Abstract
Dysregulation of innate and adaptive intestinal immune responses to bacterial microbiota is supposed to be involved in pathogenetic mechanisms of inflammatory bowel diseases (IBDs). We investigated expression of Toll-like receptor 2 (TLR2), TLR4, and their transmembrane coreceptor CD14 inbiopsy samples from patients with IBD and in non-inflamed gut mucosa from controls. Small intestine and colon samples were obtained by colonoscopy from patients with Crohn's disease (CD), ulcerative colitis (UC), and controls. Immunohistochemical analysis of cryostat sections using polyclonal and monoclonal antibodies specific for TLR2, TLR4, and CD14 showed a significant increase in TLR2 expression in the terminal ileum of patients with inactive and active UC against controls. Significant upregulation of TLR4 expression relative to controls was found in the terminal ileum and rectum of UC patients in remission and in the terminal ileum of CD patients with active disease. CD14 expression was upregulated in the terminal ileum of CD patients in remission and with active disease, in the cecum of UC patients in remission and with active disease, and in rectum of UC patients with active disease. Hence, dysregulation of TLR2, TLR4, and CD14 expression in different parts of the intestinal mucosa may be crucial in IBD pathogenesis.
I
Dysregulation of the intestinal immune response to bacterial flora is suggested to play a crucial role in the development of IBD with loss of normal physiological regulatory mechanisms of the local immune system. Perhaps a breakdown of oral tolerance to environmental antigens/commensal gut bacteria may be involved in the pathogenic mechanisms (Duchmann et al. 1995; Tlaskalova-Hogenova et al. 2004; Swidsinski et al. 2005). Animals serving as models of human IBD reared under germ-free conditions did not develop the disease and confirmed the important role of normal microbiota (Hudcovic et al. 2001; Elson et al. 2005).
Characteristic, highly conserved bacterial components, e.g., lipopolysaccharides (LPS), are recognized by innate immunity cells through pattern recognition receptors (PRRs) (TLRs, NODs, NaLPs, etc.). Toll-like receptors (TLRs) are molecules identifying selective bacterial patterns. The TLR family comprises human homologs of the Drosophila Toll protein, which play an essential role in the immune response to microbial infection. TLR2 has been shown to signal the presence of bacterial lipoproteins, lipoteichoic acids, peptidoglycan, and zymosan. Moreover, evidence suggested that human TLR2 interacts with CD14 and forms a LPS receptor complex. TLR4 is the predominant receptor for LPS from gram-negative organisms. MD2 protein has been shown to form a complex with TLR4 and is required for surface expression and LPS-regulated activation of TLR4. LPS is opsonized by lipopolysaccharide-binding protein (LBP) and recognized by CD14 on the macrophage. CD14 is a glycosylphosphatidyl inositol (GPI)-anchored receptor, which is not capable of generating a transmembrane signal. Nascent LPS–LBP– CD14 complex activates TLR4, which signals through adaptor protein MyD88 and serine kinase IL-1R-associated kinase 4 (IRAK4) and another adaptor protein TNF receptor-associated factor 6 (TRAF6). This finally results in activation of NF-κB and MAP kinases and triggers cytokine production (Aderem and Ulevitch 2000; Akira and Hemmi 2003; Kawai and Akira 2006).
There are few papers mentioning the expression levels of TLR2 and TLR4 in terminal ileum of patients with IBD (Cario and Podolsky 2000; Hausmann et al. 2002; Toiyama et al. 2006). In previous studies, only samples of colon were examined. Furthermore, data regarding expression levels of TLR2 and TLR4 by colonic intestinal epithelial cells have been controversial. Expression levels of mRNA purified from tissues of healthy persons have been shown to be the highest for TLR3, 4, 5, and 7 (Fusunyan et al. 2001; Zarember and Godowski 2002). Expression of TLR2, which recognizes molecular patterns in gram-positive organisms, has been reported at the level of mRNA isolated from intestinal epithelial cell (IEC) lines. However, TLR2 protein expression has not been confirmed in colonic epithelial cells (Cario and Podolsky 2000; Cario et al. 2002; Hausmann et al. 2002; Melmed et al. 2003). Our material involves the terminal ileum of patients with CD and UC. Thus, the present study is aimed at characterizing the expression of TLR2, TLR4, and their transmembrane coreceptor CD14 in the intestinal mucosa obtained from different parts of the intestine, including the terminal ileum from patients with UC and CD, and comparing it with controls.
Materials and Methods
Patients and Tissue Samples
A total of 155 intestinal tissue samples from 59 different patients undergoing colonoscopy were obtained for our study at the Institute of Clinical and Experimental Medicine (IKEM), Prague, Czech Republic (Table 1). Informed consent was obtained from all patients, and the protocol was approved by the Human Studies Committee of the IKEM. Each patient had a confirmed diagnosis by standard endoscopic and histological criteria, and disease activity was graded according to the Harvey–Bradshaw index (HBI) or Mayo score (Best 2006). Control samples were taken from patients who underwent colonoscopy because of colon cancer screening examination or polypectomy; all had normal endoscopic findings without macroscopic and microscopic evidence of inflammatory or neoplastic disease. Biopsies were collected from patients with active IBD and patients who were in remission and had no signs of active disease at the time of sampling. Grading of mucosal inflammation (remission or active disease) was performed by an expert gastroenterologist, who specializes in IBD, according the following criteria: CD and UC remission (HBI <3, Mayo score <4, respectively), CD and UC active (HBI >8, Mayo score >6, respectively). All specimens were taken from areas of rectum, cecum, and terminal ileum.
Immunohistochemistry
Fresh tissue samples were frozen in liquid nitrogen and stored at −80C until further processing. Cryostat sections (6 μm) were extracted with ice-cold acetone for 5 min and eluted with PBS with Tween. TLR2 and TLR4 expression was detected using rabbit polyclonal IgG and goat polyclonal IgG antibodies (Abs) [diluted 1:10 in human serum albumin (HSA); Santa Cruz Biotechnology, Heidelberg, Germany] for 1 hr at room temperature. Specificity of these Abs was confirmed by Medvedev et al. (2002) and Wang et al. (2002). Nonspecific binding of antibodies was blocked for 20 min at room temperature with normal swine or rabbit serum (Gibco, KRD Ltd.; Prague, Czech Republic) diluted 1:50 in PBS or horse serum (Vector Laboratories; Burlingame, CA) according to the manufacturer's instructions. In negative controls, primary Abs were substituted with 20% HSA (diluted 1:50; Sevapharma, Prague, Czech Republic) in PBS or preimmune goat or rabbit serum (Gibco). Biotinylated swine anti-rabbit (SwaR) or rabbit anti-goat (RAG) IgG Abs (Dako; Brno, Czech Republic) were used as the second step (1:50 in PBS; 1 hr at room temperature). CD14 and CD68 expression was detected using mouse monoclonal IgG Abs (Exbio; Prague, Czech Republic; Dako) (1:50 in HSA) for 1 hr at room temperature. Mouse immunoglobulin (MIgG) (1:50 in PBS) was used in negative controls. Biotinylated horse anti-mouse (HaM) IgG (Vector Laboratories) was used according to the manufacturer's instructions as a second-step antibody. Avidin/biotin peroxidase complex (ABC Elite; Vector Laboratories) was applied for 30 min. Horseradish peroxidase activity was visualized by treatment with 3,3′-diaminobenzidine–H2O2 reagent for 4–7 min. Finally, sections were lightly counter-stained with Harris’ hematoxylin. In the first experiments, blocking of endogenous peroxidase (Epx) was performed with 0.5% H2O2 in methanol (10 min at room temperature). However, this treatment highly suppressed expression of tested epitopes, chiefly membranous antigens CD14 and CD68. The final assessment was therefore based on comparison of the results obtained using specific antibody and control samples treated by HSA and MIgG. Sections were analyzed under an Olympus BX 40 microscope equipped with an Olympus DP 70 camera (Olympus; Tokyo, Japan).
Study population and tissue samples
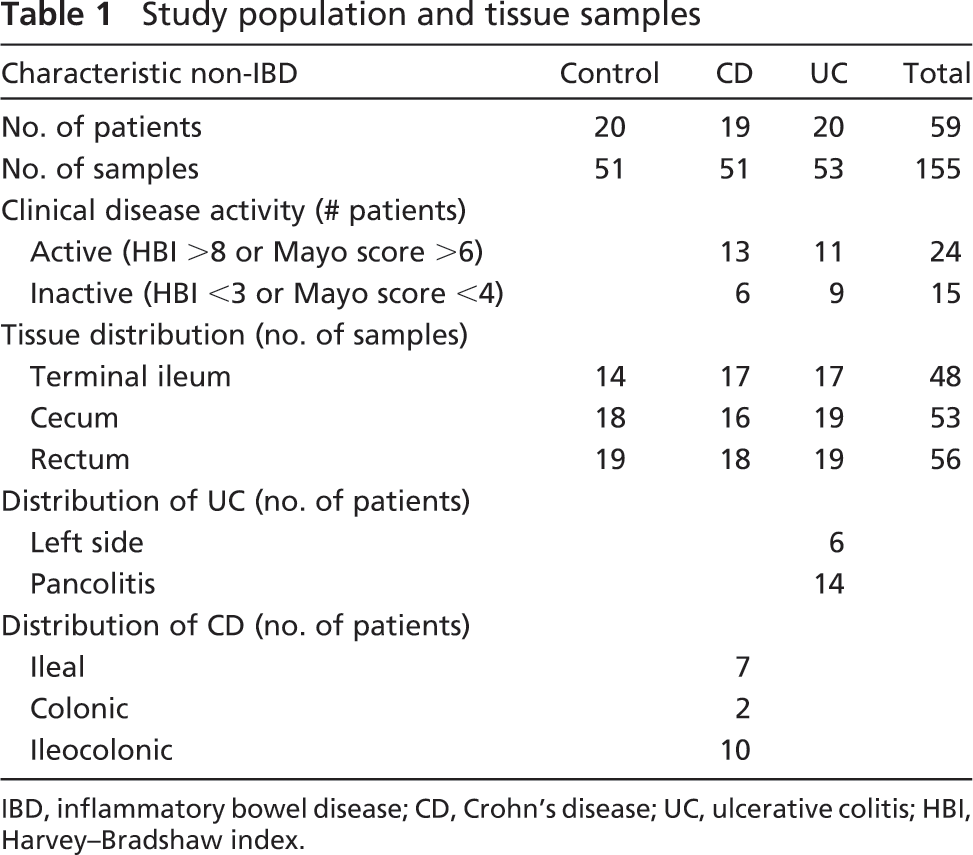
IBD, inflammatory bowel disease; CD, Crohn's disease; UC, ulcerative colitis; HBI, Harvey–Bradshaw index.
For fluorescence microscopy, FITC-labeled CD14 mouse anti-human MAb (Dako) was applied. CD68 epitope was detected by two-step immunofluorescence with the TR-labeled goat anti-mouse IgG (Jackson ImmunoResearch Laboratories; West Grove, PA). Confocal microscope was used for detection (SPE; Leica, Wetzlar, Germany). Digitized images were then revised using Adobe Photoshop 6 (Adobe Systems; Mountain View, CA). Pathologists regularly analyzed at least two sections per sample. Degree of expression of TLR2, TLR4, and CD14 in the specimens was blindly evaluated by a pathologist, independently confirmed by a second examiner, and graded microscopically as follows: for TLR2 and TLR4: 0, the same as background; 0.5, close to background; 1, well-marked positivity; 1.5, focally enhanced; 2, strong positivity; 2.5, very strong positivity; for CD14: 0, no cells presented; 0.5, sporadic single cells; 1, scattered single cells; 1.5, scattered cells with discrete clusters; 2, large groups or clusters of cells; 2.5, dense dissemination.
Data are expressed as mean ± SD of the total of about gained values (Table 2). Statistical analyses were performed using Kruskal–Wallis and Mann—Whitney tests. Differences were considered significant when p<0.05.
Results
TLR 2 Expression
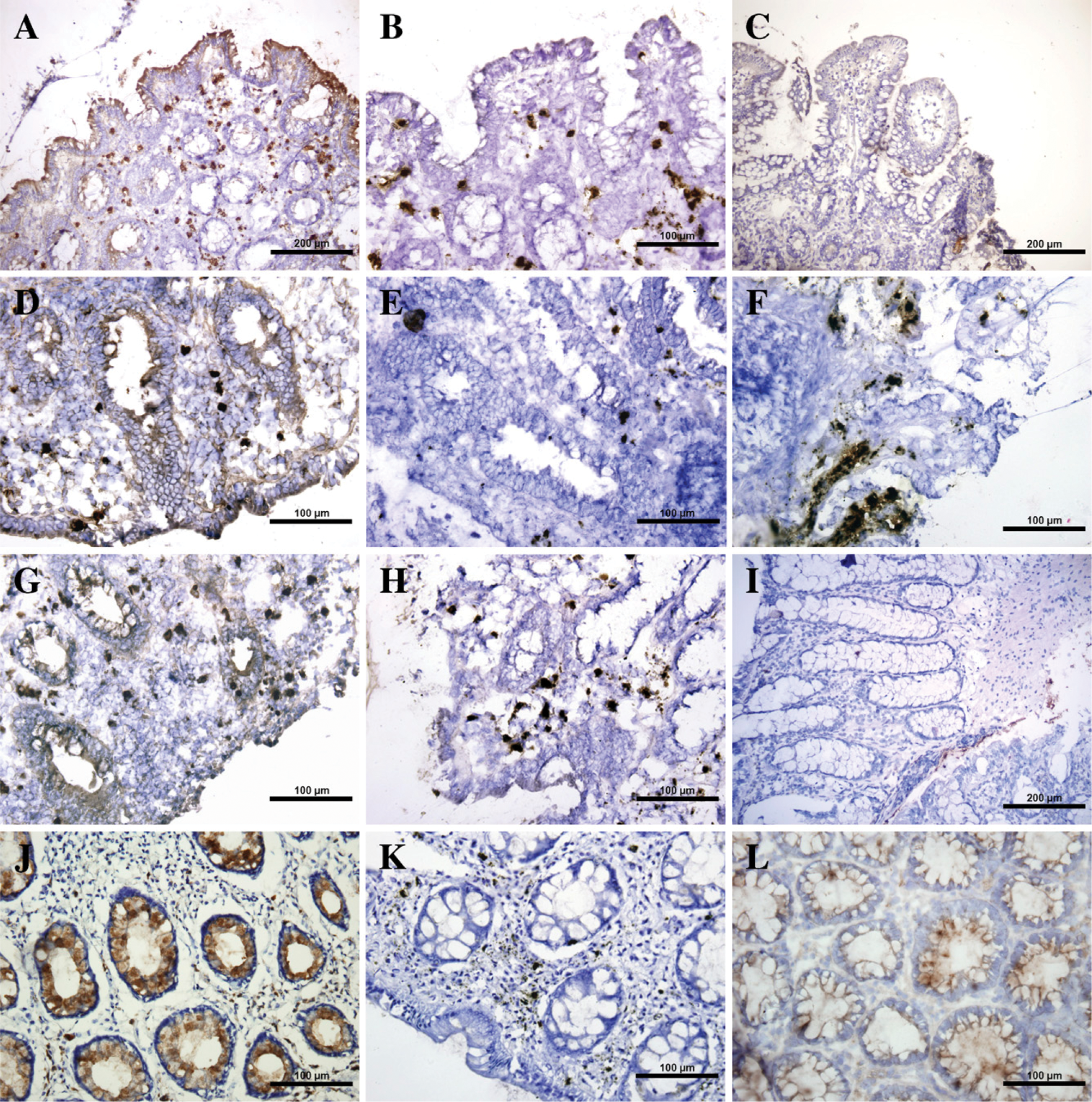
Control samples of normal mucosa were only weakly positive for TLR2 except cecum, where the presence of positive epithelial cells was comparable with that in IBD patients. In the specimens obtained from patients with inflamed mucosa, TLR2-positive epithelial cells were accumulated on the mucosal surface rather than in the crypts (Figure 1A). The lamina propria (LP) remained negative or rarely displayed sporadic single cells. Both macroscopically inflamed and non-inflamed areas of mucosa from the cecum and rectum as well as terminal ileum showed cells positive for TLR2. It is noteworthy that statistically significant upregulation of TLR2 protein expression was observed in the ileal epithelium from UC patients with inactive (p=0.0256) and active disease (p=0.0078) as compared with the normal intestine (Table 2; Figures 1A–1C).
TLR4 Expression
Epithelial TLR4 cell expression was usually stronger than TLR2 expression. It was abundantly positive in colonic epithelium of all UC and CD patients (often but not always stronger on the surface than in the depth of crypts), whereas in the terminal ileum the expression was less evident. In both TLR2 and TLR4, expression was more apparent in the surface epithelium, mainly in samples with weaker positivity. In a case with strong positivity, crypts were equally or more involved. TLR4-positive epithelial cells were also detected in non-IBD colonic mucosa (Figure 1L), whereas minimally positive cells were found in the terminal ilea of controls. LP was close to negative as in TLR2. Differences were processed and reached statistical significance in the terminal ileum (p=0.0228) and rectum (p=0.0087) of UC patients in remission (Table 2; Figures 1G–1J) and in the terminal ileum (p=0.0106) of CD patients with active disease (Table 2; Figure 1D).
Expression of TLR2, TLR4, and CD14 in patients with active and inactive UC and CD compared with controls
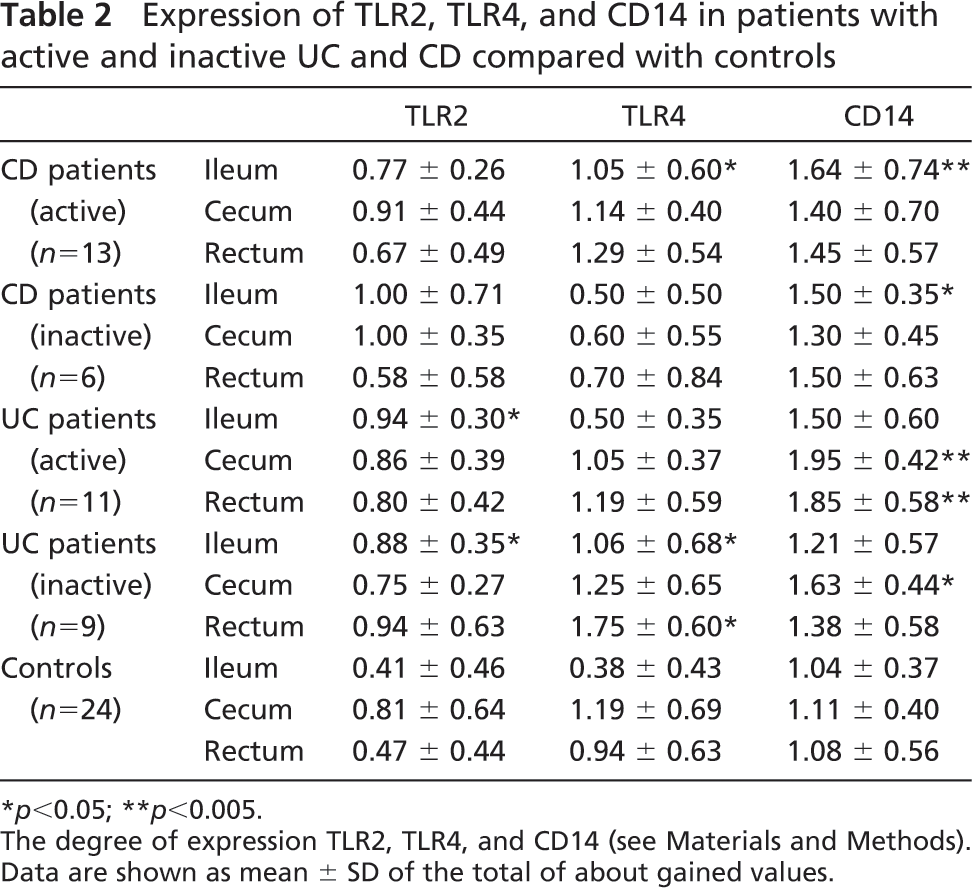
∗ p<0.05; ∗∗p<0.005.
The degree of expression TLR2, TLR4, and CD14 (see Materials and Methods).
Data are shown as mean ± SD of the total of about gained values.
(
CD14 Expression
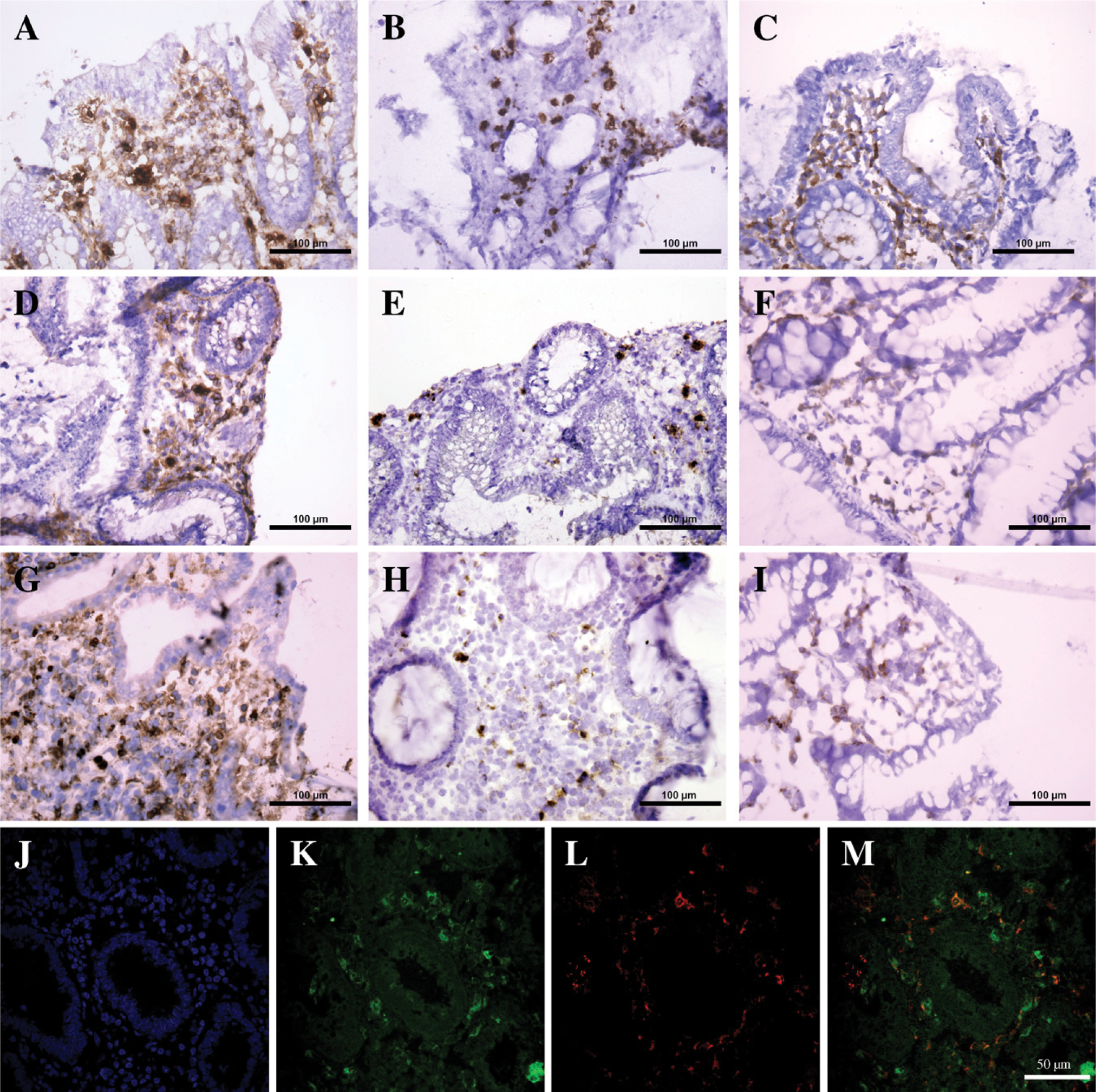
Many mononuclear cells in the LP displayed strong membranous expression of CD14 in both UC and CD patients, whereas epithelial cells remained negative. The membranous positivity was highly different from the coarse granular product of endogenous peroxidase; this latter was also evident in control sections. Positivity of a lesser degree was noticed in non-IBD controls (Figures 1C, 1F, and 1I). Scattered positive cells were also seen in the mucosal muscular layer. Significant upregulation of CD14 expression was identified in the terminal ileum of CD patients in remission (p50.0170) and also those with active disease (p=0.0056) (Figure 1A). Moreover, significant upregulation was found in the cecum of UC patients in remission (p=0.0129) and with active disease (p=0.0001) and in the rectum of UC patients with active disease (p=0.0039) as compared with non-IBD tissues (Table 2; Figures 1D and 1G). To confirm the exclusive or highly preferential expression of CD14 epitope on macrophages, we used an antibody specific for the typical macrophage marker CD68. The latter antibody matched the results yielded by CD14 (Figures 2J–2L). In fluorescence microscopy, about one half of the fluorescence cells showed coexpression of CD14 and CD68 (Figure 2M).
Interestingly, in our study the extent of colitis in UC and the distribution of inflammation in CD have no influence on expression of TLRs and CD14. Only two cases from IBD did not match the mean expression. Infiltration of CD14-positive macrophages in the LP was significantly more intense in the ileal samples of active CD, which differed from the higher density in the cecum and rectum of UC.
Discussion
In the present study we describe an unexpected finding of a significant increase of TLR2 expression present in the non-impaired (by inflammation) part of the gut, in the terminal ileum of patients with UC, as compared with controls. Statistically significant upregulation of TLR4 expression was observed in the terminal ileum and rectum of UC patients and in the terminal ileum of CD patients as compared with healthy controls. CD14 expression was upregulated in the terminal ileum of CD patients and in the cecum and rectum of UC patients when compared with controls. Cario et al. affirm that IECs of normal non-diseased mucosa constitutively express TLR3 and TLR5. On the other hand, in their study TLR2 and TLR4 were described to be present with low intensity. We focused our interest on three different parts of the bowel, which can play different roles in the development of disease and expression of TLRs. We have demonstrated low levels of TLR2 protein expression in the non-diseased sections of terminal ileum and rectum of controls. However, TLR2-positive cells were found in samples of healthy cecum in numbers comparable to those from the cecum of IBD patients. We may explain this discrepancy by the fact that healthy cecum is a part of the human bowel containing a high density of bacteria combined with low motility (peristalsis) (Savage 1999; Tlaskalova-Hogenova et al. 2004). The mucosal barrier is therefore attacked here by potentially pathogenic bacteria and their cell wall components, which are recognized by TLRs. In the present study, significant increase in TLR2 expression was observed in the terminal ileum in patients with UC as compared with controls, even if macroscopic inflammation or backwash ileitis was lacking.
Interestingly, TLR2- and TLR4-positive IECs were found in the terminal ileum in the area of the gut not usually affected by inflammation in these patients. Moreover, TLR2 expression is significantly increased, regardless of whether assessed in active or inactive disease of UC patients, and TLR4 expression is detected in UC patients in remission. A possible explanation of this unexpected finding could be that this upregulation reflects the activation of innate immunity cells by bacterial components, until now unknown, present also in the proximal part of the gut of UC patients. Another explanation is that immunosuppression may influence bacterial colonization or bowel peristalsis in the terminal ileum. Thus, it seems that the terminal ileum can sensitively react to “potentially pathogenic” but still undiscovered components of microbiota by immunological mechanisms impairing homeostasis in the distant part of the gut, i.e., colonic tissues.
Moreover, we found statistically significant upregulation of TLR4 expression in biopsy samples from the ileum and rectum of UC patients in remission and in the terminal ileum of CD patients with active disease. Recent studies (Cario and Podolsky 2000; Abreu et al. 2002; Hausmann et al. 2002) have described low TLR4 expression by IECs in healthy human biopsies and significantly increased expression of TLR4 in patients with IBD. These results corroborated our data because we found high TLR4 expression in samples of inflamed IBD tissues (terminal ileum of CD patients and ileum and rectum of UC patients). Large quantities of luminal LPS are usually well tolerated within the healthy intestine. In mouse model, it was also show that downregulation of TLR4 expression is responsible for LPS tolerance. Some authors supposed that other mechanisms along with it could be involved in this process (Nomura et al. 2000). However, increased TLR4 expression in IBD patients may be the consequence of impaired host tolerance toward luminal LPS antigens (Duchmann et al. 1995; Lange et al. 1996). Immunohistochemical findings have been confirmed by studies measuring TLR expression on mRNA level (Abreu et al. 2002; Hausmann et al. 2002; Melmed et al. 2003; Furrie et al. 2005). Our own experiments in this direction are in progress and are the subject of another publication (Zanvit et al., unpublished data).
(
Monocytes recruited to normal intestinal mucosa are characterized by downregulation of the proinflammatory phenotype and functional profile. On the other hand, monocytes recruited to the inflamed mucosa do not undergo such stringent downregulation and retain their proinflammatory potential (Smith et al. 2005). Other authors showed that nearly a third of LP macrophages in the inflamed mucosa of patients with IBD express CD14 (Rugtveit et al. 1994; Grimm et al. 1995; Rogler et al. 1997). In our study, cells possessing CD68 (macrophage marker) exhibited strong expression of CD14 in the LP in both UC and CD patients in comparison with controls. Significant upregulation of CD14 expression on macrophages was identified in areas affected by IBD (terminal ileum of active CD patients and CD patients in remission, cecum of UC patients in active and remission stage, and rectum of UC patients active disease) as compared with non-IBD (control) samples. However, positivity of a lesser degree was also noticed in non-IBD controls. We did not observe CD14 in intestinal epithelial cells. Recent studies showed that human enterocytes in vivo do not express membrane CD14 at a level detectable by immunohistochemistry (Pugin et al. 1993; Schumann et al. 1994; Grimm et al. 1995).
In conclusion, there is increasing evidence that intestinal microflora play a crucial role in induction and maintenance of inflammation in patients with IBD. IECs form the first line of contact between the host and the normal microbiota and provide a connection with the immune system via cells in the LP. Moreover, the intestinal epithelium is an active participant in the mucosal immune response through its expression of proinflammatory genes, secretion of inflammatory cytokines, and recruitment of inflammatory cells operative against pathogenic bacteria and their products (Autschbach et al. 2002; Tlaskalova-Hogenova et al. 2002). PRRs, which are situated also on epithelial cells and other innate immunity cells, participate in recognition of various characteristic bacterial components. Correct recognition and adequate response against commensal bacteria are essential for healthy function and maintenance of the mucosal barrier.
IBD is the result of concerted actions of innate immune signals such as the binding of LPS to TLR4, as well as adaptive immune signals such as INF-γ produced by Th1 cells. There are numerous data demonstrating the presence of T cells directed to bacterial components of microflora, suggesting loss of tolerance to gut microflora (Duchmann et al. 1995, 1999). T regulatory (Treg) cells are effective in the prevention and downregulation of IBD animal models (Singh et al. 2001). CD4+CD25
Footnotes
Acknowledgements
This work was supported by the Czech Science Foundation (projects no. #310/03/H147 and #303060974); Grant Agency of Academy of Sciences (project #S500200572); Ministry of Education, Youth and Sports of the Czech Republic (project #2B06155); and Institute of Microbiology (project #AV0Z50200510).
