Abstract
The Slug transcription factor plays an important role in epithelial-mesenchymal transformation during embryogenesis and is expressed in adult tissues during carcinogenesis. By detecting expression of a Slug–β-galactosidase fusion protein, we have now demonstrated that Slug is also re-expressed in a variety of normal tissues in the adult mouse. Slug is expressed at relatively high levels in patchy stretches of basal cells in stratified and pseudostratified epithelium, including skin, oral mucosa, esophagus, stomach, rectum, cervix, and trachea. Slug is also found at variable levels in fibroblasts and stromal smooth muscle cells in many tissues. Sites of more intense Slug expression in mesenchymal tissues include cartilage, kidney glomeruli, lung, ovary, and uterus. Therefore, Slug expression is not restricted to the period of embryonic development or to pathological processes. The pattern of localization to basal cells in various epithelia suggests that Slug may play a role in the cell migration that occurs during continual renewal of these tissues. (
P
Snail family members play a key role in the epithelial–mesenchymal transformation (EMT) that occurs during normal embryonic development (reviewed in Hemavathy et al. 2000a; Manzanares et al. 2001; Nieto 2002). In the chicken, Slug is expressed in cells undergoing EMT during gastrulation, neural crest emergence, somitogenesis, and limb development (Nieto et al. 1994; Ros et al. 1997; Sefton et al. 1998). Slug appears to be essential for EMT in the heart of the chicken and for neural crest induction and migration in both the chicken and
In the mouse embryo, Slug protein is expressed at high levels in craniofacial mesenchyme, developing bone, the wall of the stomach and intestines, and the mesenchymal components of many organs, including the kidney, lung, and submandibular salivary gland (Oram et al. 2003). Slug expression is also seen in the outflow tract of the heart, the endocardial cushions, and the heart valves. Slug is expressed in a few embryonic epithelial structures, including the vibrissae and hair follicles. By days 17–18 of gestation, Slug levels decline markedly. Although Slug expression decreases after differentiation, Slug mRNA can be detected in many adult mammalian tissues. A single RNA transcript of 2.1 kb is found in the lung, liver, testis, kidney, heart, and skeletal muscle of adult mice (Jiang et al. 1998b; Savagner et al. 1998). In humans, a 2.2-kb Slug transcript is expressed at relatively high levels in heart and placenta and at lower levels in liver, skeletal muscle, kidney, and pancreas (Cohen et al. 1998). The function of Slug in normal adult tissues has not been identified.
In this study we demonstrate that Slug expression in normal tissues of the adult mouse is widespread but not universal. Several of the sites of Slug expression in the adult are sites at which the protein is found in the embryo. Somewhat unexpectedly, Slug is also found in clusters of basal cells of various stratified and pseudostratified epithelia in the adult mouse, suggesting a role in the normal renewal of these tissues.
Materials and Methods
Mice
In Slug-lacZ mice, the Slug locus has been inactivated by an in-frame insertion of the β-galactosidase gene into the zinc-finger coding region of the Slug gene (Jiang et al. 1998a). The β-galactosidase insertion inactivates Slug but results in the production of a Slug-lacZ fusion protein bearing 120 amino acids of the amino terminal region of the Slug protein under the control of the Slug promoter. Slug expression in these mice can be monitored by detecting β-galactosidase activity (Jiang et al. 1998a). For the present studies, heterozygous Slug-lacZ males on a mixed C57BL/6 and 129 background were obtained from the original colony of Slug–lacZ mice generated by Jiang et al. (1998a) and were mated to homozygous wild-type females (C57BL/6 or 129 inbred background) to produce heterozygous and homozygous wild-type offspring in a 1:1 ratio. Genotypes of the mice were readily determined by PCR, as follows: DNA was isolated from small segments of tail using the QIAmp DNA Mini kit (Qiagen; Valencia, CA). DNA was amplified using two sets of primers, one specific for the wild-type Slug locus (5′-AGAACGGAAGCTGTTTGGCTG-3′, 5′-CTATTTGGTTGGTAAGCACATGAG-3′) and one specific for the Slug-lacZ locus (5′-TCGCCTTCTATCGCCTTCTTG-3′, 5′-CTATTTGGTTGGTAAGCACATGAG-3′). Forty cycles of amplification were carried out in a volume of 20 μl at an annealing temperature of 60C in a mixture containing 10 μl lysis buffer (50 mM KCl, 10 mM Tris-HCl, pH 8.3, 2 mM MgCl2, 0.45% Tween-20, 0.45% NP40), 1 μl PCR buffer (166 mM (NH4)2SO4, 670 mM Tris-HCl, pH 8.8, 1 mg/ml fraction V bovine serum albumin), 1 mM additional MgCl2, 200 μM each dNTP, 125 ng of each primer, 0.5 μg of DNA, and 1 U of Taq polymerase (Qiagen). Amplification with the primer set specific for the wild-type locus yielded a 590-bp fragment, while amplification with the primer set specific for the modified Slug locus produced a 430-bp fragment. Animal breeding, care, genotyping, and use were conducted in accordance with all relevant local, state, and federal guidelines.
Detection of β-Galactosidase Activity
Slug–lacZ heterozygotes and wild-type mice were sacrificed humanely by CO2 inhalation. Tissue samples were embedded in OCT, frozen in liquid nitrogen, and sectioned at 10 μm. Sections were air-dried for 30 min, then stained for 24 hr at 37C in a humidified chamber using the Beta-Gal staining kit (PanVera; Madison, WI) as recommended by the supplier, except that the substrate (5-bromo-4-chloro-3-indolyl-β-d-galactoside) was employed at twice the recommended concentration. Slides were counterstained with 0.1% nuclear fast red in 5% (NH4)2SO4, dehydrated, and mounted. Negative controls included tissues from wild-type mice that did not express β-galactosidase and tissues from Slug–lacZ mice incubated in the absence of substrate. No β-galactosidase activity was detected in either case (not shown).
Preparation of Skin Sheets
The two sides of the ear were separated and placed in cold 25 mM EDTA in PBS. After tissues were incubated for 1 hr on ice, cartilage was removed from the underside of the skin using fine forceps. The skin was cut into strips 3–4 mm wide and incubated in fresh 25 mM EDTA in PBS for 90 min at 37C. The epidermis was then peeled from the dermis. Epidermal sheets were rinsed in PBS for 30 min and placed in the Beta-Gal staining solution for overnight incubation at 37C. The next day, the sheets were mounted in glycerol/gelatin.
Immunohistochemistry for β-Galactosidase
Immunohistochemistry for β-galactosidase was performed to confirm findings with histochemical techniques. Frozen sections were rinsed in PBS and fixed briefly in acetone. Peroxidase and protein block (DAKO; Carpinteria, CA) were applied as recommended by the supplier. Primary antibody diluted 1:500 in antibody diluent (DAKO) was applied for a 60-min incubation at room temperature. The antibody employed was a polyclonal rabbit anti-β-galactosidase antibody kindly provided by Dr. Achim Gossler of the Institute for Molecular Biology, Hannover Medical School (Hannover, Germany). After rinsing, secondary antibody, avidinbiotin complex reagents, and diaminobenzidine chromogen were applied as recommended by the producer of the VectaStain Elite kit employed (Vector Laboratories; Burlingame, CA). Slides were counterstained with nuclear fast red, dehydrated, and coverslipped. Negative controls included tissues from wild-type mice that did not express Slug–β-galactosidase and tissues from Slug-lacZ mice incubated in the absence of primary antibody. No β-galactosidase immunoreactivity was detected in either case (not shown).
Results
Previous studies have used in situ hybridization to detect expression of Slug mRNA in mice (Nieto et al. 1994; Jiang et al. 1998a; Locascio et al. 2002). Detection of Slug protein expression in mice has been accomplished by histochemistry for β-galactosidase in the same strain of heterozygous Slug-lacZ mice used for the present studies. These mice produce a Slug–β-galactosidase fusion protein (Jiang et al. 1998a; Oram et al. 2003). The Slug–β-galactosidase knock-in allele not only retains the native Slug promoter but also includes the first 120 amino acids of the Slug protein. Therefore, the nuclear localization signal of the protein is intact (Jiang et al. 1998a). Mice heterozygous for a Slug–β-galactosidase knock-in allele appear phenotypically normal (Jiang et al. 1998a). The expression pattern of the Slug–β-galactosidase fusion protein accurately represents the expression pattern of Slug mRNA (Jiang et al. 1998a), and histochemistry for β-galactosidase is more sensitive than ISH for demonstration of patterns of Slug expression (Oram et al. 2003).
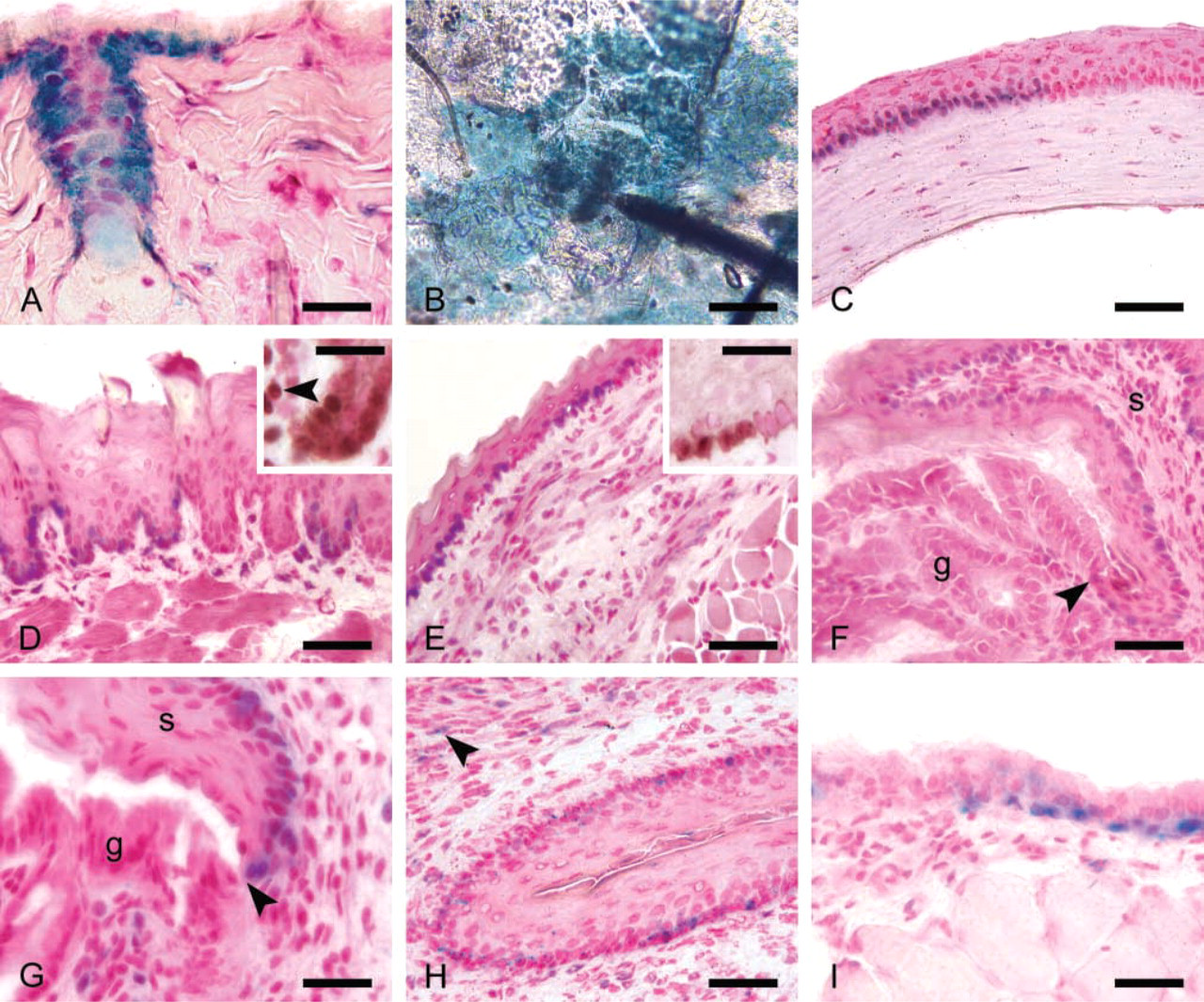
Slug localization in stratified and pseudostratified epithelium. (
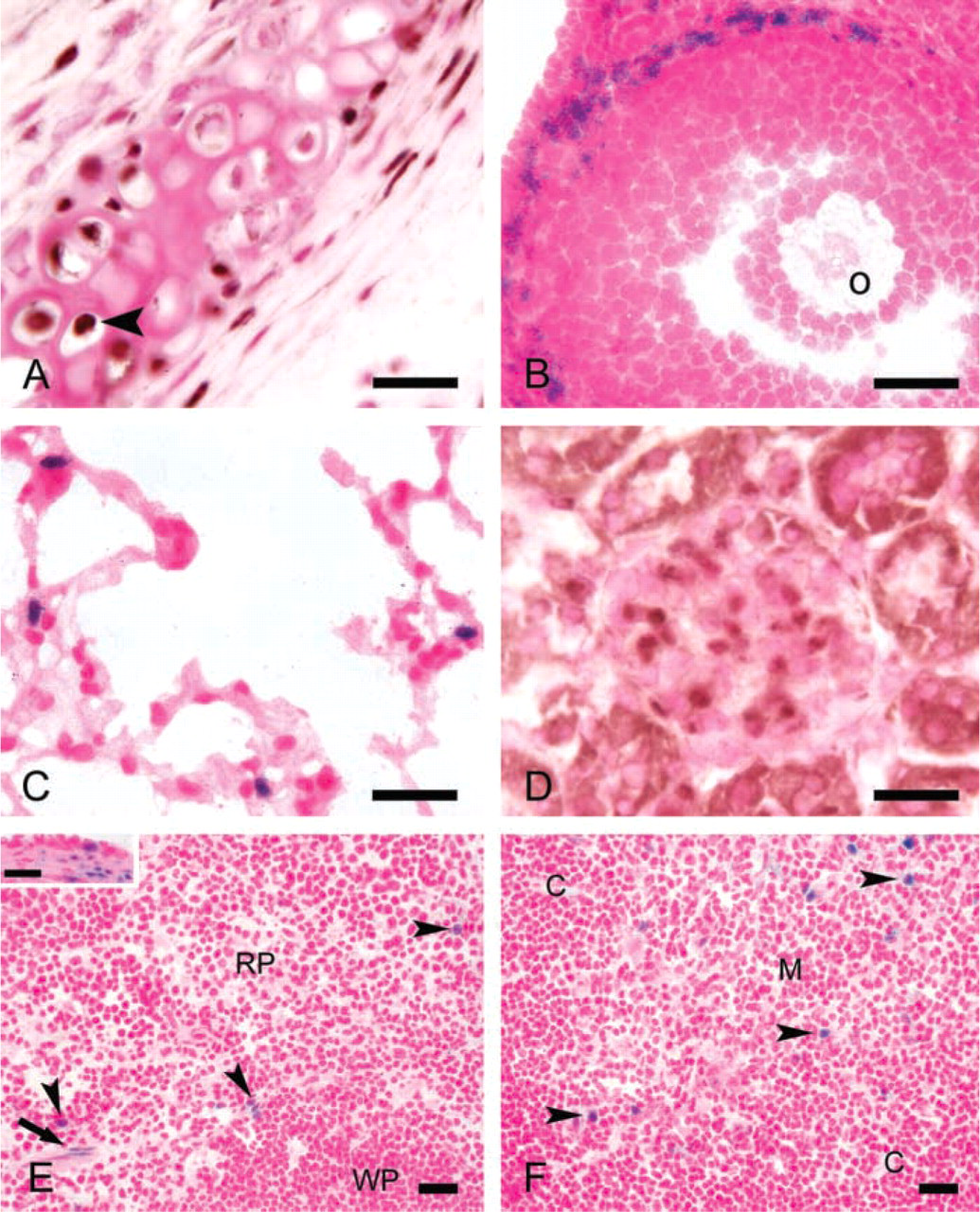
Slug expression in various other tissues. (
In the present studies we confirmed the patterns of β-galactosidase expression revealed by histochemistry in several organs using IHC techniques. IHC for the β-galactosidase protein also proved to be a sensitive and accurate measure of Slug expression. As expected for a fusion protein containing the nuclear localization signal of a transcription factor, the Slug–lacZ fusion protein was localized to cell nuclei. Slug was widely expressed in stratified and pseudostratified epithelium. Nuclear Slug protein was detected in epithelium of the skin, cornea, esophagus, stomach, rectum, trachea, and cervix (Figure 1). In the skin, Slug expression was seen primarily in hair follicles (Figure 1A) and in groups of basal cells in the interfollicular epidermis. Slug-expressing interfollicular cells were usually clustered around hair follicles (Figure 1B). A similar pattern of Slug expression in large clusters of basal epithelial cells was seen in the cornea (Figure 1C), tongue (Figure 1D), esophagus (Figure 1E), stomach (Figure 1F), and rectum (Figure 1G). Slug expression was restricted to the stratified squamous epithelium of the stomach and rectum, and ceased abruptly at the border between the squamous and columnar epithelium at both sites (Figures 1F and 1G). Slug expression in the stratified squamous epithelium of the cervix and vagina was observed in small numbers of scattered basal epithelial cells (Figure 1H).
In general, Slug expression was limited to stratified squamous epithelium. No expression was observed in the columnar epithelium of the gut, the lining epithelium of the uterus, airway or alveolar epithelium in the lung, or the transitional cell epithelium of the urinary bladder. Furthermore, the epithelia of solid organs, including the liver, pancreas, kidney, and salivary glands, did not express Slug. However, basal cells within the psuedostratified columnar epithelium of the trachea expressed readily detectable Slug (Figure 1I).
We detected widespread although not universal Slug expression in mesenchymal tissues throughout the body, including some fibroblasts and stromal smooth muscle cells. Slug expression was particularly prominent in chondrocytes (Figure 2A), smooth muscle cells of the uterus and cervix (Figure 1H), and thecal cells of the ovary (Figure 2B). Modest Slug expression was demonstrated in the lung, kidney, spleen, and thymus. In the lung, Slug was expressed in scattered interstitial cells with round to oval nuclei (Figure 2C). Slug-expressing cells in the kidney were localized to the glomeruli (Figure 2D). The identity of these glomerular cells could not be determined. However, their polygonal morphology suggested that they were not vascular endothelial cells. Slug expression in the spleen and thymus was confined to small numbers of scattered cells of unknown identity (Figures 2E and 2F). Slug expression was not detected in the brain, heart, skeletal muscle, retina, or melanocytes. For comparison, patterns of Slug expression identified in previous studies are summarized in Table 1.
Comparison of Slug expression in human and murine adult and embryonic tissues a

—, none; +, low; ++, moderate; +++, high.
Discussion
It has been suggested that Slug expression during chicken development facilitates the release of cells from epithelial structures, thus enabling them to migrate (Sefton et al. 1998). This is in keeping with the postulated role of Snail family transcription factors in EMT-like processes that occur during carcinoma progression in the adult (Cano et al. 2000; Hemavathy et al. 2000a; Savagner 2001). In adult epithelium, Snail family members appear to modulate EMT by controlling the expression of proteins that comprise intercellular adhesion structures.
For example, expression of Snail is elevated in melanomas, in oral squamous cell carcinomas, and in some breast tumors. In these tumors, levels of Snail and E-cadherin expression are inversely correlated (Poser et al. 2001; Yokoyama et al. 2001; Blanco et al. 2002). Snail binds directly to the E-box within the E-cadherin gene to repress its expression (Batlle et al. 2000; Cano et al. 2000). Although it is likely that there is some redundancy between the functions of Slug and Snail, Snail clearly regulates E-cadherin expression in mouse keratinocytes, whereas Slug appears to play an insignificant role in this process (Cano et al. 2000). Recent studies in breast tumor cells, however, indicate that both Slug and Snail can repress endogenous E-cadherin expression and that this process is mediated by binding to E-box elements (Hajra et al. 2002). Slug may also modulate desmosomal and adherens functions. Overexpression of Slug in the NBT-II bladder carcinoma cell line stimulates the first stages of growth factor-induced EMT, including desmosomal dissociation and cell spreading (Savagner et al. 1997).
Table 1 summarizes the sites of Slug expression that have been identified in previous studies of fetal and adult mice and humans. In the present study we observed widespread expression of Slug in adult mesenchymal cells such as fibroblasts, smooth muscle cells, and chondrocytes. This is in keeping with the previous identification of Slug mRNA in many adult human tissues (Cohen et al. 1998) and with high expression of Slug in the gut wall and developing bone in the mouse embryo (Oram et al. 2003). In addition, the pattern of Slug expression we observed in hair follicles has also been reported in mouse embryos (Oram et al. 2003). However, prominent Slug expression in clusters of basal epithelial cells in stratified squamous and pseudostratified columnar epithelium of the integumentary, gastrointestinal, urogenital, and respiratory systems was unexpected. Such focal aggregates of Slug-expressing cells may represent the expanding progeny of individual precursor cells, and formation of these foci may be part of the normal renewal process in stratified and pseudostratified epithelium. Expression of Slug in these cells may be related to the migratory capabilities of expanding epithelial cell populations. Furthermore, fibroblasts retain some migratory capabilities, and therefore Slug expression in some of these cells may also reflect transient migratory behavior. Evidence to support a potential role for Slug in the migratory behavior of adult cells comes from the demonstration that c-kit-expressing hematopoietic cells from Slug knockout mice have a reduced ability to migrate in response to stem cell factor (Perez-Losada et al. 2002).
These observations expand the potential contributions of Slug to regenerative processes in adult tissues. More work is needed to define Slug-controlled transcriptional programs and their implications for the adult organism. In particular, it will be important to determine the environmental signals that stimulate Slug expression and to identify the immediate targets of Slug transcriptional regulation. Based on the regulation of Snail family factors by growth factors during development (reviewed in Nieto 2002), it is likely that these growth factors will also play an important role in modulating Slug expression in adult tissues. Furthermore, direct transcriptional regulation of E-cadherin by Snail and Slug (Batlle et al. 2000; Cano et al. 2000; Hajra et al. 2002) suggests that Slug may be important in regulating the expression, localization, and function of the component proteins of intercellular adhesion structures.
Footnotes
Acknowledgements
Supported by funds from the Department of Veterinary Biosciences, The Ohio State University (DK) and by National Institutes of Health grants #HD34883 (TG) and #CA89216 (DK).
