Abstract
The animal model Caenorhabditis elegans was employed to study polyphenol- and humic substances-induced hormetic changes in lifespan. A detailed insight into the underlying mechanism of hormesis was uncovered by applying whole genome DNA microarray experimentation over a range of quercetin (Q), tannic acid (TA), and humic substances (HuminFeed®, HF) concentrations. The transcriptional response to all exposures followed a non-linear mode which highlighted differential signaling and metabolic pathways. While low Q concentrations regulated processes improving the health of the nematodes, higher concentrations extended lifespan and modulated substantially the global transcriptional response. Over-represented transcripts were notably part of the biotransformation process: enhanced catabolism of toxic intermediates possibly contributes to the lifespan extension. The regulation of transcription, Dauer entry, and nucleosome suggests the presence of distinct exposure dependent differences in transcription and signaling pathways. TA- and HF-mediated transcript expression patterns were overall similar to each other, but changed across the concentration range indicating that their transcriptional dynamics are complex and cannot be attributed to a simple adaptive response. In contrast, Q-mediated hormesis was well aligned to fit the definition of an adaptive response. Simple molecules are more likely to induce an adaptive response than more complex molecules.
Keywords
INTRODUCTION
In previous experiments, we identified that Caenorhabditis elegans (Maupas, 1900) actively migrates into some unfavorable environments and in doing so suffers from symptoms of stress such as loss of energy, activation of oxygen, and induction of stress genes (Menzel et al. 2005; Steinberg et al. 2007; Menzel et al. 2011). Many of these responses are transcriptionally controlled (Menzel et al. 2005, 2012) and demand an increase in energy levels (Cazenave et al. 2006). However, the animals benefit from this seemingly absurd behavior: exposure to low or intermediate concentrations of humic substances, which were shown to be stressors, extends their lifespan significantly, whereas high concentrations did not differ from the control (Steinberg et al. 2007; Menzel et al. 2011). Hence, by definition, this dose-response relationship is hormetic in nature. In C. elegans longevity can be induced via several chemical and physical triggers, which supports the notion that this nematode is ideally positioned to study hormetic effects (Yashin et al. 2001; Cypser and Johnson 2002; Hartwig et al. 2009; Le Bourg 2009).
One central paradigm of hormesis is that the “adaptive response” is characterized by biphasic dose responses that display analogous quantitative features such as the amplitude and range of the stimulatory response (which are either directly induced or the result of compensatory biological processes following an initial disruption in homeostasis) (Calabrese and Baldwin 2002). However several recent papers (reviewed by Steinberg et al. (2008)) have challenged this paradigm and suggest that low and high dose exposure are characterized by different pathways. In order to identify the major regulatory and metabolic pathways, we compared the whole-genome transcriptomes of nematodes exposed to three different polyphenolic substrates with increasing complexity at the molecular and structural level, namely quercetin, tannic acid, and humic substances (HuminFeed®, HF), a commercial humic preparation. Initially, we defined the differentially expressed genes (DEGs), many of which displayed concentration dependent changes in expression. Subsequent investigations included gene ontology (GO; Ashburner et al. (2000)) and Kyoto encyclopedia of genes and genomes (KEGG; Kanehisa (2002)) pathway analyses. In concert, these bioinformatic pipelines allowed the identification of gene classes and pathways which are significant over-representations in the treated nematodes.
This paper relies predominantly on published data (Menzel et al. 2012; Pietsch et al. 2012), but focuses in particular on the identification of putative mechanisms underlying hormetic dose-responses, an aspect that was previously not covered in depth. Overall, this study provides evidence that polyphenols and HSs induce longevity via a hormetic mode of action. Moreover, polyphenols and HSs induce distinct pathways at different concentrations which reject, at least in part, the notion that the adaptive response is the major underlying mechanism.
MATERIAL AND METHODS
The methods are described in full in Pietsch et al. (2012) and Menzel et al. (2012). Here, we provide only a brief overview.
Nematodes
N2 wild type C. elegans were maintained on nematode growth medium (NGM) plates using Escherichia coli OP50 as a food source (Brenner 1974). Approximately 10,000–15,000 untreated larval nematodes per concentration and trial were transferred onto control and treatment plates (50, 100, and 200 μM Q; 100, 200, and 300 μM TA; 200 and 2,000 μM HF as dissolved organic carbon, DOC) and incubated at 20°C for 4 days for the polyphenols and, for HF, at 25 °C to the young adult stage (3 days) or an older adult stage (11 days). Nematodes were fed every third day by replenishing the bacterial suspension and a fresh preparation of humic substances. Each condition was cultivated in triplicate.
RNA preparation
Total RNA of each individual condition (n = 3) was isolated using an innuSPEED issue RNA Kit (AnalytikJena, Jena, Germany), which included an improved homogenization step with a Speed-Mill (AnalytikJena, Jena, Germany) and the removal of genomic DNA through an initial spin filter column step. The RNA quality and quantity was analyzed spectroscopically (NanoDrop 1000, ThermoScientific, UK) and by means of Agilent's Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA). All RNA-samples showed no signs of degradation (RNA integrity numbers ≥ 7.5).
DNA microarrays
The processing of each RNA sample was performed with a MessageAmp™Premier RNA Amplification Kit (Ambion, Austin, TX, USA). We used the GeneChip® C. elegans genome platform (Affymetrix, Santa Clara, CA, USA), representing 22,548 different transcripts. To conduct the microarray hybridization experiments, we followed the specifications from Affymetrix's GeneChip® kit. The scans were conducted by means of a GeneChip® scanner 3000 7G (Affymetrix). Triplicate GeneChips® were run for each condition.
Data interpretation and statistical analysis
Pre-processing of DNA microarray raw data included probe-specific background correction, summarization of probe set values, and normalization using the GCRMA algorithm with CARMAweb 1.4, an R- and Bioconductor-based web service for microarray data analysis (Rainer et al. 2006). For selection of DEGs, an unpaired t-test was performed followed by a significance analysis of microarray test including a calculation that estimates the false discovery rate (FDR), which was set to a non-stringent level of <12.5%. DEGs exceeding a fold change of 1.25 and returning a p-value (t-test) below 0.05 were further analyzed with respect to their functional clustering (Grigoryev et al. 2004). Only the genes, which fulfill these criteria (fold change ≥1.25, p-value ≤ 0.05, and FDR <12.5%) were included in the analysis. This analysis was conducted using the functional annotation clustering tool of the Database for Annotation, Visualization, and Integrated Discovery (DAVID; Huang et al. (2007)). The resultant annotation clusters were ranked according to the statistical significance of cluster enrichment. The microarray data is MIAME compliant and has been deposited in the National Center for Biotechnology Information's Gene Expression Omnibus, GEO (http://www.ncbi.nlm.nih.gov/geo, accession numbers GSE35354 and GSE35360).
Representation factor
Due to huge sample numbers, HF-treatments were run in two different series. To assess the level of overlap between different conditions, we calculated the representation factor (RF) in order to explore the fold enrichment. The RF identifies the level of enrichment (of individual transcripts) between gene lists (Kim et al. 2001). RF were considered significant when RF>1.
Validation of DNA microarray data by qRT-PCR
qRT-PCR analyses were conducted with samples from control, 200 μM quercetin and 200 μM tannic acid treated nematodes. The qRT-PCR validation for the HuminFeed® microarray was already published (Menzel et al., 2012). The RNA was isolated as described previously and was then reverse transcribed into cDNA with the aid of the M-MLV (Moloney Murine Leukemia Virus) reverse transcriptase (Promega, USA). Quantitative real-time amplification was performed in a MyiQ single color qPCR detection system (BIO-RAD, Germany) using the double-stranded DNA intercalating fluorescent agent EvaGreen for amplicon detection. Each reaction consisted of the qPCR Green Core Kit (Jena Bioscience, Germany), 200 nM of each primer pair, and cDNA template equivalent to 5 ng RNA starting material. The relative expression of the target genes was calculated by means of the comparative 2−ΔΔCT method (Livak and Schmittgen 2001) and act-1 and cdc-42 were used as reference genes. The list of primers is given in Supplementary File 2 as well as the validation of the reliability of the microarray platform via qRT-PCR.
Polyphenols and humic substances
Tannic acid (TA) and quercetin (Q) were purchased from Sigma-Aldrich, Taufkirchen, Germany. HuminFeed® (HF, Humintech GmbH, Düsseldorf, Germany) was made by an alkaline extraction process of highly oxidized lignite (for a detailed and comparative physicochemical analysis see (Meinelt et al. 2007)). HF was used for practical reasons and by no means as an advertisement for this product. Our experiments used the same HF batch as previously chemically characterized by Meinelt et al. (2007). TA, Q and HF were added to the NGM and the OP50 bacteria with final concentrations ranging between 25 and 2,000 μM. The final concentrations of each substance were chosen according to its solubility, to published information and/or preliminary tests performed in our laboratory. Equal amounts of solvent (final concentration of 0.3% [v/v] DMSO; Applichem, Darmstadt, Germany) were used in all conditions except HF.
RESULTS
Quercetin
Quercetin-mediated mean and median lifespan changes are shown in Figure 1: while 50 μM Q evoked no significant change in lifespan, 100 and 200 μM Q prolonged the mean lifespan by approximately 10%. Higher concentrations (250 μM Q) reduced the lifespan by 7% and therefore can be considered to be toxic. Thus, we utilized two lifespan prolonging concentrations (100 and 200 μM), and a non-effective (50 μM) pre-longevity dose to study the global transcriptomic profile of Q exposure.

Concentration-dependent variation of mean and median lifespan quercetin, tannic acid and HuminFeed® were tested for their ability to modulate the lifespan of C. elegans at 20°C. The percentage variations of the mean (upper row) as well as the median (lower row) lifespan compared to control are illustrated. The error bars represent the SEM, and differences in the mean lifespan were considered significant at ∗ p < 0.05 and ∗∗ p < 0.005, respectively (log-rank test). Each data point represents the average of at least 2 independent trials with a total of at least 171 nematodes per concentration. Further details are presented in the Supplementary File 1. The data are modified from Pietsch et al. (2011, with permission from Springer), Saul et al. (2011, with permission from the American Chemical Society), and Steinberg et al. (2007, courtesy of Northeastern University, Boston, MA).
Venn diagrams (Figure 2) summarize the level of overlap between doses. The number of DEGs increased markedly at the lifespan modulating doses of 100 and 200 μM. At 50 μM Q exposure, more genes were up-regulated than down-regulated, but the opposite was observed at 100 and 200 μM. The majority of DEGs overlapped in 100 and 200 μM exposures, thus suggesting the presence of common response pathways. DEGs in 50 μM-treated nematodes differed strongly from those in the 100 and 200 μM treatments. In contrast, the expression profiles in response to the two latter concentrations resembled each other in identity and magnitude. The doubling in dose (from 50 to the 100 μM) increased the number of DEGs by a factor of 3.3, which was further increased by a marginal 1.1 fold in the 200 μM-treatment. A further discrepancy between the non-longevity provoking concentration 50 μM and both longevity-mediating doses was the proportion of up- and down-regulated transcripts: three times more genes were up-regulated than down-regulated at 50 μM, a ratio that was reversed in the higher concentrations. Furthermore, the high percentage (37 %) of transcripts exclusively regulated at 50 μM suggests the presence of a distinct mode of action, possibly leading to the observed variance in life extension.

Over-represented genes. The Venn diagrams show the overlap of significantly up (left) -or down (right) -regulated genes (fold change >1.25 or <0.8) per concentration in nematodes exposed to quercetin (modified from (Pietsch et al. 2012), courtesy of Frontiers in Genetics).
Classifying DEGs by GO-term analyses (Figure 3), highlighted that over-represented GO-terms derived from the up-regulated transcripts are shared between 100 and 200 μM exposures and included chromatin assembly, lipid metabolic processes, monooxygenase, and nucleosome activities. GO-terms from down-regulated genes comprised nervous system development, regulation of multicellular organism growth, Dauer entry, regulation -of transcription, -of response to stimulus, -of cell communication, -of biological quality, -of locomotion, and -of programmed cell death.

Overview of significant over-represented GO-terms from differentially expressed genes (DEG) following exposure to quercetin. Shown is the fold change for DEGs in the respective list in relation to the overall count in significantly enriched GO-terms. Pale and deep colors correspond to different cut-offs.
KEGG-analysis returned a multitude of pathways (Figure 4). Over-represented KEGGs in 50μM Q included pathways involved in the metabolism of amino acids, glutathione, and xenobiotics as well as fatty acid elongation. At higher exposures seven (100 μM) and eight (200 μM) KEGG-pathways were found to be over-represented, notably most (six) were present at both concentrations (Figure 4) with an analogous mode/direction of regulation, i.e., either repressed or induced. Pathways include metabolism of amino acids, xenobiotics or drugs, transport processes (lysosome), and signal transduction processing (Wnt and transforming growth factor-β (TGF-β signaling). DEGs from amino acid metabolism and the lysosome displayed heterogeneous expression levels, transcripts belonging to the signaling pathways were predominantly repressed; in contrast almost all genes associated with drug/xenobiotic metabolism were found to be induced.
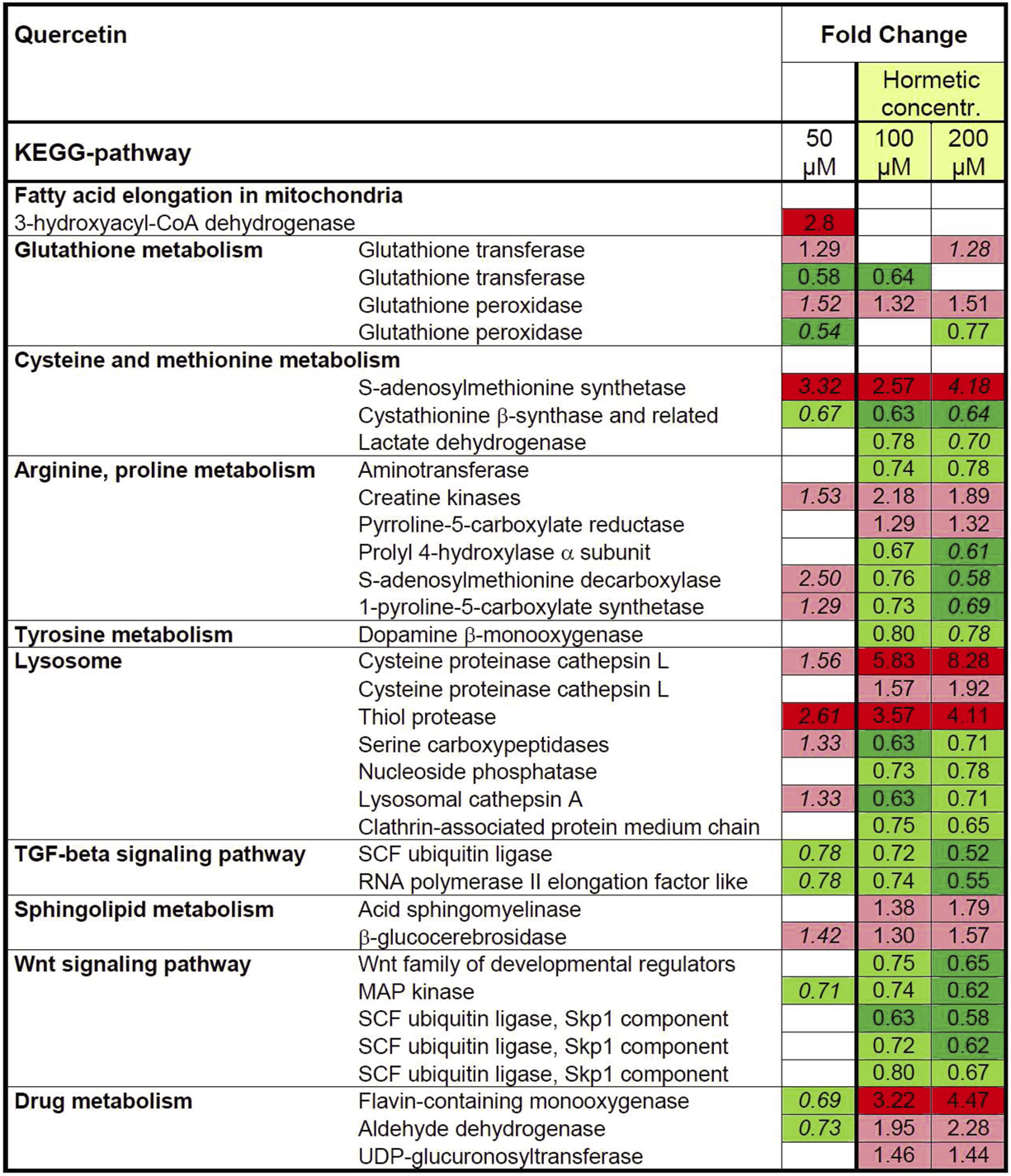
Overview of significant over-represented KEGG terms from differentially expressed genes (DEG) following exposure to quercetin. Shown is the fold change for DEGs in the respective list in relation to the overall count in significantly enriched KEGG terms. Repetition of gene names indicates the presence of different loci. Red and green labels define up- and down-regulated genes; respective pale and deep colors correspond to different cut-offs. The numbers in italics show the Fold Changes values of 50 μM Q, however lacking statistical robustness.
Tannic acid
For TA, the most effective concentration in terms of lifespan extension was 100 μM, which resulted in an increase of 18%, an effect that was still significant but notably less pronounced at 200 μM (8%) and was absent at 300 μM (Saul et al. 2010). 400 μM reduced the mean lifespan (Figure 1). Hence, we used two life-extending concentrations (100 and 200 μM), and in addition one post-effective concentration (300 μM).
As with Q, exposure to TA resulted in a dose dependent increase in DEGs which was most pronounced at the highest concentration tested (Figure 5), the dose that did not affect longevity. More genes were up-regulated than down-regulated in the lifespan-prolonging exposures 100 and 200 μM, a ratio that was reversed at 300 μM where more genes were down-regulated (Figure 5). Striking was the proportion of DEGs that responded exclusively to 300 μM (61%), a substantial proportion compared to 100 μM (2%) or 200 μM (6%).

Over-represented genes. The Venn diagrams show the overlap of significantly up (left) -or down (right)-regulated genes (fold change >1.25 or <0.8) per concentration in nematodes exposed to tannic acid (modified from (Pietsch et al. 2012), courtesy of Frontiers in Genetics).
In longevity-inducing TA exposures, significant GO-terms were linked to muscle contraction, neurotransmitter transporter activity and cytoskeleton, embryonic development ending in birth or egg hatching, positive regulation of biological process, and chromatin (Figure 6). The dynamic response patterns of Q exposure were not observed in the TA treatments. Although several genes were modulated, no KEGG-pathways could be linked to the most potent lifespan extending concentration, 100 μM TA (Figure 7). For 200 μM TA, only two KEGG-pathways were found to be over represented, both derived from amino acid metabolism. Analysis of the 300 μM TA gene list returned six KEGG-pathways, notably DNA replication, mismatch repair, and Ubiquitin-mediated proteolysis.

Overview of significant over-represented GO-terms from differentially expressed genes (DEG) following exposure to tannic acid. Shown is the fold change for DEGs in the respective list in relation to the overall count in significantly enriched GO-terms. Pale and deep colors correspond to different cut-offs.
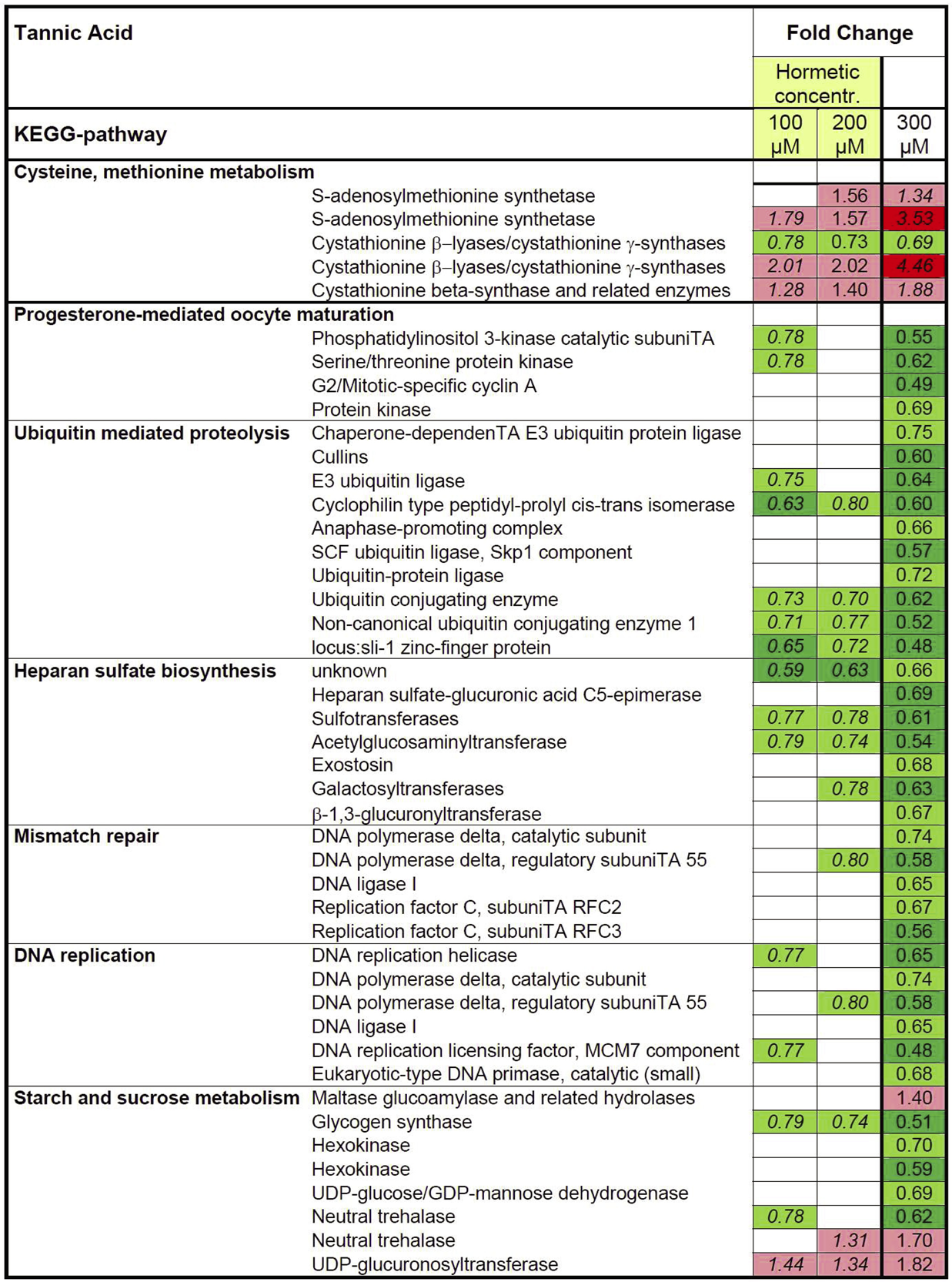
Overview of significant over-represented KEGG terms from differentially expressed genes (DEG) following exposure to tannic acid. Shown is the fold change for DEGs in the respective list in relation to the overall count in significantly enriched KEGG terms. Repetition of gene names indicates the presence of different loci. Red and green labels define up- and down-regulated genes; respective pale and deep colors correspond to different cut-offs. The numbers in italics show the Fold Changes values, however possibly lacking statistical robustness.
Humic substances
Previously, we were able to show that low HF concentrations (up to 800 μM) prolonged the lifespan of C. elegans, whereas exposure to 2,000 μM HF resembled control conditions (Steinberg et al. 2007) (Figure 1). This non-linear dose-response relationship therefore lacked concentrations that exert adverse effects, however, toxic effects induced by high doses of HSs have been documented in the past (Cheng et al. 2003; Bittner et al. 2006; Cazenave et al. 2006; Matsuo et al. 2006; Andersson et al. 2010). Our global transcriptomics includes lifespan extending (200 μM DOC) and non-lifespan extending (2,000 μM DOC) concentrations.
The Venn diagrams (Figure 8) show that, due to the low threshold (a minimum fold change in transcript expression of 1.25); many genes were classified as DEGs, especially in young adults. However, the extended incubation time of 11 days was characterized by a substantial decline in the number of DEGs, in particular down-regulated genes. An extensive overlap was observed between the two concentrations (Figure 8).

Over-represented genes. The Venn diagrams show the overlap of significantly up- or down-regulated genes (fold change >1.25 or <0.8) per concentration in nematodes exposed for 3 days (left) or 11 days to HuminFeed® (right) (modified from (Menzel et al. 2012), courtesy of Frontiers in Genetics).
The GO-analysis produced a multitude of significantly enriched terms, many represented by the same genes across and within the three GO-domains: biological process, cellular component, and molecular function. A selection of the 25 most striking terms are given in Figure 9. Young adult nematodes were much more responsive than old adults and their response was also more consistent. Lysosomal processes, defense response as well as lipid and fatty acid metabolism were found to be enriched in the group of up-regulated genes. Moreover, HF induced the expression of genes coding for constituents of the cuticle and cytoskeleton. The persistently strong induction of the cellular components pseudopodium and extracellular region is caused by a comprehensive up-regulation of various major sperm proteins (MSP). HF may contribute to slowing down the reproductive development of C. elegans.

Overview of significant over-represented GO-terms from differentially expressed genes (DEG) following exposure to humic substances. Shown is the representation factor for DEGs in the respective list in relation to the overall count in significantly enriched GO-terms. Red and green labels define up- and down-regulated genes; respective pale coloring corresponds to p<0.05, deep colors to p<0.001 (from Menzel et al. (2012), courtesy of Frontiers in Genetics).
In young adults, HF exposure at 200 μM DOC induced 10 KEGG pathways and 16 KEGG pathways at 2,000 μM DOC (Figure 10). Particularly noticeable was the induction of fatty acid metabolism, in particular arachidonic acid (AA) and sphingolipid metabolism, as well as the up-regulation of lysosome related genes. In the 11 day old HF-exposed worms, only four significantly enriched KEGG pathways were modulated, which were limited to the lower concentration of HF (200 μM DOC). Besides the persistent induction of the AA metabolism, HF (200 μM DOC) was marked by a pronounced up-regulation of the biotransformation machinery, which includes glutathione and cytochrome P450 (CYP) pathways. It should be noted that HF (2,000 μM DOC) also induced biotransformation associated genes, however, because of the large number of DEGs, they were not found to be significantly enriched. The validation of the microarray via qRT-PCR is shown in Menzel et al. (2012).
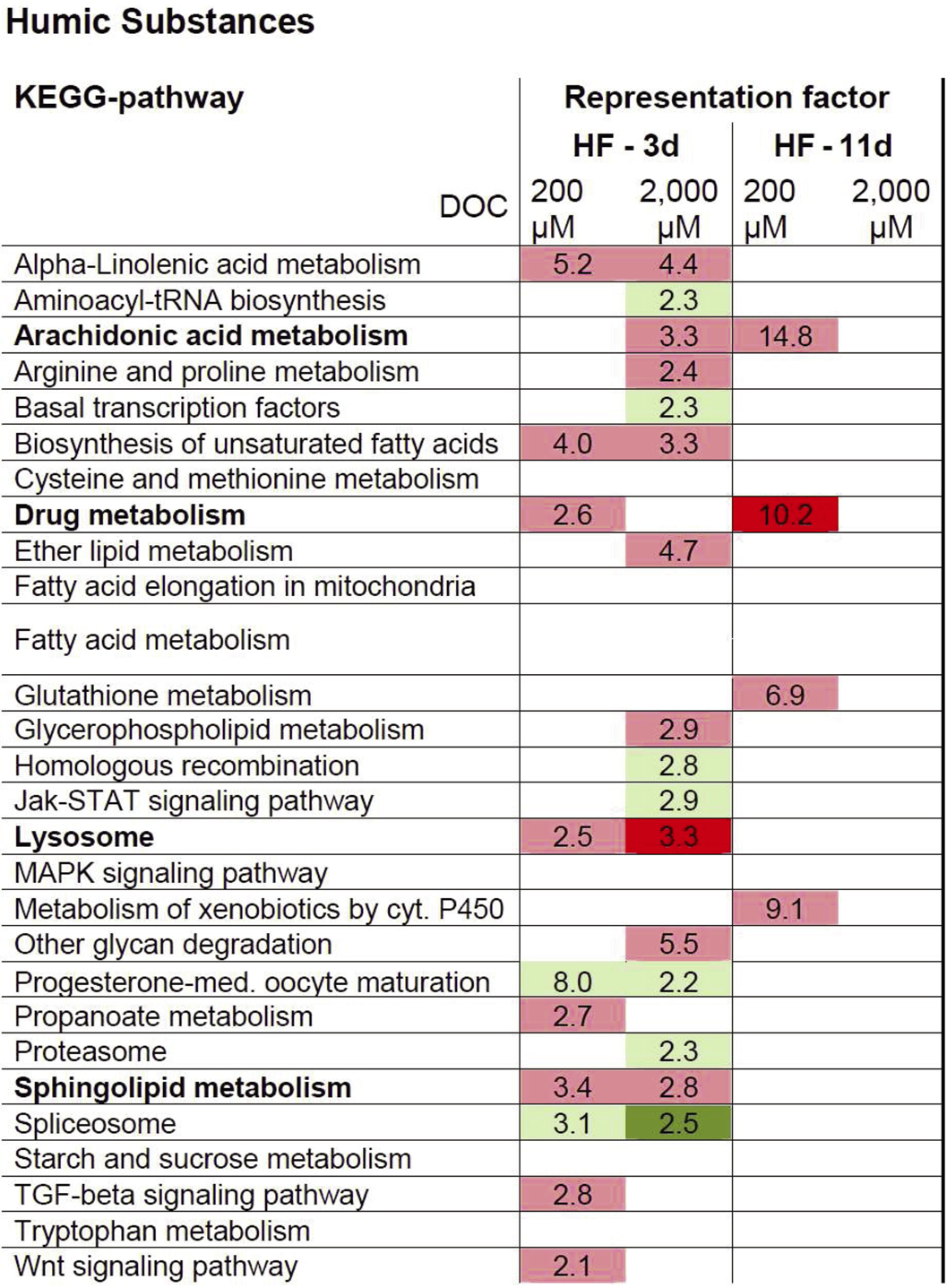
Overview of significant over-represented KEGG terms from differentially expressed genes (DEG) following exposure to humic substances. Shown is the representation factor for DEGs in the respective list in relation to the overall count in significantly enriched KEGG terms. Red and green labels define up- and down-regulated genes; respective pale coloring corresponds to p<0.05, deep colors to p<0.001 (from Menzel et al. (2012), courtesy of Frontiers in Genetics).
DISCUSSION
It remains to be confirmed whether hormesis is the direct result of an adaptive response. It is assumed that hormesis is characterized by similar quantitative features with respect to amplitude and range of the stimulatory response that are either directly induced or the result of compensatory biological processes following an initial disruption in homeostasis (Calabrese and Baldwin 2002). The term ‘adaptive response’ implies that low- and high-dose exposures should activate analogous response or defense pathways and in consequence the low-dose exposure trains the defense systems to cope with future potentially adverse exposures. Differential gene expression analysis may prove a promising tool to evaluate this descriptive hypothesis. Indeed, there is an increasing body of evidence that transcript expression profiles of low-dose exposures differ from higher doses, including studies on radiation (Ding et al. 2005; Sokolov et al. 2006; Abou-El-Ardat et al. 2012), gravity (Allen et al. 2007), or chemicals. For instance, Toyoshiba et al. (2006) analyzed low and high-dose effects of acetaminophen (paracetamol) administered to rats. The authors identified two gene interaction networks clearly segregated by the two doses: at lower doses, oxidative stress signaling pathways did not interact with apoptosis-related genes, but did in the higher doses. In a more environmentally relevant context, Gong et al. (2007a, b) uncovered a hormetic increase in reproduction in earthworms exposed to trinitrotoluene. This effect was mirrored by specific transcriptional responses at the low-dose exposure. Recently, Celorio-Mancera et al. (2011) showed a transcription based hormetic response in a generalist herbivore challenged by the toxic plant secondary metabolite gossypol: at low levels, genes involved in energy acquisition (such as β-fructofuranosidases) were up-regulated in the gut, and genes involved in cell adhesion were down-regulated in the body. High levels of gossypol affected transcripts predicted to be integral to the membrane or associated with the proteasome core complex. Oxidoreductase activity-related genes were observed to be significantly altered in both tissues at the highest gossypol dose. The authors concluded that hormesis may be due to a lower induction threshold of growth-promoting, stress-coping responses and a higher induction threshold of detoxification pathways that are costly and cause collateral damage to the cell. These examples indicate that hormesis is not limited to a simple adaptive response but accompanied by a complex transcriptional activation of interwoven pathways. In line with this argument is the recent plea by Piña and Barata (2011) to increase our understanding of transcriptomic responses to high and low doses.
We were able to show for three natural xenobiotics (for terminology, see also Li et al. (2007)) that all act in a hormetic fashion concerning lifespan in C. elegans and, in two of the three chemicals, low concentrations induce pathways that are distinct from those activated at high concentrations, findings which do not support the adaptive response theory. However, Q-mediated longevity modulation does align well with the adaptive response mode (Figures 3, 4). In detail: The number of DEG in 50, 100, and 200 μM Q suggests that the action of Q in C. elegans follows a non-linear mode: low concentrations (50 μM) regulate processes which potentially improve the health of the nematodes (e.g. the gene classes gsts, peroxidases, lysozymes; GO-term: oxidoreductase activity; KEGG pathway: glutathione metabolism and metabolism of xenobiotics by cytochrome P450). In consequence, though insufficient to entend lifespan, 50 μM Q induces enhanced thermal stress resistance (Pietsch 2011). Concentrations of 100 and 200 μM Q promote a longevity phenotype, and the global transcriptional response of the worms changed substantially. The over-representation of DEGs related to biotransformation (gene-classes: gsts, ugts, cyps; KEGG pathways: drug- and xenobiotic metabolism) indicate that enhanced catabolism of toxic intermediates contributes to the lifespan extension. Thus, Q action resembles predictions made by the green theory of aging (Gems and McElwee 2005), and is further supported by an additional enrichment of hsp expression levels in 200 μM-treatment (Pietsch 2011). However, it is likely that further mechanisms contribute to Q induced longevity, as indicated by the over-representation of pdz genes, transcription factors and vitellogenins. The GO-term (regulation of transcription, Dauer entry, nucleosome and others) and KEGG pathway (Wnt, TGF-β) results suggest the presence of extensive alterations in signaling pathways and transcription. Moreover, the alterations of the amino acid metabolism and certain degradation processes in the lysosome may contribute to the longevity phenotype. Overall, both hormetic Q concentrations appear to induce common basic longevity-controlling pathways; thus, Q longevity modulation fits best to the concept described as the adaptive response.
With TA, longevity is possibly linked to the amino acid metabolism and modulated by the TGF-β and the p38 MAPK pathways. DAF-12, which integrates TGF-β and Insulin-like downstream signaling, and genetic players of the p38 MAPK pathway therefore may constitute crucial regulators. Although some similarities in the longevity-controlling pathways between Q and TA prevaile, the lowest TA concentration (100 μM) with the highest lifespan extension does not display any distinct pathways, neither by GO-term nor by KEGG-pathway evaluation. Consequently, one may question whether this hormetic response really reflects an adaptive response.
The transcriptional profiles of HF most closely resembles the results from TA-treated nematodes. However, a significant proportion of genes up-regulated after 3 days are down-regulated after 11 days. This result demonstrates the dynamic nature of the transcriptome during the HF-mediated aging process. Despite the observed similarities, the mode of action of HF is complex and heavily influenced by the negative regulation of TGF-β signaling as well as a predicted increase in lysosomal activity. At the lower concentration (200 μM DOC), HF regulates processes which potentially promoted growth and improved health of the nematodes (e.g. lysozymes, cuticle development; GO-term: oxidoreductase activity; KEGG pathway: α-linolenic acid metabolism, drug metabolism, TGF-β signaling pathway; progesterone-mediated oocyte maturation) and thereby likely induces longevity, however with a delay in reproductive development. At the higher concentration (2,000 μM DOC), longevity is not observed but the global transcriptional response changes. Over-represented DEGs can be linked to lipid, protein, glycan degradation, and ATP decomposition (GO-term: lipase, ceramidase, and ATPase activity; KEGG pathways: proteasome, glycan degradation) indicating that the integrity of the body is challenged.
CONCLUSION
The dose-response of hormesis has been observed in a diverse range of biological systems (Calabrese and Baldwin 2003; Calabrese 2005; Calabrese and Blain 2011; Calabrese and Mattson 2011). Hence, it is unlikely that one single hypothesis is universally applicable (Celorio-Mancera et al. 2011). By exploring three natural xenobiotics within the model organism C. elegans, we show that hormesis emerges as one of two types of distinct and specific transcriptional responses to chemically-mediated stress. The occurrence of an adaptive response is seemingly dependent on the molecular characteristics of the chemical. Simple molecules, such as quercetin are more likely to induce an adaptive response than more complex molecules, such as tannic acid or humic substances.
Footnotes
ACKNOWLEDGMENTS
We gratefully acknowledge supports from Deutsche Forschungsgemeinschaft (grants STE 673/16 and STE 673/18) and we thank two anonymous reviewers for their constructive comments. Finally, we declare that there is no conflict of interests and that experiments comply with the current laws of the countries (Germany, U.K.) in which the experiments were conducted.
