Abstract
The purpose of this study was to determine and characterize the biodegradation of reactive dyes on cotton jersey fabrics buried in soil. Four commonly-used reactive dyes, C.I. Reactive Black 5 (RBlk5), C.I. Reactive Red 198 (RR198), C.I. Reactive Blue 49 (RB49), and C.I. Reactive Orange 35 (RO35), were used in this study. Degradation products were characterized and proposed based on high-performance liquid chromatography (HPLC) coupled with high-resolution mass spectrometry (HRMS). A chemical method and an enzymatic digestion were developed and used to remove the reactive dye from the control and degraded cotton fabrics. Hydrolyzed reactive dyes and reactive dyes having cellobiose units were synthesized and used as standards for comparison in this study. The possible degradation mechanism pathways of the reactive dyes bonded to cotton fabrics depends on the chemical structure of each dye.
Introduction
Nowadays, environmental pollution is one of the most significant problems that the world is facing. Unlike the natural organic compounds that are readily degraded into the environment, some synthetic chemicals are extremely resistant to biodegradation by native microorganisms. 1
Over 100,000 dyes have been generated worldwide with an annual production of over 7 × 105 metric tons since the production of the first synthetic dye, “mauevine,” in 1856. 2 The application of synthetic dyes is also widespread, including textile dyeing, paper printing, color photography, pharmaceutical, cosmetic, food, and leather industries. 3
In the textile industry, reactive dyes are widely used on cotton, rayon, and other cellulosic fabrics, due to their high solubility and stability, good diffusivity and dye permeability, and good colorfastness. 4 Moreover, most of the synthetic dyes are toxic and highly resistant to degradation due to their complex chemical structures, such as benzidine and other aromatic compounds. 1
Textile biodegradation is considered when textiles are buried in soil and the
microorganisms present in it take part in the degradation of textiles. The
biodegradation of textiles is expected to be influenced by factors like
crystallinity, hydro-phobicity, hydrophilicity, degree of polymerization of the
textile materials, soil composition, and the species of microorganisms
present.
5
The
biodegradation of reactive dyes on cotton fabrics is not fully understood, due to
the behavior that these organic molecules have during decomposition. However, the
possible degradation mechanism pathways of the synthetic dyes bonded to textile
fabrics will depend on the chemical structure of each dye. The motivation for this
study is to understand the biodegradation of four reactive dyes (RBlk5, RR198, RB49,
and RO35) commonly used on cotton fabrics. This will be accomplished by using a
chemical method (treatment) and an enzymatic digestion of fabrics dyed with reactive
dyes previously degraded in soil and analyzed with high-performance liquid
chromatography-high-resolution mass spectrometry (HPLC-HRMS). Mass spectrometry (MS)
is the main analytical technique used for this project due to its unequaled
sensitivity, detection limit, speed, and diversity of its applications. MS is a
technique that helps identify the amount and type of chemicals present in a sample
by measuring the mass-to-charge (
Experimental
Materials
Reactive Dyes
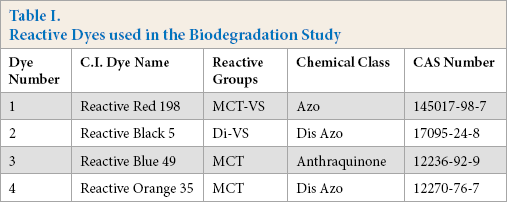
Reactive dyes (Table I) were provided by Cotton Incorporated and used without further purification. Dye solutions were prepared by dissolving 1 mg of the commercial dye in 1 mL of 50:50 (v/v) methanol/deionized (DI) water.
Reactive Dyes used in the Biodegradation Study
Dyed Fabrics and Degraded Samples
The dyeing was performed at the Dyeing and Finishing Application Laboratory at Cotton Incorporated (Cary, NC, USA), by the following steps. Fabric (1370 g) was loaded into an OptiLab dyeing machine at a 15:1 liquor ratio. The salt (sodium sulfate) was then added at a rate of 55 g/L (1130 g of salt for a total of 21 L) and circulated for 5 min at 37.8 °C. Next, 27.4 g of dye at 2.0% on weight of goods (owg) was added and circulated for 15 min at 60 °C. After that, a total of 267 g of soda ash (sodium carbonate) was added at a rate of 13 g/L and circulated for 45 min. Ten, 10 g of acetic acid was added at a rate of 0.5 g/L and circulated for 10 min after decreasing the temperature to 48.9 °C. Next, the fabric was circulated for 10 min after increasing the temperature to 93.3 °C. Finally, the fabric was cooled to 82.2 °C, the solution drained, the bath filled with cold water, and circulated for 2 min.
All reactive dyes were applied by dyeing a single knit 28 cut jersey fabric (100% cotton ring-spun yarn) with 2.0% owg dye. The dyed fabrics were cut into 2 × 2 cm fabric strips. For each dyed sample, three strips were biodegraded for 45 days, which was carried out at the Department of Fiber Science and Apparel Design at Cornell University, under controlled laboratory conditions according to ASTM D5988-037 Standard Test Method for Determining Aerobic Biodegradation in Soil of Plastic Materials or Residual Plastic Materials After Composting. The dyed fabrics and degraded samples are shown as comparisons in Fig. 1.

Dyed and dyed and degraded fabric samples (from top to bottom: cotton, RO35, RBlk5, RR198, and RB49, respectively).
Chemicals
For LC-MS experiments, methanol, acetonitrile (HPLC grade, Sigma-Aldrich) and purified water (18.2 Ω, Milli-Q System, Purelab Ultra) were used. Sodium hydroxide (NaOH), hydrochloric acid (HCl), acetic acid (CH3COOH), cellulase, and dimethylformamide (DMF, (CH3)2NCOH, ACS grade), from Sigma-Aldrich were used. In addition, ACS grade sodium sulfate, cellobiose, ethyl acetate, ammonium formate, formic acid, 1-butanol, ethanol, NH4OH, and pyridine were purchased from Sigma-Aldrich.
Procedures
Synthesis and Preparation of Analytical Standards of Reactive Dyes
Hydrolyzed Dye Standards Synthesis
These experiments were carried out to develop standards for potential hydrolyzed dye products generated by dye removal by acids and bases from fabric samples. It is important to mention that there are no commercially-available hydrolyzed dye standards. Based on our studies, it was proved that alkali treatment can remove the greatest amount of dyes compared to solvent wash (remove dye from the surface) or acidic treatment. Thus, in this work, alkali treatment of reactive dyes was used. Sodium hydroxide was tested and selected as the chemical of choice for this study due to the detection of hydrolyzed reactive dye by HRMS and minimal dye degradation observed under different conditions. Briefly, we described the general alkali method applied to RBlk5 (two vinyl sulfone groups) and RR198 (one monochlorotriazine, and one vinyl sulfone group) to synthesize the reactive dye hydrolyzed standard.
For RBlk5, 0.0507 g of RBlk5 dye powder was weighed and added to an 80-mL beaker. Ten, 50 mL of 0.15% NaOH was added to the beaker to obtain a 0.001 M dye solution. The solution was gently stirred at 60 °C for 3 h. Finally, the hydro-lyzed product was monitored by thin layer chromatography (TLC) and confirmed by HRMS. The TLC solvent system used for RBlk5 was composed of isopropanol/1-butanol/ethyl acetate/water at a ratio of 2/4/1/3, respectively. After completion of the reaction, the bath was stirred and neutralized with 1 N HCl to pH 7. For RR198, the procedures were the same except that the temperature was held to 80 °C for 3 h instead of 60 °C. The TLC solvent system used for RR198 sample elution was composed of 1-butanol/ethanol/NH4OH/pyridine/ water at a ratio of 8/2/8/4/3, respectively.
Dye Purification
The synthesized hydrolyzed standards were concentrated by a rotary evaporator; the water in the synthesized product was evaporated at a temperature of 60 °C. After evaporation of the solvent, the solid precipitate in the flask was purified by the following procedure. DMF (15 mL) was added to the sample in the flask, the solution was stirred and filtered to remove salt, and the filtrate was collected and transferred to a beaker. Ethyl acetate (50 mL) was added dropwise to the sample and all the purified dye precipitated out of the solution. Ten, the upper solvent layer was decanted, and the sample was filtered to remove excess solvent, and finally, the purified dye standard was obtained. Fig. 2 shows the precipitation and purification experimental set up for hydrolyzed standard RBlk5.
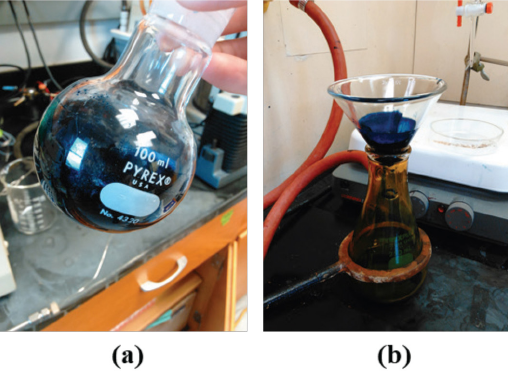

Precipitation and purification of RBlk5 hydrolyzed standard sample. (a) shows the synthesized sample after solvent evaporation using a rotary evaporator, and (b) shows the desalination and purification experimental set up.
Enzymatic Digestion Dye Standards
Synthesis
The enzymatic digestion standards were also synthesized for all dyes. The synthesis procedure was as follows. The dye (0.01 mol) and cellobiose (0.05 mol) powder were individually dissolved in 50 and 20 mL of purified water, respectively. The pH of the cellobiose solution was adjusted to 7 by adding a 0.375 M NaOH solution. The initial pH of the dye solution was 3.75 and it was adjusted to pH 7 by adding 60 μL of 1.5% NaOH solution. The dye solution was then heated to 60 °C and the neutralized cellobiose solution was added dropwise. The reaction was held at 60 °C for 2 h with constant stirring. A total of 1.20 g cellobiose powder was added to complete the reaction. The final molar ratio between the dye and cellobiose was 1 to 15. The reaction was monitored by TLC. The reaction was cooled to room temperature (RT) and neutralized to pH 7 with 1 N HCl. No purification was done on these samples to have a mixture of enzymatic digestion standards. Standards were analyzed by LC-DAD (liquid chromatography-diode array detection)-MS, by confirming their exact mass and λmax value. The detailed analytical procedure is described later.
Chemical Treatment Method Development for Removal of Reactive Dyes from Fabrics
The alkali (base) chemical treatment was optimized and applied to all control and degraded samples. A brief procedure for RBlk5 and RR198 dyed samples is described below as a reference. Before treatment, the degraded fabrics were washed three times with different solvents (2-mL water, 2-mL methanol, and 2-mL acetonitrile) in turn to remove impurities that may interfere with treatment. Sampling was carried out by three replicates from different sections of the degraded and control fabrics.
The following standard operating procedure (SOP) of chemical treatment was carried out on control and degraded fabrics. The fabric samples were weighted to 3 mg because of the limited quantity of degraded fabrics; a 1 mL of 1.5% NaOH solution was added to a 5-mL Fisher glass vial, then the vial was heated to 80 °C for 1 h with constant stirring. After 1 h, the sample was removed from the heating source, stirred to cool down, and then neutralized by adding 300 μL of 1 M HCl solution. Finally, the neutralized solution of the sample was filtered with a polytetrafluoroethylene (PTFE) syringe filter and transferred into HPLC vials for LC-MS analysis.
Enzymatic Digestion Method Development for Control and Degraded Samples
Biodegradation can occur in a landfill or in a laboratory-controlled experiment. 8 During biodegradation of cellulosic fiber, the long-chain polymer of cellulose is digested to small monomers or dimers of glucose and cellobiose units. When reactive dyes are covalently bonded to cellulosic fibers, it is assumed that one or two glucose units will be bonded with dye during biodegradation. For this reason, an enzymatic method was developed to remove the reactive dye from the fabric. An optimized enzymatic method was applied to prewashed control and degraded fabrics. Fig. 3 shows the visual appearance of the digested samples after enzymatic treatments.

The visual appearance of the samples after enzymatic digestion treatments. (a) RBlk5 at 0 days (control black), (b) RBlk5 at 45 days degraded sample, and (c) RR198 at 0 day sample (control red).
A brief description of the method is as follows. First, a prewash for control and degraded samples was performed by adding 3 mg of fabric to each sample vial, followed by the additio of 1 mL of water and gently shaking by hand. The water was decanted and saved for later analysis. Second, 1 mL of metha- nol was added, washed as before, and the solvent was removed and saved for analysis. Third, 1 mL of acetonitrile was added and followed the same procedure as before (all the washes were analyzed by HRMS for the presence of any chemicals that may be present in the surface of the fabrics). Finally, the washed fabric samples were dried at RT and were ready for enzymatic treatment.
After prewashing, the general procedure for the enzymatic method was carried out as follows. A fabric sample (3 mg) was weighed and transferred into a vial. Ten, 100 μL of 3 M NaOH solution was added to the vial, the vial was placed in a grip seal bag, and placed in a container containing ice for 4 h. Ten, the NaOH solution was removed and 500 μL of 0.5 M acetic acid was added and incubated for 1 min; acetic acid was then removed and 1.5 mL of buffer solution (0.1 M sodium acetate, pH 5 with acetic acid) was added and kept for 1 min; the buffer solution was removed and 1 mL of enzyme solution (90-g cellulase in 50-mL buffer) was added. The vials were sealed and placed in a shaking bath for 24 h at 50 °C. Finally, the vials were removed from the shaker and sonicated for 30 s, the solutions were filtered and transferred into HPLC vials for LC-DAD-MS analysis.
For the case of RR198, RBlk5, RB49, and RO35, 515, 620, 600, 430, and 254 nm wavelengths were used for DAD analysis, respectively. Three replicates from different sections of the degraded and control fabrics were taken for reproducibility purposes.
Analytical Methods HPLC-HRMS
All solutions were analyzed using an Agilent 1200 LC SL HPLC with DAD coupled to an accurate 6520 Quadrupole Time-of-flight (Q-TOF) mass spectrometer (Agilent Technologies) with an electrospray ionization (ESI) source. 6
HPLC separation was achieved using a Zorbax Eclipse Plus C18 (2.1 × 50 mm, 3.5 μm) column with a Zorbax Eclipse Plus C18 narrow bore guard column (2.1 × 12.5 mm, 5 μm) at 40 °C. The mobile phases were water (A) with 20-mM ammonium formate and formic acid (pH = 4), and 70/30 methanol/acetonitrile (B). The flow rate of the mobile phase was 0.5 mL/min with an injection volume of 10 μL.
A gradient method was used for all the analysis and is as follows: 3% B from 0 to 1 min, 3%–60% B from 1 to 1.5 min, 60%–90% B from 1.5 to 7 min, holding at 90% B from 7 to 9 min, and 3% B at 9 to 9.5 min. A 4-min post run of 3% B before the next run was performed. The DAD detector analyzed at a spectral range of 200 to 800 nm. The main wavelengths for absorbance analyses were set to 254, 515, 610, 620, and 660 nm.
The Q-TOF mass spectrometer was operated in negative electrospray ionization
(ESI) mode at high resolution (4 GHz) with a resolving power ranging from 9700
to 18,000 for the
Results and Discussion
Synthesis and Preparation of Hydrolyzed Analytical Standards of Reactive Dyes
The confirmation of the final product was carried out both by TLC and MS. Two
solvent systems were used for the elution of samples in TLC for RBlk5, RR198,
RB49, and RO35 respectively, as described in the experimental section. Fig. 4 shows the TLC and MS
confirmation of the hydrolyzed standard of RBlk5. Here, the spot named ST is the
starting dye, which has multiple components, and the spot denoted as AR is the
after reaction final product, which has only one component, and was used as the
hydrolyzed standard. The cross spot represents the mixture of starting dye and
the reaction product. The MS spectra at
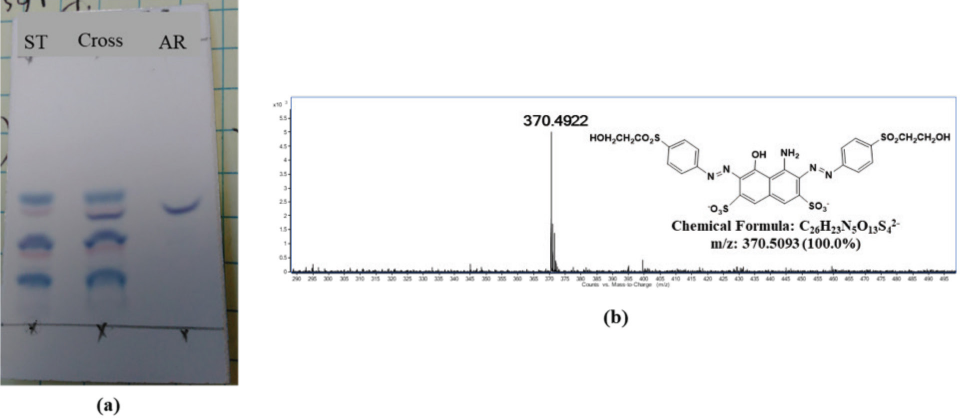
The confirmation of hydrolyzed standard synthesis of RBlk5. (a) TLC confirmation of single hydrolyzed product formation, and (b) MS confirmation of mass of the hydrolyzed standard.
Similarly, Fig. 5 shows the synthesized hydrolyzed standards of RR198. From the TLC analysis, it was observed that some starting dye remained on the after-reaction products. The reaction parameter optimization or separation would be needed for this synthesis reaction. The same conditions were used to successfully synthesize the hydrolyzed standards of RB49 and RO35.
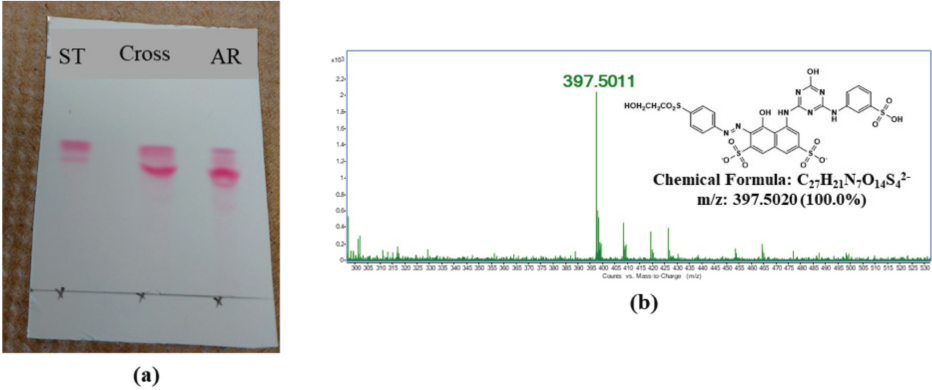
The confirmation of hydrolyzed standard synthesis of RR198. (a) TLC confirmation of the hydrolyzed product formation, and (b) MS confirmation of mass of the hydrolyzed standard.
Chemical Treatment Method Development for Removal of Reactive Dyes from Fabrics RR198
Application of the optimized chemical treatment (0.15% NaOH, 80 °C, 1 h) to
control RR198 fabric removed the hydrolyzed form of the dye from the fabric. The
hydrolyzed form of dye at
Dyes Detected and Identified from Fabrics via Mass Spectrometry from Chemical Treatment
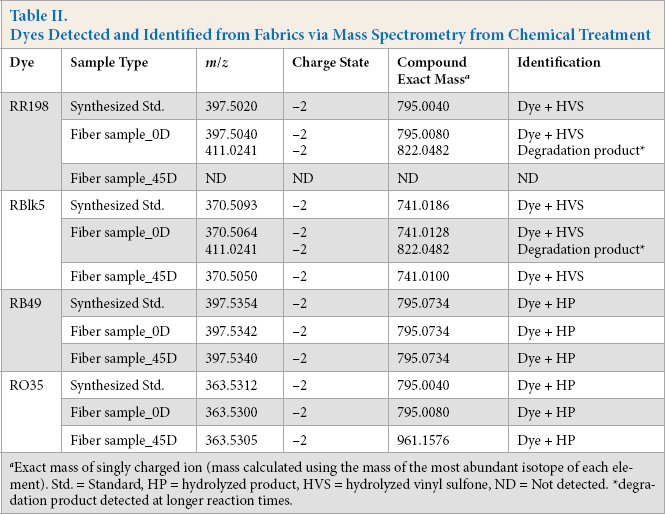
Exact mass of singly charged ion (mass calculated using the mass of the most abundant isotope of each element). Std. = Standard, HP = hydrolyzed product, HVS = hydrolyzed vinyl sulfone, ND = Not detected. *degra-dation product detected at longer reaction times.
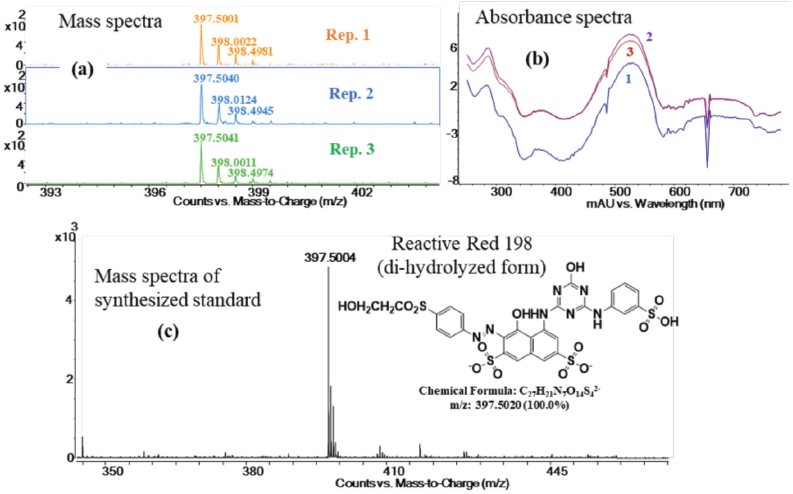
The comparison of removed dye via chemical treatment from control RR198
fabric and the synthesized standards. (a) The mass spectra of the
replicate experiments identify the doubly charged hydrolyzed
deprotonated dye ion at
RBlk5
The same alkali method used on RR198 fabrics was used on RBlk5 control fabrics.
The results showed that the hydrolyzed form of dye from the fabric was extracted
and detected. The hydrolyzed form of dye at
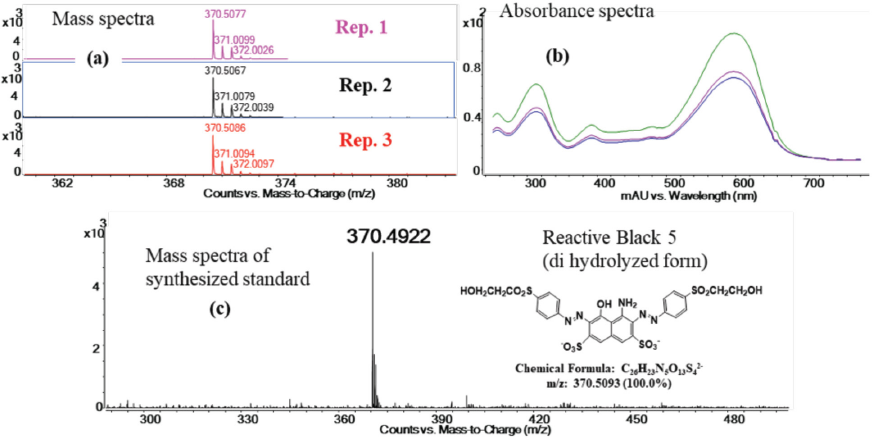
The comparison of removed dye via chemical treatment from control RBlk5
fabric and the synthesized standard. (a) The mass spectra of the
replicate experiments identify the doubly charged hydrolyzed
deprotonated dye ion at
RB49 and RO35
The same alkali method used previously was used on RB49 and RO35 control fabrics.
The results showed that the hydro-lyzed form of dye from the fabric was
extracted and detected. The hydrolyzed form of RB49 with an
In summary, these results from chemical treatment clearly indicate the strength and weakness of this treatment. Because of the covalent bonding between fiber and reactive dye, it would be hard to extract the dye from the fabrics using a conventional method like a solvent extraction. To overcome this covalent bonding, a harsh chemical such as sodium hydroxide was used. According to the results, the chemical treatment could successfully remove the fixed dye from the fabric as expected, since the hydrolysis products of the different dyes were observed in the treatment of both control and degraded fabrics. This effective removal of the reactive dye also comes with a sacrifice as certain reactive dyes showed a loss of functional groups after treatment.
Enzymatic Digestion Method Development for Control and Degraded Samples
Enzymatic Digestion of Control Samples of RBlk5
After analyzing digested samples by LC-DAD-MS, high molecular weight digested
compounds were observed. The DAD chromatogram, with a λmax at 610
nm, of the digested products is shown in Fig. 8a. This figure indicates that the
digested product mainly absorbs in the blue region and Fig. 8b shows the absorbance spectrum of
this product at 610 nm. The MS characterizes the digested product as a
doubly charged ion at
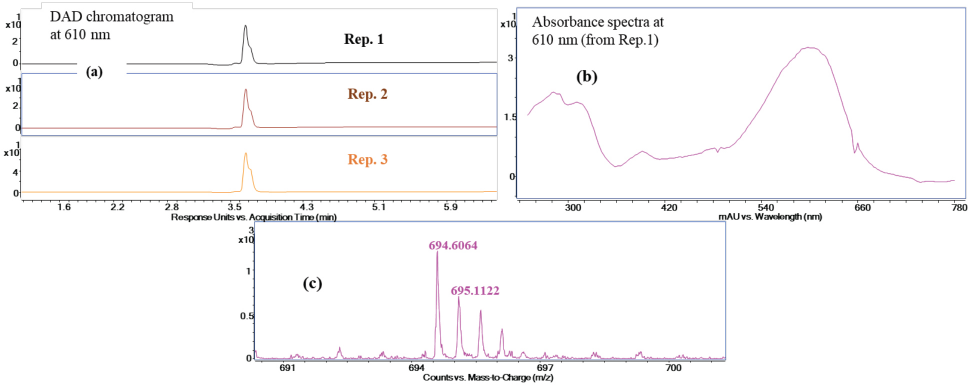
Enzymatic digestion of RBlk5 control dyed fabric. (a) The DAD chromatogram at 610 nm for three replicate experiments, (b) the absorbance spectrum of replicate 1 at a retention time of 3.67 min, and (c) the mass spectrum of the digested product.
This ion was characterized as the dye attached with two cellobiose molecules
at two ends of the reactive sites. In the digested samples, another compound
was identified at
Mass Spectrometry Results of the Enzymatic Treatment of Reactive Dyes and Synthesized Standards a
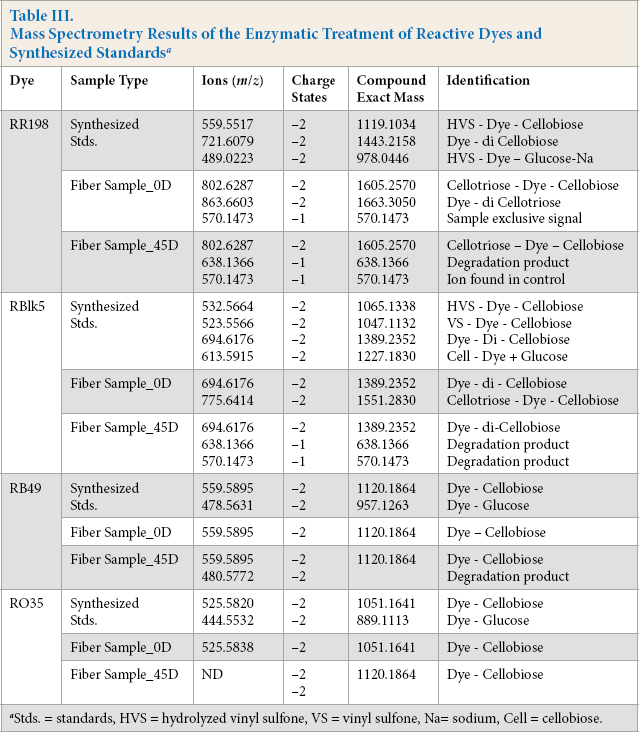
Stds. = standards, HVS = hydrolyzed vinyl sulfone, VS = vinyl sulfone, Na= sodium, Cell = cellobiose.
Enzymatic Digestion of Control Samples of RR198
The replicate enzymatic digestion experiments were carried out on the control
dyed fabrics of RR198. Fig.
9a shows the DAD chromatogram, with λmax = 515 nm,
during the digested treatment. The DAD chromatogram had three distinct peaks
at a retention time of 3.51, 3.53, and 3.76 mins (Fig. 9a). Three peaks had the absorbance at
515 nm at different intensities (Fig. 9b). Fig.
9a shows three peaks in the DAD chromatogram, where two main
peaks were from dye attached to high mass sugar moieties. The ion at
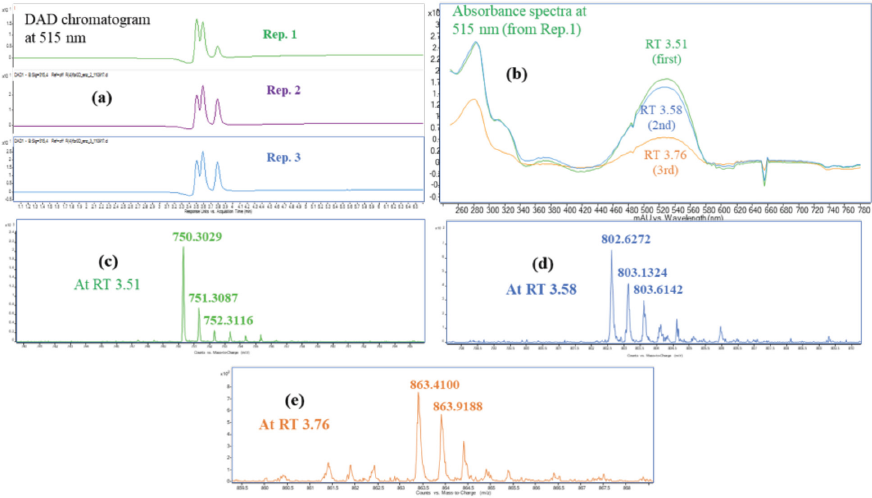
Enzymatic digestion of RR198 control dyed fabric. (a) The DAD chromatogram at 515 nm for three replicate experiments, (b) the absorbance spectra of replicate 1, which had three peaks, and (c), (d), and (e) the mass spectra of the digested products at different retention times based on the DAD chromatogram.
Enzymatic Digestion of Control Samples of RB49 and RO35
The same SOP used previously was used for RB49 and RO35. The digestion results
for RB49 showed the dye bonded to a cellobiose unit
(
Application of Enzymatic Digestion Method to Degraded Samples
According to mass spectral results, all digested products from the control dyed
fabrics were observed in the degraded samples, except for RO35, where no dye
related signals were detected. Additionally, four degradation products with
The ion with m/z values of 639 and 638 are the possible degradation products for RR198 and RBlk5 respectively (the structures are proposed in Fig. 10). Table III shows the summary of the analysis of the enzymatic digestion for 0 and 45 day samples, along with their synthesis standards. As shown in Tables II and III, there were significant differences between the control and degraded fabrics by comparing the signals of detected compounds.
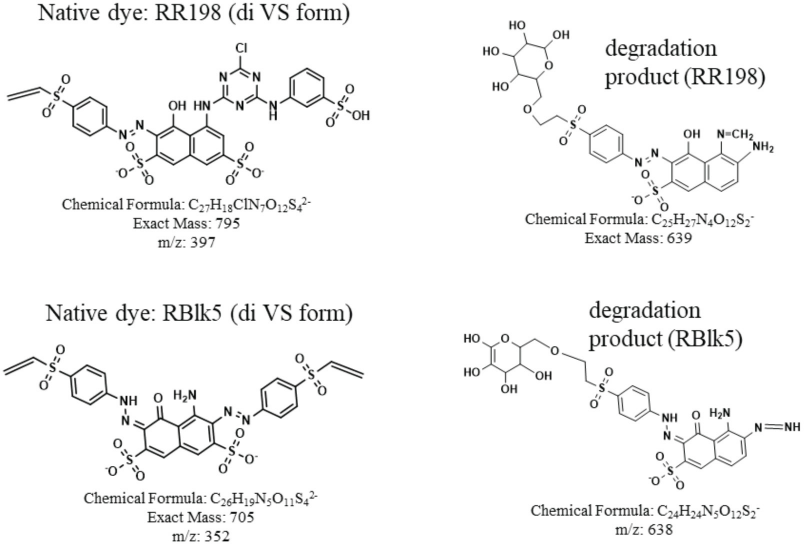
Enzymatic digestion of both RBlk5 and RR198 45 days degraded samples and the proposed structures of the degradation products.
An ion with
In the case of RO35, as mentioned earlier, no identifiable signals related to the dye were detected on the Q-TOF mass spectrometer. This may be due to the poor enzymatic digestion of this dyed fabric, based on the control samples' results. Further studies are needed to understand the behavior of this dye during digestion.
In summary, we discussed the assessment of the enzymatic digestion method for fabrics dyed with reactive dyes RBlk5 and RR198. The digested samples were compared to synthesized model compounds, which contained a cellobiose unit bonded to the reactive dye. This method is milder than the chemical treatment and can be used to complement or add additional information on the biodegradation of dyes. The optimized enzymatic treatment was able to digest the cotton fabric and removed RBlk5, RR198, and RB49. The biode-graded samples analysis showed biodegradation products from these dyes. The structures of these products have been proposed based on exact mass measurements.
Conclusions
The study of dyed fabric biodegradation in the soil helps us to have a better understanding of how selected reactive dyes behave under these conditions. Based on the high-resolution mass spectrometry results, it was clear that each reactive dye will follow different degradation pathways. However, it was established that the chemical treatment with NaOH was useful for removing all reactive dyes from the fabric to obtain their hydrolyzed products with minimal damage to the dye structure.
The synthesized standards for chemical treatment and enzymatic treatment were characterized by TLC and HRMS. These standards were useful for the characterization of potential degradation products. Each standard aided in tuning instrument conditions for chromatographic and mass spectrometric setups.
According to the results, the chemical treatment successfully removed the fixed dye from the fabric as expected, since hydrolysis forms of the different dyes were observed in the treatment of both control and degraded fabrics.
The assessment of the enzymatic digestion method for fabrics having reactive dyes RBlk5, RR198, and RB49 was discussed, which was confirmed to digest the cotton fabric and removed RBlk5, RR198, and RB49. During the experiment, the observed decolorization on all dyed samples indicated successful dye removal. For RBlk5, RR198, and RB49, HRMS confirmed there were multiple sample-exclusive signals, which indicated possible degraded dye fragments. However, for RO35, no degradation signals were detected; it is likely that enzymatic digestion did not work as observed with the other reactive dyes, with the dye structure being the likely issue.
By using HPLC-HRMS, all possible signals were detected. Mass spectrometry analysis and characterization proved to be an excellent tool to study the biodegradation of reactive dyes in cotton fabrics in soil. These results are the initial steps of understanding the degradation of reactive dyes under these conditions. Proposed structures of possible degradation dye products have been presented, which can give us a better idea of what can be leaching into the soil. A study of soil analysis will be presented in the future.
Footnotes
Acknowledgment
The authors would like to thank the generous support by Cotton Incorporated Grant No. 18-238, and Cotton Research and Development Corp. Grant No. 2018-143. We would also like to thank Dr. Margaret Frey from Cornell University for the degradation of the dyed cotton fabrics.
