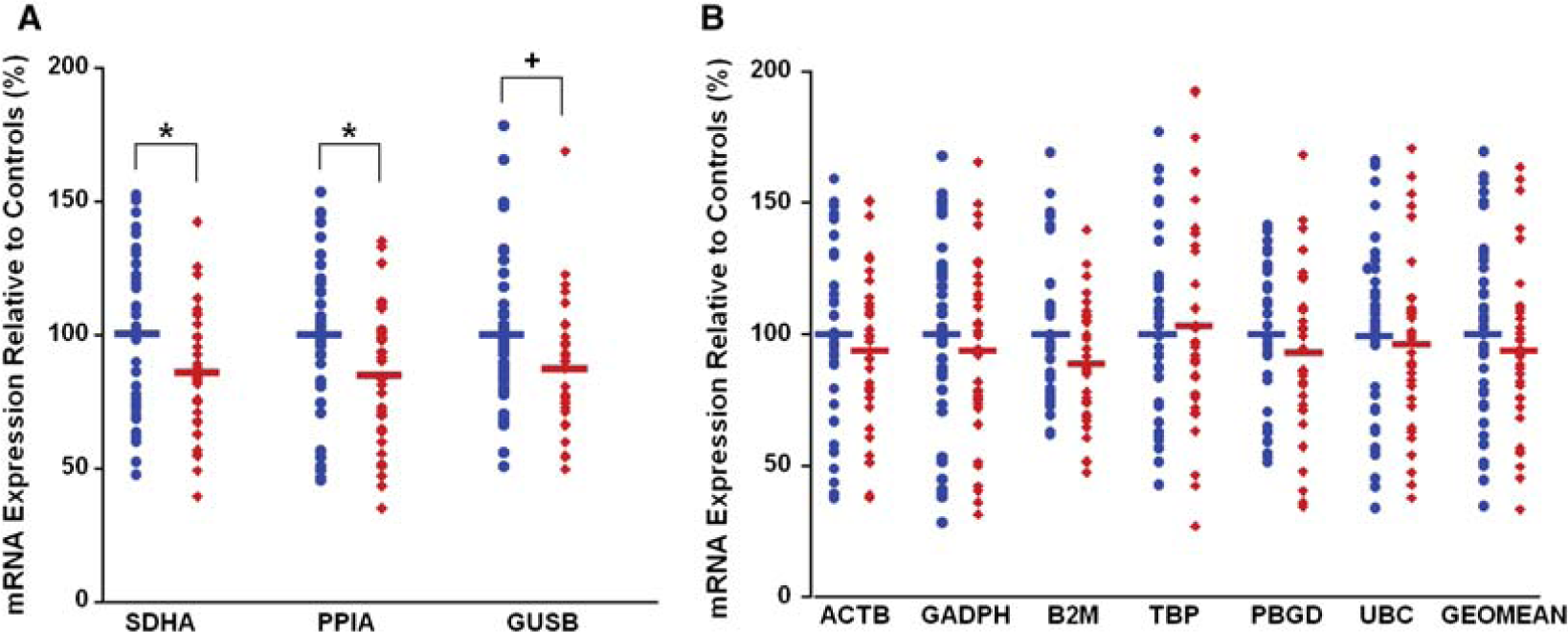
mRNA expression of housekeeping genes in control and schizophrenia dorsolateral prefrontal cortex (DLPFC). mRNA expression was measured by quantitative reverse transcription-polymerase chain reaction and all cases expressed as a percentage of the control mean for that gene. (A) A significant reduction in expression of SDHA, and cyclophilin (PPIA) and strong trend in GUSB mRNAs was found in schizophrenia patients compared to controls (14.4%, p = 0.016; 14.9%, p = 0.015; and 12.4%, p = 0.052, respectively; paired t-test). (B) mRNA expression levels for all remaining housekeeping genes that did not significantly differ between diagnostic groups, including the geomean(4) (all p > 0.34).
Housekeeper coefficients of variance
Effects of hemisphere and gender on geomean(4)
We found that there was a main effect of hemisphere on the geomean(4) by two-way ANOVA (F = 6.26, df = 1, 69, p = 0.021), but no overall effect of diagnosis (F = 6.26, df = 1, 69, p = 0.228) and no interaction effect (F = 6.26, df = 1,69, p = 0.989). Both people with schizophrenia and controls had significantly more expression of the housekeeper geomean(4) on the left hemisphere as compared to the right. When we covaried for factors that correlated with the geomean(4) (age, pH and RIN), we found that the main effect of hemisphere was similar (F = 3.86, df = 1,66, p = 0.053) and the effect of diagnosis and the diagnosis × hemisphere interaction remained non-significant. We did not detect a main effect of gender or diagnosis or an interaction between diagnosis and gender on the levels of geomean(4) (all F < 1.3, all p > 0.260).
Consideration of clinical variables
The patients with schizophrenia in the present cohort had 4-45 years duration of illness, which was positively correlated with daily chlorpromazine equivalent neuroleptic dose (CPZ; r = 0.38, p = 0.020, n = 37) and the last recorded dose of CPZ (r = 0.38, p = 0.027, n = 37). Total brainweight of people with schizophrenia and/or schizoaffective disorders was negatively correlated with daily CPZ (r = −0.34, p = 0.038, n = 37; Figure 5A), duration of illness (r = −0.35, p = 0.031, n = 37; Figure 5B), and lifetime CPZ estimates (r = −0.39, p = 0.018, n = 37). The relationship of brain volume with duration of illness and lifetime CPZ estimates reached trend levels of significance (p = 0.060–0.084, n = 37). The only control mRNA that showed a statistically significant correlation with any medication measure was UBC with the last recorded CPZ (r = −0.36, p = 0.048, n = 31). Duration of illness did, however, negatively correlate with SDHA and ACTB (r > −0.37 p = 0.028, n = 35; r = −0.38, p = 0.022, n = 36, respectively) mRNA expression, and with the geomean(4) (r = −0.35, p = 0.034, n = 36; Figure 5C). Because these same dependent variables correlated with age in the people with schizophrenia only, we ran partial correlations with age and found that only SDHA mRNA retained a relationship with duration of illness at a trend level (r = −0.32, p = 0.066).
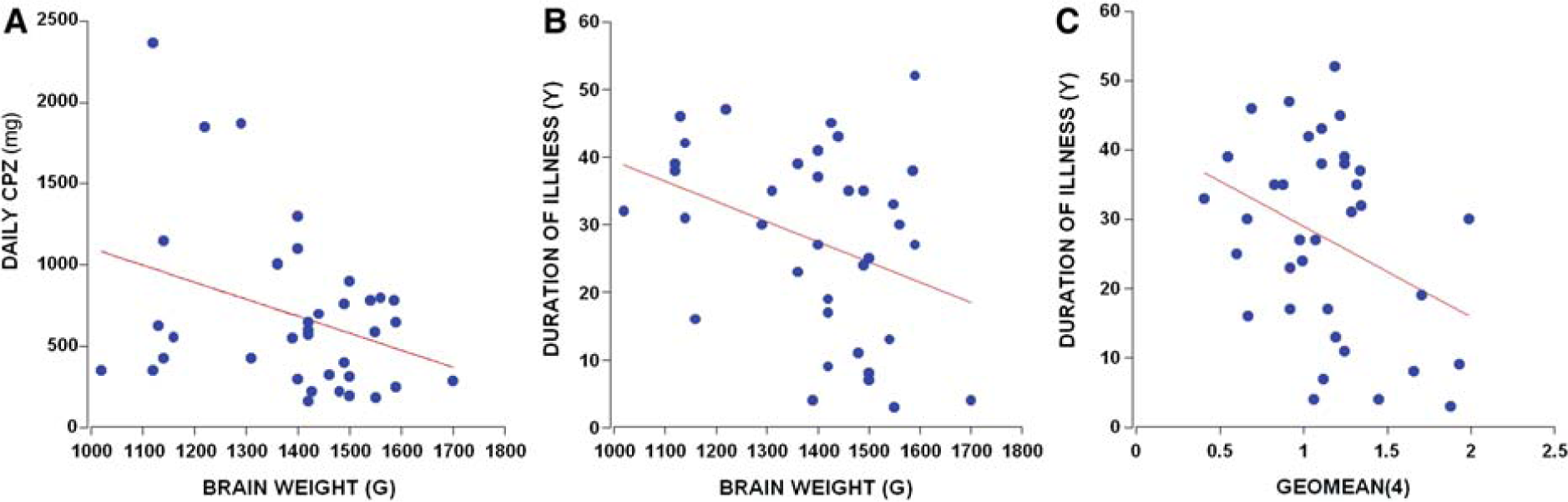
The clinical variables that correlated with brainweight were (A) daily chlorpromazine equivalent neuroleptic dose (CPZ) (r = −0.34, p = 0.038, n = 37) and (B) duration of illness (r = −0.35, p = 0.031, n = 37). (C) Duration of illness negatively correlated with the geomean(4) (r = −0.35, p = 0.034, n = 36).
Power analysis
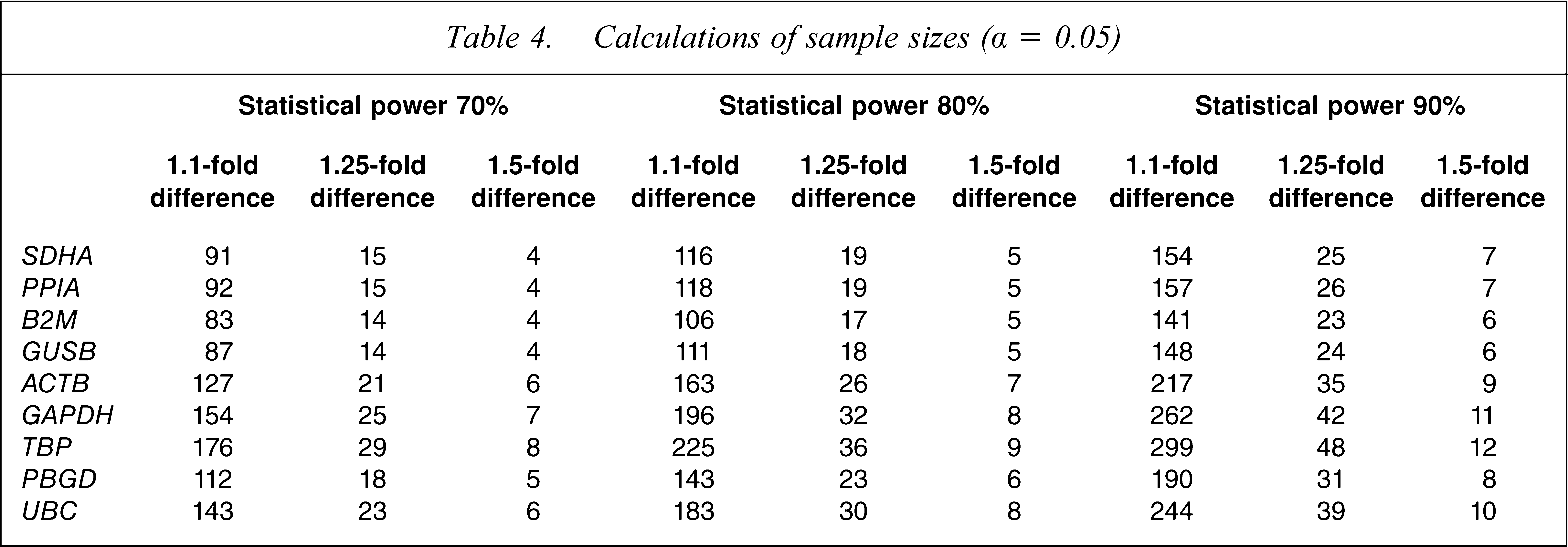
Power calculations, performed using means and standard deviations of housekeeping genes in controls and patients with schizophrenia, were used to determine the sample sizes required to reject a null hypothesis of no difference in housekeeping gene expression between diagnostic groups. Power analysis indicated that the present cohort contained a sufficient sample size to detect a 1.5-fold difference in the mRNA expression of all housekeeping genes between patients and controls, with 90% statistical power (α = 0.05), requiring between six and 12 samples per group to detect this fold change. Additionally, we had sufficient sample size to detect a 1.25-fold change in expression for all of the genes at 80% power and for six of nine genes at 90% power (Table 4). To detect a 1.1-fold difference with 90% statistical power would require 141–299 samples per group or, relaxing stringency to 70% statistical power, 83–176 samples per group (Table 4).
Calculations of sample sizes (α = 0.05)
Discussion
Some consensus concerning the effects and relative importance of premortem and postmortem factors on human brain RNA quality is emerging. We find, as others do, that RIN is a powerful indicator of mRNA levels and appears to be the most critical factor for matching schizophrenia cases and controls [6,26,27]. But because RIN measures are available only after RNA isolation, we found that tissue pH can be used as a general guide to select potential controls to match to cases prior to RNA isolation [26,27]. Because cerebellar and cortical pH correlate, determining cerebellar pH may be a valuable first step in evaluating the suitability of cases for RNA analysis [26,28], but high tissue pH does not always indicate good RNA [26,27], and low pH does not necessarily yield low-quality RNA. The current cut-off for what is an acceptable RIN varies considerably, with some studies suggesting that a moderate amount of degradation and a RIN as low as 3.8–4.0 may be usable [6,8,29]. Also, it should be mentioned that RINs are typically higher after total RNA is purified by columns, but this purification step would exclude some small RNAs of interest such as microRNAs, and thus columns were not used in the present study. Ultimately, the cut-off for a usable RIN would depend on criteria set by the end user depending on the assay used and the molecule of interest, and thus, will vary. Earlier studies emphasized the importance of matching for tissue pH and RIN, and we have prioritized tissue pH, RIN, and age as variables to control for in assembling a cohort of schizophrenia and normal brains that are pair-selected within ranges, creating a closely matched and well-powered cohort.
Most brain banks attempt to collect cases with the shortest PMI possible, although researchers have challenged the importance of PMI in determining RNA quality [6,30–33]. A number of studies suggested that PMI has no correlation or only a weak correlation with RNA quality [10,26–28]. Studies that have examined human tissue collected during neurosurgery for epilepsy with minimal delay shows that the freshly acquired brain tissue pH and RIN are comparable or even lower to that collected after death [10,26]. In agreement with these published findings, we found no correlation between PMI and most of the RNA quality measures in the DLPFC [26,34], with only one slight negative correlation with SDHA as previously described [6]. A microarray study in mouse brain tissue identified >1000 transcripts that are degradation susceptible (labile) with increasing PMI, but they found that increased PMI does not affect GADPH mRNA levels [35]. Other studies in human brain suggest that another housekeeper control gene, B2M, is highly labile [36,37]. On one hand, B2M may serve to correct for degradation levels if used to normalize genes of interest; on the other hand, we found that B2M is a fairly unstable housekeeper control gene and thus, we would not recommend it as a good control for schizophrenia studies. Other studies in humans suggest that some housekeeper transcripts are slightly influenced by PMI, especially in the hippocampus [6], although this may not be the case for all hippocampal housekeeper RNAs [28]. Thus, although effects of PMI need to be evaluated for particular transcripts of interest through correlational analyses, PMI is unlikely to be a major determinant of RNA or transcript quality and may be helpful but not critical to consider when matching samples.
In contrast, it has been well demonstrated that tissue pH influences mRNA preservation in postmortem tissue [6,32,38,39]. In the present study we found that the majority of housekeeper RNAs assayed and the geomean(4) correlated strongly with tissue pH. Studies that have measured the same housekeeper controls have also found positive correlations with brain tissue pH [6,40]. Some previous reports suggest that a pH cut-off (pH <5.9–6.0) can be used as a screen to choose which brains from which to extract RNA and analyse further [4], or that tissue pH could be used as a critical lower threshold for transcriptome analysis (pH range 6.54–6.8) [28,41]. Because most postmortem samples used here and in other current cohorts span a large pH range [6,26–28], with as many as 39–56% of subjects from psychiatric groups having a pH <6.5 [8] and as many as 28% of the present current sample, it would not be reasonable to exclude this large number of cases. Thus it is of critical importance to either pair-match for, or group (stratum) match for tissue pH. Several research groups have noted that pH is not always a reliable predictor of RNA quality and we find that the individual with the lowest brain pH (5.7) had a RIN of 6.2. Hence, we agree with the recommendation that RNA isolation and RIN quality assessment should be completed on all samples before precious material is excluded from cohorts based on tissue pH alone [6,27].
The importance of agonal state in postmortem research has been increasingly recognized over the past several decades [36,41–44]. We found that the longer the agonal state (prolonged death) the lower the brain pH; but we did not find an overall effect of agonal state on RIN. This agrees with studies that show that as agonal state scores increase, the tissue pH decreases [27,28,45], but RIN may not differ significantly with agonal state [27]. We did find a lower value for geomean(4) with higher agonal state scores. Prolonged agonal state can downregulate some mRNAs (energy metabolism genes), while increasing others (stress response and inflammation control genes) [45], suggesting that agonal state is an important factor to consider when matching samples or groups of samples.
Studies of the effects of aging on housekeeper gene expression in the human brain have led to the conclusion that age at death influences mRNA expression, especially when individuals span a large age range [6,40]. In the present study, individuals spanned six decades of life with an average age of 51, which is comparable to several other cohorts used to study people with schizophrenia compared to controls [6,46]. One of the surprising findings of the present study was that five of nine housekeeper control genes and the geomean(4) negatively correlated with age in the schizophrenia patients only. We also found that women but not men with schizophrenia had significantly smaller brains with increasing age. Lighter brains in both sexes with schizophrenia has been reported previously [47], but the neurobiological underpinnings of this are not understood. We also found that overall brainweight and brain volume negatively correlated with the mean daily dose of neuroleptic medication and duration of illness, suggesting that the longer someone was ill or the higher the dose of daily neuroleptics, the lower the brainweight and the smaller the brain at death. Patients with a longer duration of illness tended to have higher average lifetime antipsychotic dose, higher antipsychotic dose at the time of death, and lower housekeeper mRNA expression. Because the clinical measure of duration of illness is linked to age at death, we attempted to control for the effect of age by using partial correlation and found that SDHA mRNA correlated negatively with duration of illness at a trend level. The fact that the correlations between duration of illness and housekeeper gene mRNAs are no longer statistically significant is difficult to interpret because age is not independent from duration of illness and thus some of the true variance coming from the duration of illness will be subtracted in the partial correlation for age. This is also true for earlier reports that partialled out the effect of age and failed to find an effect of duration of illness on brainweight using meta-analysis [47]. The fact that older people with schizophrenia with a longer duration of illness have smaller brains and have decreased control mRNAs suggests that the longer someone lives with schizophrenia the greater the chance that they have accumulated deleterious impacts on their brain. This observation, although made on cross-sectional data across the lifespan, fits well with volumetric studies showing decreased grey matter volume over time in people suffering from chronic schizophrenia [48,49]. The present data suggest that some of these deleterious effects may be linked to antipsychotic medication effects, or at the very least are not prevented by current treatment strategies. Recent reports suggest that chronic antipsychotic use (both typical and atypical) causes a 10% decrease in brainweight and does in fact decrease cortical brain volume in controlled stereological studies of non-human primates [50,51]. This volume reduction was linked to a significant reduction in astrocytes and oligodendrocytes, a finding that parallels the reduction in astrocytic and oligodendrocyte markers found in some [52–55] but not all [56,57] postmortem studies of people with schizophrenia.
In the present study we found that two housekeeper genes, β-actin and GAPDH mRNAs, do not differ between people with schizophrenia compared to control as previously found [58,59], and the present data suggest that both are particularly stable mRNAs and useful housekeeping control genes for postmortem human brain studies. It should be emphasized, however, that the utility of GAPDH as a control varies from brain collection to brain collection, because some laboratories find GAPDH to be stable and to not differ in people with schizophrenia as compared to controls, as we have done [59,60], while others suggest that it is one of the least stable and thus, least reliable control genes [22,61]. We found that three putative housekeepers, GUSB, PPIA, and SDHA, are lower in people with schizophrenia. PPIA, and SDHA mRNA differences reached statistical significance and both of these housekeeper mRNAs may be regulated by cellular stress. PPIA encodes a member of the peptidyl-prolyl cis-trans isomerase (PPIase) family, which is a cyclosporin binding protein and can accelerate the folding of proteins. PPIA mRNA is not typically reduced in patients with schizophrenia in other cohorts [60,62–64]. Thus, the reductions in PPIA mRNA that we have found may represent increased power to detect a difference (approx. 10% reduction) or may represent a finding specific to the present cohort. The other transcript we found to be decreased, SDHA, encodes a major catalytic subunit of succinate-ubiqui-none oxidoreductase, part of the mitochondrial respiratory chain complex II on the mitochondrial inner membrane. Mutations in SDHA have been associated with a form of mitochondrial respiratory chain deficiency known as Leigh syndrome and with late-onset optic atrophy [65,66]. Dysfunction of mitochondrial respiration can lead to cognitive decline in attention and mental flexibility because respiratory chain diseases most often affect muscle and the central nervous system (CNS) [67]. CNS symptoms can include epilepsy, demyelination, dementia and psychosis. Oxidative stress related to alterations in metabolism linked to mitochondrial distress has been observed in schizophrenia [43,68–70]. GUSB, or glucuronidase-β, mRNA encodes a protein that is required to degrade glycosaminoglycans (i.e. heparin sulphate), and genetic changes in GUSB can result in deafness, behavioural deficits and mental retardation [71]. Thus, although certain genes are chosen to represent the integrity of the transcript pool and are thus used as housekeeper control or reference genes, they can have a critical function in brain as evidenced when mutated. The question regarding whether slightly lowered reference gene transcript levels have any functional significance requires further investigation. The present study suggests that GUSB, PPIA and SDHA mRNAs are not optimal choices for reference genes in schizophrenia studies.
Taken together, the present data suggest that although only a small amount of variance in RNA quality or housekeeper mRNA level is explained by any one demographic (age) or tissue characteristic (pH), their effects are consistent across many laboratories and represent known confounders/covariates that can be controlled for when constructing postmortem cohorts. We suggest that matching diagnostic groups minimally for age, pH and RIN may help to resolve differences in gene expression that occur in schizophrenia as either a cause, a compensation or a consequence of the disease state [17].
Footnotes
Acknowledgements
Since its establishment, Schizophrenia Research Institute (SRI) has supported schizophrenia research across a broad range of disciplines and has also been a major supporter of the New South Wales Tissue Resource Centre, an infrastructure facility that collects, stores and distributes fixed and frozen postmortem human brain tissue for projects related to schizophrenia. This work was supported by Schizophrenia Research Institute, utilizing infrastructure funding from NSW Health, University of New South Wales School of Psychiatry, and Prince of Wales Medical Research Institute. We thank Dusan Hadzi-Pavlovic for statistical advice.
