Abstract
CD163 is a member of the scavenger receptor cysteine-rich superfamily restricted to the monocyte/macrophage lineage and is thought to be a useful marker for anti-inflammatory or alternatively activated macrophages. In this study we used mass spectrometric analysis to determine that the antigen recognized by the antibody AM-3K, which we previously generated as a tissue macrophage-specific monoclonal antibody, was CD163. An anti-inflammatory subtype of macrophages stimulated by dexamethasone or interleukin-10 showed strong reactivity for AM-3K and increased expression of CD163 mRNA. Immunohistochemical staining of routinely processed pathological specimens revealed that AM-3K recognized a specialized subpopulation of macrophages. In granulomatous diseases such as tuberculosis, sarcoidosis, or foreign body reactions, tissue macrophages around granulomas, but not component cells of the granulomas such as epithelioid cells and multinucleated giant cells, showed positive staining for AM-3K. In atherosclerotic lesions, scattered macrophages in diffuse intimal lesions were strongly positive for AM-3K, whereas foamy macrophages in atheromatous plaques demonstrated only weak staining. We therefore suggest that, in routine pathological specimens, AM-3K is a useful marker for anti-inflammatory macrophages because these cells can be distinguished from inflammatory or classically activated macrophages. Because AM-3K cross-reacts with macrophage subpopulations in different animal species including rats, guinea pigs, rabbits, cats, dogs, goats, pigs, bovine species, horses, monkeys, and cetaceans, it will have wide application for detection of CD163 in various animals.
Keywords
CD163 is a monocyte/macrophage-restricted membrane protein belonging to the scavenger receptor cysteine-rich (SRCR) domain family (Backé et al. 1991; Pulford et al. 1992; Law et al. 1993; Högger et al. 1998a; Buechler et al. 2000; Lau et al. 2004). Its function has remained unknown until recently, when CD163 was identified as a scavenger receptor for haptoglobin-hemoglobin complexes (hemoglobin scavenger receptor) (Kristiansen et al. 2001). Because CD163 expression is induced by interleukin (IL)-6, IL-10, and glucocorticoid (Sulahian et al. 2000; Schaer et al. 2001) but is down-regulated by lipopolysaccharide and interferon-γ (IFN-γ), it is thought to be one of the molecules associated with an anti-inflammatory macrophage phenotype (Buechler et al. 2000; Sulahian et al. 2000; Hintz et al. 2002; Yoon et al. 2003).
We previously generated AM-3K as a monoclonal antibody specific for tissue macrophages by using human alveolar macrophages as an immunogen (Zeng et al. 1996a). In immunohistochemical studies, AM-3K reacted with tissue macrophages including Kupffer cells, alveolar macrophages of the lung, microglial cells, and sinus macrophages in lymph nodes. By means of immunoelectron microscopy, AM-3K antigen was found on the macrophage cell surface membrane (Zeng et al. 1996a). AM-3K was reactive not only with human macrophages but also with macrophages from various animal species including rats, guinea pigs, rabbits, cats, dogs, goats, pigs, bovine species, horses, monkeys, and cetaceans (Zeng et al. 1996b; Yamate et al. 2000; Ide et al. 2001; Kawashima et al. 2004). Although the immunohistochemical and ultrastructural data for AM-3K antigen (Zeng et al. 1996a) indicate similarities to CD163, the molecular nature of AM-3K antigen was not clear.
In the present study, we determined AM-3K antigen to be CD163 by means of mass spectrometric analysis. To understand the relationship between CD163 expression and macrophage differentiation, we investigated AM-3K reactivity of various differentiated and activated monocyte-derived macrophages, either classically or alternatively activated, in vitro. We also ascertained the distribution of the antigen recognized by AM-3K (CD163) in various pathological conditions by using routinely processed paraffin-embedded tissue samples.
Materials and Methods
Affinity Chromatography
Because many AM-3K-positive macrophages were found in the spleen, the antigen recognized by AM-3K was purified from spleen extract by means of affinity chromatography. Human spleen obtained at autopsy was homogenized in Tris-buffered saline (TBS). After centrifugation of samples at 40,000 rpm for 1 hr, separated cell nuclei and cell membrane pellets were homogenized in TBS containing 10 mM CHAPS (3-cholamidopropyl-1-propane sulfonate; Dojin, Kumamoto, Japan). Samples were centrifuged at 15,000 rpm for 30 min, after which supernatants were incubated with AM-3K or isotype-matched non-relevant control antibody, RM-4 (an anti-rat macrophage, IgG1, κ) (Iyonaga et al. 1997), coupled to cyanogen bromide-activated Sepharose (Amersham Bioscience; Piscataway, NJ). Beads were washed three times in TBS containing 20 mM CHAPS, and bound proteins were eluted with 50 mM glycine-HCl (pH 2.5) by using polystyrene columns (Pierce; Rockford, IL). Eluate was immediately neutralized by addition of 1 M Tris (pH 8.5).
Sodium Dodecyl Sulfate-Polyacrylamide Gel Electrophoresis (SDS-PAGE) and Mass Spectrometry
Proteins were separated by SDS-PAGE under reducing or non-reducing conditions by use of 2-mercaptoethanol (Sigma; St Louis, MO). Deglycosylation was performed with N-glycosidase (Roche Diagnostics; Mannheim, Germany) as described previously (Högger et al. 1998b). Protein bands were stained with either Coomassie blue (Invitrogen; Carlsbad, CA) or silver staining kit (Amersham), according to the manufacturer's protocol. Protein resolved by SDS-PAGE with deglycosylation and visualized by Coomassie blue was excised from the polyacrylamide gel and digested in gel with trypsin. The sample was desalted by Zip Tips C18 (Millipore; Bedford, MA) and was analyzed via electrospray ion trap mass spectrometry (Amersham). A higher energy tandem mass spectrometry scan was obtained after the initial scan for verification of peptide identification. The amino acid sequences obtained from the analysis were subjected to a search for sequence similarity against the Mascot search engine.
Immunoprecipitation and Western Blotting
To bind antibodies to Protein G Sepharose beads (Amersham), 10 μg each of AM-3K, RM3/1 (anti-CD163; BMA Biomedicals, Augst, Switzerland) or RM-4 (control antibody) was incubated with protein beads for 1 hr. The beads binding antibodies were coupled with 20 mM dimethyl pimelimidate (Pierce) for 30 min at room temperature (RT). Then the reactions were blocked with 0.2 M glycine buffer (pH 8.0) for 2 hr at RT. Spleen protein was prepared as described above and incubated for 2 hr with AM-3K-, RM3/1-, or RM- 4-coupled beads. The beads were washed three times in TBS containing 10 mM CHAPS or 1% NP-40, and samples including spleen protein as positive control were processed on 5-20% gradient gel and subjected to Western blotting under non-reducing conditions. The primary antibodies (AM-3K and RM3/1) were used at a final concentration of 0.4 μg/ml. Blots were developed with an enhanced chemiluminescence substrate (Amersham).
Cell Culture
Peripheral blood cells were obtained from the Kumamoto Red Cross Blood Center (Kumamoto, Japan) and centrifuged via Lymphoprep (Geiner; Oslo, Norway) to obtain peripheral blood mononuclear cells, which were resuspended in Dulbecco's Modified Eagle's Medium (Sigma) supplemented by 10% fetal bovine serum (FBS) and were then plated in 96-well microtiter plates for 2 hr at 37C in 5% CO2 and 95% air. Non-adherent cells were removed by gentle washing with PBS; remaining monocytes were cultured up to 5 days to induce differentiation and maturation into macrophages. Before assay, cultured cells were washed once with PBS and then cultured for 24 hr in RPMI 1640 plus 10% FBS supplemented with various combinations of 100 ng/ml lipopolysaccharide (LPS; Sigma), 200 nM dexamethasone (DXA; Banyu Pharmaceutical, Tokyo, Japan), 10 ng/ml IL-10, 10 ng/ml IL-4, and/or 25 ng/ml INF-γ (Peprotech; Rocky Hill, NJ).
Cell Enzyme-linked Immunosorbent Assay (ELISA)
CD163 expression on human macrophages was evaluated with the aid of a cell ELISA as previously described (Andreesen et al. 1988; Möst et al. 1997). Cells cultured in a 96-well plate at a density of 5 × 105 cells per well were fixed with 2% paraformaldehyde for 10 min on ice and then kept at −30C until use. Cells were incubated with 5% normal sheep serum for 20 min to block nonspecific reactions and were subsequently reacted with AM-3K or RM-4 hybridoma supernatant overnight on ice. After excess antibody was washed out with PBS, samples were incubated with horseradish peroxidase (HRP)-labeled sheep anti-mouse antibody (Amersham) for 60 min. Plates were washed and 3,3′,5,5′-teramethylbenzidine developing solution (KPL; Gaithersburg, MD) was added. After sufficient color development, the same volume of 0.5 M H2SO4 was added to stop the reaction. Plates were read via a microplate reader at 450 nm. The background signal (isotype-matched control antibody RM-4) was subtracted, and signals for each well were normalized with the optical density of non-stimulated control wells. Statistical comparisons between groups were made by using an unpaired Student's t-test.
Immunocytochemistry
Cultured macrophages on chamber slides (Nalge Nunc; Roskilde, Denmark) were fixed with 2% paraformaldehyde for 10 min on ice and kept at −30C until use. After inhibition of endogenous peroxidase activity as described previously (Isobe et al. 1977), cells were incubated with 5% normal sheep serum for 20 min and then reacted with AM-3K hybridoma supernatant overnight on ice. After excess antibody was washed out with PBS, samples were incubated with HRP-labeled sheep anti-mouse antibody (Amersham) for 60 min. The reaction was visualized by use of diaminobenzidine (DAB; Dojin).
Reverse Transcriptase-Polymerase Chain Reaction (RT-PCR)
RNA was extracted from cultured macrophages with RNeasy Mini Kit (Qiagen, Hilden, Germany) and made into cDNA with Omniscript RT Kit (Qiagen) according to the manufacturer's instructions. PCR was performed with HotStarTaq PCR Kit (Qiagen). Cycling conditions consisted of a single denaturing step at 94C for 15 min followed by 33 cycles (CD163) or 28 cycles (β-actin) at 94C for 30 sec, 55C for 30 sec, and 72C for 30 sec, followed by a final extension step at 94C for 10 min. The primers were as follows: CD163 primers: 5′-CAAGAAGATTTAGGAATCATTG-3′ (forward) and 5′-CAAGAAGATTTAGGAATCATTG −3′ (reverse); and β-actin primers: 5′-ATCACCATGGCAATGAGCG −3′ (forward) and 5′-TTGAAGGTAGTTTCGTGGAT −3′ (reverse).
Immunohistochemical Evaluation of Paraffin-embedded Tissues
Various tissues were obtained from the files of the Department of Cell Pathology, Kumamoto University, Kumamoto, Japan. The diagnoses of these samples were established on the basis of morphological and clinical features. Tissue samples were fixed in 10% neutral-buffered formalin and embedded in paraffin. After sections were deparaffinized in xylene and rehydrated in a graded ethanol series, antigens were retrieved by incubation with proteinase K (DAKO; Glostrup, Denmark) for 5 min. Endogenous peroxidase activity was inhibited as described above, after which sections were incubated with 5% normal goat serum for 20 min and then reacted with AM-3K (supernatant) or PG-M1 (anti-CD68, diluted 1:200; DAKO) at 4C for 6 hr. For the staining with an anti-CD163, clone10D6 (Novocastra; New Castle, UK), deparaffinized sections were pretreated with microwave oven for 15 min and then processed the same as for AM-3K. After excess antibody was washed out with PBS, samples were incubated with HRP-labeled goat anti-mouse antibody (Nichirei; Tokyo, Japan) for 60 min. The reaction was visualized by use of the DAB substrate system (DAKO). Slides were counterstained with hematoxylin.
Results
Characterization of the Antigen Recognized by AM-3K Using SDS-PAGE analysis, the molecular mass of this antigen was 130 kDa under non-reducing conditions and 150 kDa under reducing conditions (Figure 1A, Lanes 1 and 2). When AM-3K antigen was deglycosylated by incubation with N-glycosidase, a band shift of 20-30 kDa was detected (Figure 1A, Lane 3), which indicated that the antigen is a glycosylated protein. After preparative SDS-PAGE, a gel slice containing the antigen was excised and prepared for mass spectrometry. Trypsin digestion produced two peptide sequences (LEVFYNGAWGTVGK, EAEFGOGTGPIWLNEVK) that coincided with those of CD163 protein (No. 739-752 and No. 981-997, respectively).
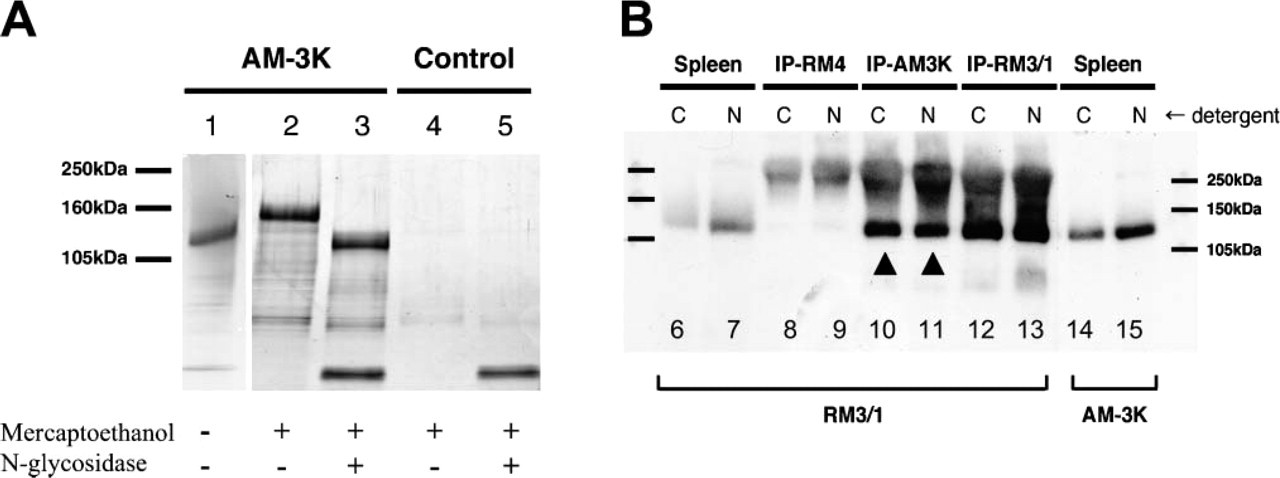
Characterization of antigen recognized by AM-3K. (
To ensure that AM-3K recognized CD163 proteins, Western blotting was performed with proteins immunoprecipitated by AM-3K, RM3/1 (anti-CD163), and RM-4 (non-relevant control). RM3/1 recognized proteins precipitated by AM-3K and RM3/1 but not that precipitated by RM-4 (Figure 1B). With untreated spleen protein, both RM3/1 and AM-3K recognized 130-kDa protein (Figure 1B). These observations indicate that AM-3K recognized the same antigen (CD163) that RM3/1 recognized.
AM-3K Preferentially Recognized an Anti-inflammatory Macrophage Phenotype
To specify the reactivity of AM-3K with various activated macrophage subpopulations, human monocyte-derived macrophages were treated with IFN-γ, IFN-γ + LPS, IL-10, IL-4, or DXA. In the cell ELISA (Figure 2A) and immunohistochemical staining assays (Figure 2B), AM-3K strongly reacted with macrophage populations stimulated with IL-10 or DXA, populations thought to have an anti-inflammatory phenotype. In contrast, macrophages stimulated with IL-4 or classically activated macrophages stimulated with IFN-γ showed very weak reactivity with AM-3K, and stimulation with IFN-γ + LPS further reduced AM-3K expression (Figures 2A and 2B). These reaction profiles were consistent with mRNA expression of CD163 (Figure 2C).
Immunohistochemical Reactivity of AM-3K with Various Types of Macrophages in Paraffin Sections
Table 1 summarizes the reactivities of AM-3K and PG-M1 (anti-CD68) in various macrophage-related diseases. In granulomatous disorders such as tuberculosis and sarcoidosis, epithelioid cells (ECs) and multinucleated giant cells (MGCs) were strongly positive for CD68 (Figures 3A and 3D), one of the most commonly used macrophage markers. However, these cells were negative or only weakly positive for AM-3K, although macrophages surrounding granulomas were strongly positive for this antibody (Figures 3B and 3E). These reaction patterns of AM-3K were consistent with those of 10D6 (Figures 3C and 3F), a recently reported anti-CD163 antibody applicable to paraffin sections.
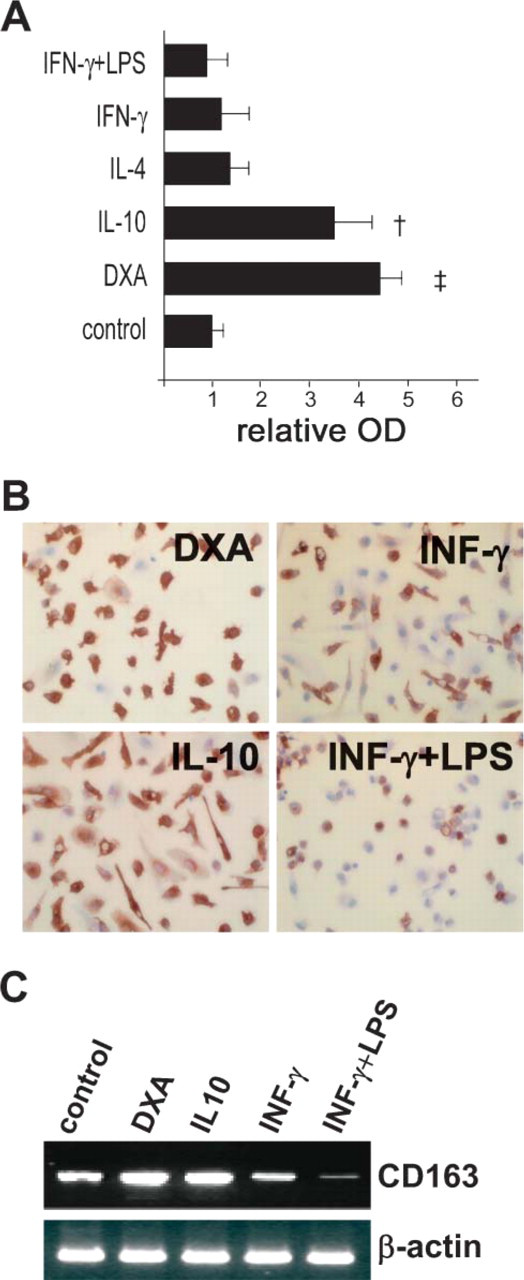
Macrophages stimulated with anti-inflammatory molecules interleukin-10 (IL-10) and dexamethasone (DXA) showed strong reactivity against AM-3K and increased expression of CD163 mRNA. Monocyte-derived macrophages cultured with DXA and IL-10 had higher reactivity for AM-3K in the cell ELISA (
Immunohistochemical reactivities of AM-3K (CD163) and PG-M1 (CD68) in macrophage-related disorders
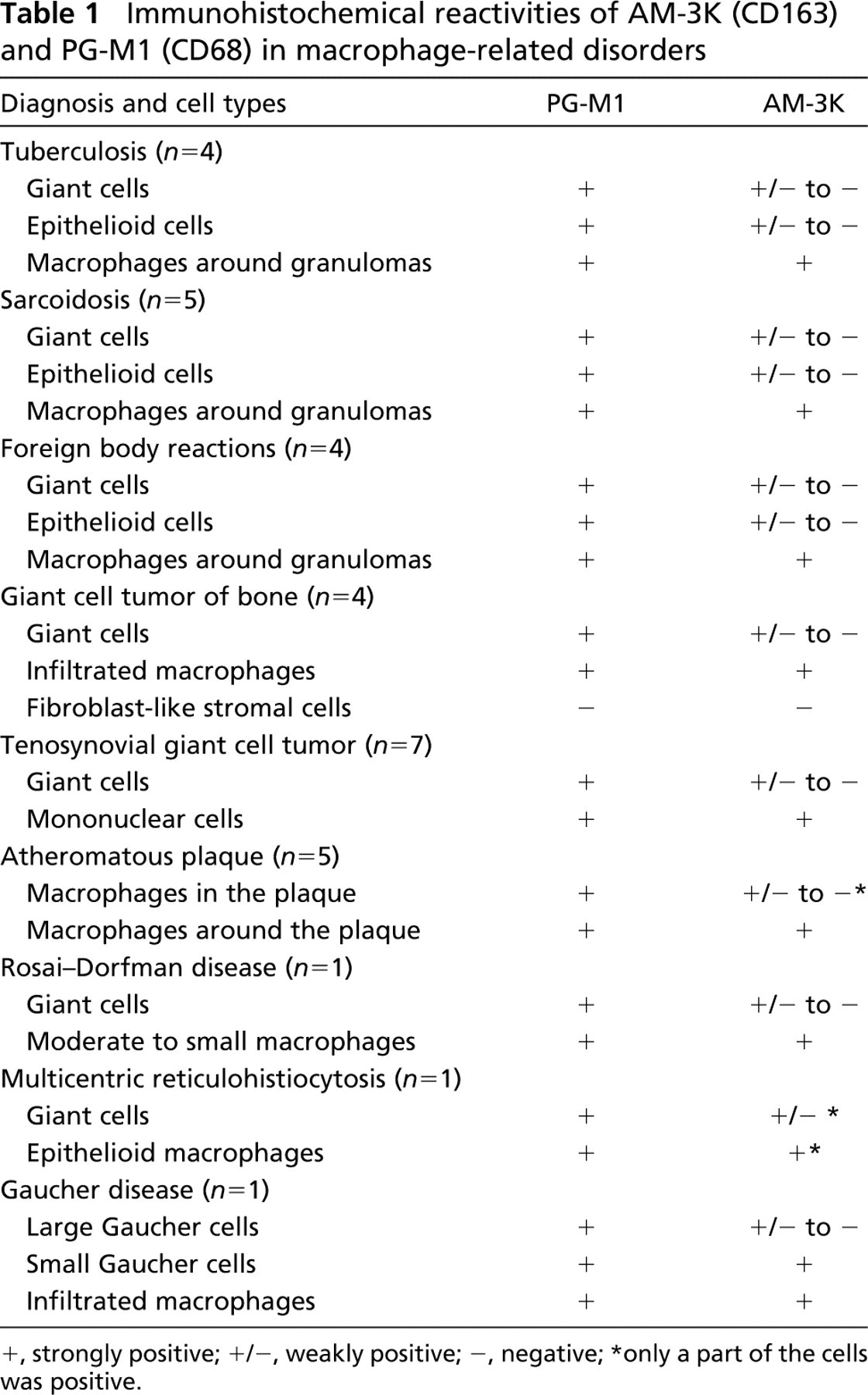
, strongly positive;
, weakly positive;
, negative;
only a part of the cells was positive.
In giant cell tumor (GCT) of bone and in tenosynovial GCT, osteoclast-like MGCs and infiltrated macrophages were positive for CD68 (Figure 4A), whereas AM-3K selectively reacted with infiltrated macrophages but not with osteoclast-like MGCs or fibroblast-like stromal cells (Figure 4B). Most lipid-laden foamy macrophages in atherosclerotic plaques were stained strongly positive for CD68 (Figure 4C). By contrast, macrophages in the plaque surface or those with less foamy transformation surrounding atheromatous core were positive for AM-3K (Figure 4D). In a case of Rosai-Dorfman disease, small and medium macrophages were stained positive for both CD68 and AM-3K. Large macrophages and MGCs were intensely positive for CD68 (Figure 4E) but only weakly positive for AM-3K (Figure 4F). Tingible body macrophages in lymphoid follicles were negative for AM-3K (data not shown). In multicentric reticulohistiocytosis, both epithelioid macrophages and MGCs were positive for CD68 (Figure 4G) and AM-3K (Figure 4H); however, the number of AM-3K-positive macrophages was smaller than that of CD68. In the staining of Gaucher disease, CD68 showed strong reactivity for most Gaucher cells. On the other hand, AM-3K selectively stained smaller Gaucher cells with lesser accumulation of glucocerebrosides (Figure 4J).
Discussion
Heterogeneity of macrophages has been discussed with regard to different responses to various microenvironmental stimuli. Macrophage activation in response to proliferating microbes and IFN-γ has been well characterized; this kind of activation has been called classical activation. Recent studies have described alternative activation: that induced by Th2-type cytokines such as IL-4 and IL-13. Such macrophage subpopulations show different types of receptor expression and cytokine and chemokine production. For instance, scavenger receptors A and B, mannose receptor, CCL18, CCL16, and arginase were specifically expressed by alternatively activated macrophages, whereas tumor necrosis factor-α (TNF-α), IL-1β, IL-6, IL-12, and inducible nitric oxide synthase (iNOS) showed restricted expression by these macrophages (Goerdt and Orfanos 1999; Gordon 2003; Mosser 2003; Mantovani et al. 2004; Taylor et al. 2005). Although expression of CD163 protein and mRNA may be related to alternative activation, it is downregulated by IL-4 and IL-13 (Sulahian et al. 2004). Because CD163 is upregulated by anti-inflammatory reactants such as IL-10 and DXA, expression of CD163 may regulate the anti-inflammatory nature of macrophages.
CD163 is a monocyte/macrophage-restricted membrane protein belonging to the SRCR domain family (Law et al. 1993). It functions as an endocytic receptor and binds hemoglobin in a complex with haptoglobin, which leads to removal of the hemoglobin-haptoglobin complex from plasma (Kristiansen et al. 2001). CD163 expression is induced by the anti-inflammatory mediators IL-6, IL-10, and DXA, whereas the proinflammatory cytokines IFN-γ and TNF-α suppress its expression (Buechler et al. 2000; Sulahian et al. 2000). CD163 is thought to be one of the most specific markers for macrophages (Radzun et al. 1987; Backé et al. 1991; Pulford et al. 1992; Van den Heuvel et al. 1999; Lau et al. 2004). It is found on resident tissue macrophages of almost all tissues except splenic white pulp macrophages and germinal center tingible macrophages (Radzun et al. 1987; Zeng et al. 1996a; Lau et al. 2004).
AM-3K is a monoclonal antibody raised against human alveolar macrophages. Although immunohistochemical data indicated that the reactivity of AM-3K is very similar to that of CD163, the molecular nature of the antigen recognized by AM-3K had not been determined (Zeng et al. 1996a). The present study clearly demonstrated by mass spectrometry that the immunoprecipitated precipitated antigen recognized by AM-3K is identical to CD163.
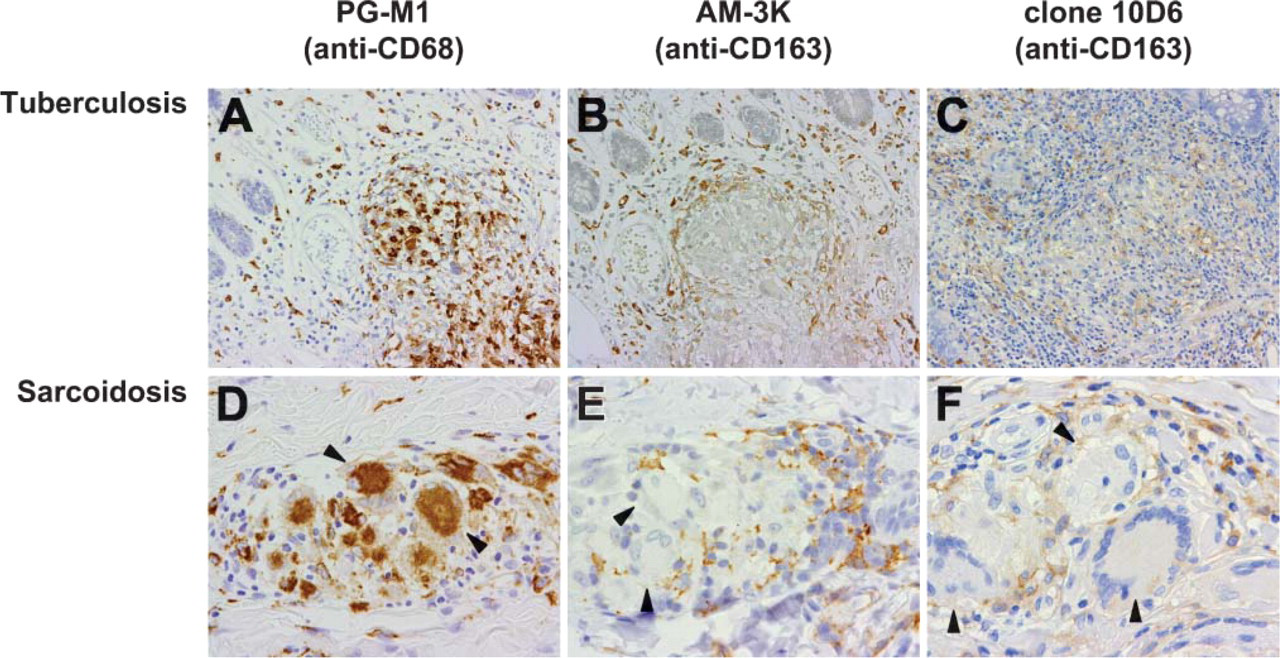
Immunohistochemical staining of typical cases of tuberculosis (
With routinely processed paraffin-embedded tissue sections, AM-3K labeled specific subpopulations of macrophages. For example, in granuloma-forming chronic inflammation, AM-3K labeled macrophages surrounding granulomas but failed to recognize ECs and MGCs (Figures 3B and 3E). Also, in GCTs of bone (Figure 4B) and in tenosynovial GCTs, AM-3K failed to label MGCs. Because CD163 is believed to be a molecule characterizing a subset of the anti-inflammatory phenotype, such a reactivity pattern may reflect different activation states of the MGCs and the surrounding mononuclear macrophages.
Recently, a new anti-CD163 antibody, clone 10D6, has been reported to be applicable to paraffin-embedded tissues (Lau et al. 2004; Nguyen et al. 2005). By using clone 10D6, Lau et al. (2004) reported that CD163 expression was restricted largely to disorders with a monocyte/macrophage derivation including chronic myelomonocytic leukemia, histiocytic sarcoma, and Rosai-Dorfman disease. With the same antibody, Nguyen et al. (2005) found staining for CD163 in Rosai-Dorfman disease, histiocytic sarcoma, littoral cell angioma, and Langerhans cell histiocytosis. They also demonstrated that acute myeloid leukemia with monocytic differentiation (FAB subtype M5), as well as a majority of tenosynovial GCTs, were positive for CD163. We have confirmed that the reactivity of AM-3K in granulomatous diseases is consistent with that of 10D6 (Figure 3).
ECs and MGCs are characteristic features of granulomas in tuberculosis and sarcoidosis. In tuberculosis, these cells produce large amounts of various cytokines such as IFN-γ and TNF-α (Myatt et al. 1994; Fenhalls et al. 2002) and iNOS, which play an important role in killing of bacteria (Hernandez-Pando et al. 2001). Granulomatous lesions in sarcoidosis have elevated levels of various cytokines and anti-inflammatory molecules such as IL-2, IL-12, TNF-α, IFN-γ, and iNOS (Myatt et al. 1994; Minshall et al. 1997; Moodley et al. 1999). These observations suggest that ECs and MGCs in granulomatous lesions in tuberculosis and sarcoidosis have properties identifying them as classically activated macrophages. Sandor et al. have described the heterogeneity of granuloma formation, i.e., Th1-type cytokines including IFN-γ induce classical activation of macrophages in type 1 granulomas (as in tuberculosis and sarcoidosis) and Th2-type cytokines such as IL-4 and IL-13 induce alternative activation of macrophages in type 2 granulomas (Sandor et al. 2003). Our present findings showed that ECs and MGCs in granulomas were negative for AM-3K and 10D6, whereas macrophages around granulomas were strongly positive for these antibodies. These results indicate that the classically activated macrophages (M1) that form the granulomas express a lesser amount of CD163, in contrast to macrophages around the granulomas. MGCs in GCTs of bone have been shown to secrete M1-type cytokines such as IL-1β, TNF-α, and IL-6, and IL-1β and TNF-α regulate induction of matrix metalloproteinase-9 from MGCs (Rao et al. 1999a,b; Gamberi et al. 2004). In this study we showed that MGCs in GCTs of bone were negative for AM-3K. We therefore suggest that MGCs in GCTs of bone could be characterized as M1 macrophages, whereas infiltrated macrophages are alternatively activated macrophages (M2) because of high expression of CD163.
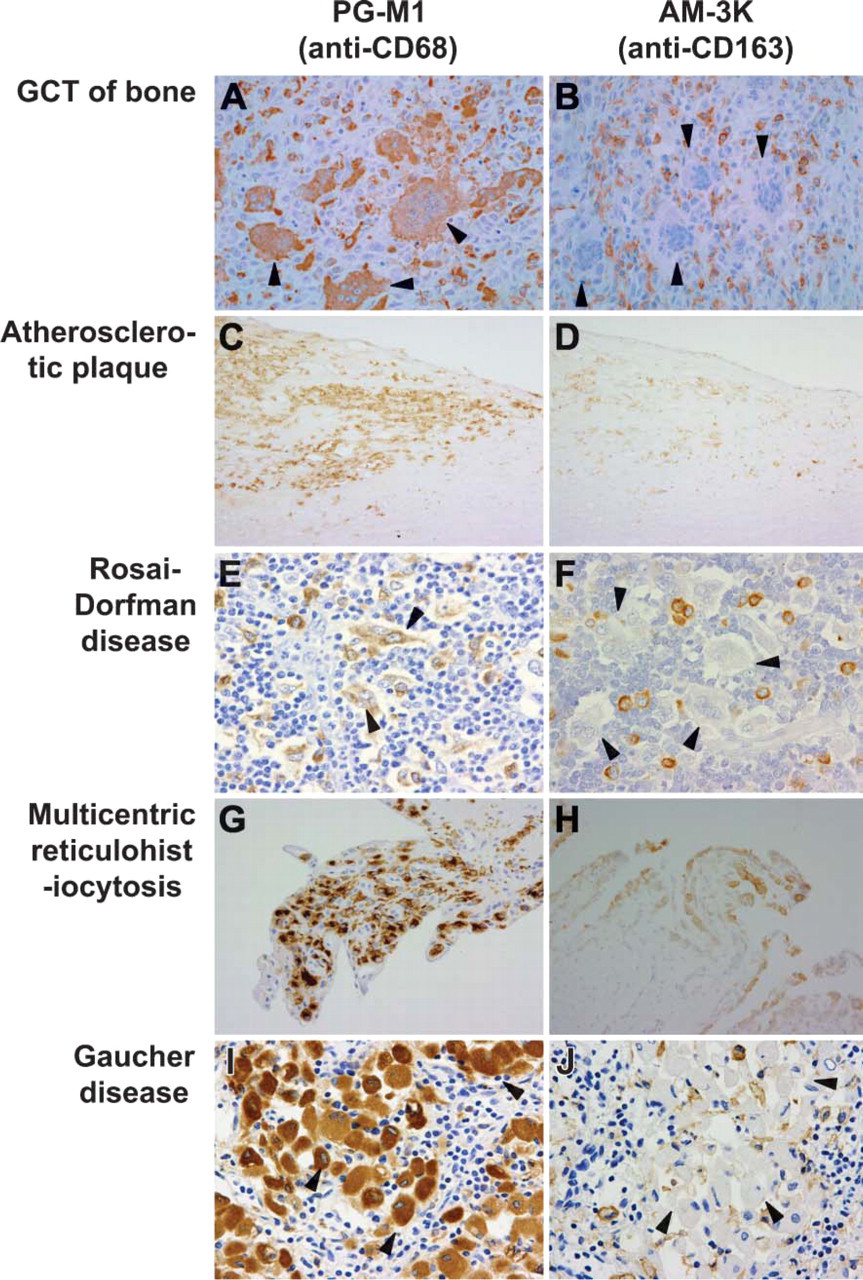
Immunophenotypic features of various macrophage-related disorders showing different reaction patterns for PG-M1 (A,C,E,G,I) and AM-3K (B,D,F,H,J). (
Atherosclerotic lesions in coronary arteries contain many infiltrated macrophages with or without accumulated lipid, and we observed that these cells produced little CD163 protein. Macrophages isolated from atheromatous plaque are known to secrete IL-1β and TNF-α (Tipping and Hancock 1993). In addition, oxysterols, which are biologically active molecules generated during oxidation of low-density lipoprotein and highly expressed by macrophages in plaque, induce macrophages to produce IL-1β in vitro (Rosklint et al. 2002). These observations suggest that macrophages infiltrating the plaque were M1 macrophages. Our observations in this study also demonstrated low reactivity of foamy macrophages for AM-3K, which indicated that foamy macrophages in plaque differentiated to become M1 macrophages.
Gaucher disease is a lysosomal storage disorder that is caused by deficiency of the lysosomal enzyme glucocerebrosidase. Massive amounts of glucocerebrosides accumulate in macrophages, and these glycolipid-laden macrophages are called Gaucher cells. Boven et al. (2004) observed the expression of CD163 on Gaucher cells using frozen sections of two cases of Gaucher disease. They found small and intermediate-sized Gaucher cells expressed CD163 but large Gaucher cells did not. Such observation corresponds well with our observation with AM-3K. Boven et al. (2004) indicated that CD163-positive Gaucher cells resemble alternatively activated macrophages because they did not express proinflammatory cytokines such as TNF-α and monocyte chemoattractant protein-1 but express several other markers for alternatively activated macrophages such as CC-chemokine ligand 18 and interleukin-1 receptor antagonist. In Rosai-Dorfman disease and multicentric reticulohistiocytosis, a restricted population of macrophages was positive for AM-3K comparing to CD68. Because we have observed only one case each for these rare histiocytic disorders, additional cases should be analyzed to delineate the expression and function of CD163 in these diseases.
Our present data, together with the previous reports (Lau et al. 2004; Nguyen et al. 2005), indicate that CD163 is a reliable macrophage marker antigen in immunohistochemical studies. Compared with CD68, one of the most commonly used macrophage markers, CD163 is considered to be a more specific marker for macrophages, especially a lineage with anti-inflammatory properties.
AM-3K possesses an additional advantage compared with the other CD163 antibodies. It is applicable not only to paraffin-embedded human tissues but also to those obtained from various animal species. In our previous studies, AM-3K was successfully used to label macrophage subpopulations in various animal species including rats, guinea pigs, rabbits, cats, dogs, goats, pigs, bovine species, horses, monkeys, and cetaceans (Zeng et al. 1996b; Yamate et al. 2000; Ide et al. 2001; Kawashima et al. 2004). About 30-50% of pulmonary and peritoneal macrophages obtained from cats and dogs were reactive for AM-3K (Yamate et al. 2000). In pathological specimens, AM-3K recognized infiltrated macrophage subpopulations in hepatic and renal fibrosis and in encephalomalacia of dogs (Yamate et al. 2000; Ide et al. 2001). It is interesting to note that many AM-3K-positive macrophages were present in the early stage of renal fibrosis (Ide et al. 2001). Because alternatively activated macrophages play an important role in tissue repair and subsequent fibrosis (Gordon 2003), appearance of AM-3K-positive macrophages in canine hepatic and renal fibrosis may be correlated with the disease process.
In summary, we clearly demonstrated that AM-3K, a previously reported macrophage-specific antibody, recognizes CD163. Because AM-3K is applicable to paraffin sections and is cross-reactive with macrophages in most experimental animals, it will be a useful tool for the detection of CD163-positive alternatively activated macrophage subtype in humans and many animal species.
Footnotes
Acknowledgements
We thank Dr. Hisato Saito (Institute of Molecular Embryology and Genetics) for helpful discussions during the course of this work and Mr. T. Nakagawa, Mr. O. Nakamura, and Ms. M. Komohara for technical assistance.
