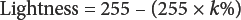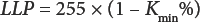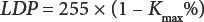Abstract
Seed coat fragments (SCFs) in cotton fabrics are defects that negatively impact the value and quality of cotton if they are visible. The ability to accurately and rapidly count SCFs is important to efficient quality control and for research into improved cotton wet processing. Innovative scouring/bleaching using N-[4-(triethylammoniomethyl)benzoyl]butyrolactam chloride (TBBC) under neutral conditions produces satisfactory whiteness, but inferior performance in SCF removal. In this study, an image-based program was developed to quantitatively analyze SCFs based on their appearance on bleached cotton fabrics. With optimized thresholds, the program was capable of counting SCFs in fabric images semi-automatically, with a diagnostic sensitivity of 94.7% and an accuracy of 94.1%. Compared to the conventional method, image-based counting gave high accuracy and sensitivity.
Introduction
Seed coat fragments (SCFs), usually black or brown in color, can cause defects that negatively impact the value and quality of cotton if they are visible. 1 SCFs are removed via mechanical processing procedures such as hackling, combing, and lint-cleaning, 2 followed by chemical processing including desizing, scouring, and bleaching.3–5 In the context of mechanical processing, certain seed fragments are so small that they can hardly be removed completely. 6 Hence, most tiny SCFs are decolorized or decomposed via chemical processing rather than being removed.3,4 Conventional chemical processing removes SCFs in boiling sodium hydroxide with strong agitation,6–9 but also consumes large quantities of water, energy, and causes a series of environmental problems. 10 Therefore, developing alternative, environmentally-benign processing for cotton preparation were of particular interest in past decades, with many of them based on enzymes and bleach activators.7,9–13
Our recent work showed that an innovative scouring/bleaching system, using N-[4-(triethylammoniomethyl)benzoyl] butyrolactam chloride (TBBC), was capable of removing natural coloring matter present in greige cotton and produced satisfactory whiteness, absorbance, and soft hand while consuming minimal energy and water. 10 However, it did not remove SCFs adequately. This limitation was also found in other bioprocessing methods.7,8,11,14
The inability to remove all SCFs is due in part to their intricate entanglement with fabrics, as well as their complex chemical compositions.2,8,15 Seed coats have a water impermeable cuticle that limits the penetration of liquids and reagents. Wax and pectin present on greige cotton are hydrophobic and consequently reduce the effectiveness of fabric wet processing. Additionally, complex pigmentation cannot be easily decolorized without both oxidative and reductive bleaching. 3 In view of the importance of SCF removal and ongoing research to develop environmentally-benign preparation processes for cotton, a facile and rapid evaluation procedure for quantitative analysis of SCFs in a given cotton fabric is necessary.
Current methods of counting SCFs are mostly based on manual visual counting, 16 which relies on the operator being able to distinguish overall size and shape of SCFs as well as their different hue, which is prone to inconsistency.17,18In addition, a number of psychophysical factors, including visual acuity, the subject's attention span, and diligence during test performance, can affect the assessments. Furthermore, manual counting is affected by evaluation environments, for instance, illumination conditions. Therefore, accurate and reliable assessments that do not depend on operator and evaluation environment are needed for standard SCFs counting.
Image-based technology has effectively counted particles, 19 cells, 20 and bacteria 21 in the field of biomedical research. Image-based technology applied to SCF counting previously demonstrated its benefits—convenience and efficiency. Bel and Xu developed an Advanced Fiber Information System to automatically sort SCFs in fabrics, which was much faster than hand sorting. 17 Later, they devised a specific automated fabric inspection system to count dark specks in four varieties of greige cotton fabrics by examination of the L*a*b* color value distribution in a fabric image. 18 Commercially-available image-based software has been applied to evaluate the percent SCFs after bioprocessing. 22 However, these methods require special software and, in the latter case, demonstrated less reproducibility. In this paper, we describe an image-based Matlab program developed to analyze the morphology of SCFs, with the primary goal of developing a new method of counting SCFs that had higher efficiency and accuracy.20,23,24
Experimental
Materials and Equipment
Chemicals
Cationic bleach activator TBBC was synthesized and purified to > 97%.25,26 Plain weave 100% cotton greige fabric (122 g/m) was purchased from Testfabrics. Fabric was pre-sized with 100% starch sizing agent. Hydrogen peroxide (H2O2, 35% w/w) was purchased from the Sigma-Aldrich Co. Aquazym Prime 12000 L (alpha-amylase) was provided by Novozymes. Sodium dihydrogen phosphate monohy-drate (NaH2PO4?H2O) and disodium hydrogen phosphate (Na2HPO4) were obtained from Fisher Scientific. The auxiliaries used were nonionic wetting agents (Triton X-100, Sigma-Aldrich), chelating agent (Dekol N-SN, BASF), and peroxide stabilizer (Prestogen N-D, BASF). 10
Equipment
All images were acquired by an optical scanner (Epson Expression 1000XL) interfacing with Adobe Photoshop CS6 and Matlab (R2011a) for image processing. An X-Rite Spec-tralite III light box with calibrated daylight simulation (6500 K) was used for all visual assessments.
Methods
Sample Preparation
Greige cotton fabric samples containing varied number of SCFs were treated using four different TBBC concentrations for scouring/bleaching. Samples were treated with amylase desizing and bleach activator-activated bleaching in an AHIBA Texomat laboratory dyeing machine at a liquor-to-goods ratio of 20:1. Enzyme (1 g/L) and surfactant (1 g/L) were added in the bath for a 30 min desizing, followed by 30 min activated bleaching using TBBC at four concentration levels (1 g/L, 2 g/L, 3 g/L, and 5 g/L). The molar ratio between TBBC and H2O2 was 1:1.2. The bath solution was made using 200 mL of 0.1M phosphate buffer with a pH value of 7.2 ± 0.05. A varied number of SCFs were present in the four standard fabrics.
Microscopy
Fabric samples taken after processing were examined with a stereo microscope (Nikon SMZ-1000 Zoom Stereo Microscope). Individual SCFs were isolated, fixed on a glass slide using distilled water, and covered with a glass slide. Microscopic examinations of SCFs were carried out using a polarizing microscope (Nikon Eclipse 50 i POL).
Image-Based SCFs Counting
Image Analysis
A scanner (Epson Expression 1000XL) was used to capture fabric images with a physical resolution of 300 pixels/in. Fig. 1 shows a sample image in grey scale mode. SCFs can be differen-tiated from non-SCFs in an image, such as textures and creases, by judicial selection of the lightness of pixel range. 25
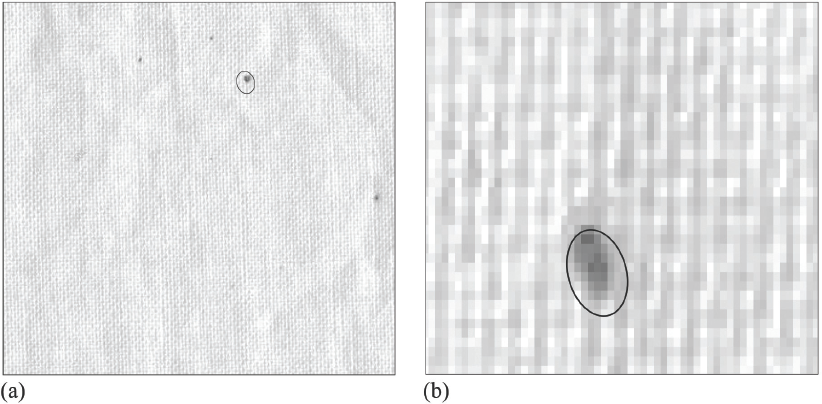

Image of seed coat fragments (SCFs) in cotton fabrics: (a) scanned image of cotton fabrics (resolution: 300 pixels/square inch and (b) stacks of pixels belonging to the SCFs in image (a).
As seen in Fig. 1a, SCFs appear considerably darker than the fabric and vary in size and hue. A magnified view of image Fig. 1a (Fig. 1b) reads the SCF in the image as a stack of pixels that fall within a specific lightness range, with the number of pixels in the stack determining the shape and area of the SCF. Therefore, counting SCFs is essentially the process of counting dark specks. According to a similar principle in previous research,21,26 we developed a two-dimensional thresholding algorithm based on the lightness as well as the number of pixels.
The lightness of pixels in a grey mode image was expressed by k%, which represents the proportion of black subtracted from perfect whiteness, assuming the perfect whiteness value is 255. 27 The lightness of pixel was readily obtained using Eq. 1.
A k% value was determined for each pixel of interest using Adobe Photoshop CS 6.
SCF Counting
In this study, a Matlab program was used to automatically analyze SCFs across the fabric. Lightness of light pixel (LLP) and lightness of dark pixel (LDP) are the key thresholds defining the lightest and darkest pixels belonging to the SCFs and were calculated by Eqs. 2 and 3.
Kmin and Kmax are the k values of the lightest and darkest pixels, respectively, belonging to the SCFs. Four “standard” cotton fabrics (A–D) containing varied number of SCFs were used for the selection of the LLP and LDP. The threshold k% was determined by visual assessment; it was proposed that the specks containing the number of pixels equal to or greater than 3 that fell within the appropriate lightness range were regarded as SCFs.
Matlab Program Image Processing Algorithm
First, all pixels in the image were enumerated and the coordinate of one pixel was saved if it had a lightness within the defined threshold limits. The program included a loop to put these saved pixels into stacks based on their coordinates (Fig. 2a). The flow chart in Fig. 2b describes the image process. In the loop, the first saved pixel was put into a new stack. The coordinates of the second pixel were compared with those of the first pixel. If its x or y coordinates were both within six units of the first pixel, this second pixel was put into the first stack. Otherwise, it was put into a new stack. This procedure was repeated for all pixels in the image. The program compared each pixel with the last one to determine whether they could be put into the same stack or not.
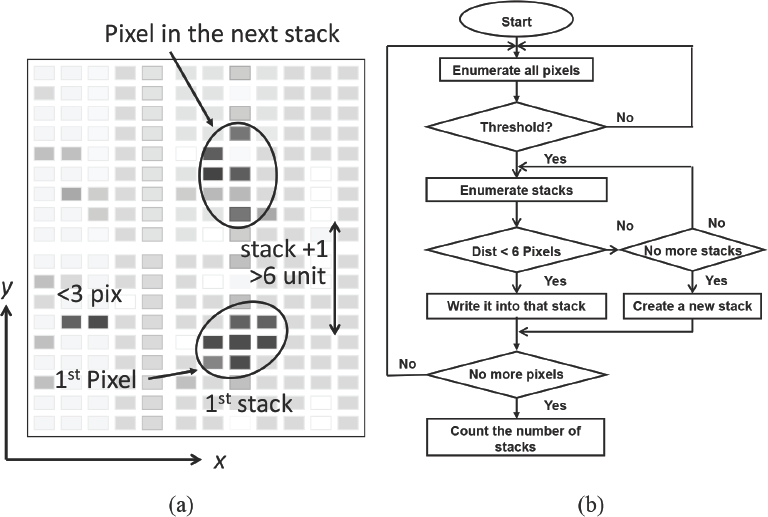
(a) Analysis of pixels and stacks in a simulated image. The x and y axes represent the pixels’ coordinates. (b) flow chart of the image processing algorithm.
Finally, the program enumerated all the stacks and out-putted the calculated number of stacks. If the number of points in one stack was less than or equal to the minimal size (3 pixels) for a dark point, the stack was not counted, as this would be too small to be a visible SCF. If the darkest pixel in a stack was higher than the LDP, it was also removed. The resultant total number of stacks was the total SCFs counted.
Manual Counting
To compare the overall performance of image-based reading and conventional manual counting, standard cotton fabrics, as well as three unknown fabrics, were evaluated by five observers. Each assessment of the same fabric was performed three times, with the intervals between tests being at least 24 h. Manual counting was performed under the same conditions defined by AATCC Evaluation Procedure (EP) 9-2011 (Visual Assessment of Color Difference of Textiles). 28
Results and Discussion
Microscopy of SCFs
Visible and polarizing microscopy images (Fig. 3a–i) demonstrate that the SCFs varied in size, shape, and hue. They were fragments that may or may not be entangled with fibers. Hue was determined by the distribution and chemical compositions of the SCFs.3,15 Distribution of pigmentation, in either seed coats or inner layer of immature fibers, is shown in Figs. 3g–i.
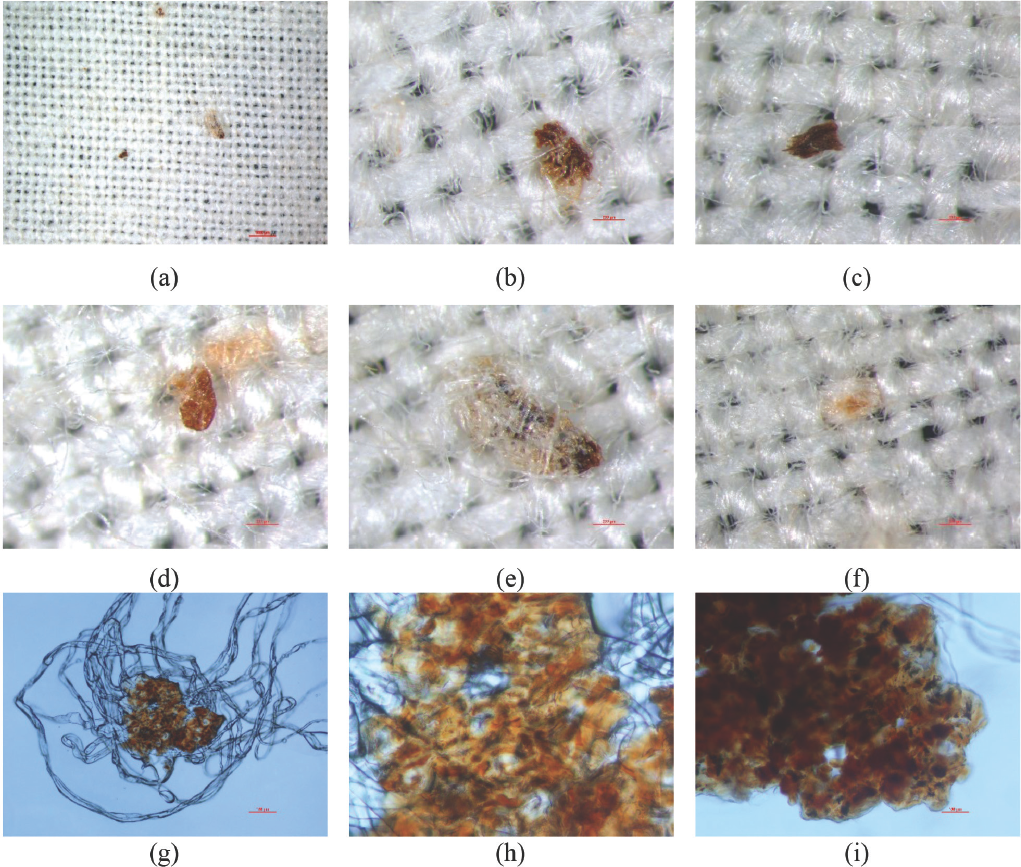
Microscopy images of SCFs in cotton fabrics: (a) cotton fabrics (1000 μm, 10x magnification), (b, c) seed coat fragments (200 μm, 60x magnification), (d, e) SCFs entangled with fibers (200 μm, 60x magnification), (f) a cluster of brown fibers (200 μm, 60x magnification), (g) isolated seed fragments entangled with a cluster of fibers (polarizing microscopy, 100 μm, 100x magnification), and (h, i) distribution of pigments in immature fibers (polarizing microscopy, 100 μm, 400x magnification).
Lightness Range of SCFs
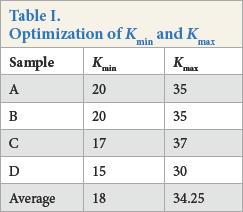
To select a reasonable range of SCFs lightness, a lightness range (Kmin and Kmax) was obtained after averaging those of “standard” fabrics (A–D) by judicial selection using Adobe Photoshop CS 6 (Table I). The corresponding LLP and LDP values were 209.10 and 167.67, respectively. Further optimization found that a combination of LLP = 210 and LDP = 180 values permitted rational SCF counting in this study.
Optimization of Kmin and Kmax
Performance Validation
To determine if the program gave reasonable counts and to evaluate the variability of the method, visual assessment was conducted on the post-processed image. SCFs in the post-processed image were classified into four categories: 26
True positives (TP) = SCFs that were correctly detected
True negatives (TN) = non-SCFs that were rejected
False positives (FP) = non-SCF specks that were detected as SCFs
False negatives (FN) = SCFs that were not detected
For example, if the original image of standard fabric D (Fig. 4a) had 16 SCFs by visual assessment, 12 of them were correctly detected by the program as true positives (TPs) (Fig. 4b). The other 4 SCFs that were missed by the program were false negative (FNs).
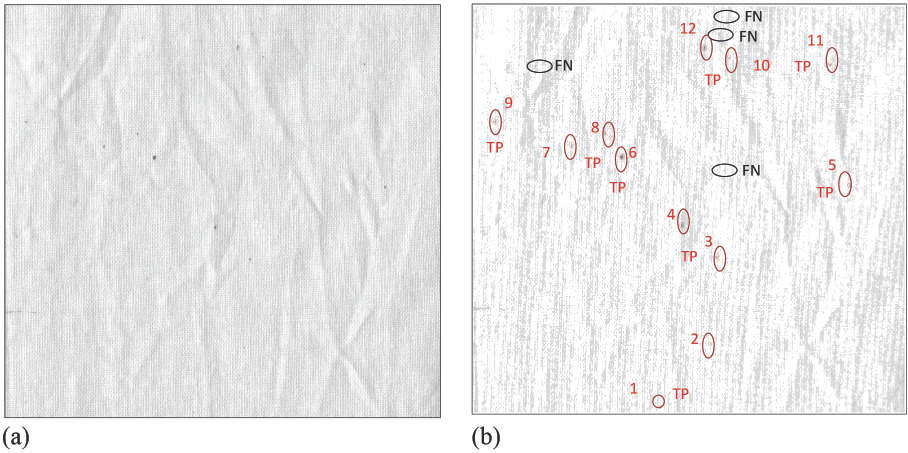

(a) Image analysis of imaging-based counting method (The program was made to output a statement such as: “Tere are 12 SCFs, i.e., 2341 SCFs/ m2.)” (b) detected SCFs are circled in red. Note: the smaller red circles were generated by the program, while the black circles and big red circles are marked manually by the authors for demonstration: (FN: SCFs are not detected, TP: SCFs detected by the program).
A validation process was conducted for the seven samples (standard samples A–D and unknown samples 1–3) by examining two parameters: sensitivity (the percent actual SCFs detected) and diagnostic accuracy (the overall estimate of the counting program accuracy) are defined in Eqs. 4 and 5.21,26
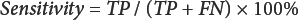
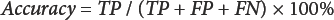
As shown in Table II, results from the validation process indicate that the optimized program gave comparatively high overall sensitivity (94.7%) and diagnostic accuracy (94.1%).
Validation of Performance of Image-Based Counting
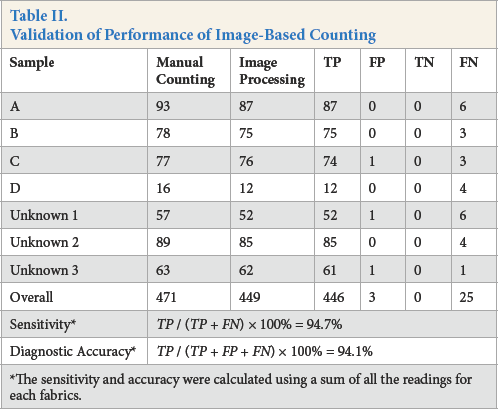
The sensitivity and accuracy were calculated using a sum of all the readings for each fabrics.
Comparison of Image-Based and Manual Counting
Manual counting of standard fabrics (A–D) showed intra-variability by the same observers. Compared to optimized imaging-based counting, manual counting gave results that were varied among the observers (inter-observer variability, Fig. 5a) and varied for the same observers (intra-observer variability, Fig. 5b). Averaged reading of manual counting results from five observers gave large variances (Fig. 5c). This variability (reproducibility and potential undercounting) may be caused by a number of psychophysical factors, including visual acuity, the observer's independent understanding of the SCF definition, and the subject's attention span and diligence at the time the test was performed.
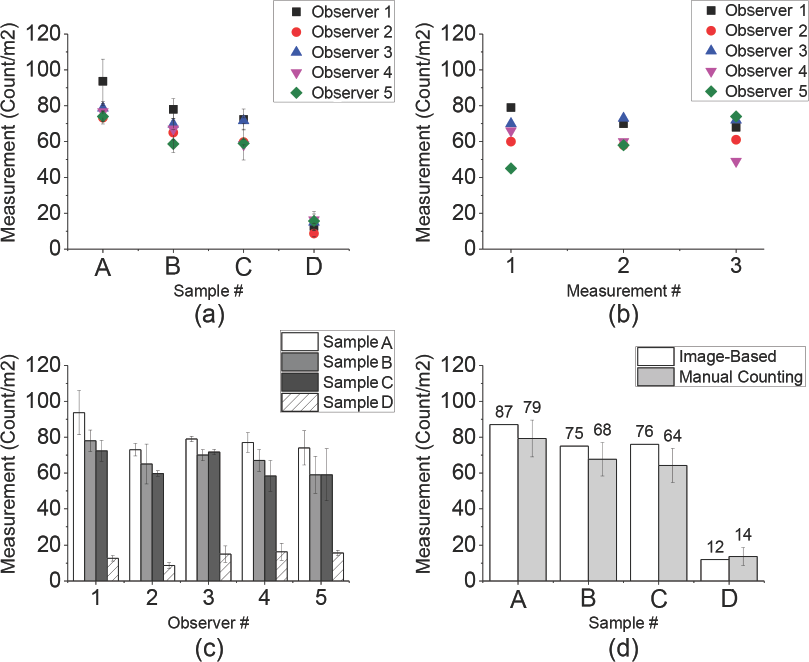
Inter-observer and intra-observer variability of manual counting. (a) inter-observer variability in the same sample, (b) intra-observer variability in different measurements, (c) variability of manual counting, and (d) comparison of image-based counting with those obtained by manual counting.
Conclusions
In this limited study of four fabrics, an imaging-based seed coat fragments (SCFs) counting program with optimized thresholds of lightness and darkness was capable of counting SCFs on cotton fabrics with a sensitivity and diagnostic accuracy of 94.7% and 94.1%, respectively. The results of manual counting performed by different observers showed that observers had a tendency to undercount SCFs, especially when the number of SCFs was high, and that manual counting lacked both repeatability (for a given observer) and reproducibility (between observers).
While more in-depth study is needed, the results given by imaging-based counting gave quite high accuracy and sensitivity. The method provided the potential to set an industrial standard for counting SCFs on fabrics prior to dyeing. More work is needed on the reproducibility of the image-based counting method and further optimization of the algorithm is required.
Footnotes
Acknowledgements
Cotton Incorporated is gratefully acknowledged for partial funding of this work. Many thanks also to everyone in David Hinks’ research group and to the test observers: Guan Wang, Sha Fu, Yongxin Wang, Wenwen Zhang, and Xiaoyan Tang. Also, we thank Judy Eldson for her assistance in obtaining the microscopic images of SCFs, Peter Bloomfield for his suggestions on the statistical methodology of performance validation, Peter Hauser for reviewing this article, and Jeffery Krauss for laboratory support.