Abstract
Industrial testing for bacterial growth on commercial substrates is highly efficient, with high-throughput means for quantifying growth. However, current antifungal testing is mostly subjective, with quantitative tests reliant on labor-intensive colony counting. We evaluated the use of a commercially available most probable number (MPN) system for quantifying growth of mold in a modified version of the AATCC Test Method (TM) 100. Using the MPN system, we were able to detect up to a 3.7-fold log reduction in mold growth between antifungal treated and untreated fabric samples. Using agar plate counts in parallel, we reproduced the trends observed using the MPN system. The trends in log reduction of growth correlated well and confirm that the MPN system satisfactorily substitutes for plate counts in evaluating antimicrobial technologies.
Introduction
The antimicrobial textile industry is dependent on standard methods from international organizations (e.g., ISO, ASTM, and AATCC) to determine the effectiveness of a treatment on a textile. These textile methods originated as qualitative methods evaluating a zone of inhibition of microbial growth created by the treated product. While qualitative testing methods do provide a basis for determining the ability of an antimicrobial to provide efficacy, these methods do not efficiently determine the actual reduction provided on the garment. To compare competitor products, it becomes important to quantitatively determine the bioburden on a garment at the end of a testing period. To meet the industry need for quantitative methods, a multitude of quantitative bacterial methods were introduced and now include but are not limited to: AATCC Test Method (TM) 100, 1 JIS L 1902, 2 ISO 20743, 3 and ASTM E2149. 4 These methods have drastically altered the textile industry by providing a platform to evaluate multiple antimicrobials using a common methodology.
In addition to antibacterial performance, many textiles must demonstrate antifungal activity. The ability to inhibit numerous fungal species becomes important when dealing with environments conducive to mold growth. This would include shipping textiles throughout the supply chain, from manufacturing origin to final point of sale. Fungal species that can degrade a garment during transport include organisms that are capable of breaking down cellulosic material, such as Aspergillus niger and Penicillium sp., and are of importance on naturally occurring fibers or fiber blends. 5 A textile can also play a key role in suppressing skin microflora that are often associated with disease, 6 such as Trichophyton mentagrophytes, the causative agent of athlete's foot. To date only two antimicrobial agents are registered for this use: quaternary-ammonium silane and copper threads.
While antibacterial standard methods have evolved throughout the years to meet a growing industry demand, antifungal test methods have remained largely unchanged. Currently, there are only a handful of recognized industry fungal standard methods and all are qualitative in nature, such as AATCC TM 307 and ASTM G21. 8 Qualitative fungal test methods will always be important for evaluating the functionality of a treated textile, but are limited in the breadth of antimicrobials that can be tested and differentiated. Currently there is a draft method being considered by the ISO technical committee for textiles (TC 38) that would be able to quantitatively determine the impact of an antifungal treatment on a textile. 9 This would allow the industry to better understand all available antimicrobial technologies in the context of fungal inhibition and provide a common platform for the comparison of antimicrobials.
In this study, we examined a methodology similar to the proposed ISO fungal standard and the AATCC TM 1001for bacteria. The antifungal sensitivity of the method was evaluated side by side with the AATCC TM 30, Test III 7 and was found to have increased sensitivity and a reduced turnaround-time. We were able to demonstrate that a simple modification of AATCC TM 100 allows for the quantification of A. niger and T. mentagrophytes. In addition, we show that both the traditional plate count method and the high-throughput most probable number (MPN) system were able to generate comparable results.
The bioMérieux MPN system was designed as a rapid most probable number enumeration system for the food industry. This system was easily integrated into the antimicrobial industry with the flexibility to spike samples and enumerate at many time points. The MPN system uses a general heterotrophic media containing a fluorescent indicator which enumerates colony forming unit (CFU) values of of a 48-well plate across 3 different dilutions. The resulting counts can subsequently be used to make percent and log reduction calculations.
This work demonstrates a methodology that would allow for the quantification of fungal inhibition by antimicrobials, thereby providing a basis for the comparison of antifungal technologies.
Experimental
Strains and Culture Conditions
A. niger van Tieghem (ATCC 6275) or T. mentagrophytes (Robin) Blanchard (ATCC 9533) were rehydrated from lyophilized stock following instructions from ATCC (American Type Culture Collection). The rehydrated conidia were inoculated onto appropriate media for the production of conidia (i.e., Sabouraud Dextrose Agar (SDA) plates for A. niger and Potato Dextrose Agar (PDA) plates for T. mentagro-phytes). Conidia were harvested in sterile distilled water with 0.05% Triton X-100 after 7 days growth at 30 °C and adjusted to a final concentration of 1–3 × 106 colony forming units (CFU)/mL. Potato dextrose agar (PDA) plates were used for plate quantification of CFU.
Most Probable Number System
The TEMPO bioMérieux system was selected for evaluation of an automated MPN system. This system uses a proprietary vital dye indicator that is has been validated in the food and medical industry.10–12 It served as a platform for analysis of the most probable number system for evaluation of fungal species.
For each assessment, dehydrated media for yeast mold (YM) cards containing a fluorescent vital dye and required media was used. The media was re-hydrated with 3 mL of sterile water, followed by the addition of a 1-mL aliquot from the sample. The vial was vortexed for 10 s and subsequently placed into a YM card using a vacuum. The cards were incubated at the specified temperature and time. After incubation, the fluorescent wells were recorded and a most probable number (mpn) assigned by the software. These numbers were compared to traditional plate count numbers as described above.
Textile Construction and Structure
Two 100% polyester textiles were evaluated over the course of the study: 95 g/m2 of polyester microfiber and 150D plain weave polyester. Each was treated topically with an antifungal before testing.
Establishment of Inoculum Level
To establish the correct inoculum level for testing, an A. niger conidial suspension (at a starting concentration of 1.39 × 107 CFU/mL) was used to prepare a 10-fold dilution series from 106 to 103 CFU/mL in Sabouraud Dextrose Broth (SDB). Per AATCC TM 100, 1 mL of each inoculum dilution was inoculated onto previously sterilized polyester fabric in 100 mL jars and incubated at 30 °C for 24 h. To enumerate the resulting growth, the samples were diluted 1:1000 and 1:100, respectively. The diluted 106 and 105samples (100 μL) were each plated on PDA plates in duplicate, as was 100 μL of the original 103 inoculum sample. For the 104 inoculum sample, 10 μL was plated on PDA in duplicate. All inoculum plates were incubated at 37 °C for 18 h and counted.
Previous studies have demonstrated that Trichophyton species can exhibit up to an 80% loss of conidial germination for microconidia preparations from various media. 13 There-fore, for T. mentagrophytes, a concentrated microconidial preparation was diluted to 1.0–3.0 × 105 CFU/mL for use in the modified AATCC TM 100. This concentration of spores yielded final inoculum counts of 2 × 104 ± 1 × 104 CFU/mL (data not shown).
Antimicrobial Efficacy Test
For the antimicrobial efficacy test, AATCC TM 100-Assessment of Antibacterial Finishes on Textile Materials was followed, with the following exceptions. The conidial inoculum was adjusted to 2.0 × 104 ± 1.0 × 104 CFU/mL (A. niger) or 2.0 × 105 ± 1.0 × 105 CFU/mL (T. mentagrophytes) in Sabouraud Dextrose Broth (SDB) plus 0.05% Triton X-100. Following inoculation of the samples with 1 mL of the spore suspension each, the inoculated samples were incubated at 30 °C for 42 ± 2 h. This incubation time frame was selected based on the recommendation from the ISO draft standard and because A. niger has been shown to reach maximum biomass on solid surfaces in the 42-h time frame. 14 Following the inoculations, 1 mL of the inoculum was added to 99 mL of Letheen neutralization broth, mixed vigorously, and 1 mL of this diluted inoculum was plated on PDA in duplicate. The inoculum PDA plates were then incubated at 37 °C for 18–24 h (A. niger) or 24–72 h (T. mentagrophytes); colonies were counted following incubation to verify the inoculum concentration.
The test samples were recovered after incubation by adding 100 mL of Letheen neutralization broth to each sample and shaking vigorously for 120 ± 60 s using a wrist-action shaker (Burrell Scientific). Following shaking, 1 mL of each sample was removed and added to separate vital dye indicator jars for use with the MPN system to give a final dilution factor of 1:400 as described elsewhere. 15 The samples were then loaded into MPN YM cards per manufacturer's instructions, incubated at 25 °C for 72 h (A. niger) or 168 h (T. mentagrophytes), and enumerated. Additionally, 1 mL of each sample was also inoculated onto PDA plates for enumeration by plate counting to con-firm the MPN results. To compare the utility of the quantification protocol to qualitative antifungal methods, swatches of the same samples were also subjected to testing according to AATCC TM 30-2004, Test III.
Results and Discussion
To more accurately assess the ability of antimicrobial-treated articles to resist fungal growth, we used the MPN system in conjunction with a modified AATCC TM 100, and A. niger as the test organism. First, we wanted to establish the correct inoculum level for testing. As shown in Fig. 1, a target of 104 CFU/mL gave a consistent reading for both replicates compared to the inoculum count, with all calculated CFU readings falling in the same log range. For this reason, we chose 2.0 × 104 ± 1.0 × 104 CFU/mL as our target inoculum range.
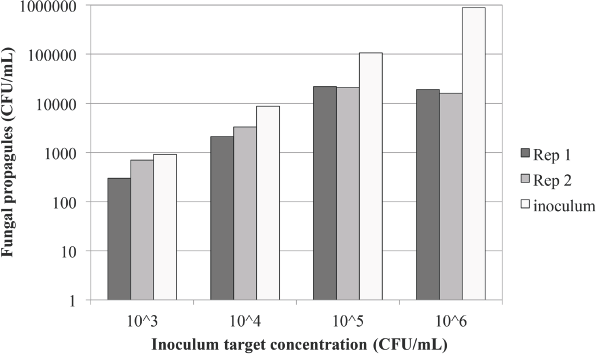
Consistency of propagule measurement. Target CFU concentrations/mL were determined by hemocytometer counts of prepared spore suspensions. Concentrations were empirically determined using standard plate count methods. A target CFU/mL of 104 provided reproducible results and was chosen as the target concentration for subsequent testing.
Next, we sought to determine the consistency of the MPN system for enumerating growth as compared to the standard plate counting method. In multiple experiments, we enumerated CFU counts using both the MPN system and standard plate counting on PDA. CFU counts were consistent between both enumeration methods over a range from 102–105 CFU (Fig. 2A), with an R 2 value of 0.993. Given that the spread of values varied more at the lower end of the counts, we narrowed our range to between 103–105 CFU, and the correlation improved to an R 2 value of 0.9951 (Fig. 2B). Therefore, the MPN is adequate for approximating the actual CFU values for samples.
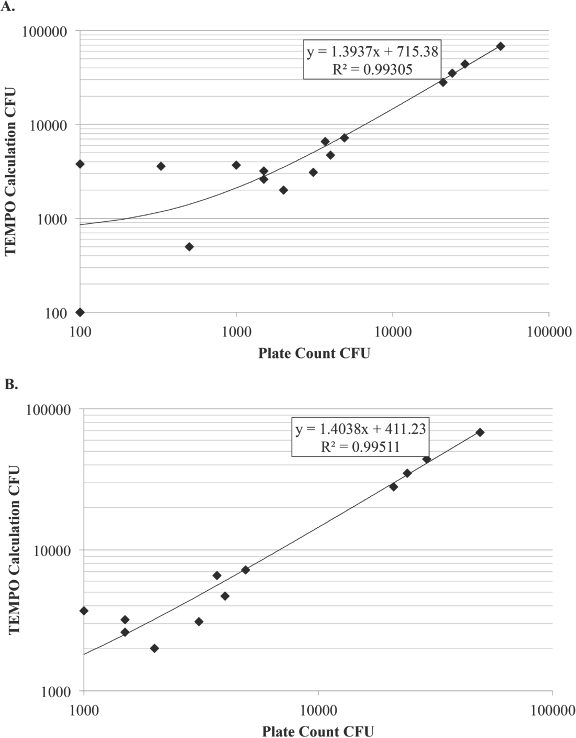
Correlation of MPN CFU to plate CFU. To determine the correlation between a most probable number system (MPN) and a standard plate count method, a dilution series was performed and enumerated using both methods. The resulting numbers were then graphed and an R 2 value was calculated. The lower detection limit for test was 102 CFU/mL and the upper limit was set to 105 CFU/mL. Across the range (102–105) there was a correlation of R 2 = 0.993 (A). However, when the range was narrowed (B) to eliminate the lower range (103–105) the correlation increased to R 2 = 0.995. Testing was conducted in triplicate.
Additionally, we compared our modified AATCC TM 100 to the current standard antifungal test for textiles, AATCC TM 30, Test III. The methods differ in how the test is conducted and in assessment of activity. AATCC TM 100 introduces a liquid inoculum, carried in a nutritive diluent, directly onto the test surface. The test surface and inoculum are incubated at the indicated time and temperature. Following incubation, the samples are removed, neutralized, and enumerated. The results are a quantitative assessment of surviving organisms. This is in contrast to the AATCC TM 30, Test III procedure. Samples are placed on nutritive agar saturated with spores of A. niger and inoculated with additional spores of A. niger. The scoring scheme is subjective and is based on whether the observed growth is microscopically (score = 1) or macroscopically (score = 2) visible. Additionally, percent coverage of growth is estimated, but this again is subjective in nature due to the shapes of samples subjected to testing and the scorer's assessment of the percent coverage. Samples that score 2 for macroscopic growth only at their edges provide excellent test cases to compare the subjective scoring of the TM 30, Test III to the objective calculation of log-reduction in growth provided by TM 100.
As shown in Table I, both treated and untreated articles scored 2's in TM 30, Test III, and therefore did not demonstrate antifungal activity using this test. When these same samples were assessed using the modified TM 100, greater than 2-log reductions in growth were observed for the treated articles corresponding to a greater than 99% reduction in growth of the fungus. These treated articles therefore demonstrated antifungal activity using the modified TM 100, whereas no antifungal activity was found using Test Method 30, Test III.
Comparison of Quantitative and Qualitative Fungal Test Methods
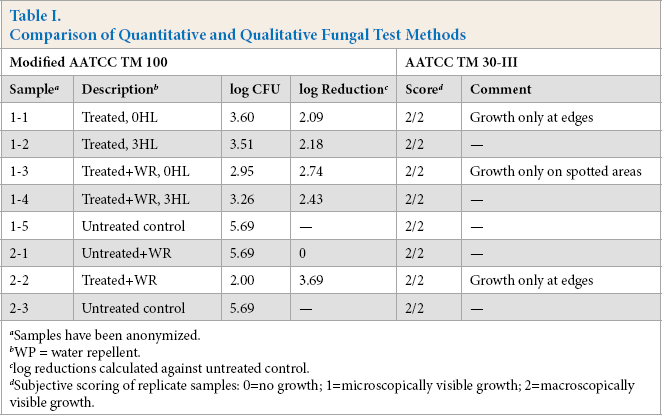
Samples have been anonymized.
WP = water repellent.
log reductions calculated against untreated control.
Subjective scoring of replicate samples: 0=no growth; 1=microscopically visible growth; 2=macroscopically visible growth.
Finally, we used this modified AATCC TM 100 to assess the ability of a treated article to inhibit the growth of T. menta-grophytes. As shown in Table II, antimicrobial treatment of polyester fabric inhibited fungal growth below the detectable limit for a greater than 1.7-fold log reduction. This log reduction increases to greater than 2.5-fold if the inoculum is used for the comparison.
Assessment of Antifungal Efficacy against T. mentagrophytes
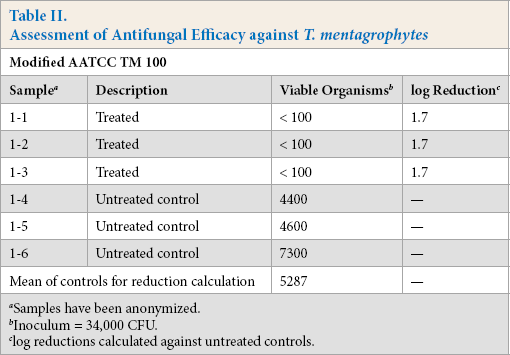
Samples have been anonymized.
Inoculum = 34,000 CFU.
log reductions calculated against untreated controls.
Conclusion
We have demonstrated a modified AATCC TM 100 test that is able to confirm antifungal activity for antimicrobial-treated articles that would otherwise demonstrate limited activity using the standard TM 30, Test III. The modified TM 100 provided objective measurement of antifungal activity and decreased test turn-around time from 7 days to 5 days. The MPN system is able to accurately measure the presence of fungi in samples, and is comparable to counting CFU by hand using dilution plating. Future work includes adapting this method for use with other fungi that may be of interest to the textile industry.
Footnotes
Acknowledgements
The authors wish to thank Joshua Smith at bioMérieux for his helpful discussions regarding mold growth and the MPN YM card system. The authors would also like to thank Duane Centola at Microban for critical critiques and expertise.
