Abstract
Evolved functions of integrin-αvβ6 include roles in epithelial cell-extracellular matrix protein interactions and in the binding and activation of latent TGF-β1. Integrin-αvβ6 is also exploited as a receptor by foot-and-mouth disease virus (FMDV) and may play a significant role in its transmission and pathogenesis. The ovine β6 integrin subunit was cloned and sequenced (EMBL accession no. AJ439062). Screening of normal ovine tissues by RT-PCR and immunocytochemistry confirmed that integrin-αvβ6 is restricted to sheep epithelial cells. Integrin-αvβ6 expression was detected in epithelia of the airways, oral cavity, gastrointestinal tract, kidney, sweat glands, hair follicle sheaths, and the epidermis of pedal coronary band (PB) but not of normal skin. Consistent with FMDV tropism, integrin-αvβ6 was detected within the basal layers of the stratified squamous epithelium of the oral mucosa and PB. In addition, integrin-αvβ6 appears to be constitutively expressed in the normal airways of both cattle and sheep. The latter finding suggests that ruminant airway epithelium presents a highly accessible target for initiation of infection with FMDV by inhalation.
Keywords
I
A number of viruses belonging to the Picornaviridae family have evolved to take advantage of RGD-binding integrins as receptors. The RGD motif is present on the foot-and-mouth disease virus (FMDV) capsid protein VP1 (Logan et al. 1993) and has been shown to mediate infection of susceptible cells in vitro (Mason et al. 1994). Four RGD-dependent integrins, αvβ1, αvβ3, αvβ6, and αvβ8, have been shown to function as receptors for FMDV in vitro (Berinstein et al. 1995; Jackson et al. 2000,2002,2004). Although the relative contribution of each of these integrins to the transmission and pathogenesis of foot-and-mouth disease is not known, the predominantly epithelial distribution of integrin-αvβ6 in primates (Breuss et al. 1993,1995) is consistent with the tropism of FMDV in ungulates (Grubman and Baxt 2004). Moreover, Monaghan et al. (2005) recently published evidence of integrin-αvβ6 expression by cells within the stratified squamous epithelium of the tongue, ventral soft palate, interdigital skin, and coronary band of cattle, where FMDV lesions are most commonly observed.
In the current study we describe the cloning and sequencing of ovine integrin-β6 and the subsequent screening of sheep tissues for integrin-β6 gene expression by RT-PCR. The presence and location of the integrin-αvβ6 heterodimer within each tissue was determined using immunofluorescence and confocal microscopy. Our findings confirm that integrin-αvβ6 expression is restricted to epithelial cells in the sheep, and that it is expressed at sites of known FMDV replication including the basal layers of the stratified squamous epithelium of the oral mucosa and pedal coronary band (PB). We also show for the first time that integrin-αvβ6 is expressed by both sheep and cattle airway epithelia in the apparent absence of inflammation. Importantly, constitutive expression of integrin-αvβ6 by respiratory epithelial cells, which are readily accessible to inhaled aerosols, suggests that the lung is a likely portal of entry for FMDV.
Materials and Methods
Sample Collection
All experimental procedures involving laboratory animals were approved by The University of Edinburgh's Biological Services Ethical Review Committee and were performed under license, as required by the UK's Animals (Scientific Procedures) Act 1986. Samples were obtained from clinically normal, outdoor-reared, adult crossbred ewes, two 15-day-old Holstein-Friesian bull calves, and a Holstein-Friesian heifer. Animals were euthanized by IV injection of sodium pentobarbital (Euthatal; Merial Animal Health Ltd., Harlow, UK). Tissues were removed at necropsy and inspected for any evidence of ongoing or recent pathological changes. Freshly isolated tissue samples (≤0.4 cm3) were collected into RNAlater (Ambion; Huntingdon, UK), maintained at 4C overnight, and transferred to −20C for long-term storage. Duplicate tissue samples (≤1 cm3) were also mounted on cork blocks using optimal cutting temperature (OCT) compound (BDH Laboratory Supplies; Dorset, UK) and snap frozen in dry-ice-cooled isopentane. In the case of lung samples, the right caudal diaphragmatic lobe was inflated via the major airway with OCT compound diluted 1:1 with pH 7.4 PBS prior to mounting and snap freezing as described above. Lung samples from the left caudal lobe were collected into RNAlater (Ambion) for RNA isolation. Frozen sections (8 μm) were prepared and stained with Harris' hematoxylin and eosin (H and E).
Cloning and Sequencing of the Ovine Integrin-β 6 Subunit
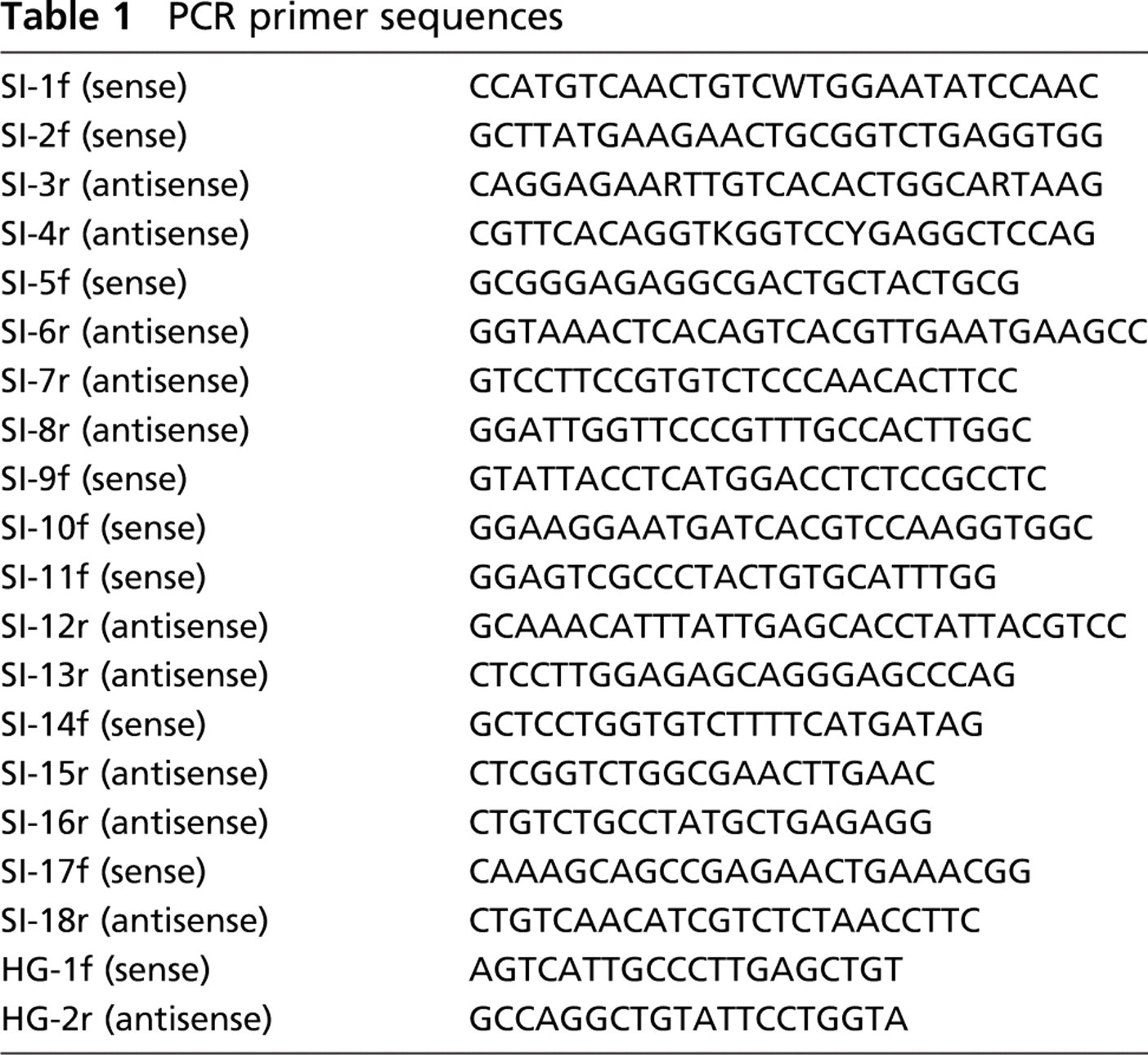
Cloning and sequencing were as described previously (McAleese et al. 1998). Samples of ovine bronchus (n=3) were homogenized on ice in 2 ml of Tri-Reagent (Sigma-Aldrich; Poole, UK) using a tissue homogenizer and total RNA isolated using chloroform extraction and precipitation with isopropanol as described previously (Wastling et al. 1998). cDNA was amplified using primers SI2f and SI3r (Table 1), based on well-conserved regions in known sequences (human, J05522; mouse, AF115476; and guinea pig, J05522). Degenerate positions were introduced in SI3r where there was uncertainty. PCR amplification product SI2f-SI3r was suitable for direct sequencing and provided ~500 bp of sequence information. Gene-specific primers SI5f, SI6r, and SI7r (Table 1) were designed using this data. Three more primers based on the previously known sequences were also made: SI8r located near the 3î end; SI9f and SI10f located toward the 5î end (Table 1). PCR amplification products SI5f-SI8r and SI9f-SI6r were suitable for direct sequencing, followed by SI10f-SI13r. The 3î end was obtained by two, part-nested, PCR amplifications using the gene-specific forward primers SI11f and SI14f, together with an oligo(dT)-anchor primer. For the 5î end, a 5î3î RACE (Rapid Amplification of cDNA Ends) kit (Roche Molecular Biochemicals; Lewes, UK) was used, following manufacturer's instructions. Total RNA was reverse transcribed using the gene-specific primer SI13r. The dA-tailed cDNA was amplified twice by PCR using the 5î dT-anchor primer supplied and the gene-specific primers SI15r and SI16r. PCR product for the 3î and 5î ends were cloned into pCR4-TOPO for sequencing.
PCR primer sequences

Analysis of Integrin-β 6 Expression and Distribution in Ovine and Bovine Tissues
Samples (n=3) obtained from the following ovine tissues were screened for integrin-β6 expression by RT-PCR and immunocytochemistry: gingivae, tongue, pharynx, abomasum, jejunum, colon, rectum, mesenteric lymph node, bronchial lymph node, kidney, liver, spleen, bronchus, lung parenchyma, skin, and PB. Samples (n=1) of gingivae, tongue, pharynx, bronchus, lung parenchyma, abomasum, jejunum, colon, rectum, skin, and PB were also obtained from a Holstein-Friesian heifer. Total RNA was isolated from RNAαlater-fixed samples (Ambion) of gingivae, tongue, PB, and skin using Tri-Reagent (Sigma-Aldrich), as described previously. For all other tissues, samples were homogenized in RLT buffer (Qiagen; Crawley, UK) using a Mini Bead-Beater-8 (Stratech Scientific; Soham, UK), and total RNA was isolated and purified using an RNeasy mini kit as directed by the manufacturer (Qiagen). One μg of DNA-free RNA was reverse transcribed using 2.5 mM (dT)15 and Promega (Southampton, UK) reverse transcriptase; 1/20th volume was amplified by PCR using integrin-β6-specific primers (Table 1, SI2f and SI3r) or for the housekeeping gene ATP-synthase (Table 1, HG1f and HG2r) with equivalent quantities of non-reverse-transcribed RNA as negative controls. Amplifications were carried out for 30 sec at 94C, 30 sec at 60C, and 30 sec at 72C for 35 thermocycles for integrin-β6, and 30 thermocycles for ATP-synthase in a final magnesium concentration of 1.5 mM at pH 8.3.
Cryostat sections (8 μm) of frozen samples were mounted on Snow Coat X-tra charged slides (BDH Laboratory Supplies), air dried for 10 min under forced air, and fixed for 10 min at 21C in PBS (pH 7.4) containing 2% paraformaldehyde. Sections were then washed with staining buffer (pH 7.4, PBS supplemented with 4 mM MgCl2 and 0.5% Tween 80) (Sigma-Aldrich), blocked for 15 min with staining buffer + 10% heat-inactivated normal sheep serum (NShS) and incubated for 1 hr with mouse IgG2a anti-integrin-αvβ6 (clone 10D5; Chemicon Europe, Chandlers Ford, UK) or control mouse IgG2a (clone OX34; Serotec, Kidlington, UK) diluted to 0.5 μg/ml in staining buffer + 10% NShS. Sections were washed with PBS + 4 mM MgCl2 and incubated for 30 min with Alexa Fluor-488-conjugated rabbit anti-mouse IgG (Invitrogen; Paisley, UK) diluted to 2 μg/ml in staining buffer + 10% NShS. Sections were then washed with PBS + 4mM MgCl2 and incubated for an additional 30 min with Alexa Fluor-488-conjugated donkey anti-rabbit IgG (Invitrogen) diluted to 2 μg/ml in staining buffer + 10% NShS. Sections were counterstained for 10 min at 21C with staining buffer containing 0.5 μg/ml of propidium iodide, washed with PBS + 4 mM MgCl2, and mounted with No. 1.5 glass coverslips using Mowiol (pH 8.5) mounting media (EMD Biosciences; San Diego, CA).

cDNA sequence and deduced amino acid sequence for ovine integrin-β6. The transmembrane region is underlined.
Imaging and Microscopy
Integrin-β6 and ATP-synthase-specific PCR products were run on ethidium bromide-stained 1.6% agarose gels and visualized using a Molecular Imager FX Pro Plus (Bio-Rad Laboratories; Hemel Hempstead, UK).
The degree of integrin-αvβ6-specific labeling was assessed in three discontiguous sections from each frozen tissue block (two blocks per sample). Labeling intensity was classified as follows: negative-where there was no detectable specific labeling, even at the highest magnification [x63 1.4 numerical aperture (NA) oil immersion objective lens]; weak-where specific labeling was only visible at x63; moderate-where specific labeling was detected with a x20 dry 0.6 NA objective lens; strong-where specific labeling was detected with a x10 dry 0.32 NA objective lens; and very strong-where specific labeling was readily visible with a x10 dry 0.32 NA objective lens. Representative images were acquired using an MRC-600 confocal laser-scanning microscope (CLSM; Bio-Rad Laboratories) mounted on an Axiovert 100 inverted microscope (Carl Zeiss; Welwyn Garden City, UK). Fluorophores were excited and imaged sequentially using the 488-nm (Alexa Fluor-488) and 568-nm (propidium iodide) lines from a 15-mW Kr/Ar laser (Bio-Rad Laboratories).
Images were prepared for publication using Object-Image (Vischer et al. 1994) and Photoshop (Adobe Systems, Uxbridge, UK). Object-Image is a public domain software package based on NIH Image (Rasband and Bright 1995), developed by Norbert Vischer (The University of Amsterdam, Amsterdam, The Netherlands) and is freely available via the Internet at http://simon.bio.uva.nl/object-image.html.
Results
Pathology
All ovine and bovine organs used in this study were examined and deemed free of gross pathological changes at necropsy. In addition, H and E staining revealed no evidence for significant ongoing or recent inflammation in any of the frozen sections used for integrin-αvβ6-specific immunocytochemistry.
Cloning and Sequencing of Ovine Integrin-β 6
A total of 2862 bp of nucleotide sequence was obtained, including 2364 bp of open reading frame (Figure 1; EMBL accession no. AJ439062). The sequence of the coding region was confirmed by sequencing clones obtained using the gene-specific primers 17f (bases-24 to −1) and 18r (bases 2500-2523), from the 5î and 3î noncoding regions, respectively. SI18r was chosen some distance from the stop codon to avoid a TA-rich region. Bronchial epithelial RNA samples from three different animals were used, and the sequences obtained were found to be almost identical. Relative to sheep 1 (Figure 1), RNA from sheep 2 (clone L17-4) had one base change resulting in a change in the deduced amino acid sequence (Asn 629 to Ser). Sheep 3 (clone L15-1) had two base changes resulting in changes in the deduced amino acid sequence (Asn 100 to Ser, Asn 235 to Ser). Comparison of the ovine integrin-β6 nucleic acid sequence with those published for other species indicates homologies of 97% with bovine, 88% with human and guinea pig, and 84% with mouse (data not shown)
Tissue-specific Transcription of Integrin-β 6
Primers SI2f and SI3r (Table 1) were used to screen for integrin-β6 expression in a variety of ovine tissues from three sheep and a Holstein-Friesian heifer (Figure 2). These primers span an intron and produce PCR amplification products of 532 bp from cDNA and ~1200 bp from DNA. A similar pattern of integrin-β6 expression was observed in all three sheep and in the bovine samples (Figure 2). Integrin-β6 mRNA was consistently detected, albeit at varying levels, in samples that contained epithelium, but there was no evidence of integrin-β6 transcription in ovine liver, spleen, or mesenteric and bronchial lymph nodes (Figure 2). To confirm their specificity for integrin-β6, SI2f and SI3r primer sequences were compared with equivalent regions in related integrin subunits (β1, u10865, and x07979; β3, j02703; and β5, af468059, and j05633). In addition, RT-PCR amplification products obtained from skin and coronary band were sequenced and gave identical results to those for airway epithelium (data not shown).
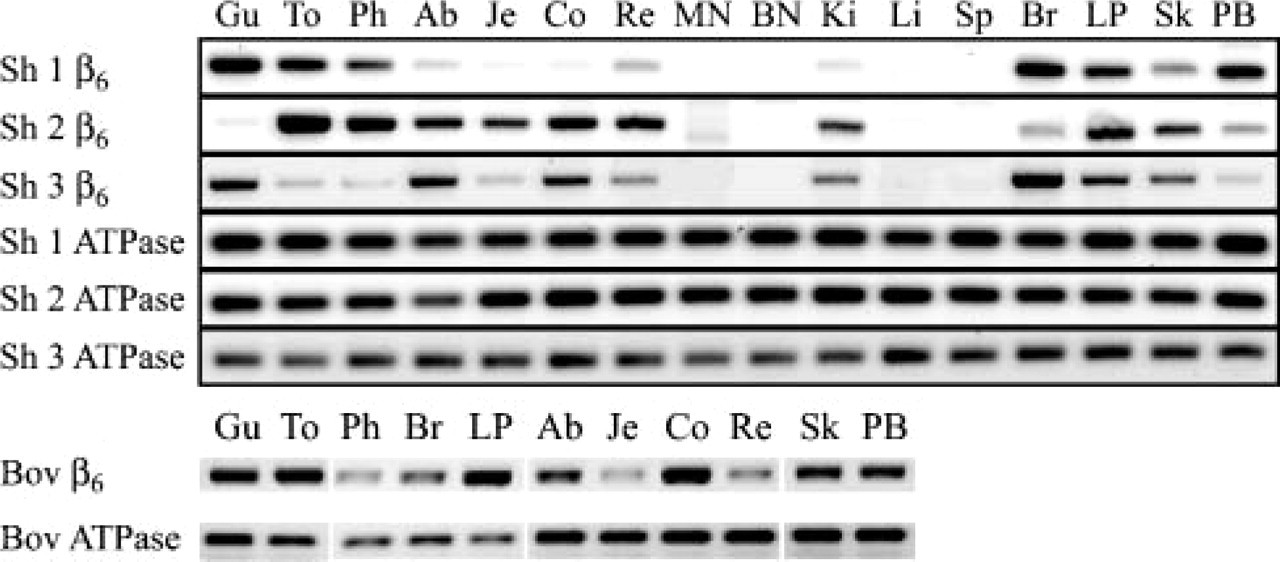
RT-PCR analysis of integrin-β6 transcription. Samples of the following tissues were screened for integrin-β6 and ATP-synthase transcription by RT-PCR: gingivae (Gu); tongue (To); pharynx (Ph); abomasum (Ab); jejunum (Je); colon (Co); rectum (Re); mesenteric lymph node
Immunocytochemical Localization of Integrin-α vβ6 in Ovine Tissue
The mouse monoclonal antibody 10D5 recognizes a divalent cation-dependent epitope present in both human and murine integrin-αvβ6 (Huang et al. 1998). Images of ovine lung sections probed with 10D5 in the presence or absence of MgCl2 are shown in Figure 3. The signal-to-noise ratio obtained with 10D5 was substantially improved using tertiary donkey anti-rabbit IgG Alexa Fluor-488 antibodies (data not shown). There was minimal evidence for specific staining when labeling was performed in standard PBS containing 0.5% Tween 80 (Figure 3A). However, well-defined labeling of the membranes of airway epithelial cells was observed when the staining buffer was supplemented with 4 mM MgCl2 (Figure 3B). In addition to being divalent cation dependent, the epitope targeted by 10D5 is readily denatured by methanol, even at −20C (data not shown).
A variety of tissues were screened for the presence of integrin-αvβ6 using 10D5. The results obtained from three different sheep are presented in Table 2 along with representative images of selected tissues (Figure 4). Integrin-αvβ6-specific staining was restricted to epithelial cells in all of the tissues examined and was remarkably consistent between different sheep. In agreement with the RT-PCR data (Figure 2), integrin-αvβ6 was not detected by immunocytochemistry in samples from liver, spleen, or mesenteric and bronchial lymph nodes (Table 2). Weak, but nonetheless specific, integrin-αvβ6 staining of stratified epithelium was present throughout the oral cavity with more intense staining observed on gingivae and tonsillar crypt epithelial cells (Table 2; Figures 4A-4D). In the remainder of the gastrointestinal tract, 10D5 labeling was most prominent in the glandular epithelium of the abomasum, where it increased in intensity from the basal to superficial layers (Table 2; Figure 4E). The glandular epithelium of the rectum exhibited a similar, although less intense, integrin-αvβ6 staining pattern (Table 2; Figure 4H). In contrast to RT-PCR (Figure 2), immunocytochemistry yielded only sporadic evidence of localized integrin-αvβ6 expression in the jejunum and colon (Table 2; Figures 4F and 4G). With the exception of the alveolar lining cells, integrin-αvβ6 was detected on the surface of epithelial cells throughout the lung (Table 2; Figure 4J). Based on 10D5 labeling intensity, integrin-αvβ6 expression appears to be increased in smaller airways, reaching maximal levels in terminal bronchiolar epithelium (Table 2). Integrin-αvβ6 specific staining was also observed on distal convoluted tubular epithelial cells in the kidney. There was little evidence for integrin-αvβ6-specific staining in the stratified epithelium of normal skin, although it was present in sweat glands and detected at relatively high intensity in hair follicles (Table 2; Figure 4K). By contrast, the stratified epithelium of PB epidermis exhibited strong integrin-αvβ6-specific staining (Figure 4L).
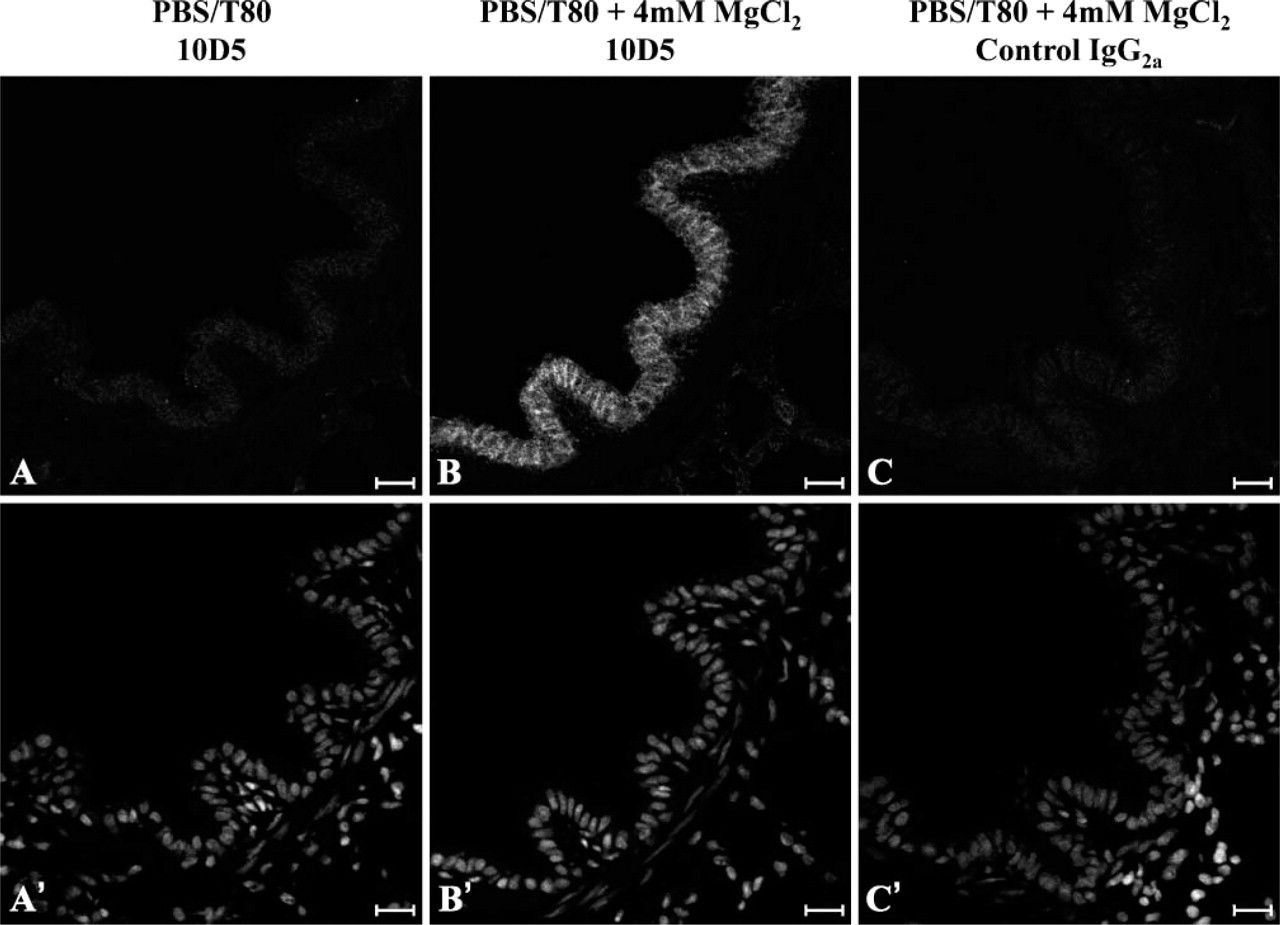
The integrin-αvβ6-specific monoclonal antibody, 10D5, recognizes a divalent cation-dependent epitope on the surface of ovine epithelial cells. Serial sections of snap-frozen, paraformaldehyde-fixed, ovine lung were incubated with 10D5 in the absence
Integrin-αvβ6 immunoreactivity in various ovine tissues
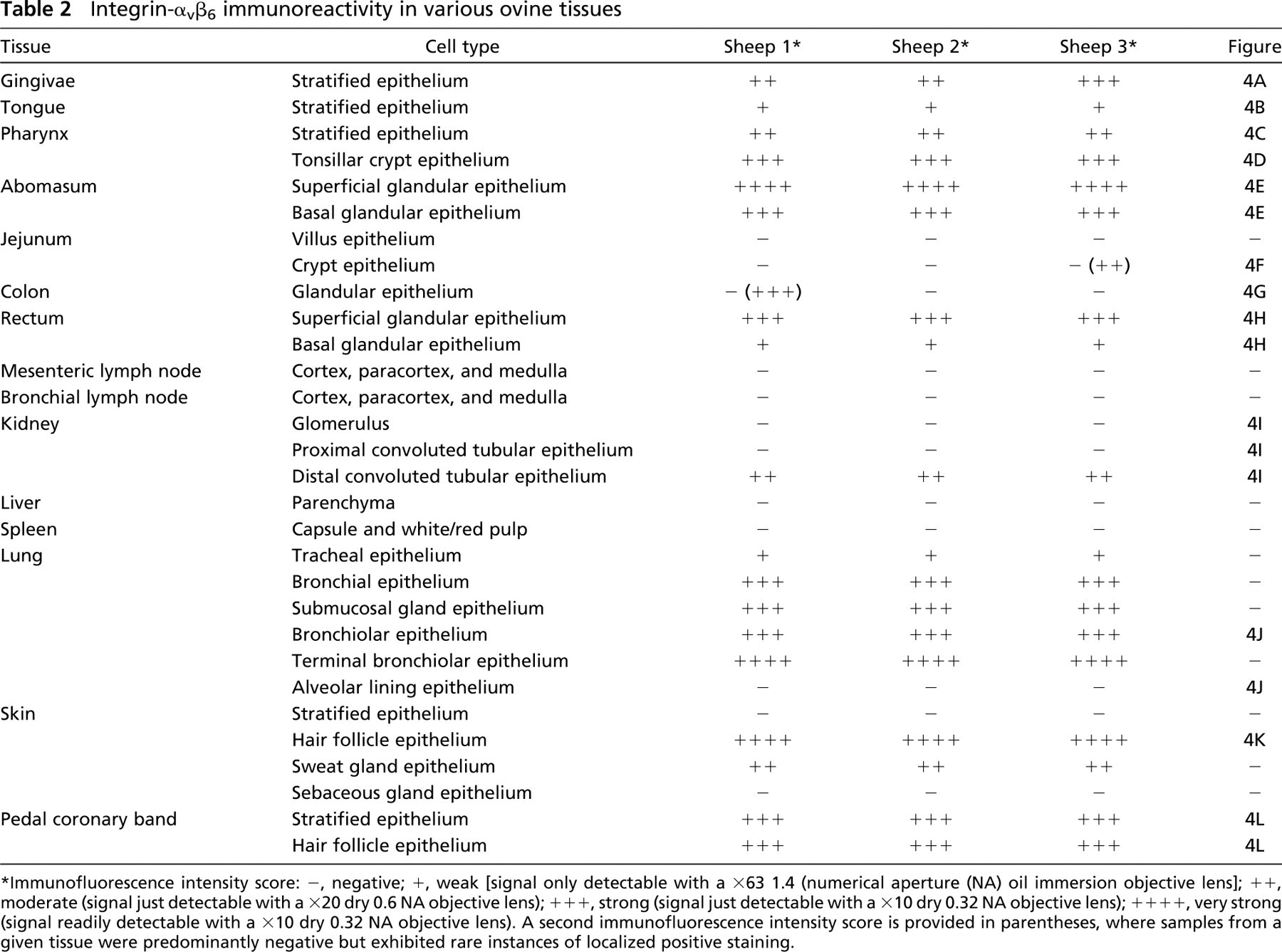
∗ Immunofluorescence intensity score: -, negative; +, weak [signal only detectable with a x63 1.4 (numerical aperture
Immunocytochemical Localization of Integrin-α vβ6 in Bovine Lung
Snap-frozen sections of lung from 15-day-old bull calves (n=2) were labeled with 10D5 (Figure 5). Integrin-αvβ6-specific staining exhibited a similar distribution to that observed in sheep and was present on the surface of epithelial cells lining bronchi (Figure 5A) and bronchioles (Figures 5C and 5E) throughout the lungs of both calves. Specific staining was not detectable in tracheal epithelium of either calf and, like sheep, there appears to be a progressive decrease in integrin-αvβ6 expression proximally toward mainstem bronchi and trachea.
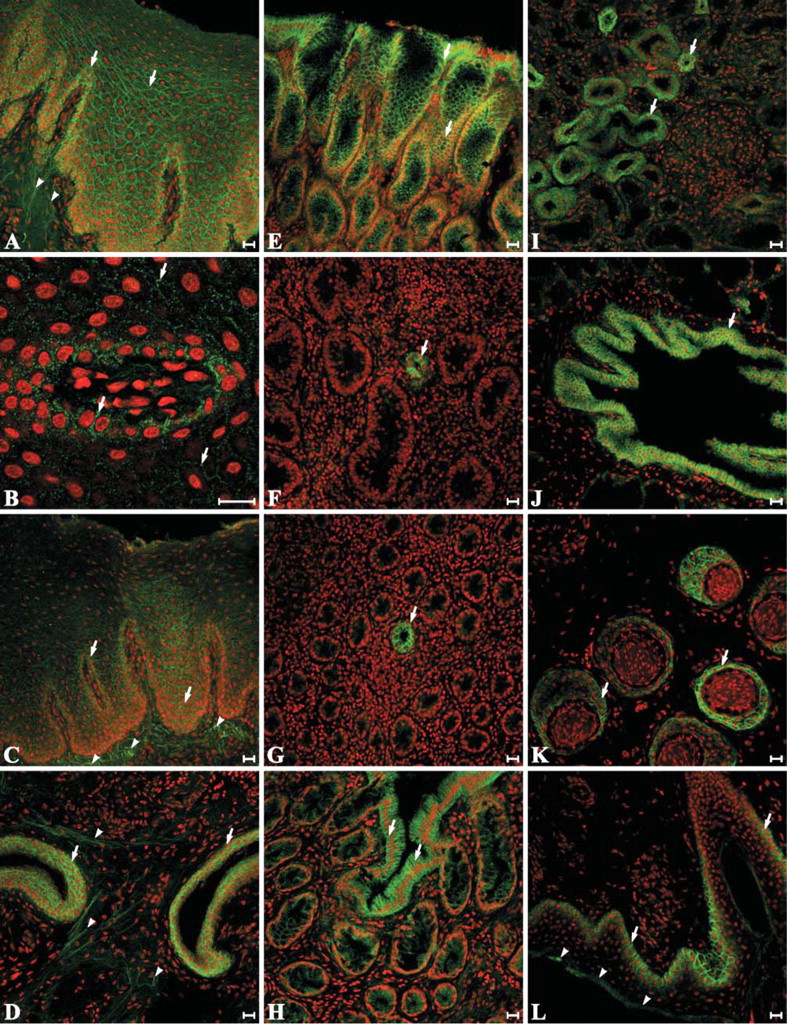
Immunofluorescent labeling of integrin-αvβ6 in ovine tissues. Snap frozen, paraformaldehyde fixed, sections of ovine tissues were indirectly labeled with Alexa fluor-488 (green) using an integrin-αvβ6 specific primary antibody, 10D5. Nuclei were counterstained with propidium iodide (red). Specific integrin-αvβ6 labeling is indicated with arrows. Arrowheads, where present, identify auto-fluorescent connective tissue fibers. Individual panels are shown for
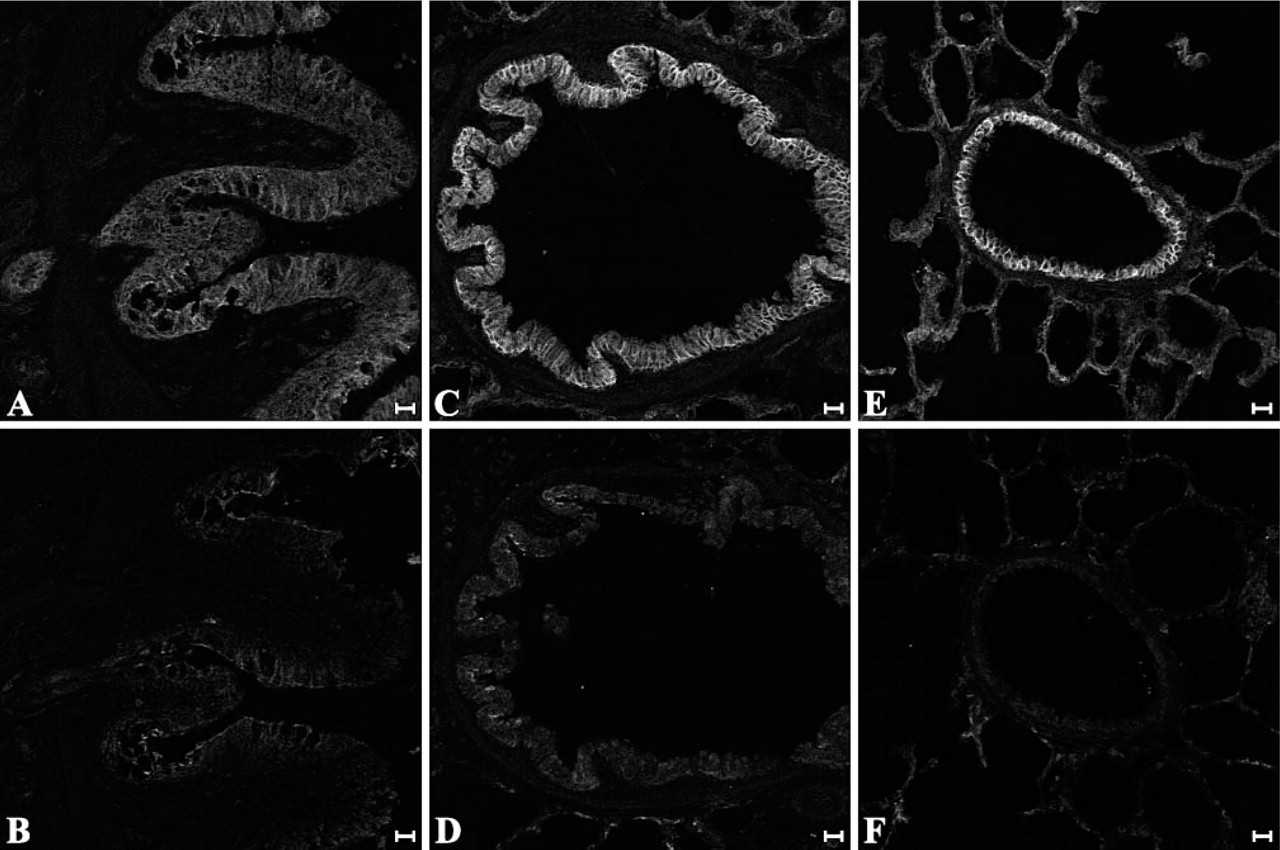
Immunofluorescent labeling of integrin-αvβ6 in bovine tissues. Sections of snap-frozen, paraformaldehyde-fixed lung from 15-day-old calves were incubated with 10D5
Discussion
Cloning and sequencing of ovine integrin-β6 subunit (Figure 1) revealed close homology (~88%) with its human counterpart (Sheppard et al. 1990), which has previously been shown to convey susceptibility to FMDV infection when expressed in the normally non-permissive cell line SW480 (Jackson et al. 2000). Integrin-β6 pairs exclusively with integrin-αv (Hynes 2002) and analysis of integrin-β6 distribution in rhesus macaque and human tissues suggests that integrin-αvβ6 is expressed exclusively by epithelial cells (Breuss et al. 1993,1995). Data presented here show that integrin-β6 transcription is similarly restricted to epithelia in ruminants (Figure 2), and that the integrin-αvβ6 heterodimer is expressed on the surface of epithelial cells within these tissues (Table 2; Figure 4 and Figure 5).
Ruminants appear to differ significantly from primates and rodents in that integrin-αvβ6 appears to be constitutively and quite strongly expressed in the airways (Figure 4 and Figure 5). Integrin-β6 is either not expressed or is expressed at very low levels in rhesus macaque and human lung under normal conditions but is upregulated in inflamed lung (Breuss et al. 1995). Similarly, expression of integrin-αvβ6 in pulmonary tissues in the mouse occurs in response to inflammatory stimuli (Munger et al. 1999). In contrast, ovine integrin-β6 was cloned from mRNA extracted from apparently normal bronchial epithelium, and relatively high levels of integrin-β6 transcription and integrin-αvβ6 immunoreactivity were consistently detected in ovine airway respiratory epithelium, in the apparent absence of inflammation (Figure 2; Table 2). Similarly, the presence of integrin-αvβ6 on the surface of bronchial and bronchiolar epithelium from normal 15-day-old calves (Figure 5) is strongly indicative of constitutive expression at this site and entirely consistent with the hypothesis that bronchiolar respiratory epithelium is the initial site of FMDV replication in cattle infected by aerosol challenge (Brown et al. 1996).
Although RT-PCR analysis of ovine integrin-β6 transcription was generally well corroborated by immuno cytochemistry, there was only sporadic evidence for integrin-αvβ6-specific immunofluorescence in the ovine jejunum and colon (Table 2; Figure 4), despite the fact that integrin-β6 mRNA was consistently detected in these tissues (Figure 2). This may be due either to strong transcription and high turnover of protein with limited surface expression or to patchy, localized integrin-αvβ6 expression in the jejunum and colon (Figures 4F and 4G), which is more readily detected by RT-PCR than by immunocytochemistry. Alternatively, posttranscriptional regulation of integrin-αvβ6 expression, including regulation of αv and β6 subunit dimerization, may also contribute to this apparent discrepancy. These findings are consistent with our studies of mouse jejunum where constitutive expression detected by RT-PCR was associated with low-level protein detection, which was apparently unaffected by inflammatory stimuli (Knight et al. 2002).
Integrin-β6 transcripts were detected throughout the upper gastrointestinal tract (Figure 2), including the oral mucosa where, once infection is established, productive FMDV lesions are usually found (Grubman and Baxt 2004). Consistent with a study of bovine tongue and ventral soft palate (Monaghan et al. 2005), integrin-αvβ6-specific labeling of stratified epithelium from ovine gingivae, tongue, and pharynx was most intense within the basal layers (Figure 4), a pattern which is remarkably similar to the distribution of FMDV within the epithelium of the tongue and pharynx of experimentally infected cattle (Prato Murphy et al. 1999; Zhang and Alexandersen 2004; Monaghan et al. 2005). FMDV lesions are also commonly observed at the PB of infected animals (Grubman and Baxt 2004), and integrin-β6 mRNA was present within samples of PB from all three sheep. Immunocytochemistry revealed relatively strong integrin-αvβ6 expression within the basal layers of PB epithelium but not of normal skin epithelium (Table 2; Figure 4). This is remarkably similar to integrin-αvβ6 expression in bovine interdigital skin and PB (Monaghan et al. 2005) and may explain the tropism of FMDV for the PB once it has disseminated systemically from the primary sites of infection. Integrin-αvβ6 also appears to be constitutively expressed in sweat glands in ovine skin (Table 2), which has not been described in primates (Breuss et al. 1993,1995) but is likely to have limited significance in the context of FMDV. RT-PCR analysis of samples from a Holstein-Friesian heifer indicates that integrin-β6 expression is similarly distributed in cattle (Figure 2).
As discussed above, FMDV exploits integrin-αvβ6 as a receptor (Jackson et al. 2000; Monaghan et al. 2005), and this is the first study characterizing expression of this integrin in sheep. Data presented here indicate that integrin-αvβ6 expression is restricted to epithelial cells in the sheep and, in agreement with studies conducted in cattle (Monaghan et al. 2005), occurs at sites of known FMDV replication. Importantly, integrin-αvβ6 is expressed at high levels by tonsillar crypt epithelium in the sheep and by airway epithelium in both sheep and cattle. These are both sites in which classical FMDV lesions have yet to be described. However, the very large surface area represented by airway epithelium and the high infectivity of FMDV suggests that this could represent the most likely portal of entry for the virus, which is consistent with studies showing the presence of virus in the bovine respiratory system within 24 hr of infection (Brown et al. 1996).
Footnotes
Acknowledgements
This work was funded by the Norman Salvesen Emphysema Research Trust and the Wellcome Trust (Grant #060312).
DNA sequencing was performed by the DNA Sequencing Service, The University of Durham. We thank Dr. Mike Wilkinson (GlaxoSmithKline) for the donation of the BIORAD MRC600 confocal microscope.
