Abstract
The recently identified gene
A
Although mice are used extensively as a model system to study organ physiology and pathophysiology, little information is currently available regarding the expression of
The translation product of full-length
Materials and Methods
Animals and Collection of Tissues
Animal protocols were approved by the institutional animal care and use committees at the University of Cincinnati and the University of Florida, where appropriate. Glands were excised from mice euthanized by exsanguination after carbon dioxide narcosis.
Generation of Muc19-EGFP Knockin Mice
Genomic DNA to create the 5' and 3' homologous arms of the targeting vector were generated by PCR of bacterial artificial chromosome (BAC) DNA (clone E11, RPCI-22 BAC Library from 129S6/SvEvTac female mouse; BACPAC Resources, Oakland, CA) using the following primers: 5' arm, forward primer, 5'-CAAAAGCACACCCTCAGATAACG-3' and reverse primer, 5'-CAATGTTGAAGTCTCCTCCAGGC-3'; 3' arm, forward primer, 5'-GCCCTTTGAGCAGGAGCAATG-3' and reverse primer, 5'-CGAGTGTAAGTGTGCCCTGAAGTTG-3'. The 5' arm PCR product (5431 bp) was cut with PstI and EaeI (4961-bp final product), and the 3' arm PCR product was cut with SacI and HindIII (2462-bp final product). The 5' arm was coupled to EGFP from pEGFP-N3 (BD Biosciences, San Jose, CA; GenBank accession number U57609). A NotI site immediately downstream of the EGFP coding region was first filled in using T4 polymerase followed by ligation of the resultant blunt ends. The 5' arm plus EGFP with its own stop codon, polyadenylation signal, and SV40 poly A was inserted 5' to the upstream loxP site of the selection cassette PGKneolox2DTA (Soriano 1997; kindly provided by Dr. Rulang Jiang) that encodes for neomycin under control of the phosphoglycerate kinase promoter. Sequence from the STOP cassette, pBS302 (kindly provided by Dr. Rulang Jiang; GenBank accession number U51223) was inserted immediately 3' of the downstream loxP site. This cassette was added to help prevent transcriptional read-through to more downstream exons encoding
The vector was cloned into Electro 10 Blue cells (Stratagene; La Jolla, CA) and sequenced, and the purified plasmid DNA was linearized with PvuI. Linearized plasmid was electroporated into 129S6/SvEvTac embryonic stem (ES) cells at the University of Cincinnati Gene Targeted Mouse Service. ES cell clones (after G418 selection) were screened initially by PCR using two primer sets to identify appropriately targeted cells. Primer Set 1 targets genomic sequence 43 bp upstream of the 5' arm (5'-GGCAGTTCACGATTGTCTGTGCAAGC-3') and within PGKneo (5'-CCTGCGTGCAATCCATCTTGTTCAATGGC-3'). Primer Set 2 targets 39 bp downstream of the 3' arm (5'-TGCTTCCCTGGACATTGTTCCTTTGC-3') and within the SV40 polyadenylation signal from pBS302 (5'-GTGCCTTGACTAGAGATCATAATCAGCC-3'). PCR products were verified by direct sequencing from the 5' and 3' ends. Positive ES cell clones identified by PCR were further verified for correct vector integration by Southern analysis. Templates for both 5' and 3' outside probes (308 bp and 332 bp, respectively) were generated by PCR from genomic DNA using the following primers: 5' probe (PvuII probe), 5'-TGCCATCTCTAATAGTTTCAGTGC-3' and 5'-CTCTTGAAGGGGACACCCTAAACC-3'; 3' probe (SphI probe), 5'-TTCGCGATTCACAAGTGTGCATG-3' and 5'-GCACATTAATTAGCTTGGAGCAC-3'. PCR reactions were run in Taq PCR Master Mix (Qiagen; Valencia, CA) with the following conditions: 3 min at 98C; 35 cycles of 30 sec at 94C, 30 sec at 60C, and 30 sec at 72C, followed by 3 min at 72C. Southern blots were performed with random primed probes, and genomic DNA was cut with either PvuII or SphI using standard techniques. Appropriate-sized bands were obtained in all cases (not shown).
Three ES cell clones were expanded and injected into C57Bl/6 blastocysts, resulting in a total of 32 chimeric mice. Chimeric males were mated to Black Swiss (NIHBS) females. One male breeder produced 31 agouti heterozygous F1 progeny, from which four breeder pairs were established at the University of Florida and interbred to generate wild-type and heterozygous animals for EGFP expression experiments. Appropriate targeting of these breeders was confirmed by PCR as described above for ES cell lines (not shown). Tail genomic DNA samples (Wizard Genomic DNA Purification Kit; Promega, Madison, WI) from the resultant pups were used for genotyping by PCR with allele-specific primers to generate bands of 595 bp (wild type) and/or 856 bp (targeted allele). Primers used were: wild type forward, 5'-GCTGCTTTGCTTTCAGTTTTGTAGTG-3'; wild type reverse, 5'-GTCTGGCTGGTTTTGTCTCATACTC-3'; EGFP reverse, 5'-TCACCTTGATGCCGTTCTTCTG-3'. PCR reactions were run in the FailSafe PCR buffer system (Epicentre; Madison, WI). Conditions were: buffer F, 3 min at 94C, 30 cycles of 30 sec at 94C, 30 sec at 55C, and 1 min at 72C, followed by 3 min at 72C. Products were resolved in 1.0% agarose gels made in Tris-acetate-EDTA (TAE) buffer with 0.1% ethidium bromide.
RT-PCR Assays
To prepare cDNA, frozen glands were homogenized directly in TRIzol Reagent (Invitrogen; Carlsbad, CA) using a Mini Bead Beater 8 (BioSpec Products; Bartlesville, OK) for 90 sec in the presence of ~500 mg of silicone carbide beads (1 mm in size). Total RNA was isolated using standard protocols and treated with DNase I using Ambion's DNA-free Reagent Kit (Applied Biosystems; Foster City, CA). Removal of genomic DNA from DNase I-treated RNA (1 μg) was verified by the inability to amplify at 40 cycles the specific 253-bp mouse microsatellite marker, D1Mit46 (UniSTS 116254), using forward primer 5'-AGTCAGTCAGGGCTACATGATG-3' and reverse primer 5'-CACGGGTGCTCTATTTGGAA-3'. RNA purity was assessed by capillary electrophoresis (Agilent 2100 Bioanalyzer; Agilent Technologies, Inc., Santa Clara, CA). RNA integrity numbers ranged from 8.7 to 9.7. RNA (5 μg) was reverse transcribed with random primers using the high-capacity cDNA archive kit (Applied Biosystems), and the resultant cDNA was purified with the QIAquick PCR purification system (Qiagen; Valencia, CA). RNA and cDNA were quantified using the Quant-iT RNA assay kit or the Quant-iT dsDNA HS assay kit with the Qubit fluorometer (Invitrogen).
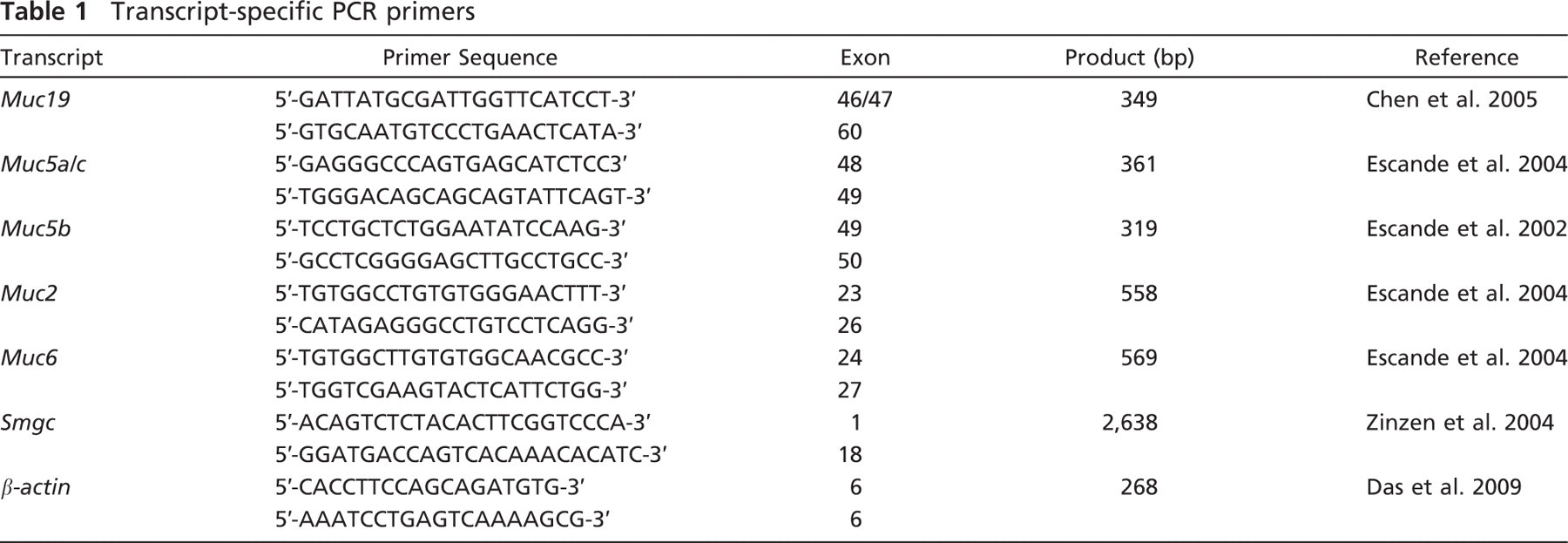
Standard conditions for PCR reactions (10–50 ng cDNA) were: 3 min at 94C; 30–37 cycles of 1 min at 94C, 1 min at 55C, and 1 min at 72C, followed by 10 min at 72C. Products were resolved in 3% agarose gels made in TAE buffer with 0.1% ethidium bromide. Transcript-specific primers (Invitrogen) are shown in Table 1. All reactions were run in Qiagen's Taq PCR Master Mix, except for
Transcript-specific PCR primers
DNA Sequencing
Sequencing was performed at the University of Florida Interdisciplinary Center for Biotechnology Research using an Applied Biosystems 3100 Genetic Analyzer ABI and Big Dye version 3.1 chemistry.
Immunohistochemistry
Whole or minced tissues were fixed for 5 hr in 4% paraformaldehyde in PBS and stored at 4C in 1% paraformaldehyde in PBS. Sections (5 μm) were deparaffinized and treated for 10 min in 5% urea containing 50 mM β-mercaptoethanol at 95C to unmask antigenic sites. Sections were then probed with rabbit anti-Muc19 or preimmune serum (1:300 dilution) as described previously (Fallon et al. 2003), except that we used a Vectastain ABC-HRP kit with Vector Red substrate (Vector Laboratories; Burlingame, CA), and counter-stained with hematoxylin.
EGFP Fluorescence and Confocal Microscopy
Animals were anesthetized by CO2 inhalation, the superior and inferior vena cava were exposed and cut, and then the vasculature was perfused via the left ventricle for 3 min with PBS containing 1 U/ml heparin at a rate of 5 ml/min. The perfusate was then switched to ice-cold 4% paraformaldehyde in PBS for 20 min. Tissues were excised and incubated in the same fixative for 3 hr with rocking at 4C, washed three times in PBS for 10 min, then embedded in Tissue-Tek OTC Compound (Sakura Finetek; Torrance, CA) and frozen. For brightfield and EGFP fluorescence microscopy, frozen sections (5- or 10-μm) were thawed and mounted in Vectashield Hard Set with 4',6-diamidino-2-phenylindole (Vector Laboratories). Images were captured with a Micropublisher 5.0 RTV digital camera (Q Imaging; Pleasanton, CA) using QCapture Pro software (v5.01, Q Imaging) on a Leica DM LB2 microscope (Leica Microsystems; Bannockburn, IL) equipped with a 100-W high-pressure mercury lamp and Chroma fluorescent filter cubes (Chroma Technology Corp.; Rockingham, VT).
For confocal images, frozen sections (10-μm) were thawed and incubated for 1 hr at room temperature with the nuclear stain DRAQ5 (Biostatus Limited; Leicestershire, UK) diluted 1:1000 in PBS. After washing (four times for 15 min in PBS), sections were mounted in mowiol. Images were obtained on a spinning disk confocal imaging system consisting of a CSU10 Yokagawa confocal scan head (Yokogawa Corp. of America; Newnan, GA), a Roper Cascade II EMCCD 512b camera (Roper Scientific, GmbH, Ottobrunn Germany), and an ASI X, Y, and piezo Z computer-controlled stage (Applied Scientific Instrumentation; Eugene, OR), with three lasers (491, 561, 638) for three-channel fluorescent imaging attached to a Leica DMIRB inverted microscope (Leica Microsystems). This system is controlled by the open source software package Micro-manager (http://www.micro-manager.org/). Optical slices of 0.2 μm were obtained using 491 nm ex, 520 em, (EGFP) and far-red 638 nm ex, 695 nm em (Draq5) and compiled into a three-dimensional image with ImageJ software (http://rsbweb.nih.gov/ij).
Electron Microscopy
Tissues from wild-type and Muc19-EGFP knockin mice were fixed by vascular perfusion of 2.5% glutaraldehyde in 0.1 M sodium cacodylate buffer, pH 7.4. Subsequent processing was carried out as described in Fallon et al. (2003). Thin sections were examined and photographed in a Philips CM10 transmission electron microscope.
SDS-PAGE, Western Blots, and Lectin Blots
Glandular tissues were excised immediately after death and frozen in liquid nitrogen. Mucosal scrapings were obtained from freshly isolated stomach and colon tissues and processed immediately. Tissue samples were immediately weighed, and 10–80 mg wet weight of tissue was placed in 500 μl of NuPAGE LDS sample buffer (Invitrogen). Samples were then sonicated (four 10-sec pulses at 30-sec intervals) with a Branson Digital Sonifier 250 at 30% amplitude (Branson Ultrasonic Corp.; Danbury, CT), boiled for 10 min, and centrifuged (10,000 × g for 10 min) at 4C. Supernatant aliquots of volumes equivalent to 0.1 to 3.7 mg wet weight of original tissue were applied directly to either 4–12% NuPAGE bis-Tris or 3–8% NuPAGE Trisacetate gradient gels (Invitrogen). To assay highly glycosylated glycoproteins, gels were stained with Alcian Blue, followed by subsequent silver enhancement of Alcian Blue staining as described previously (Fallon et al. 2003). Silver enhancement of Alcian Blue precedes any staining of proteins (Jay et al. 1990).
To detect SMGC and EGFP, gels were blotted onto polyvinylidene difluoride (PVDF) membranes (Sequi-Blot PVDF membrane, 0.2 μm; Bio-Rad Laboratories, Hercules, CA) and probed overnight (4C) with rabbit anti-mouse SMGC (1:10,000) (Zinzen et al. 2004). For detection of Muc19 and EGFP, gels were transferred overnight onto nitrocellulose (0.2 μm; GE Healthcare, Piscataway, NJ) in transfer buffer (20 mM Tris, 200 mM glycine, 10% methanol) using 250 mAmps of current at 10C. One blot was probed overnight (4C) with rabbit antiserum produced against purified rat sublingual mucin (1:20,000) (Man et al. 1995) and the other blot with rabbit anti-GFP (IgG fraction, 1:2000; Invitrogen). To detect antibodies, we used the WesternBreeze Chemiluminescent Western Blot Immunodetection Kit (Invitrogen) and BioMax Light film (Eastman Kodak Co.; Rochester, NY).
For lectin staining, nitrocellulose blots were probed with biotinylated lectins, either wheat germ agglutinin or peanut agglutinin (PNA) from Vector Laboratories. Blots were incubated for 1 hr in PBS + 0.2% Tween 20 (PBST). For PNA, blots were then treated with 0.1 N H2SO4 at 80C to remove interfering sialic acid residues. Blots were incubated for 1 hr with agitation in biotinylated lectin at 1 μg/ml in 10 mM HEPES, 0.15 M NaCl, 0.1 mM CaCl2, pH 7.4. After washing three times for 5 min in PBS, blots were incubated for 1 hr in alkaline phosphatase–conjugated streptavidin (1:8000) in PBST, washed three times for 10 min in PBST, followed by two 5-min washes in PBS. Lectins were detected using CDP-Star chemiluminescent substrate with Nitro-Block II (Applied Biosystems) and BioMax Light film. Protein standards included MultiMark Multi-Colored and HiMark PreStained standards (Invitrogen). Each experimental condition was performed at least three times. Digital images were obtained with a Scion Grayscale 1394 digital camera (Fotodyne).
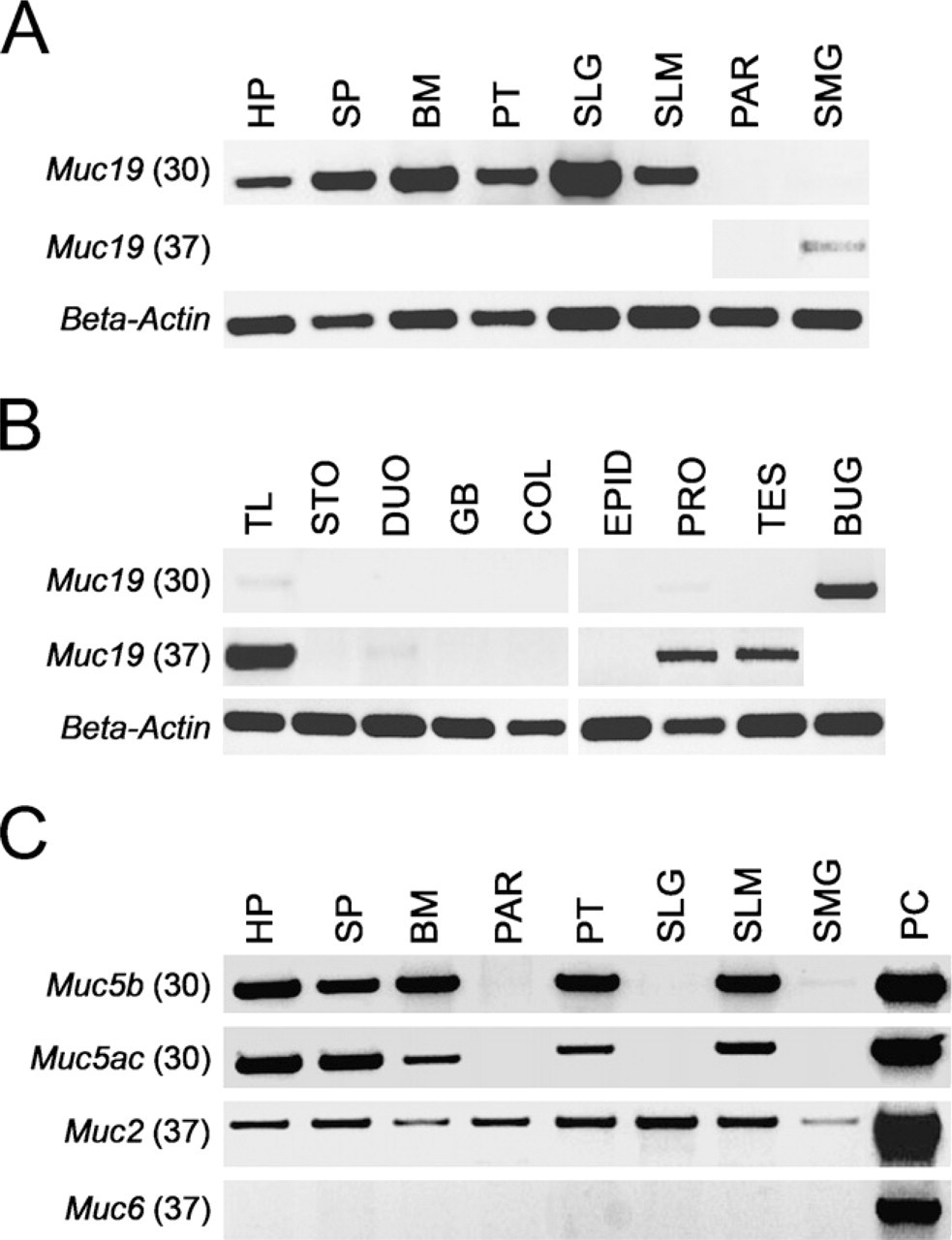
Expression of
Materials
Unless indicated, all chemicals were from Sigma Chemical Co., St. Louis, MO. All kits were used according to the manufacturers' instructions.
Tissue distribution of transcripts for gel-forming mucins in adult mice
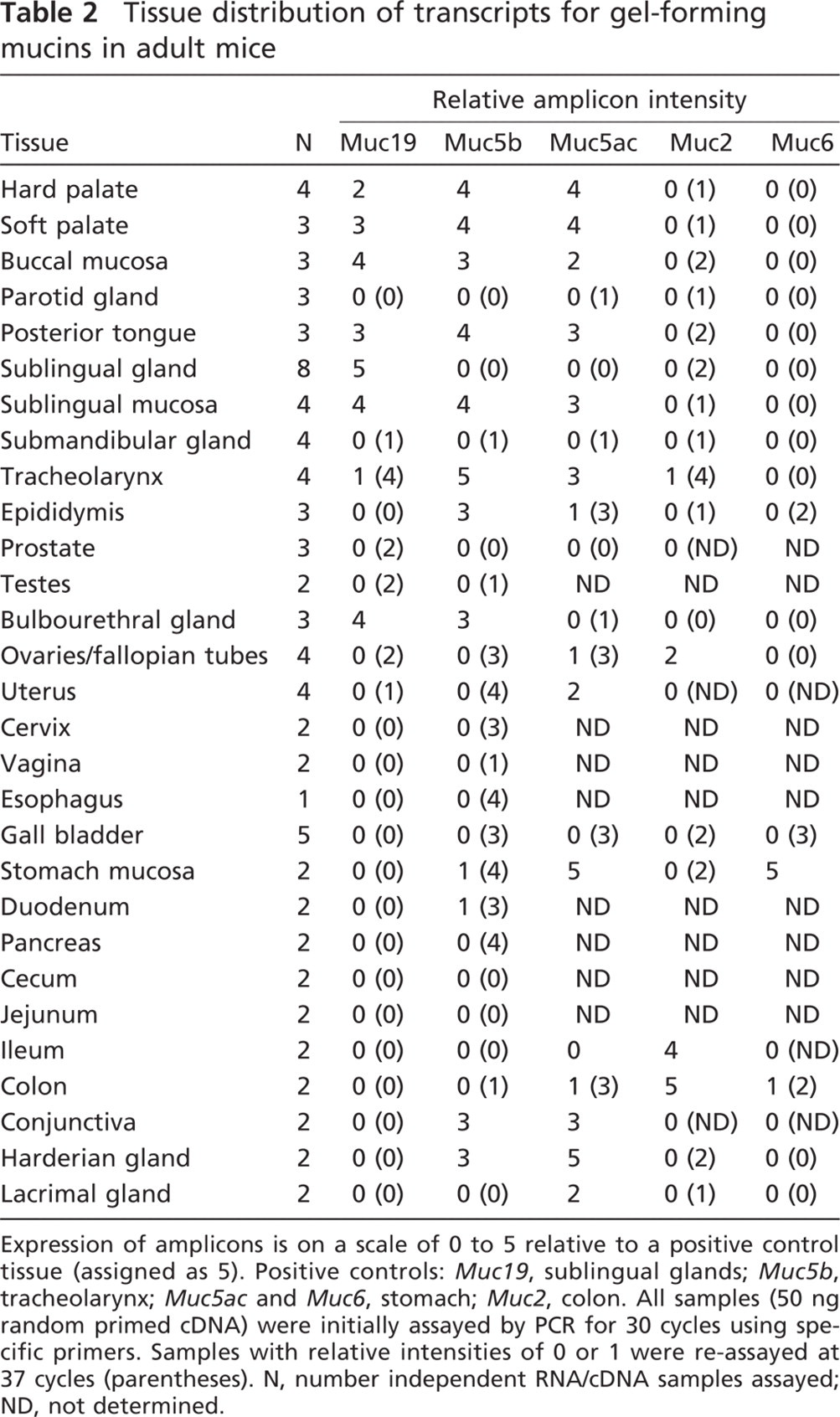
Expression of amplicons is on a scale of 0 to 5 relative to a positive control tissue (assigned as 5). Positive controls:
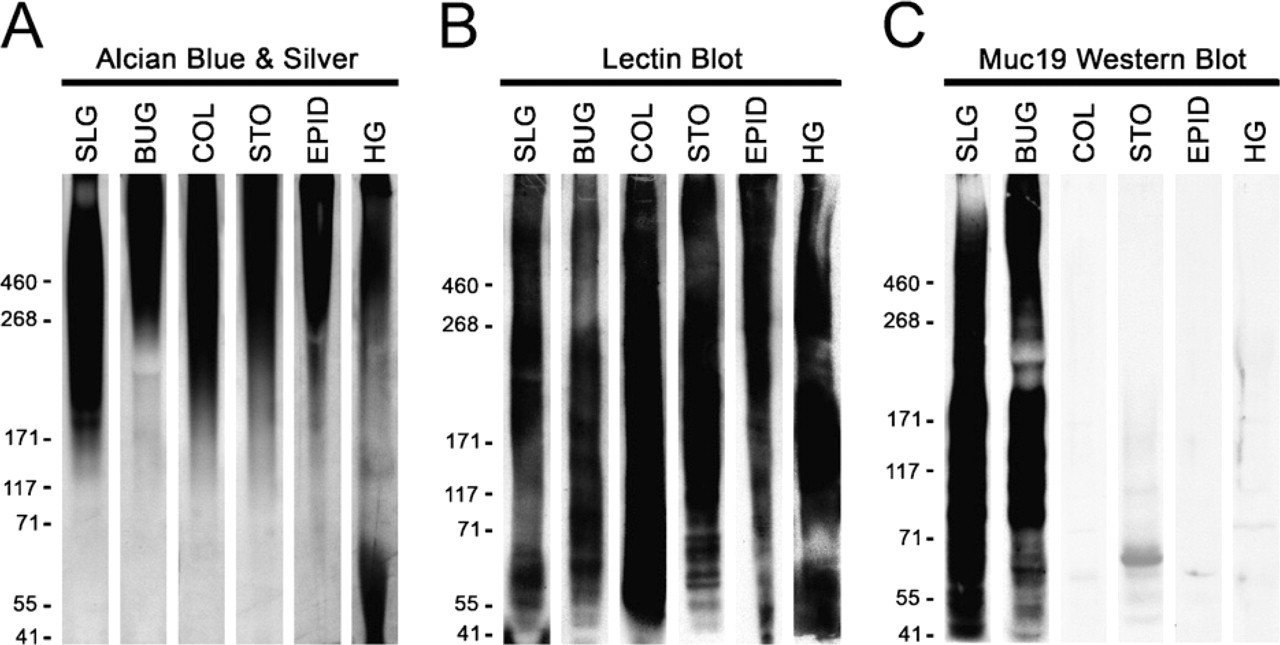
Testing the specificity against gel-forming mucins of antiserum raised against purified rat Muc19. Homogenates from select mouse tissues that express Muc19 and/or other gel-forming mucins were subjected to SDS-PAGE (3–8% gradient gels). One gel
Results
Expression of Muc19 Transcripts in Adult Mice
We initially evaluated tissue expression of
We further assayed all salivary tissues, as well as other select tissues, for transcripts to the other four gel-forming mucins,
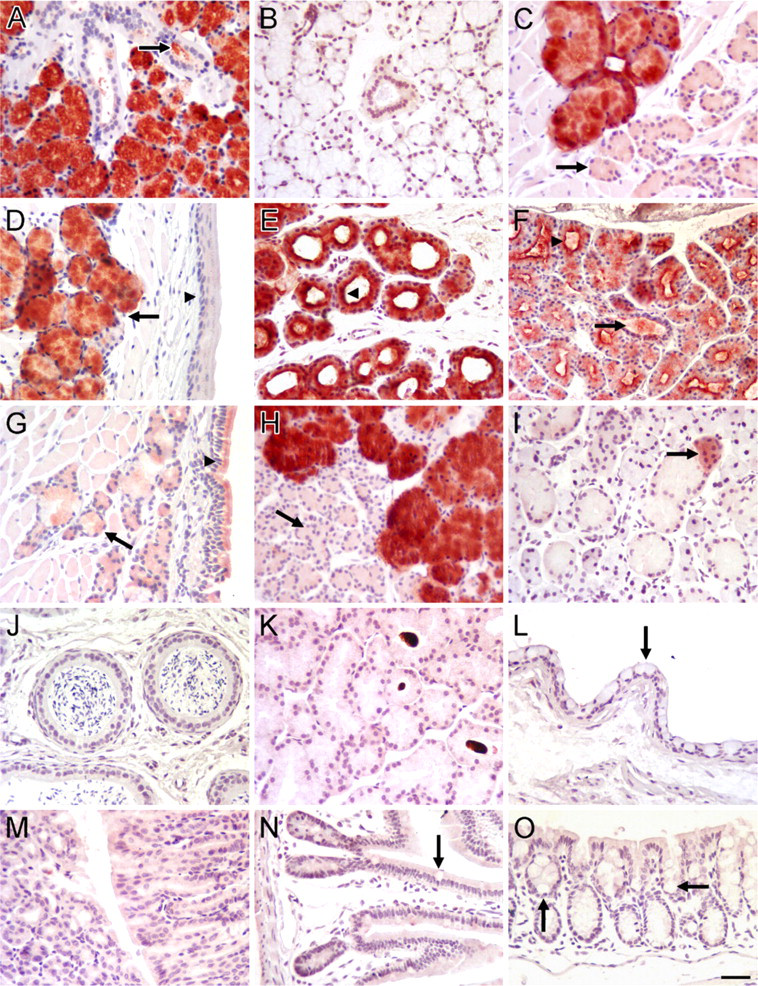
Immunohistochemical localization of Muc19 in mouse tissues.
Immunohistochemical Localization of Muc19
To assess Muc19 localization in murine tissues, we used antiserum derived previously against purified high-molecular-mass mucins from rat sublingual glands (Man et al. 1995). Although this antiserum was demonstrated previously to react with mucous cells and with high-molecular-mass glycoproteins from sublingual glands of mice (Culp et al. 2004; Das et al. 2009), we further determined its specificity for Muc19. We tested in Western blots the reactivity of the anti-serum against homogenates from tissues with undetectable Muc19 transcripts but expression of transcripts for one or more of the other gel-forming mucins: Muc2 (colon), Muc5ac (stomach and Harderian glands), Muc5b (epididymis and Harderian glands), and Muc6 (stomach). Tissue samples were homogenized directly in SDS-PAGE sample buffer with sonication and run on low-percentage (3% to 8%) gradient gels. Conditions for sonication were determined empirically not only to homogenize the tissue but also to slightly fragment endogenous high-molecular-mass glycoproteins, to allow for better penetration into the gel as well as for subsequent transfer to nitrocellulose membranes (not shown). Shown in Figure 2 are samples of homogenates after SDS-PAGE and staining with Alcian Blue, followed by subsequent silver enhancement to detect highly glycosylated glycoproteins (Jay et al. 1990). In all cases, highly glycosylated glycoproteins formed a large smear from the top of the gel to about the 171-kDa marker. The transfer of high-molecular-mass glycoproteins to nitrocellulose was verified (Figure 2B) by probing blots with lectins that react with carbohydrate epitopes of gel-forming mucins and other smaller glycoproteins (McGuckin and Thornton 2000). Upon probing identical blots of each homogenate, only high-molecular-mass glycoproteins in sub-lingual and bulbourethral glands reacted with our antiserum (Figure 2C). Components of higher mobilities also reacted with the antiserum, due mostly to smaller peptides or glycopeptides released during sonication, based upon results of initial preliminary experiments (not shown).
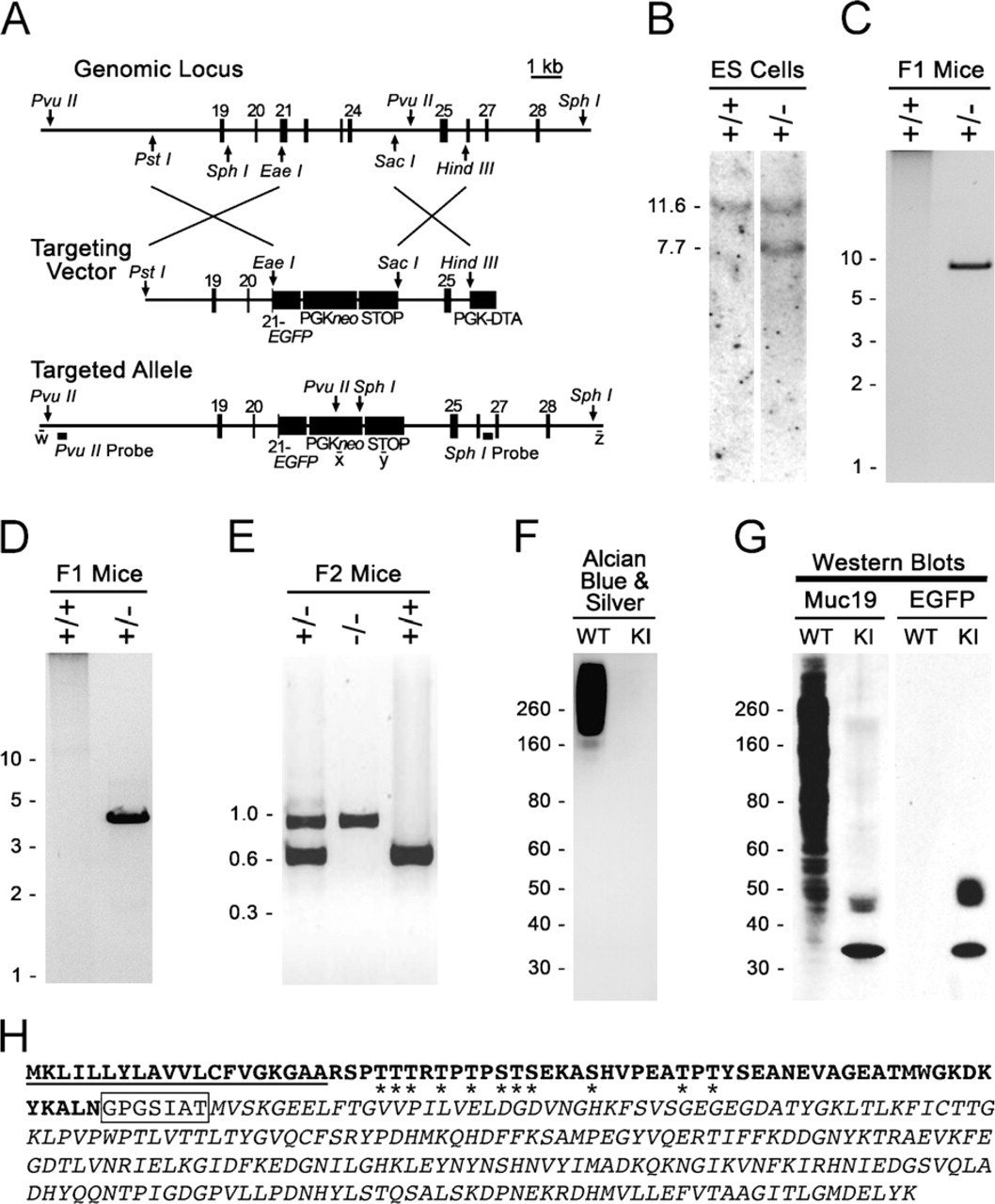
Production and initial characterization of Muc19-EGFP knockin mice.
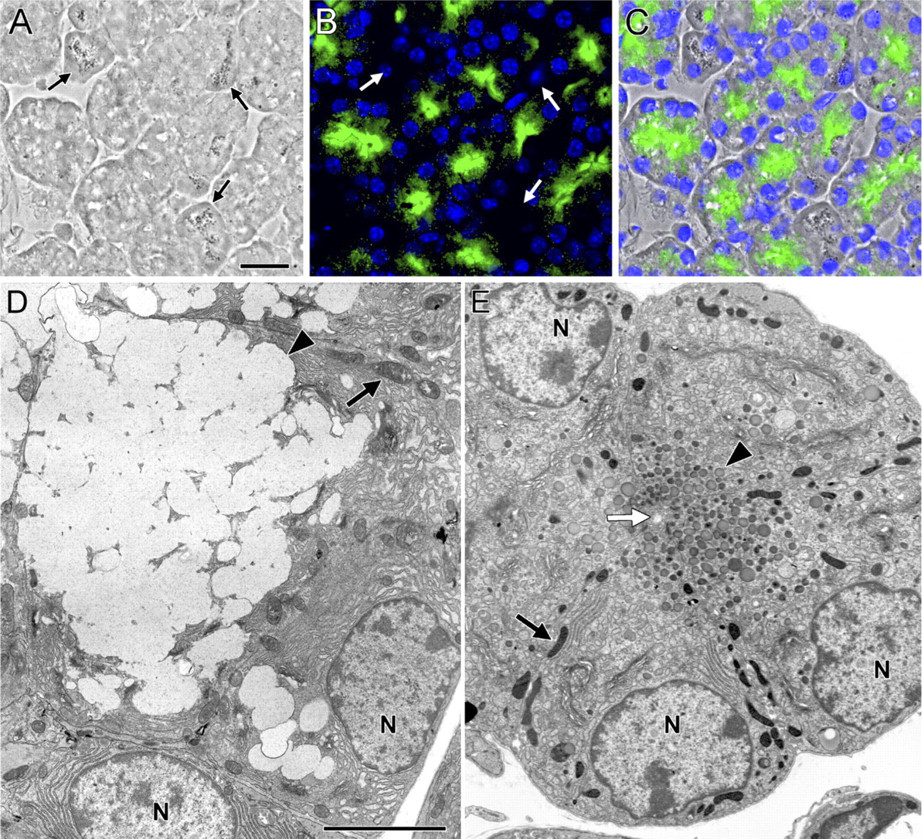
Muc19-EGFP fluorescence in sublingual mucous cells from homozygous Muc19-EGFP knockin mice and comparison of cell ultra-structure to wild-type acinar cells.
In paraffin sections of different tissues, our anti-Muc19 reacts intensely with salivary mucous cells of sublingual glands, posterior tongue, soft palate, hard palate, and buccal mucosa (Figures 3A and 3C–3F). Mucous cells in the bulbourethral glands also stain strongly (Figure 3H). Interestingly, very light staining of surface goblet cells and glandular mucous cells is observed within the nasal mucosa of the soft palate (Figure 3G). Submandibular glands contain a sparse population of small groups of lightly stained acinar cells (Figure 3I), consistent with our RT-PCR results and previous results in rats (Man et al. 1995). Non-reactive tissues include epididymis, Harderian glands, conjunctival epithelium, stomach mucosa, ileum, and colon (Figures 3J–3O).
Tissues From Muc19-EGFP Knockin Mice to Evaluate Muc19 Expression
As a complementary approach to assess tissue expression of Muc19, we created a knockin mouse model in which EGFP was inserted in-frame within exon 21 of the gene
In sublingual sections from homozygous Muc19-EGFP knockin mice, EGFP fluorescence is localized to the apical regions of nearly all tubular mucous cells but is absent in serous demilune cells (Figures 5A–5C). The conversion in expression of native Muc19 to Muc19-EGFP results in a marked transition from the typical mucous cell ultrastructure (i.e., a cytoplasm packed with large electron-lucent secretion granules; Figure 5D) to cells with small electron-dense granules in the apical cytoplasm (Figure 5E). Serous demilune cells appear unaltered, containing abundant rough endoplasmic reticulum and a few serous granules (not shown).
To evaluate tissue expression of Muc19-EGFP, we used heterozygous mice to preserve the mucous cell phenotype and thus maintain normal tissue morphology. As shown in Figure 6A, a uniform and marked green fluorescence is observed in whole sublingual glands (lower center panel) but not submandibular glands (lower right panel) from heterozygous knockin mice. As expected, sublingual glands from wild-type mice are non-fluorescent (lower left panel). In sections of sublingual glands, EGFP fluorescence is prominent in mucous cells but is absent in ductal structures (Figure 6B) and in serous demilune cells (Figure 6C). Fluorescence within mucous cells is restricted to granular structures of irregular shapes and sizes distributed throughout the cytoplasm, consistent with localization to mucous secretory granules (Figure 6D). EGFP fluorescence displays the same pattern of tissue expression as Muc19 immunoreactivity. Mucous cells of minor salivary glands are all fluorescent, including those within the sublingual mucosa (Figure 6E), posterior tongue (Figure 6F), hard palate (Figure 6G), and soft palate (Figure 6H). EGFP fluorescence is also observed throughout the mucous cell population of bulbourethral glands (Figure 6I). Mucous cells within submucosal glands of the tracheolarynx are highly inconsistent, with some cells devoid of fluorescence and others exhibiting very low intensity (Figure 6K). EGFP fluorescence is absent in samples of epididymis (Figure 6J) and colon (Figure 6L), as well as in stomach, Harderian glands, and mucous cells of the nasal airways within the soft palate (not shown).
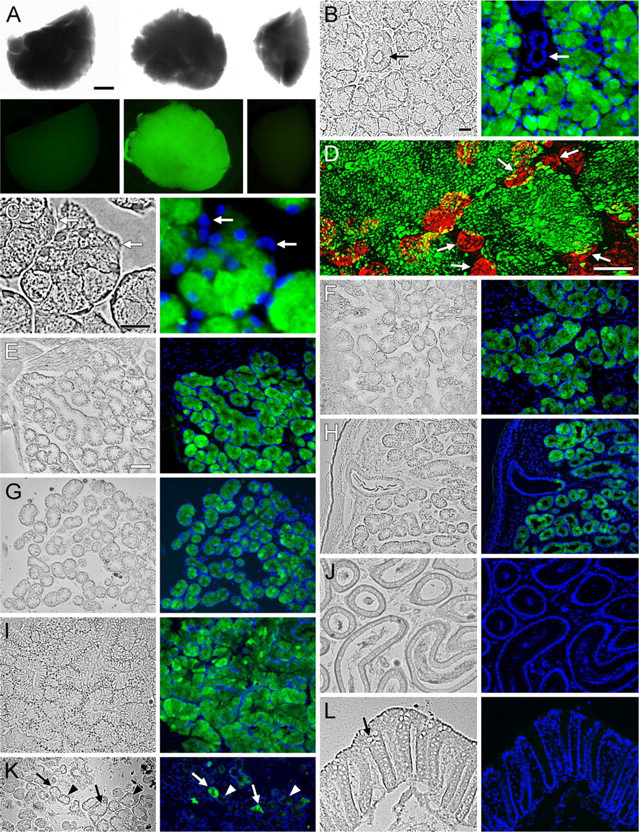
EGFP fluorescence in tissues of heterozygous Muc19-EGFP knockin mice.
Mucous Cell Phenotype in Homozygous Knockin Mice and Expression of Additional Gel-forming Mucins
As described above for Figure 5, the absence of expression of gel-forming mucins in sublingual glands results in mucous cells without a cytoplasm filled with large mucin secretion granules. In contrast to sublingual glands, RT-PCR results presented above (Table 2) suggest that mucous cells of the minor salivary glands express the gel-forming mucins Muc5b and/or Muc5ac, in addition to Muc19. Moreover, bulbourethral glands may express Muc5b, given the abundance of its transcripts. We therefore reasoned that if mucous cells of the glands from homozygous knockin mice express gel-forming mucins in addition to Muc19, then these cells would maintain the mucous cell phenotype. In control experiments with sublingual glands from homozygous knockin mice (Figure 7), we find the expected absence of intense staining for acidic mucosubstances with Alcian Blue and only sparse anti-Muc19 immunoreactivity, probably due to Muc19-EGFP fusion proteins. Minor salivary mucous glands of homozygous knockin mice, on the other hand, maintain a large cytoplasm with strong Alcian Blue staining, but display extremely light anti-Muc19 immunoreactivity. Shown in Figure 7 is an example from salivary mucous glands within the soft palate. Similar results (not shown) are found in mucous glands of the posterior tongue and the hard palate. In addition, the electron microscopic appearance of mucous glands of the soft palate is similar in wild-type and homozygous knockin mice, with typical electron-lucent mucous granules filling the apical cytoplasm (not shown). Interestingly, mucous cells of bulbourethral glands of knockin mice are noticeably smaller in size than wild-type glands and have moderate to little cytoplasmic Alcian Blue staining or anti-Muc19 immunoreactivity.
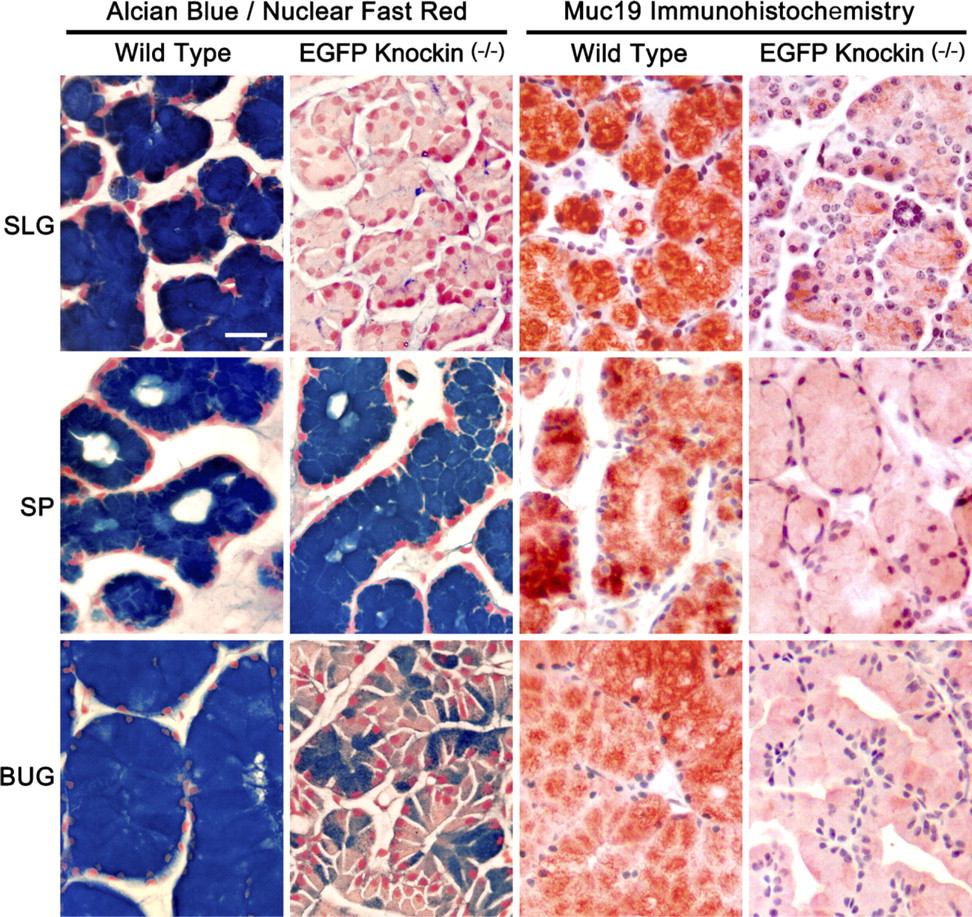
Comparisons of Alcian Blue staining with Nuclear Fast Red counterstain and anti-Muc19 immunohistochemistry in paraffin sections (7-μm) of mucous glands from wild-type and homozygous Muc19-EGFP knockin mice. SLG, sublingual gland; SP, soft palate salivary glands; BUG, bulbourethral gland. Bar = 20 μm.
Expression of Smgc in Neonatal and Adult Mice
As shown in Figure 8A, we find the restricted expression of
Discussion
From our combined analyses of adult mice, it is apparent that Muc19 is highly expressed in oral tissues containing minor salivary mucous glands and major sublingual glands. Submucosal glands of the tracheolarynx and bulbourethral glands represent the only other sites readily expressing Muc19. Although transcripts were detected in prostate, testis, ovaries/fallopian tubes, and uterus, the levels were too low to warrant further investigation. A human gene encoding MUC19 has been identified and the 3' end of
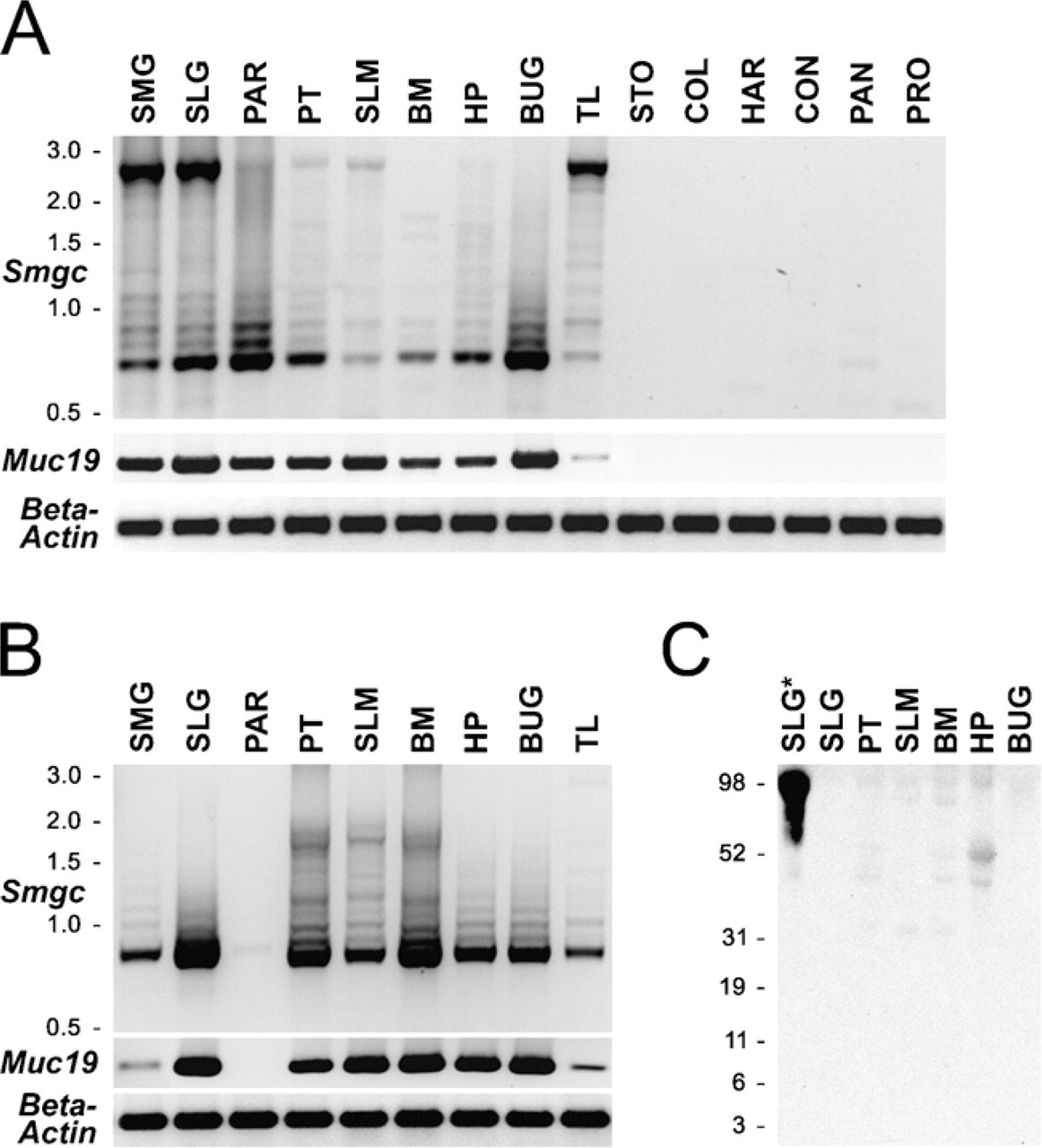
Comparative expression of
Recently, MUC19 proteins were immunolocalized to mucous cells of human middle ear epithelium (Kerschner et al. in press), a tissue we did not test. Yu et al. (2008) described the coexpression of MUC19 and MUC5AC in cells of the conjunctival epithelia and lacrimal glands of man, whereas MUC19, but not MUC5AC, is expressed in the corneal epithelium. We do not find Muc19 expression in conjunctiva or in lacrimal and Harderian glands, suggesting species differences and possibly a broader tissue distribution of MUC19 in man than in mice. In screening tissues of the human male urogenital tract, Russo et al. (2006) were unable to determine the source of MUC5B glycoproteins in seminal fluid. Transcripts for
In evaluating the contribution of Muc19 to saliva, Rousseau et al. (2008) detected Muc19 peptides by mass spectrometry in stimulated saliva from multiple mammalian species (i.e., rat, cow, pig, horse). Conversely, tryptic peptides from the predicted 5' end of human MUC19 were not detected in pooled unstimulated human saliva from six donors. These provocative results question whether MUC19 transcripts in salivary glands are indeed translated, although it is difficult to reconcile this possibility with the apparent translation of transcripts in other tissues. Alternatively, MUC19 may not contribute significantly to the unstimulated saliva fraction collected by simple drooling. It may instead preferentially coat the hard and soft oral tissues, or it may be released into saliva primarily upon stimulation, as supported by its presence in stimulated saliva samples from other mammalian species.
To test the mucin specificity of our antiserum raised against purified rat sublingual mucin, we initially utilized stomach, colon, Harderian glands, and epididymis as control tissues. Stomach expresses predominantly
The Muc19-EGFP mouse model system expresses the fusion protein in a pattern consistent with Muc19 transcripts and anti-Muc19 immunoreactivity. The fusion protein is localized to exocrine granules and may therefore serve as a marker, not only for Muc19 expression but also for exocrine secretion by mucous cells. Furthermore, the absence of Muc19 expression in sublingual glands of Muc19-EGFP knockin mice has a striking but predictable effect on mucous cell morphology and Alcian Blue staining, given the uniqueness of these cells in only expressing a single gel-forming mucin. In minor salivary glands, Muc19 expression is apparent throughout the entire population of mucous cells; yet these cells in Muc19-EGFP knockin mice continue to stain intensely with Alcian Blue. Because minor glands express significant amounts of
As presented earlier in this study, we hypothesized that SMGC may function in the cytodifferentiation of mucous and seromucous acinar cells. Our current results indicate that
Our collective results on the tissue-specific expression of Muc19 prompt speculation as to its function. The oral cavity is a major site of exposure to environmental pathogens and toxins, in addition to dietary components. A balance between host, microbial, dietary, and environmental factors is responsible for the selective establishment and maintenance of resident flora associated with the soft and hard tissues of the mouth. Subsequent invasion by more pathogenic microbial species is therefore limited by the combined actions of the resident flora and salivary constituents. Muc19 may interact with oral bacteria either to help select the resident oral flora and/or to exclude more pathogenic species. Muc19 may play a similar role in the tracheolarynx by entrapping bacteria for eventual clearance via the mucociliary escalator (Knowles and Boucher 2002). This region of the airways is probably exposed to oral bacteria through the accidental aspiration of saliva that can occur during eating or gagging. Muc19 may further function in the hydration and lubrication of the oral surfaces. Its presence in seminal fluid may also promote lubrication, although a role in bacterial interaction cannot be ruled out.
Footnotes
Acknowledgements
This study was supported by the National Institutes of Health through a RO1 grant (DE014730) to DJC.
The authors thank Sara Shah and Maya Yankova for excellent technical assistance.
