Abstract
Among tight-junction proteins, claudins, which play a key role in paracellular transport across epithelia, claudins 1 to 5 are expressed in the intestine, and changes in their abundance and/or distribution are considered to contribute to various gastrointestinal diseases. We investigated, by reverse transcription-PCR, immunoblot, and immunofluorescence analyses, which other claudin species were expressed in the mouse intestine, and whether they showed unique expression profiles. Rabbit polyclonal antibodies against mouse claudin-8, claudin-12, and claudin-15 were generated, and their specificity was verified by immunoblotting using COS-7 cells transfected with individual claudin cDNAs. Claudin-7, −8, −12, −13, and −15 appeared to be expressed in the duodenum, jejunum, ileum, and/or colon with remarkable variations in the expression levels along the intestinal tract, and had distinct subcellular localization in the intestinal epithelium. In addition, claudin-13 and −15 exhibited gradients along the crypt-surface axis of the colon. By contrast, claudin-6, −9, −10, −11, −14, −16, −18, and −19 were not observed in the intestine. Our results indicate that five additional species of claudins have very complex expression patterns along and within the intestine, and that this may reflect differences in paracellular permeable properties, providing valuable resources for studying the significance of these claudins in gastrointestinal disorders. This manuscript contains online supplemental material available at http://www.jhc.org. Please visit this article online to view these materials.
T
Claudins are 20-27 kilodalton tetraspan proteins with a short cytoplasmic N terminus, two extracellular loops, and a C-terminal cytoplasmic domain (Furuse et al. 1998a; Tsukita et al. 2001). They consist of over 20 members of a gene family in mice and humans, and expression of distinct claudins depends on the tissue and cell type (Tsukita et al. 2001; Turksen and Troy 2004). Claudins are capable of forming continuous networks of intramembranous fibrils (tight-junction strands) (Furuse et al. 1998b), thus forming the backbone of tight junctions. Moreover, several lines of evidence have indicated that claudins could be the major determinant of the barrier properties, including size and charge selectivities. This concept has been drawn from studies of knockout mice (Gow et al. 1999; Furuse et al. 2002; Nitta et al. 2003; Kitajiri et al. 2004) and inherited human diseases (Simon et al. 1999; Wilcox et al. 2001; Hadj-Rabia et al. 2004) of claudin genes, as well as overexpression experiments using wild-type and mutated claudins in cultured cell lines. In particular, the first extracellular loop, in which the position and number of charged amino acids vary widely depending on each claudin, is strongly suggested to influence the paracellular charge selectivity (Colegio et al. 2002; Van Itallie et al. 2003; Hou et al. 2005; for review, see Van Itallie and Anderson 2006).
Among claudin family members, claudins 2 to 5 and 2 to 4 are reported to be expressed in the rat and mouse intestine, respectively (Rahner et al. 2001; Tamagawa et al. 2003), and expression of claudin-1, −3, −4, and −5 is observed in the human intestine (Bürgel et al. 2002). There are also variations in the expression levels and subcellular localization of these claudins throughout the segments of the intestinal tract and/or along the crypt-to-villus axis (Rahner et al. 2001; Tamagawa et al. 2003). Furthermore, the expression and/or distribution of claudins are altered in many gastrointestinal disorders, and these changes are proposed to play important roles in the pathophysiology of such diseases (for reviews, see Clayburgh et al. 2004; Fasano and Shea-Donohue 2005). For example, expression levels of claudin-2 are elevated in Crohn's disease and ulcerative colitis, collectively known as inflammatory bowel disease (IBD), whereas those of claudin-3 and −4 are sometimes decreased or redistributed in the disease (Prasad et al. 2005). It also has been demonstrated, using the human intestinal epithelial cell line T84, that the interferon-γ-triggered internalization of claudin-1 and −4, as well as that of occludin and JAM-A, could contribute to the pathogenesis and barrier dysfunction in IBD (Bruewer et al. 2005; Utech et al. 2005; for review, see Chiba et al. 2006). In colorectal cancer, the expression of claudin-1 is frequently upregulated (Miwa et al. 2001; de Oliveira et al. 2005; Dhawan et al. 2005; Resnick et al. 2005), whereas reduced claudin-1 expression has been revealed to be associated with the recurrence and prognosis in colon cancer (Resnick et al. 2005). However, little is known about whether, in addition to claudins 1 to 5, other claudin species are expressed in the intestine. We show in the present work that claudin-7, −8, −12, −13, and −15 are differentially expressed and distributed along the mouse intestinal tract. We discuss the potential contribution of these claudins to charge-selective properties in the intestine. Our findings provide fundamental information that will help to elucidate whether and how these claudins are altered and/or involved in gastrointestinal diseases.
Materials and Methods
Animals and Tissue Preparation
C57BL/6 mice (Clea, Tokyo, Japan) were housed individually in specific-pathogen-free conditions at the Center for Animal Resources and Development, Sapporo Medical University School of Medicine. Eight- to twelve-week-old male and female mice were anesthetized with diethyl ether, and specimens of the intestine (the duodenum, jejunum, ileum, and colon), kidney, testis, bladder, and stomach were obtained. All aspects of the study were approved by the animal use and care committee of Sapporo Medical University School of Medicine.
Isolation of Claudin-12 and −15 cDNA and Construction of Their Expression Vector
Total RNA was isolated from the heart of the male C57BL/6 adult mouse using TRIZOL reagent (Gibco BRL, Grand Island, NY), and RT-PCR was performed using an RT-PCR kit (Perkin-Elmer, Branchburg, NJ) according to the manufacturer's instructions. Mouse claudin-12 and −15 cDNA fragments, in which EcoRI sites were created together with the Kozak sequence, were amplified using Pfu DNA polymerase (Stratagene, La Jolla, CA) with RT products as templates. The forward and reverse PCR primers were as follows: claudin-12, 5î-GAATTCCTGCCATGGGCTGCCGAGATGT-3î [nucleotides (nt) 286-309] and 5î-GAATTCTTAAGTGCTGTGTGAGACTACT-3î (nt 1,006-1,027); claudin-15, 5î-GAATTCTCCCAGCATGTCGGTAGC-3î (nt 233-250) and 5î-GAATTCCTACACGTATGCGTTTTTGCC-3î (nt 903-923). The PCR products were subcloned into the expression vector pD402A (a gift from Dr. D. Lohnes, Institut de Recherches Cliniques de Montreal) EcoRI site, which was located between the phosphoglycerate kinase promoter and polyadenylation site, resulting in pD402A-KmCL12 and pD402A-KmCL15. Their sequences and orientations were confirmed by dye terminator cycle sequencing (ABI Prism 377-18; Perkin-Elmer).
Analysis of Gene Expression of Claudins
For analysis of gene expression, total RNA was isolated from organs as described above, and RT-PCR was performed as previously described (Chiba et al. 1997,2003,2005; Satohisa et al. 2005). The PCR primers for mouse cDNAs were as follows: claudin-9 (GenBank accession no. AF124424), 5î-CTTTCATCGGCAACAGCATCG-3î (nt 151-171) and 5î-TCTTATCCAGTCCCGAAGCAC-3î (nt 670-690); claudin-10 (GenBank accession no. AF124425), 5î-TCGCCTTCGTAGTCTCCATC-3î (nt 35-54) and 5′-CCTTGATACTTGGTCCGAGAAG-3î (nt 620-641); claudin-11 (GenBank accession no. AF124426), 5î-TCACAACGTCCACCAATGACTG-3î (nt 68-89) and 5î-GGCACATACAGGAAACCAGATG-3î (nt 414-435); claudin-12 (GenBank accession no. NM022890), 5î-CTTTGCGGGGACTCTGCTTCC-3î (nt 364-384) and 5î-ATGAATAGGGCTGTGAGTAAGTGT-3î (nt 945-968); claudin-13 (GenBank accession no. NM020504), 5î-GTTTGGAGGGTGACCTTTCCA-3î (nt 282-302) and 5î-GAAAGTCTCTGCATACTGGGAT-3î (nt 747-768); claudin-14 (GenBank accession no. AF124429), 5î-GCTTCCTGCTTAGCTTCCTG-3î (nt 38-57) and 5î-TCCTCCTGGCAGGATAAGC-3î (nt 554-572); claudin-15 (GenBank accession no. NM021719), 5î-GCTTCTTCATGTCAGCCCTG-3î (nt 265-284) and 5î-TTCTTGGAGAGATCCATGTTGC-3î (nt 556-577); claudin-16 (GenBank accession no. NM053241), 5î-GAACCTGCGATGAGTACGAC-3î (nt 503-522) and 5î-TCCTCATGGCATAAGGGTAG-3î (nt 921-940); claudin-18 (GenBank accession no. NM019815), 5î-TGGTAGCATGGATGACTCTGC-3î (nt 385-405) and 5î-TTGCTGTCATCTGGTGTCAGG-3î (nt 670-690); claudin-19 (GenBank accession no. NM153105), 5î-TCCTCTTGGCAGGTCTCTGT-3î (nt 442-461) and 5î-CCTGGTCCTCATTCTGGTGT-3î (nt 943-962). The primers for claudins 6 to 8 and 36B4 have been described previously (Kubota et al. 2001). To make sure that amplifications were in the linear range, PCR was performed for three different cycles of between 20 and 32 cycles, depending on the genes analyzed. Aliquots of PCR products were loaded onto 1% agarose gel and analyzed after staining with ethidium bromide.
Generation of Polyclonal Antibodies
To generate rabbit polyclonal antibodies (pAbs) against claudin-8, claudin-12, and claudin-15, the following conjugated peptides, in which a cysteine residue was added at the N-terminal ends, were used as antigens. Polypeptides CSFHAEKRSPSIYSKSQY, CGMHTYSQPYSSRSRLSAI, and CTSDESDISFGKYGKNA, corresponding to the cytoplasmic domains of mouse claudin-8 [amino acids (aa) 208-224], claudin-12 (aa 216-233), and claudin-15 (aa 210-225), respectively, were synthesized and coupled via the cysteine to keyhole limpet hemocyanin. The antisera were affinity-purified with glutathione S-transferase fusion protein with the carboxyl-terminal cytoplasmic domains of the corresponding claudins. Specificity of these antibodies was verified by Western blot analysis using COS-7 cells transiently transfected with individual expression vectors containing mouse claudin-1 to claudin-8 cDNAs (Ishizaki et al. 2003), as well as those of mouse claudin-12 and −15 cDNAs. In brief, COS-7 cells were grown on 60-mm culture plates and transfected with 10 μg of each expression vector using Lipofectamine 2000 reagent (Life Technologies, Gaithersburg, MD) according to the manufacturer's protocol. Total cell extracts were isolated 24 hr after transfection, followed by Western blot analysis.
Immunohistochemistry
Ten-μm-thick frozen sections of the mouse intestine were stained with hematoxylin-eosin, or fixed in 95% ethanol for 30 min at 4C and in 100% acetone for 1 min at room temperature. After being washed three times with PBS, the sections were incubated for 1 hr at room temperature with primary antibodies, including a mouse monoclonal antibody against occludin (Zymed, San Francisco, CA), and rabbit pAbs against claudin-7 (Satohisa et al. 2005), claudin-8, claudin-12 and claudin-15. Rabbit pAbs against claudin-13 were obtained from Zymed and Aviva Systems Biology (San Diego, CA). Preimmune serum was used as controls. The sections were then rinsed with PBS and incubated for 1 hr at room temperature with appropriate secondary antibodies labeled with Alexa Fluor 488 (green) or 594 (red) (Molecular Probes, Eugene, OR). Samples were examined using a laser-scanning confocal microscope (MRC 1024; Bio-Rad, Hercules, CA), a fluorescent microscope (IX71; OLYMPUS, Tokyo, Japan) and/or a light microscope (ECLIPSE E600; Nikon, Tokyo, Japan).
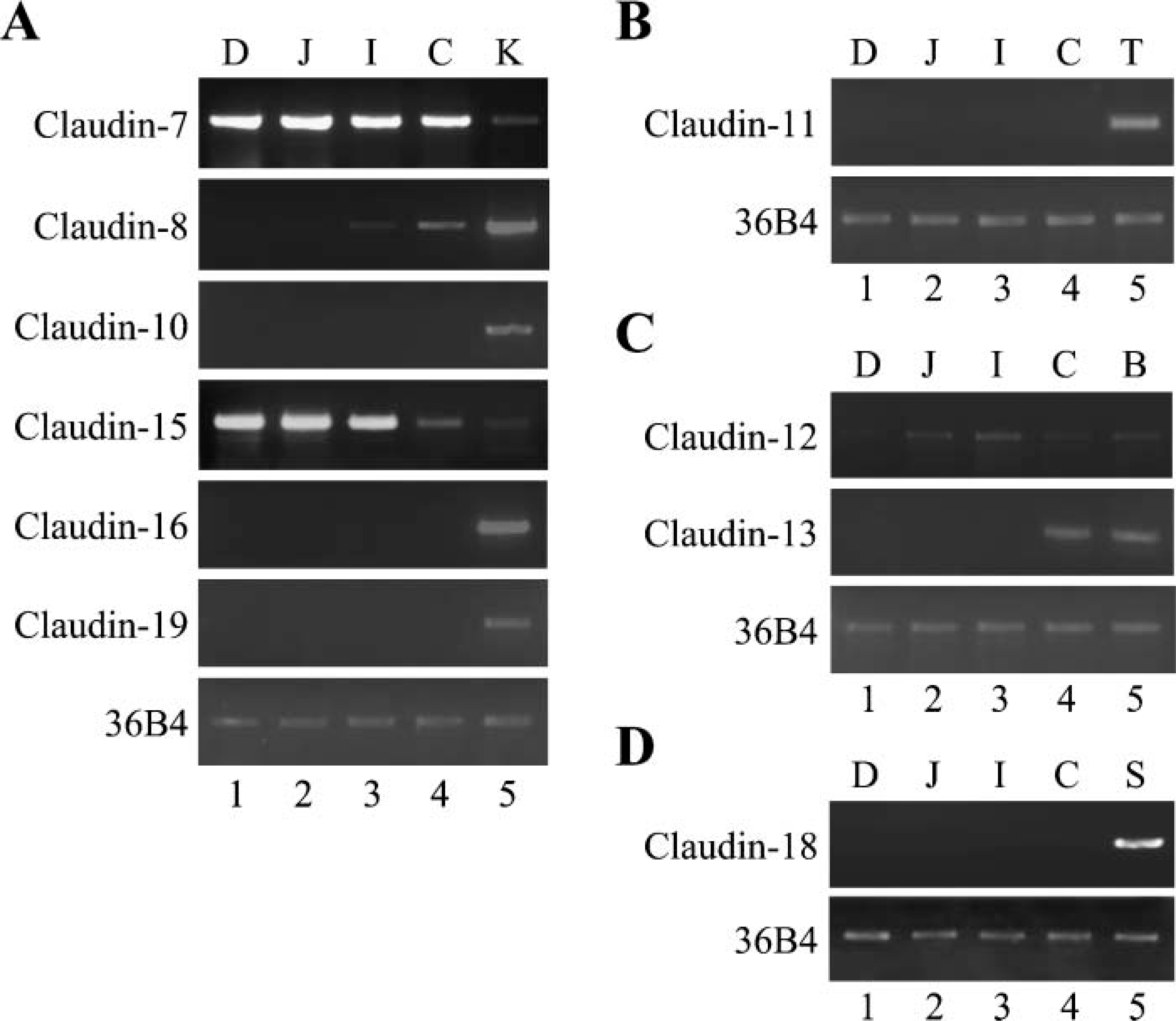
Claudin-7, −8, −12, −13, and −15 genes are differentially expressed in the mouse intestine. One μg of total RNA from each organ was subjected to RT-PCR analysis for the indicated genes. (
Gel Electrophoresis and Immunoblotting
Each segment of the mouse intestine was washed twice with PBS, sonicated for 20 sec in 2 ml of ice-cold NaHCO3 buffer (1 mM NaHCO3 and 1 mM PMSF, pH 7.5), and put on ice for 30 min. Total cell lysates were resolved by one-dimensional SDS-PAGE and electrophoretically transferred onto a polyvinylidene difluoride membrane (Immobilon; Millipore, Bedford, MA). The membrane was saturated with PBS containing 4% skim milk and 0.1% Tween 20, and incubated for 1 hr at room temperature with primary antibodies in PBS. The antibodies described above were used as primary antibodies. After rinsing in PBS containing 0.1% Tween 20, the membrane was incubated for 1 hr at room temperature with horseradish peroxidase-conjugated anti-mouse or anti-rabbit IgG (Vector Laboratories, Burlingame, CA) in PBS containing 0.3% skim milk. It was then rinsed again, and finally reacted using an enhanced chemiluminescence (ECL) Western blotting detection system (Amersham Pharmacia Biotech, Buckinghamshire, UK). The blots were stripped with Restore Western blot stripping buffer (Pierce, Rockford, IL) according to the manufacturer's recommendations and immunoprobed with a rabbit pAb against actin (Sigma; St Louis, MO). When immunoblotted signals were faint, the ECL Advance Western blotting detection system (Amersham Pharmacia Biotech) was used according to the manufacturer's instructions. Signals in immunoblots were quantified using Scion Image 1.62c (Frederic, MD).
Results
Distinct Expression Patterns of Claudin-7, −8, −12, −13, and −15 mRNAs Along the Mouse Intestinal Tract
Because claudins 1 to 5 are known to be expressed in the intestine (Rahner et al. 2001; Bürgel et al. 2002; Tamagawa et al. 2003), we first determined, by RTPCR analysis, mRNA levels of other claudins throughout the mouse intestinal tract. For these experiments, gene expression of claudins in the kidney, testis, bladder, or stomach was analyzed as a positive control (Gow et al. 1999; Niimi et al. 2001; Kiuchi-Saishin et al. 2002; Acharya et al. 2004; Luk et al. 2004). As shown in Figure 1, gene expression of claudin-7, −8, −12, −13, and −15 was detected in the duodenum, jejunum, ileum, and/or colon. The claudin-7 gene was strongly expressed in the duodenum, jejunum, ileum, and colon, and the levels were higher than those in the kidney (Figure 1A). Expression of claudin-15 mRNA was similarly detected in the duodenum and jejunum, but less and faintly observed in the ileum and colon, respectively (Figure 1A). It should also be noted that claudin-15 transcripts in these organs were expressed at high levels compared with those in the kidney. The claudin-12 gene was highly expressed in the ileum and moderately and weakly observed in the jejunum and colon, respectively, whereas few claudin-12 transcripts were detectable in the duodenum (Figure 1C). Claudin-8 mRNA, which was strongly detected in the kidney, was moderately and faintly expressed in the colon and ileum, respectively, but no transcript was detected in the duodenum or jejunum (Figure 1A). Claudin-13 transcripts were clearly observed in the colon, but not detectable in the duodenum, jejunum, or ileum (Figure 1C). In contrast, claudin-6, −9, −10, −11, −14, −16, −18, and −19 mRNAs were not expressed in the duodenum, jejunum, ileum, or colon (Figures 1A, 1B, and 1D, and data not shown). Thus, in the mouse intestinal tract, five additional species of claudins, claudin-7, −8, −12, −13, and −15, appeared to be expressed with distinct expression patterns.
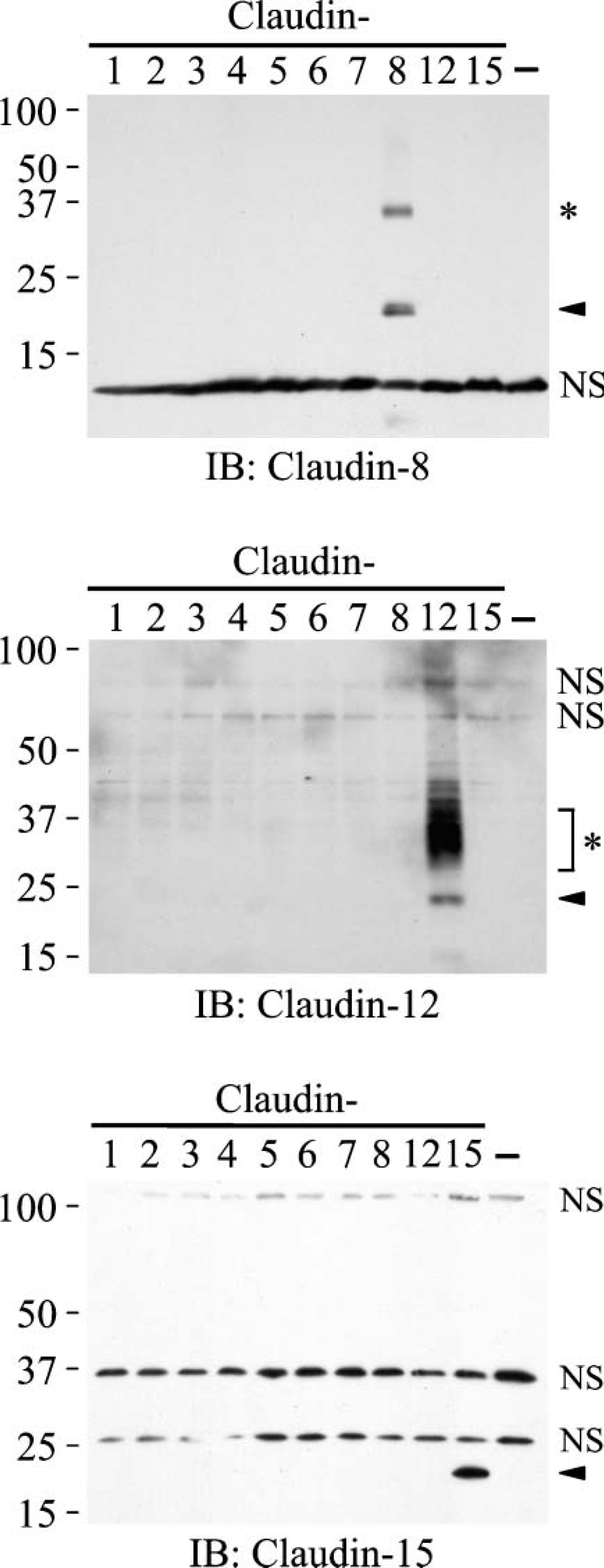
Anti-claudin-8, anti-claudin-12, and anti-claudin-15 antibodies selectively recognize the corresponding claudins. COS-7 cells were transfected with either empty or claudin-1 to claudin-8, claudin-12, and claudin-15 expression vectors. Five μg of total cell lysate was separated by SDS-PAGE and immunoblotted (IB) with anti-claudin-8, anti-claudin-12, or anti-claudin-15 polyclonal antibodies (pAbs), followed by chemiluminescence detection. The transfected mouse claudin cDNAs are indicated at the top of each lane. The mobility of molecular mass markers (kilodaltons) is indicated on the left. Arrowheads and asterisks indicate specific signals. NS, nonspecific signal.
Specificity of Anti-claudin-8, Anti-claudin-12, and Anti-claudin-15 Antibodies
We next generated novel pAbs against claudin-8, claudin-12, and claudin-15, like those against claudin-7 (Satohisa et al. 2005), because mRNAs of these claudins were expressed in the mouse intestine as described above. To check the specificity of these antibodies, COS-7 cells were transiently transfected with either individual expression vectors containing mouse claudin-1 to claudin-8, claudin-12, and claudin-15 cDNAs or an empty vector, and their lysates were subjected to gel electrophoresis and immunoblotting. As shown in Figure 2, the anti-claudin-8 pAb reacted with claudin-8, but not with claudins 1 to 7, claudin-12, or claudin-15. Similarly, the anti-claudin-12 and anti-claudin-15 pAbs selectively recognized claudin-12 and claudin-15, respectively. Additional specific bands were observed in the immunoblots using anti-claudin-8 and anti-claudin-12 pAbs (Figure 2, asterisks; 36 and 29-40 kilodaltons, respectively), suggesting that these Abs might also recognize posttranslationally modified claudins and/or dimers of the respective claudins. Claudin-8, claudin-12, and claudin-15 were not detected in COS-7 cells transfected with the empty vector.
Differential Expression of Claudin-7, −8, −12, −13, and −15 Proteins in the Mouse Intestine
We subsequently checked, by Western blot analysis, expression levels of claudin-7, −8, −12, −13, and −15 proteins in the mouse intestinal tract (Figures 3A and 3B). Claudin-7 protein was observed in the duodenum, jejunum, ileum, and colon, with no obvious differences in the expression levels. Claudin-8 protein was expressed in the colon at higher levels than that in the ileum, but hardly detected in the jejunum and duodenum. Claudin-13 protein was obviously detected in the colon, but not expressed in the duodenum, ileum, or jejunum. On the other hand, claudin-12 protein was strongly observed in the ileum, but less and weakly expressed in the jejunum and colon, respectively. In addition, claudin-12 protein was barely detected in the duodenum. Expression of claudin-15 protein was observed greatly in the duodenum and jejunum, and decreased through the ileum and colon. Hence, these five claudins showed unique profiles of expression along the mouse intestine, in good agreement with the data obtained from RT-PCR analysis.
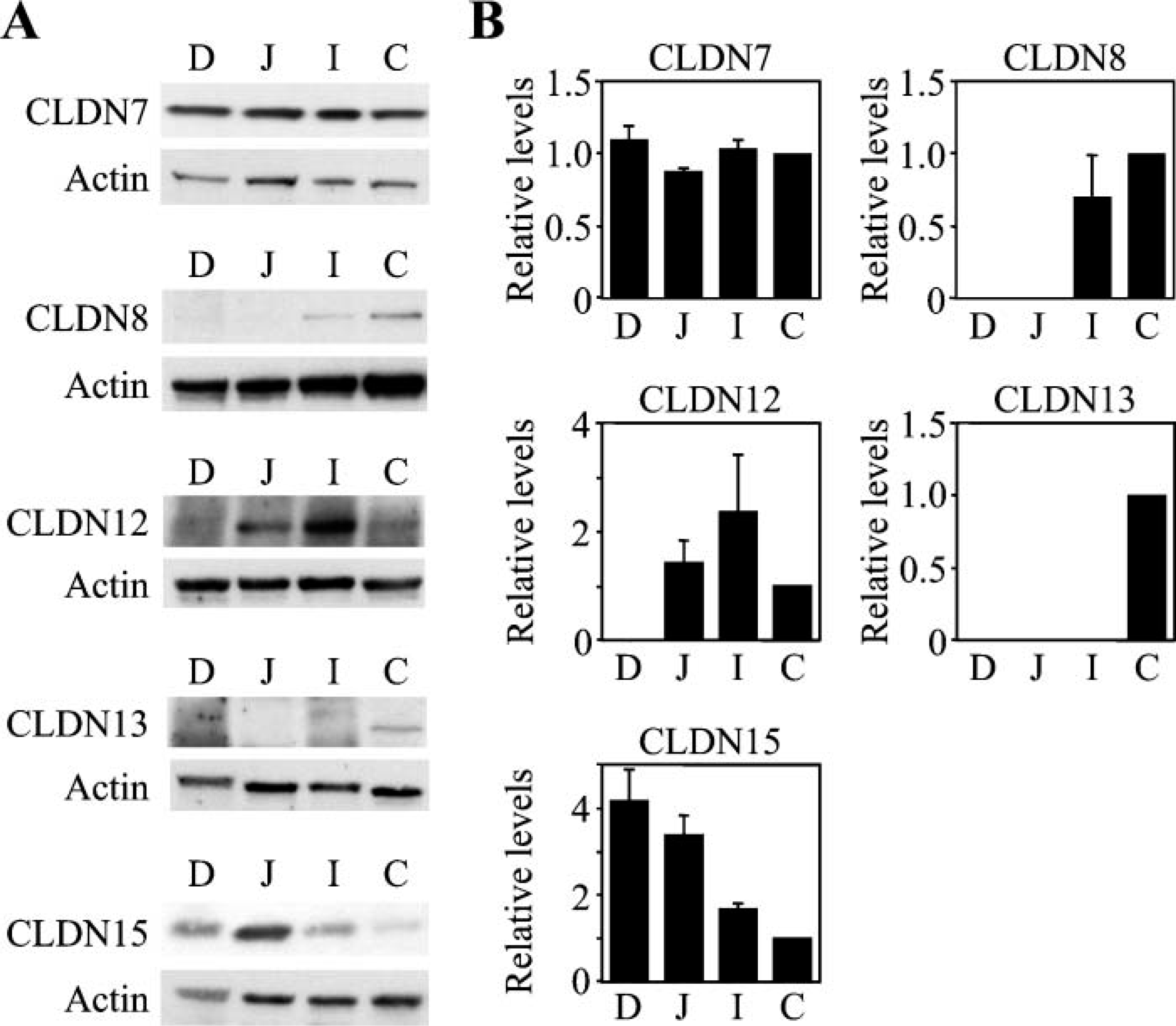
Claudin-7, −8, −12, −13, and −15 proteins are distinctively expressed in the mouse intestine. (
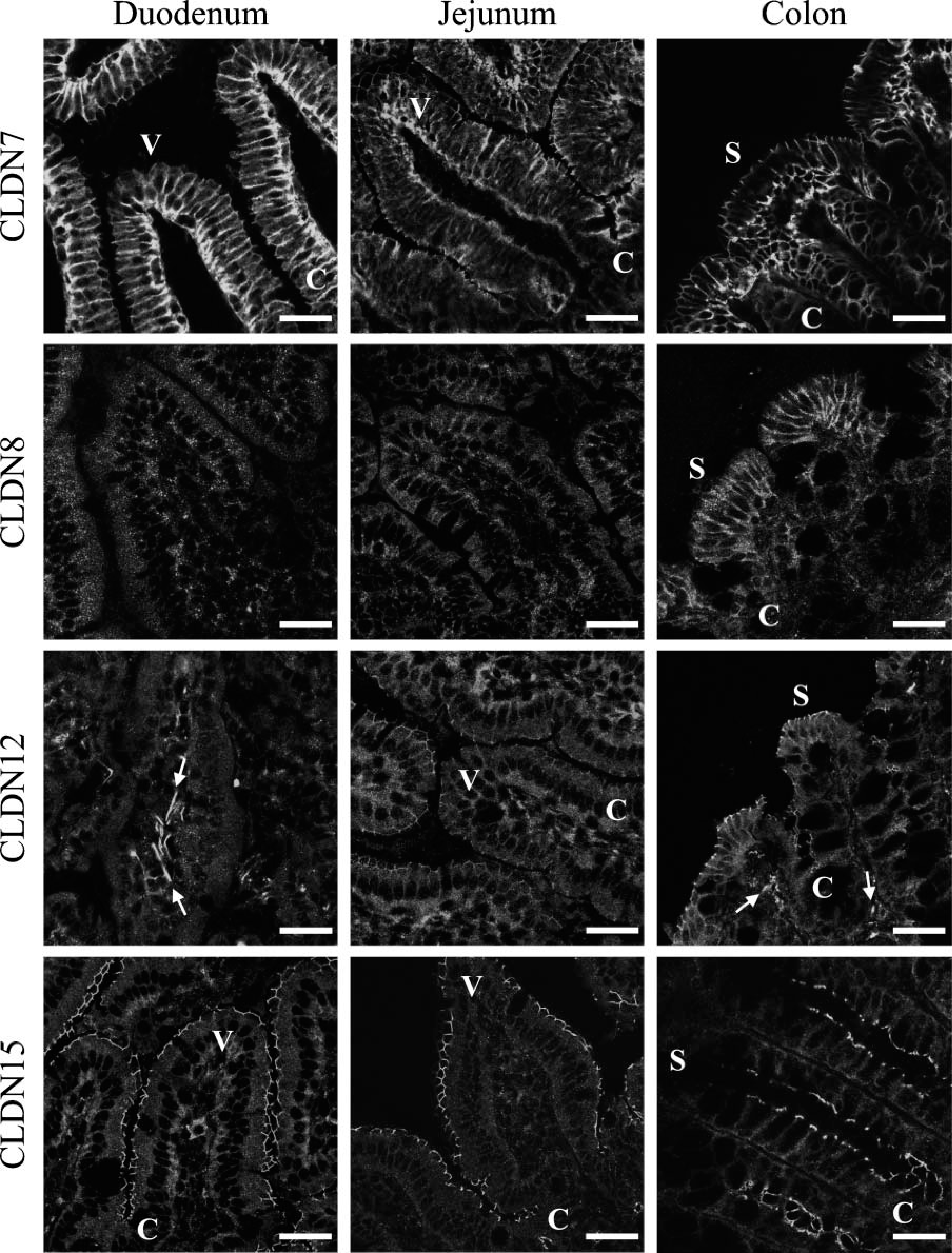
Claudin-7, −8, −12, and −15 are uniquely distributed in the duodenum, jejunum, andcolon. The duodenum, jejunum, and colon were subjected to immunostaining with the corresponding antibodies. Arrows indicate signals in vascular endothelial cells. C, crypt; S, surface epithelium; V, villus. Bar = 30 μm.
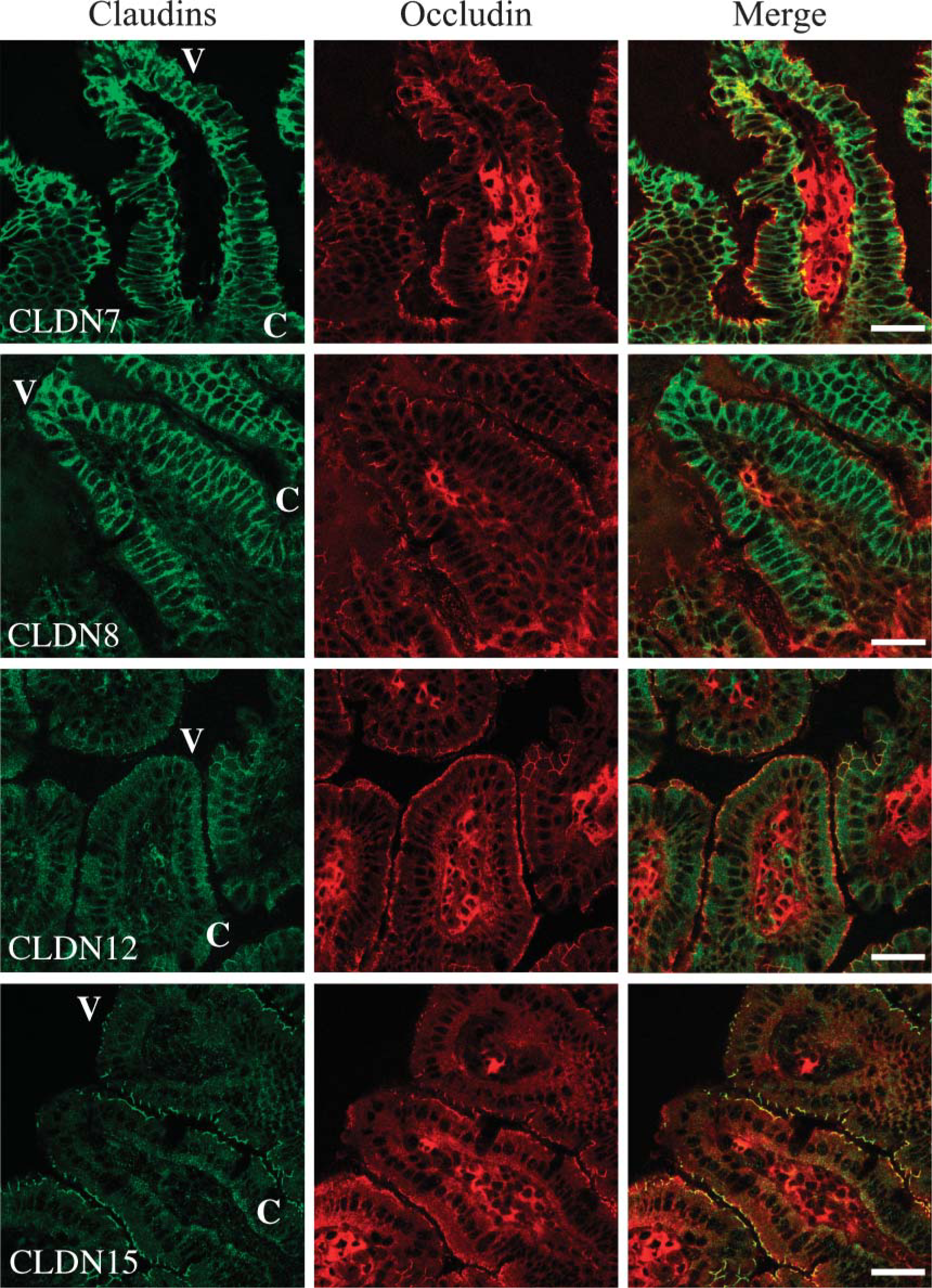
Claudin-7, −8, −12, and −15 are differentially localized in the ileum. The ileum was subjected to double immunostaining with the corresponding antibodies. C, crypt; V, villus. Bar = 30 μm.
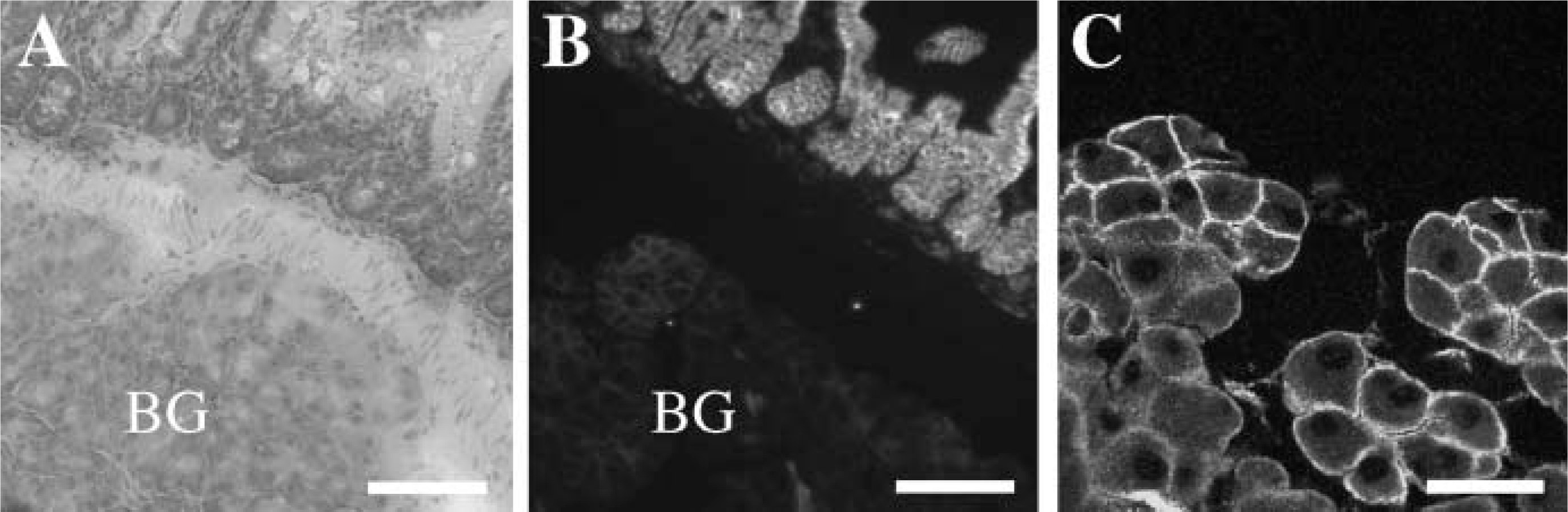
Claudin-7 is expressed not only in the crypt epithelium of the duodenum but also in epithelial cells of Brunner's glands.
Distribution of Claudin-7, −8, −12, −13, and −15 in the Mouse Intestinal Tract
We next determined, by immunofluorescent analysis, the expression and subcellular localization of claudin-7, −8, −12, −13, and −15 throughout segments of mouse intestinal tract. Claudin-7 was observed not only at the same position as occludin, but also along basolateral membranes of epithelial cells in the mouse duodenum, jejunum, ileum, and colon (Figures 4 and 5, and data not shown). In the duodenum, claudin-7 was also expressed along cell surfaces of epithelial cells in Brunner's glands, although the signal intensity was much weaker than that in the villous epithelium (Figure 6). Claudin-8 was distributed on basolateral membranes of epithelial cells in the ileum and colon, but not primarily coexpressed with occludin (Figures 4 and 5, and data not shown). Expression of claudin-8 protein was not detected in the duodenum or jejunum (Figure 4). In contrast, claudin-12 was exclusively colocalized with occludin at the apicalmost tips of lateral membranes of epithelial cells in the jejunum, ileum, and colon, but not in the duodenum (Figures 4 and 5, and data not shown). Expression of claudin-12 was also occasionally detected along cell boundaries of vascular endothelial cells of submucosal layers throughout the intestinal tract (Figure 4, and data not shown). Claudin-15 was localized at the apicalmost sites of lateral membranes of epithelial cells in the duodenum, jejunum, ileum, and colon (Figures 4 and 5). In the colon, however, claudin-15 expression appeared to be restricted to the crypts (Figures 4 and 7). Claudin-13 was not expressed in the duodenum, jejunum, or ileum, but was expressed in the colon (Figure 8, and data not shown). It was distributed at the apicalmost edges of lateral membranes of colonic epithelial cells, and the signal in the surface epithelium was strong compared with that in the crypt epithelium. Claudin-13 also seemed to be localized in basal membranes. Preimmune serum did not show positive signals (Supplemental Figure 1). Thus, claudin-7, −8, −12, −13, and −15 were differently expressed and distributed in the mouse intestinal tract.
Discussion
The aim of the present study was to determine which claudin species, other than claudins 1 to 5, were expressed in the mouse intestine, and whether these claudins exhibited unique expression profiles in terms of their expression levels and subcellular distribution along the intestinal tract. We have herein shown that claudin-7, −8, −12, −13, and −15, but not claudin-6, −9, −10, −11, −14, −16, −18, or −19, were expressed in the mouse intestine with distinct patterns of expression. Our results are summarized in Figure 9, and led to several important conclusions.
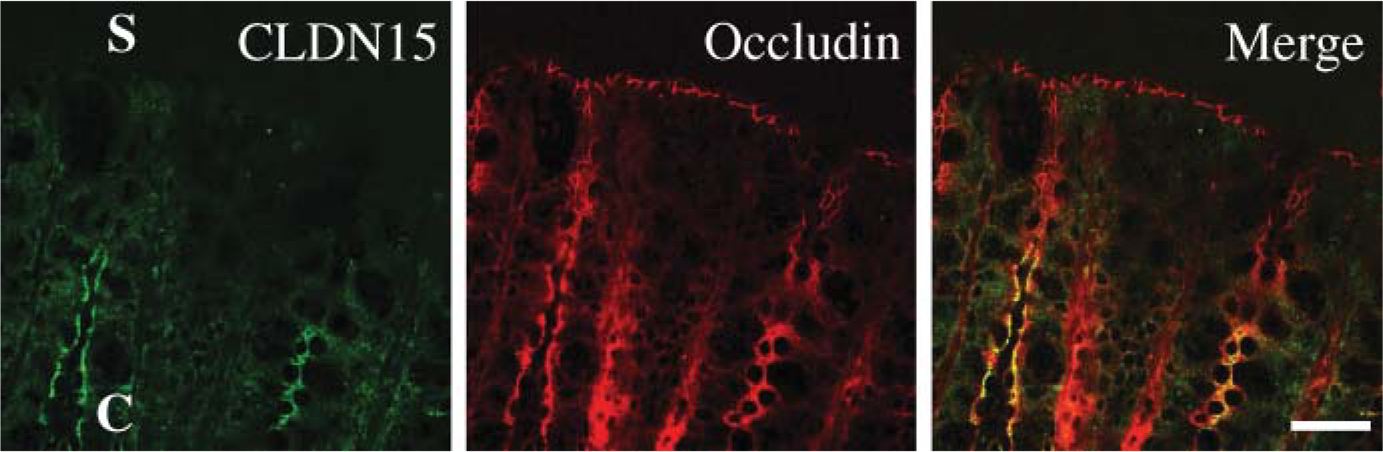
Claudin-15 is observed in the crypt epithelium of the colon, but not in the surface epithelium. The colon was subjected to double immunostaining with anti-claudin-15 and anti-occludin antibodies. C, crypt; S, surface epithelia. Bar = 30 μm.
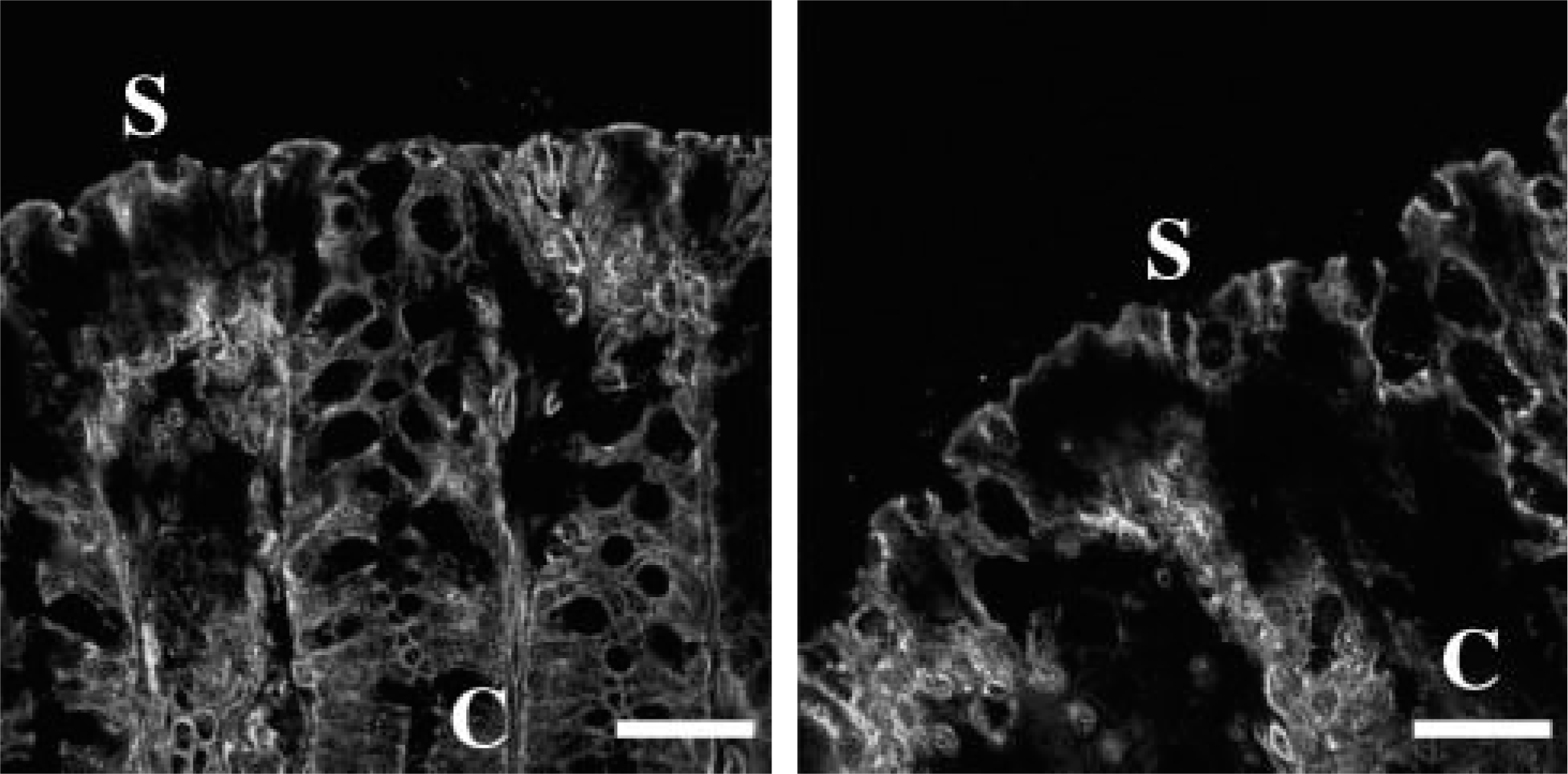
Claudin-13 is expressed in the colon. The colon was subjected to immunostaining with the anti-claudin-13 pAb. C, crypt; S, surface epithelia. Bar = 30 μm.
First, our RT-PCR, immunoblot and immunohistochemical analyses clearly indicated that there are considerable variations in the abundance of claudin-7, −8, −12, −13, and −15 throughout the segments of the mouse intestinal tract (Figure 9A). Claudin-7 was constantly expressed in the duodenum, jejunum, ileum, and colon. Claudin-15 was greatly expressed in the duodenum and jejunum, but less and only faintly observed in the ileum and colon, respectively. Claudin-12 was strongly observed in the ileum and less and weakly expressed in the jejunum and colon, but barely detected in the duodenum. In marked contrast, claudin −8 and −13 were predominantly expressed in the colon. Claudin-8, but not claudin-13, was also observed in the ileum, and no obvious expression of these claudins was detected in the duodenum or jejunum. It is strongly suggested that each claudin exhibits different charge selectivity, depending on the position and number of charged amino acids in the first extracellular loop (for review, see Van Itallie and Anderson 2006). For example, claudin-15 has two negatively charged residues (positions 53 and 64) in the latter half of this loop, and substitution of those by positive ones converts claudin-15 from cation- to anion-permissive (Colegio et al. 2002; Van Itallie et al. 2003). Similarly, claudin-7 likely acts as a paracellular pore to cations (Alexandre et al. 2005), and it contains three negatively charged amino acids within the first extracellular loop. Because claudin-12 has four negatively charged residues (positions 62, 66, 71, and 74) in the same region as claudin-15, it may also form a cation-selective pore. By contrast, claudin-8 functions primarily as a cation barrier (Yu et al. 2003), and claudin-13 might create an anion-selective pore, because it contains four positively charged amino acids in the second half of the first extracellular loop. In this respect, it is worth noting that claudin-7, −12, and −15, but not claudin-8 or −13, are expressed in the jejunum, because the rat jejunum is known to possess a cation-selective property (permeability ratio: PNa+/PCl = 10) (for review, see Powell 1981). The relative abundance of these claudins (Figure 9A) might also contribute to the charge selectivities in the ileum and colon (PNa+/PCl- = 2 and 0.63, respectively) (Powell 1981).
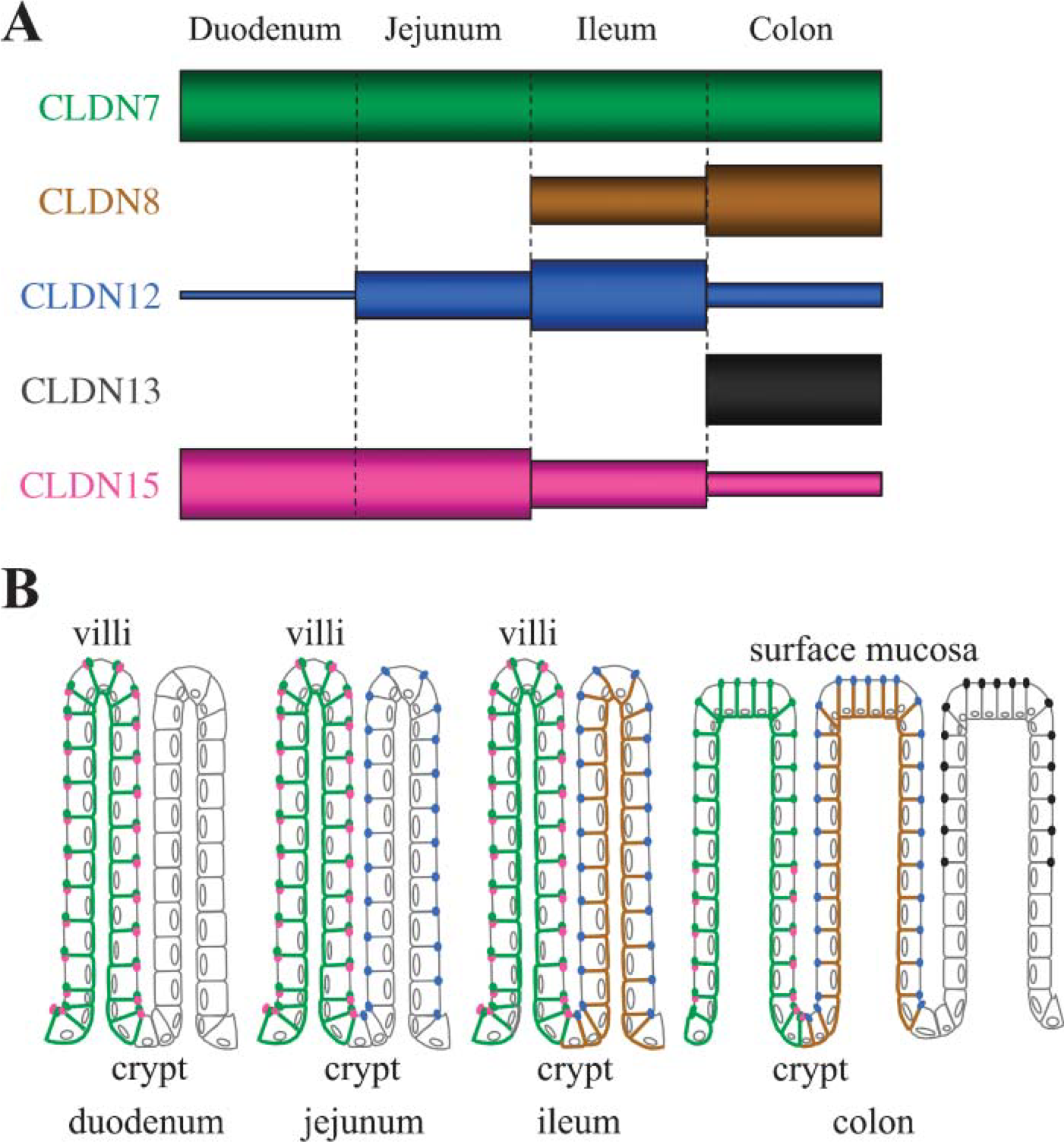
Five species of claudins, claudin-7, −8, −12, −13, and −15, show unique profiles of abundance and subcellular localization along the mouse intestine.
The second conclusion is that claudin-7, −8, −12, −13, and −15 show different subcellular localization in epithelial cells along the mouse intestine (Figure 9B). This conclusion was drawn from results using novel antibodies that specifically recognized the corresponding claudins (Figure 2, and see also Satohisa et al. 2005), and commercially available antibodies. Claudin-12 and −15 were principally colocalized with occludin at the apicalmost poles of lateral membranes of intestinal epithelial cells, although claudin-12 was not found in the epithelium of the duodenum, as described above. In the colon, claudin-13 was also localized preferentially at the apicalmost edges of lateral membranes. On the other hand, claudin-7 was expressed more strongly on basolateral surfaces of the mouse intestinal epithelium rather than on the apical tips, in good agreement with recent reports on its distribution in the human duodenum and jejunum (Ladwein et al. 2005). Similar or identical patterns of expression were observed for claudin-7 in simple epithelial cells of the lung and kidney (Coyne et al. 2003; Li et al. 2004), as well as for claudin-4 in the intestinal epithelium (Rahner et al. 2001; Tamagawa et al. 2003). In addition, we demonstrated that claudin-8 was diffusely distributed on basolateral membranes of epithelial cells in the ileum and colon, with little or no junctional staining. Basolateral membrane localization was also reported for several other claudins, such as claudin-1 in the epididymal epithelium (Gregory et al. 2001) and claudin-3 and −5 in epithelial cells of the stomach and intestine (Rahner et al. 2001). Although the role of basolateral claudins remains obscure, extrajunctional claudins might represent a storage pool for junctional claudins and alternatively contribute to cell-cell and cell-matrix adhesion or signaling (Li et al. 2004; Van Itallie and Anderson 2004). Interestingly, the cell-cell adhesion molecule EpCAM has been found to directly associate with claudin-7 on basolateral membranes (Ladwein et al. 2005), implying a novel function of basolateral claudins.
Another conclusion in the present work is that claudin-13 and −15 are expressed in gradients along the crypt-to-surface axis (Figure 9B). In the mouse colon, claudin-15 was restricted to the crypts, whereas claudin-13 expression was observed most obviously in the surface epithelial cells. In contrast, claudin-8 and −12 exhibited no evident gradient along the crypt-to-villus axis, as far as we could determine. In this sense, it is interesting to note that claudin-2 is only detected in the crypts of the rat and mouse intestine, and that claudin-4 is highly expressed in the villi (Rahner et al. 2001; Tamagawa et al. 2003). It should also be mentioned that in fetal humans, claudin-2 is detected in both crypts and villi of the jejunum but is strongly expressed in crypts of the colon (Escaffit et al. 2005). Concerning gradients within tissues, we also found that claudin-7 was expressed on cell membranes in Brunner's glands of the duodenum much less than in the crypt epithelium.
In summary, we have reported that claudin-7, −8, −12, −13, and −15 are expressed in the mouse intestinal tract with substantial variations in the expression levels. Our observations have also demonstrated that these five species of claudins show distinct subcellular distributions, namely junctional and/or basolateral localization, in the intestinal epithelium. Furthermore, we have revealed that claudin-13 and −15 exhibit gradients along the crypt-surface axis of the colon, and that claudin-7 is very weakly expressed in Brunner's glands of the duodenum compared with the villous epithelium. Taken together with the expression profile of claudins 1 to 5 in the intestine (Rahner et al. 2001; Bürgel et al. 2002; Tamagawa et al. 2003), the heterogeneity in expression of claudin-7, −8, −12, −13, and −15 might determine differences in paracellular permeable properties such as ion selectivity in each segment of the intestinal tract. Our findings are of great interest for studying the contributions of claudins to gastrointestinal diseases, including IBD and colorectal cancers.
Footnotes
Acknowledgements
This work was supported by grants-in-aid from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
The authors thank Immuno-Biological Laboratories for cooperation in the generation of anti-claudin-7, anti-claudin-8, anti-claudin-12, and anti-claudin-15 antibodies; Mr. K. Barrymore for helping with the manuscript; and Mrs. K. Takeda for her technical help.
