Abstract
The purpose of this study was to investigate in vivo monitoring of hair follicle stem cells (newborn mouse fibroblasts [NFs]) expressing enhanced firefly luciferase (effluc) (NF-effluc) using noninvasive bioluminescence imaging (BLI). Effluc gene transduction into NFs was performed by retroviral vector, and effluc messenger ribonucleic acid expression and function were evaluated by reverse transcription–polymerase chain reaction and luciferase assay. After in vivo transplantation of NF-effluc cells to generate hair follicles, BLI was performed on days 1, 3, 7, 14, and 21 after transplantation. Additionally, hair follicle generation by the implanted stem cells was investigated using microscopy. The luciferase activity of NF-effluc was 41,175-fold higher compared to that of untransfected NFs. Bioluminescence signals from the transplantation site decreased gradually over 2 weeks; then the signal plateaued. Hair follicles were confirmed at the NF-effluc cell implantation site on day 14 after transplantation. We successfully monitored hair generation by hair stem cell implantation noninvasively with optical molecular strategy in an in vivo model.
HAIR LOSS is a common problem that has a psychological impact. The causes can be hormonal changes, a poor diet and nutritional deficiencies, a variety of medications, surgery, and potentially many other factors. 1 The desire to look good makes people affected by hair loss search for treatment, which may range from clinically proven pharmaceutical treatments such as topical and oral agents to surgical intervention and transplantation after hair cloning.2–4 A number of in vivo mouse models to generate hair follicles were subsequently developed. Among them, the hair patch assay can reconstitute hair follicles by subcutaneous injection of a mixture of epidermal and dermal cells. 5 Hair stem cells play a pivotal role in hair growth regulation. The important issues for hair follicle generation include isolating the inductive dermal population, expanding their number by culture, maintaining hair inductive properties, and providing the follicles with an adequate niche and exogenous signals to enhance their interaction with epidermal cells. 6
The rapid development of molecular imaging techniques has enabled noninvasive visualization of various biologic events, such as gene delivery, endogenous gene expression, cell monitoring, signal transduction, apoptosis, tumor angiogenesis, and protein–protein interactions in living organisms.6–9 Bioluminescence imaging (BLI) has been used to study stem cell biology such as the interactions between stem cells, interactions between stem cells and their microenvironment, and the differentiation capacity of stem cells. 10 BLI, as an ideal modality for in vivo stem cell monitoring, has been frequently applied because it does not entail high levels of animal sacrifice. 11 In the field of stem cell research, these molecular imaging techniques have been used to visualize cell survival, migration, and proliferation using bioluminescent and fluorescent imaging techniques.12,13 In contrast to the fluorescent reporter genes that have been used in histology (immunofluorescence) for postmortem confirmations, BLI has been used to monitor stem cells delivered to the organs of a living small animal.14,15 BLI monitoring of mice with implanted hair stem cells might be able to monitor the cells noninvasively until hair follicles are generated without sacrificing an animal. However, no reports are available about in vivo BLI of hair stem cells showing cell viability in living subjects.
In this study, we investigated the feasibility of in vivo monitoring of hair follicle stem cells (newborn mouse fibroblasts [NFs]) expressing enhanced firefly luciferase (effluc) (NF-effluc) using noninvasive BLI in an animal hair follicle generation model.
Materials and Methods
Isolation and Culture of NF Cells
After the epithelial component was separated from the dermal component, cell preparations followed an adaptation of the procedure in Prouty and colleagues. 16 Briefly, skin was harvested from C57BL/6 (SLC, Inc., Shizuoka, Japan) neonates, and mouse dermal and epidermal layers were freshly isolated and separated by incubating overnight with 100 mg/mL of collagenase/dispase (Roche, Mannheim, Germany) at 4°C. The skin fragments were subsequently digested with 0.25% trypsin in phosphate-buffered saline at 37°C for 15 minutes to obtain single cells.
Production and Transfection of Retrovirus-Expressing Effluc Gene
Phoenix cells grown in 100 mm dishes were transduced using a calcium phosphate method with 10 μg of pRetro/LTR-effluc-IRES-Thy1.1 vector for 24 hours. The pRetro/LTR-effluc-IRES-Thy1.1 vector was kindly provided by Dr. Patrick Hwu (M.D. Anderson Cancer Center, Houston, TX). After transduction, the cells were incubated at 37°C. The supernatant containing retrovirus particles producing the effluc gene was collected 48 hours later and filtered through a vacuum-driven bottle-top filter system (Millipore, Bedford, MA). NFs were transfected with the viral supernatants in the presence of 10 μg/mL polybrene for 24 hours, and the medium was changed to fresh growth medium. Thy1.1-phycoerythrine(PE)-positive cells (NF-effluc) were sorted with CD90.1 microbeads (Miltenyi Biotec, Bergisch Gladbach, Germany).
Reverse Transcription–Polymerase Chain Reaction (RT-PCR)
NFs and NF-effluc cells were lysed using Trizol solution (Invitrogen, Carlsbad, CA), and total ribonucleic acid (RNA) was extracted according to the manufacturer's instructions. Reverse transcription was performed using a Revert-Aid First Strand cDNA Synthesis kit (Fermentas, Burlington, ON). Briefly, 2 μg of total RNA was reverse-transcribed in a final volume of 20 μL containing 1 μL of oligo(d)T primer, 4 μL of 5× reaction buffer, 2 μL of 10 mM dNTP mix, and 1 μL of MuLV reverse transcriptase (Fermentas). The effluc gene was amplified with forward (5′-GCACAAGGCCATGAAGAGAT-3′) and reverse (5′-CTTCTTGCTCACGAACACCA-3′) primers and 2 units of Taq deoxyribonucleic acid (DNA) polymerase (Takara, Shiga, Japan) using the GeneAmp PCR system (Bio-Rad, Hercules, CA). After denaturation of the samples for 1 minute at 94°C, 30 cycles of 25 seconds at 94°C, 30 seconds at 57°C, and 30 seconds at 72°C were followed by an additional 10 minutes at 72°C. Samples were separated by electrophoresis on an ethidium bromide–stained agarose gel.
In Vitro Luciferase Assay
To conduct the luciferase assay, 1 × 104 parental NFs and NF-effluc cells were seeded on 96-well white plates and cultured with Dulbecco's Modified Eagle's Medium containing 10% fetal bovine serum. After 24-hour incubation, 6 μL of D-luciferin (30 mg/mL; Biosynth International, Naperville, IL) was added to each well, and bioluminescence was measured using a microplate luminometer (Molecular Devices, Sunnyvale, CA).
In Vivo Luciferase Assay
For the in vivo imaging study, 2 × 105, 4 × 105, or 8 × 105 NF-effluc cells were subcutaneously inoculated into the left fore flank, right fore flank, and left hind flank of nude mice, respectively. BLI was performed the next day 10 minutes after an intraperitoneal injection of D-luciferin (3 mg/mouse) using the IVIS Lumina II imaging system (Xenogen, Alameda, CA).
NF-Effluc Cell Transplantation and the In Vivo Animal Model
All animal experiment protocols were approved by the Committee for the Handling and Use of Animals of Kyungpook National University. For hair reconstitution assays, cell mixtures (1 × 106 NF-effluc and 1 × 106 primary epithelial cells in a total volume of 100 μL) were injected into the hypodermis (right flank) of BALB/c nude mice (7-week-old females; SLC Inc., Hamamatsu, Japan) using a 25-gauge needle. Parent NF and epithelial cells of the same number and volume were injected into the left flank. Noninvasive BLI with D-luciferin was performed on days 1, 3, 7, 14, and 21 after cell transplantation. BLI was performed using the IVIS Lumina II imaging system. Grayscale photographic images and bioluminescent color images were superimposed using Living Image, version 2.12 (Xenogen), and IGOR image analysis FX software (WaveMetrics, Portland, OR). Bioluminescent signals were expressed in units of photons per cm2 per second per steradian (P/cm2/s/sr).
Light Microscopy and Immunohistochemistry
The skin sites at the injection of parent NFs and NF-effluc cells were harvested after 2 weeks, and hair follicle generation was observed by microscopic photography. Tissues that showed hair regeneration were removed from mice and were preserved in 10% formalin until required. All hair regeneration sites were embedded in paraffin and were sectioned at 5 μm. For immunohistologic analysis of expression of Thy1.1, the paraffin sections were incubated for 1 hour with PerCP mouse anti-CD90.1 antibody (BD Biosciences, San Jose, CA) at 1:200.
Statistical Analysis
All data are expressed as mean ± SD, and statistical significance was determined using an unpaired Student t-test; p values < .05 were considered significant.
Results
NF-Expressing Effluc
NFs were transfected with retrovirus containing the effluc gene and analyzed by flow cytometry. Following transfection, 70.9% of the cells were Thy1.1-PE-positive positive (named NF-effluc) and used for this experiment (Figure 1). RT-PCR showed that effluc messenger ribonucleic acid was expressed in NF-effluc cells but not in the parent NFs (Figure 2A).

Flow cytometry analysis demonstrated the enriched Thy1.1-PE+ cells (70.9%) in newborn mouse fibroblasts (NFs) with enhanced firefly luciferase (NF-effluc) but not in parent NFs (5.4%). PE =phycoerythrin.
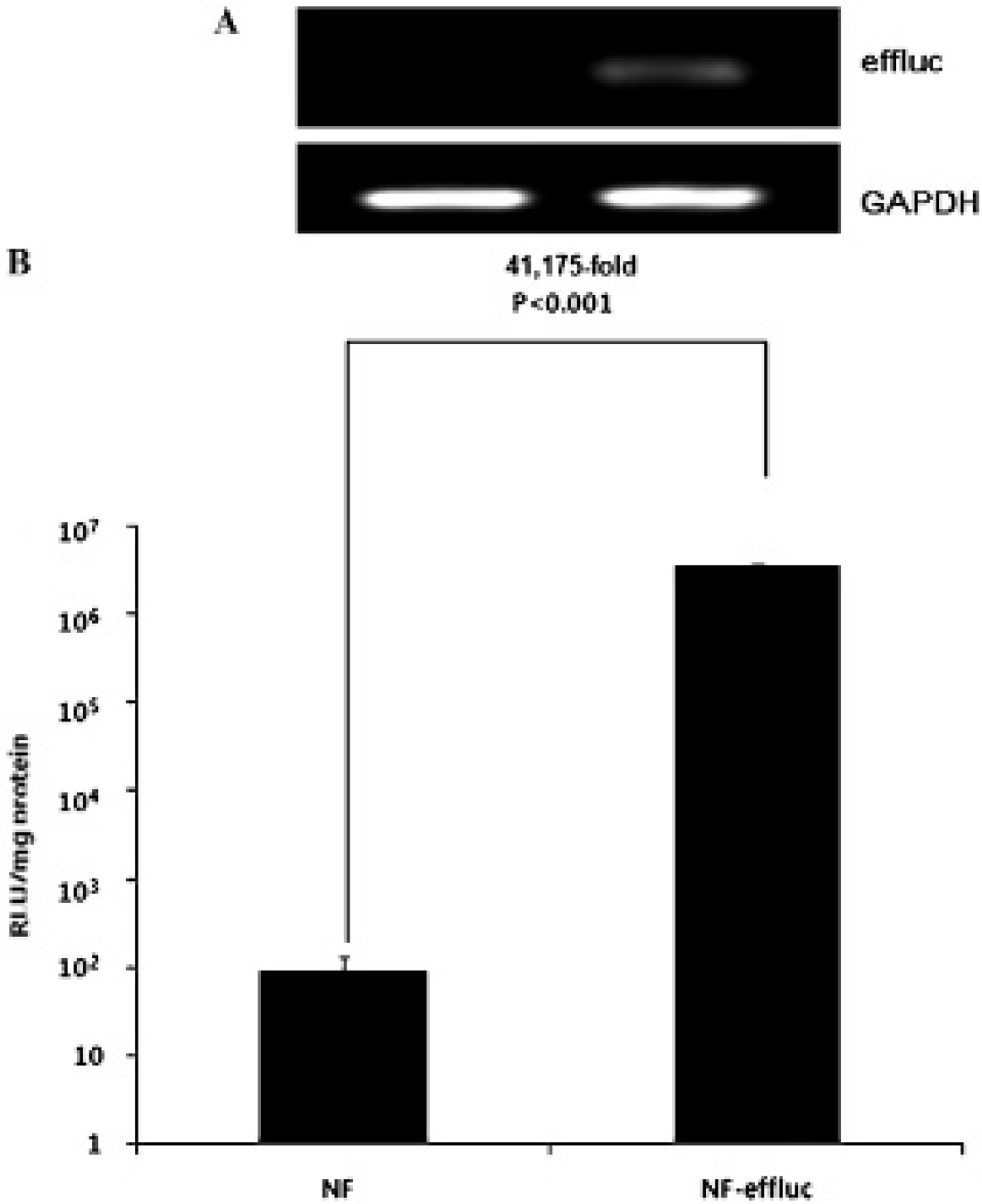
A, Reverse transcription–polymerase chain reaction analysis for enhanced firefly luciferase (effluc) in NF-effluc cells and parental newborn mouse fibroblasts (NFs). Only the NF-effluc cells demonstrated effluc expression. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as a housekeeping gene. B, Functional activity of the effluc protein in NF-effluc cells and parental NFs. The luciferase activity of NF-effluc cells was higher compared to that of parental NFs.
In Vitro Luciferase Assay
The luciferase activity of NF-effluc cells was 41,175-fold higher compared to that in untransfected parent NFs (3.6 × 106 ± 8.7 × 104 vs 89.1 ± 40.0 relative luminescence units/mg protein, p < .001) (Figure 2B).
In Vivo Luciferase Assay
BLI was acquired after inoculation of 2 × 105, 4 × 105, and 8 × 105 NF-effluc cells into the left fore flank, right fore flank, and left hind flank of nude mice, respectively (Figure 3A). BLI showed obvious signals at the inoculation sites (Figure 3B), and signal intensities were positively correlated with the number of inoculated NF-effluc cells (R2 = .9598) (Figure 3C).
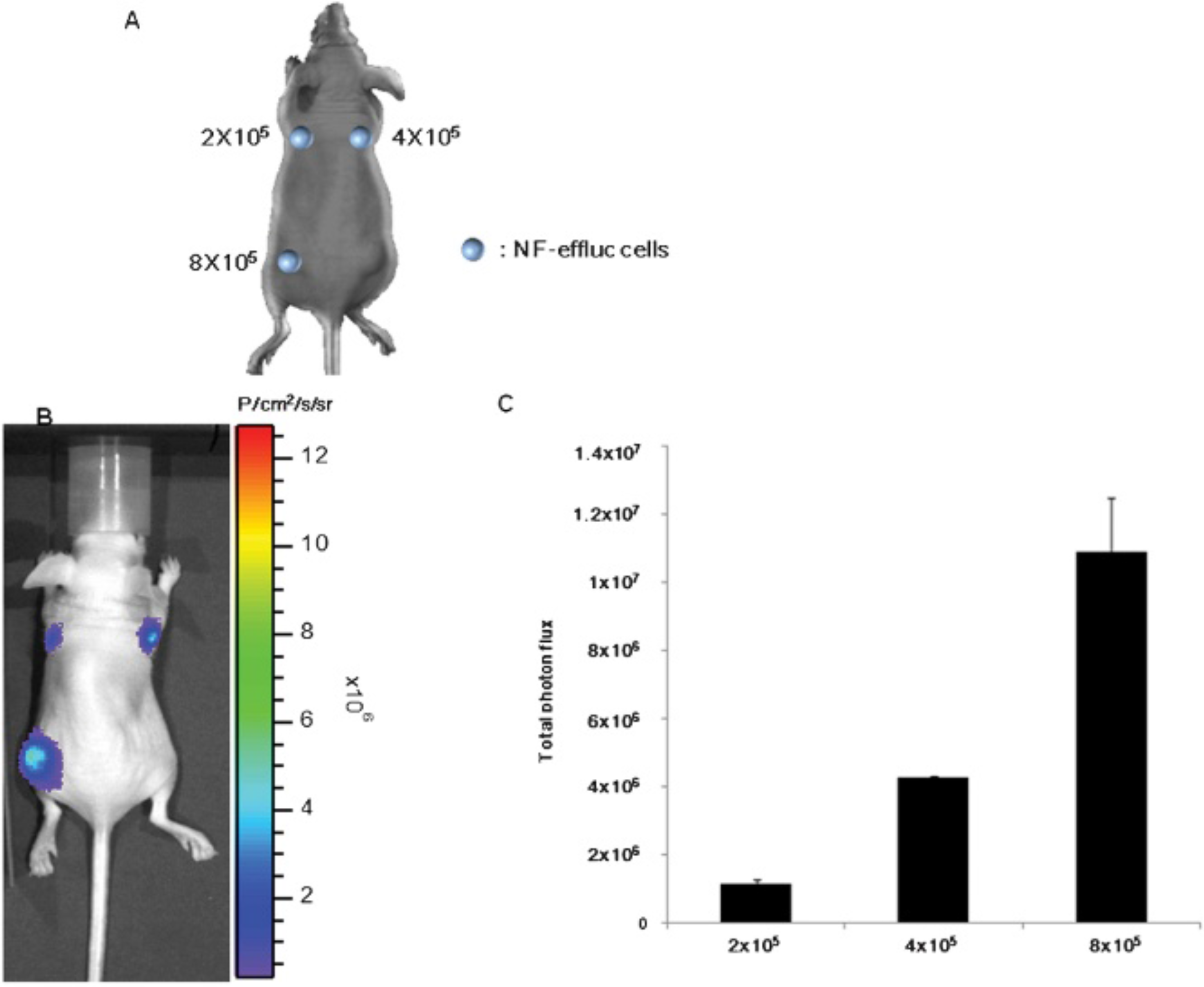
A, Bioluminescence imaging (BLI) was acquired after different numbers of NF-effluc cells were inoculated into the left fore flank, right fore flank, and left hind flank of nude mice, respectively. B, BLI showed obvious signals at the inoculation sites. C, In vivo bioluminescent signals positively correlated with the number of implanted newborn mouse fibroblasts (NFs) expressing enhanced firefly luciferase (NF-effluc).
Imaging NF-Effluc Survival in Living Mice
BLI was obtained on mice injected with the 1:1 cell mixtures of primary NF-effluc and primary epithelial cells (Figure 4A), and obvious signals were observed at the inoculation sites. Bioluminescence signals (Figure 4B) decreased gradually after the inoculation (day 1: 1.1 × 107 ± 1.3 × 106 P/cm2/s/sr; day 3: 4.7 × 106 ± 7.3 × 105 P/cm2/s/sr; day 7: 2.7 × 106 ± 5.9 × 105 P/cm2/s/sr; day 14: 1.1 × 106 ± 4.1 × 105 P/cm2/s/sr; day 21: 6.6 × 105 ± 7.8 × 104 P/cm2/s/sr).
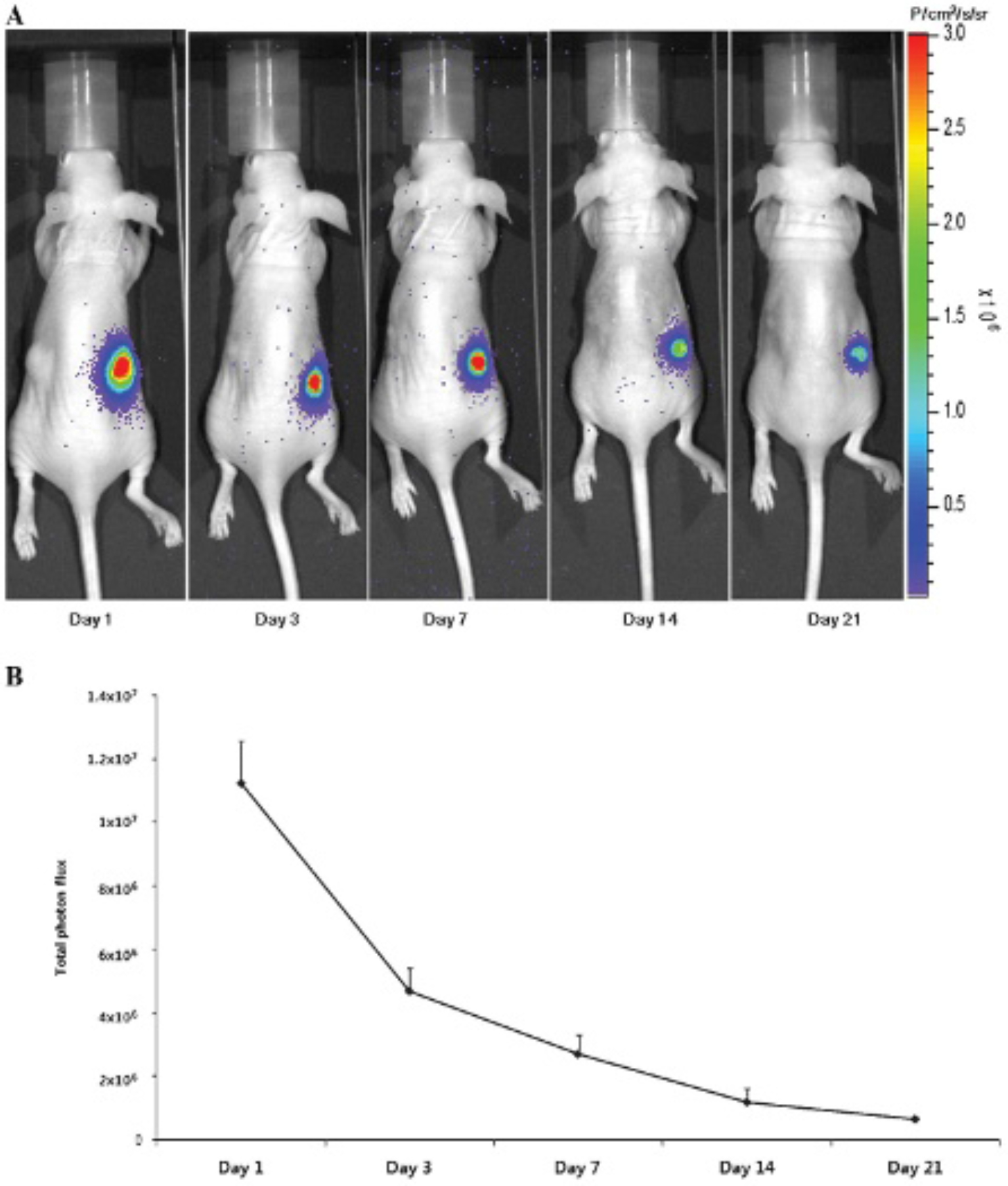
A, For the bioluminescence imaging study, cell mixtures (1 × 106 newborn mouse fibroblasts [NFs] expressing enhanced firefly luciferase [NF-effluc], 1 × 106 primary epithelial cells in a total volume of 100 μL) were inoculated into the hypodermis (on the right flank). Parent NFs and epithelial cells of the same number and volume were injected into the left flank. Bioluminescent imaging was acquired at 1, 3, 7, 14, and 21 days after the inoculation. NF-effluc-inoculated sites were well visualized following an injection of D-luciferin but not at parental NF inoculation sites. B, About 60% total photon flux decreased on day 3, 70% on day 7, 80% on day 14, and 90% on day 21 at the NF-effluc inoculation sites.
Photomicroscopy
Photomicroscopy of the hair reconstitution assay at day 14 confirmed abundant hair formation. Hair generation using a mixture of NF-effluc and epithelial cells was successful; however, less hair formation occurred compared to using a mixture of parent NF and epithelial cells (Figure 5). Immunohistochemical staining showed that CD90.1 staining (brown colors) was detected within the dermal papilla of regenerating hair follicles in a NF-effluc cell–transplanted mouse (Figure 6A, red circle). In contrast, there is no CD90.1 staining within the dermal papilla of regenerating hair follicles in the parental NF cell–transplanted mouse (Figure 6B, red circle).
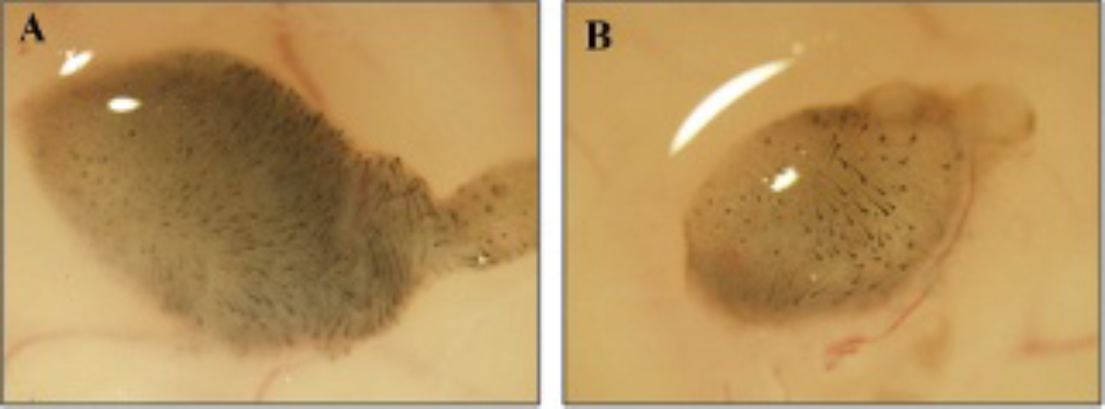
By microscopic evaluation, hair generation was demonstrated on day 14 following inoculation of newborn mouse fibroblasts (NFs) (A) or NFs expressing enhanced firefly luciferase (NF-effluc) (B). However, hair generation was sparser at the NF-effluc inoculation site compared to that at the NF inoculation site.
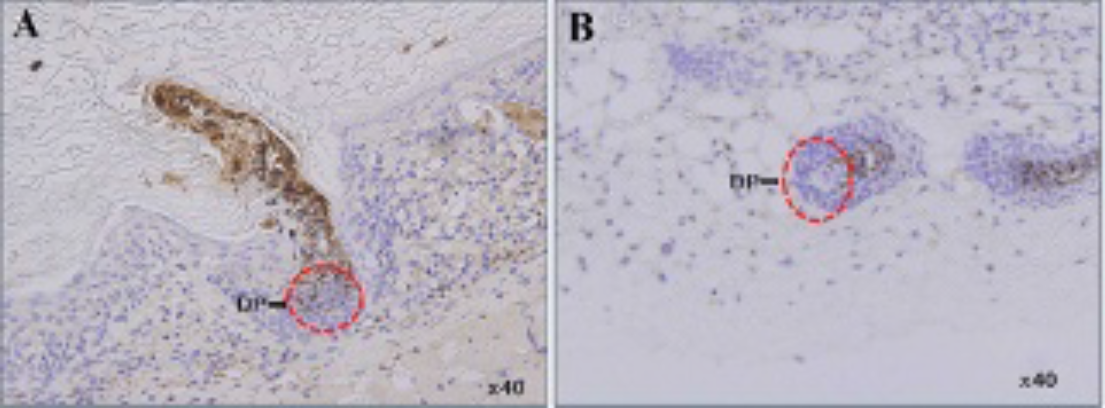
Immunohistochemical staining for analysis of Thy1.1 expression of implanted newborn mouse fibroblasts (NFs) expressing enhanced firefly luciferase (NF-effluc) in hair follicles. Tissues of hair follicles were obtained 2 weeks after implantation, and the tissue section was stained with PerCP mouse anti-CD90.1 antibody to visualize Thy1.1 expression. The red circle is the dermal papilla (DP) region of regenerating hair follicles. Brown signals were observed in the NF-effluc cells (A), but these signals were not seen in parent NFs (B).
Discussion
Many systems have attempted to form new hair follicles using hair stem cells.17–19 The hair follicle is composed of epidermal (epithelial) and dermal (mesenchymal) compartments, and their interaction plays an important role in the morphogenesis and growth of the hair follicle.20,21 Effective interaction between these two compartments is also thought to be a key process for successful generation of hair follicles. Several models have been established to study dermal–epidermal interactions to generate hair follicles. 22 Most hair follicle reconstitution assays are conducted in vivo in immunodeficient host mice using hair stem cell implantation. However, noninvasive monitoring of cells, which can provide better understanding of the in vivo fate of the implanted cells, has not been performed. The ideal imaging modality should have the characteristics of sensitive cell detection capability, accurate quantification of cell number, no dilution effect with cell proliferation, and noninvasive repetitiveness in living subjects. 23
One of the most commonly used strategies to monitor stem cells for in vivo imaging in living subjects is direct labeling.24,25 Direct cell labeling can be useful to evaluate initial cell distribution and localization as well as migration but cannot provide information about cell survival and proliferation because of persistent signals after the death of the labeled cells and signal dilution by mitotic division of the labeled cells.26,27 Direct cell-labeling protocols are easily applicable to different cell populations, and various methods have been described for in vivo detection of terminally differentiated, nonproliferating cells. When labeled cells do not proliferate, their distribution can be monitored during the physical half-life of the probe if they maintain the stability of the labeling. 28 In contrast, indirect cell labeling with reporter genes allows for accurate long-term monitoring of survival, proliferation, and migration of the labeled cells because reporter gene expression of the labeled cells is maintained in daughter cells but not in nonviable cells.29,30 Optical imaging with reporter genes is an effective means of in vivo stem cell monitoring in experimental models as it is rapid (<5 minutes), inexpensive, and noninvasive; provides single cell sensitivity; and does not involve radiation exposure.31,32 Conventional biologic methods for assessing new hair follicle formation from dissociated cells have the disadvantage of requiring postmortem immunohistologic staining, thus impeding temporal monitoring and demanding large animal numbers.
In recent years, optical imaging methods have provided new approaches for noninvasive real-time temporal monitoring of stem cell transplants allowing repetitive, longitudinal studies of the engraftment process. 33 The general limitations of optical imaging–based cell monitoring techniques include limited depth of penetration, limited quantification, and poor spatial resolution due to absorption and scatter of light. 9 However, these limitations might not pertain to hair follicle generation studies because they are usually performed on the dermoepidermal junction, just beneath the skin surface. One significant limitation related to reporter gene technique for in vivo cell monitoring could be that the reporter gene labeling (gene transduction) procedure could affect cellular traits; therefore, the biologic behaviors of labeled cells might be different from those of unlabeled parent cells.34,35 In this study, the ability to generate hair follicles in vivo was slightly impaired in NF-effluc cells compared to that in parent NFs. Development of reporter gene transduction strategies that provoke no or trivial biologic perturbation is warranted.
Hair follicles develop when dissociated dermal cells, isolated from newborn mouse skin, are injected into adult mouse truncal skin, yet very little is known about how they behave in vivo. We had achieved repetitive imaging of the hair stem cell in an in vivo animal model at different time points using bioluminescent optical imaging. The imaging was able to noninvasively visualize the fates of the implanted stem cells, including survival, death, and proliferation, which might be crucial information for a hair follicle reconstitution study. In this study, drastic reductions in signal intensity were observed within the first 4 days after NF transplantation, probably due to acute donor cell death. In a previous in vivo study of cell delivery and monitoring of stem cell viability, a drastic reduction in cell signal intensity was reported on day 1 or earlier due to acute donor cell death.36,37 It has been suggested that the dermal cells undergo apoptosis after triggering epidermal cells for hair follicle regeneration and the remaining few dermal cells actively participate in hair follicle generation. 38 In this study, the BLI signal decreased progressively over the following 3 weeks, suggesting the death of implanted dermal cells.
Many researchers have used recombinant retroviruses to achieve stable gene transfer into primary cells, 39 and once stable transfection of the cells is accomplished, the reporter gene product is detectable indefinitely in the cells, which allows for longitudinal monitoring of the cells for survival or proliferation.
Numerous molecular imaging studies related to tissue regeneration using stem cell therapy have been reported to overcome the limitations of conventional methodologies for in vivo monitoring stem cells; however, no reports are available of molecular imaging applied to visualize the viability or capability of generating hair follicles by engraftment of hair stem cells in an in vivo animal model. Bioluminescence molecular imaging might allow for a noninvasive examination of intact organs without sacrifice of experimental animals and provide consecutive imaging over a particular time period.
New hair follicle formation in embryonic skin and new hair growth in adult skin are initiated when specialized mesenchymal dermal papilla cells send cues to multipotent epithelial stem cells. 40 These findings establish a foundation for functional analyses of the roles of these genes in hair development. In the future, specific biologic activity of implanted stem cells can be investigated using another reporter gene strategy. A dermal papilla–specific promoter (eg, versican promoter) can be used to visualize differentiation of hair stem cells to dermal papilla cells in an in vivo animal model. When hair stem cells differentiate into dermal papilla cells that reconstitute the hair follicle, the specific promoter turns on in the cell and the reporter gene driven by the promoter becomes active and produces the reporter protein in the differentiated stem cells. High-throughput screening for agents or conditions that enhance hair follicle generation with appropriate differentiation of hair stem cell implantation is possible using the differentiation molecular imaging system.
Conclusion
We have established an NF-expressing bioluminescence reporter gene. Generating hair by forming follicles with NFs was feasibly and noninvasively monitored using an optical reporter gene imaging strategy in an in vivo animal model.
Footnotes
Acknowledgment
Financial disclosure of authors: This work was supported by grants (A102132, A111345) from the Korea Health Technology R&D Project, Ministry of Health & Welfare, Republic of Korea; a grant from the National Nuclear R&D Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (No.2012M2A2A7014020); a grant from the National Research Foundation of Korea (NRF) funded by the Korean government (MEST) (No. 2012-0004878, 2012-0004879).
Financial disclosure of reviewers: None reported.
