Abstract
Recent studies have identified members of the CLCA (chloride channels, calcium-activated) gene family as potential modulators of the cystic fibrosis (CF) phenotype, but differences between the human and murine CLCA genes and proteins may limit the use of murine CF models. Recently established pig models of CF are expected to mimic the human disease more closely than the available mouse models do. Here, we characterized the porcine CLCA gene locus, analyzed the expression pattern and protein processing of pCLCA1, and compared it to its human ortholog, hCLCA1. The porcine CLCA gene family is located on chromosome 4q25, with a broad synteny with the human and murine clca gene loci, except for a pig-specific gene duplication of pCLCA4. Using pCLCA1-specific antibodies, the protein was immunohistochemically localized in mucin-producing cells, including goblet cells and mucinous glands in the respiratory and alimentary tracts. Similar to hCLCA1, biochemical characterization of pCLCA1 identified a secreted soluble protein that could serve as an extracellular signaling molecule or functional constituent of the protective mucous layers. The results suggest that pCLCA1 shares essential characteristics of hCLCA1, supporting the pig model as a promising tool for studying the modulating role of pCLCA1 in the complex pathology of CF.
A
Although initially thought to represent bona fide transmembrane channel proteins (Ran and Benos 1991; Gruber et al. 1998a; Gruber et al. 2002), key CLCA proteins, including the human hCLCA1 and the murine mCLCA3, have recently been shown to be secreted, non-transmembrane proteins (Gibson et al. 2005; Mundhenk et al. 2006) that mediate a constitutively expressed endogenous chloride current when heterologously expressed in NIH/3T3, HEK293, or NCIH522 cells (Loewen et al. 2002a; Hamann et al. 2009). The electrophysiological properties of HEK293 cells heterologously transfected with cDNA of several CLCA homologs share key characteristics of the alternative chloride conductance seen in CF patients (Fuller and Benos 2002). Moreover, experimental overexpression of the murine mCLCA3 improves the intestinal pathology of CF mice and their survival (Young et al. 2007). Whether the secreted CLCA proteins directly or indirectly interact with either the CFTR protein or another as yet unidentified channel via a receptor-mediated pathway is unknown at this time. Thus, the mechanism of the suspected modulatory role of CLCA proteins in CF is far from being resolved.
Because hCLCA1 and its direct orthologs in mice and horses, mCLCA3 and eCLCA1, respectively, are abundantly expressed in goblet cell-derived mucin layers throughout the intestinal and respiratory tracts, they may serve as soluble activators of an as-yet-unidentified channel protein in the apical membrane of non-goblet cell enterocytes, respiratory cells, and other epithelial cells (Hoshino et al. 2002; Leverkoehne and Gruber 2002; Range et al. 2007). Other CLCA proteins, including mCLCA6, are coexpressed with the CFTR channel protein on the apical surface of mucous membrane epithelial cells and may directly interact with the CFTR pathway in the same cells (Bothe et al. 2008).
Several mouse models of CF have been used to explore the modulatory role of CLCA proteins in CF tissues (Brouillard et al. 2005; Leverkoehne et al. 2006). However, all mouse models available to date are regarded as less than optimal for studying the human CF pathology, owing to mouse-specific differences in their intestinal, respiratory, and pancreatic CF phenotypes (Scholte et al. 2004). The first pig models of CF have recently been introduced (Rogers et al. 2008a), and initial data indicate that their phenotype may resemble the human CF pathology more closely than the currently available mouse models do (Rogers et al. 2008b). Specifically, the macroscopic and microscopic anatomy, biochemistry, physiology, size, and genetics of pigs resemble those of humans more closely than do those of other potential models (Rogers et al. 2008c). This is particularly true for the CF-relevant tissues of the airways, intestine, and pancreas. The pig lung has already become an excellent model for studying human diseases such as bronchiolitis obliterans (Alho et al. 2007) and other diseases as well as specific therapeutic interventions (Budas et al. 2007).
Importantly, the microanatomy of the airways, including the distribution of submucosal glands, closely matches that of humans (Baskerville 1976). In addition, the longevity of pigs offers valuable opportunities for investigating bacterial and viral infections as well as other effects that become apparent over longer periods of time, similar to human CF patients. The similarity of porcine and human organs has already led to significant efforts to develop them as a source for xenotrans-plantation (Cooper et al. 2008).
For understanding the role of individual molecules in animal models and their transferability into other species, including man, however, species-specific differences are of considerable interest. Accordingly, several species-specific differences have been reported for CLCA gene family members and their direct orthologs in other species. For example, four CLCA genes exist in humans, whereas two additional consecutive gene duplications have resulted in six homologous CLCA genes in the mouse (Ritzka et al. 2003). Second, the human hCLCA3 is thought to represent a pseudogene, whereas its murine and bovine orthologs are not (Elble et al. 1997; Gruber and Pauli 1999). Third, distinct allelic variations that appear to be species-specific have been reported for the human hCLCA1 (Kamada et al. 2004) and the equine eCLCA1 (Anton et al. 2005). Fourth, the murine mCLCA6 protein is expressed in different cell types and in different subcellular structures than its direct human ortholog, hCLCA4 (Bothe et al. 2008). Moreover, the first and only porcine CLCA protein identified to date, pCLCA1 (Gaspar et al. 2000), displayed different functions and electrophysiological properties when compared with its human and murine orthologs (Loewen et al. 2002b). Thus, a detailed understanding of the porcine pCLCA1 and possible pig-specific variations in the CLCA gene family appears critical before their role as modulators of the CF phenotype can be studied and interpreted in the promising new pig models.
The aim of this study was to characterize the genomic organization of the porcine CLCA1 gene, its protein expression pattern, and its posttranslational protein modification and trafficking. The results are compared with the corresponding human and murine orthologs to disclose differences that could be relevant for the interpretation of porcine CF models.
Materials and Methods
Characterization of the pCLCA1 Genomic Structure and Other Porcine CLCA Genes
The organization of CLCA genes in mammals was evaluated by the GenBank DNA database (http://www.ncbi.nlm.nih.gov/). Porcine bacterial artificial chromosomes (BACs) notionally corresponding to the human CLCA locus were identified by comparison of the human genome with pig BAC end sequences. Subsequently, the candidate BACs were located on the porcine genome by the pig fingerprint contig map (www.ensembl.org). Four BAC clones covering the complete porcine CLCA locus, including the flanking genes ODF2L and SH3GLB1 (CH242–32G22, CH242–148M17, CH242–252F2, and CH242–483E7 with GenBank accession numbers CU695058, CU694822, CU695038, and CU469041, respectively), were obtained from CHORI BACPAC resources center (http://bacpac.chori.org/) and sequenced by the Wellcome Trust Sanger Institute (Hinxton, UK). Genes were roughly localized on the contig sequence by comparison of designated mRNA sequences from pig, human, cow, horse, mouse, and dog to the porcine BACs by BLAST search (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Predicted porcine mRNA sequences were derived from the alignment of porcine BACs and mRNA sequences from other species using BioEdit and taking into account the exon-intron structure in the different species as well as putative splicing sites in the BACs (Hall 1999). The corresponding protein sequences were deduced from the predicted mRNA sequences by in silico translation.
Phylogenetic trees of CLCA amino acid sequences from different species were generated by the PHYLIP software package (http://evolution.genetics.washington.edu/phylip.html), and nomenclature of the porcine CLCA genes was assigned by their correlation to the major branches of the trees.
Animals and Tissue Processing
Tissues from five male pigs (6 weeks old, EUROC × Pietrain), two female pigs (2 and 3 months old, mixed breed), and one male pig (7 months old, mixed breed) that had been euthanized for other reasons were included in this study. The following tissues were immersion fixed in 4% neutral-buffered formaldehyde or shock-frozen in liquid nitrogen after brief immersion in 2-methylbutane: nasal cavity, larynx, trachea, lung (three different locations: cranial left lobe, left main lobe, accessory lobe), tracheal bronchus, left principal bronchus, esophagus, stomach (glandular and non-glandular parts), duodenum, jejunum, ileum, cecum, colon, rectum, parotid salivary gland, pancreas, liver, gall bladder, kidney, urinary bladder, mandibular lymph node, spleen, heart, aorta, brain (cortex, cerebellum, medulla), eyes, skin (perineum, rooting disc, prepuce), testicle, epidymides, spermatic cord, uterus, and ovary.
Cloning and Sequencing of pCLCA1 cDNA
Total RNA was extracted from porcine rectum using the Trizol method (Invitrogen; Karlsruhe, Germany) and purified using the RNeasy Mini Kit (Qiagen; Hilden, Germany) according to the manufacturer's protocol, including a digestion with DNase I (Qiagen). Five hundred ng of total RNA was reverse transcribed (Omniscript; Qiagen) for 1 hr at 37C using random hexamer primers in the presence of 100 U recombinant RNase inhibitor (Invitrogen). PCR primers were designed (GenBank accession number NM_214148) to amplify the open reading frame (ORF) of pCLCA1 (upstream 5'-GA-TATGGGGTCATTTAGGAGTTCGCTGTTC-3'; downstream 5'-GCAGGATGGATGATTTGTTTATCT-CAGG-3'). PeqGOLD Pwo DNA polymerase (peqlab Biotechnologie GmbH; Erlangen, Germany) was used for PCR amplification (0.5 U per 25-μl reaction) and added after initial denaturation at 95C for 2 min. PCR conditions were 34 cycles at 95C for 40 sec, 61C for 40 sec, 72C for 2 min, with a time increment of 8 s per cycle, and a final extension at 72C for 10 min. PCR products were sequenced with the primers used for amplification.
T-addition to the Eco32l-digested and linearized pcDNA3.1 expression vector was performed using GoTaq Flexi DNA Polymerase (Promega; Mannheim, Germany) and 10 mM 2'-deoxythymidine 5'-triphosphate (Invitrogen) in the presence of 25 mM MgCl2 (Promega) for 30 min at 70C. A-addition to the pCLCA1 ORF was performed using the A-Addition Kit (Qiagen) and the product was ligated into the pcDNA3.1 expression vector using the QIAGEN PCR Cloning Plus Kit (Qiagen). PCR-induced sequence errors were excluded by additional sequencing of the amplification products from two further animals.
Transient Transfection of HEK293 Cells
HEK293 cells were grown at 37C in the presence of 5% CO2 in DMEM with 10% heat-inactivated fetal calf serum, 2% glutamate, and 1% penicillin/streptomycin. The cells were washed with prewarmed PBS between medium changes and transfected with the pCLCA1 ORF in pcDNA3.1 using Lipofectamine 2000 (Invitrogen). Control experiments included cells transfected with the pcDNA3.1 vector alone.
Computer-aided Sequence Analysis and Generation of Antibodies
Computational analysis of the published pCLCA1 amino acid sequence (GenBank accession number NP_999313) was accomplished using the SignalP 3.0 (Nielson et al. 1997), Kyte-Doolittle (Kyte and Doolittle 1982), SOSUI (Hirokawa et al. 1998), HMMtop (Tusnady and Simon 2001), TMPRED (Hofmann 1993), DAS (Cserzo et al. 1997), and PSORT II (Nakai and Horton 1999) software. Potential N-linked glycosylation sites were identified using the software NetNGlyc (http://www.cbs.dtu.dk/services/NetNGlyc). Three oligopeptides were synthesized based on immunogenicity prediction, two located in the N-terminal cleavage product of pCLCA1 (p1-N-1, corresponding to aa 250–264, CKEKNHNKEAPNDQN (Loewen et al. 2004) and p1-N-2, corresponding to aa 394–407, TVIKKKY-PTDGSEI) and one located in the predicted carboxy-terminal cleavage product (p1-C-1, corresponding to aa 775–788, WTAPGDDYDHGRAD). Oligopeptides were coupled to keyhole limpet hemocyanin and used for standard immunization of two rabbits each. Pre-immune sera were collected before immunization and used as controls in the immunodetection experiments. The six antisera obtained were designated p1-N-1a, p1-N-1b, p1-N-2a, p1-N-2b, p1-C-1a, and p1-C-1b. After initial testing of the antisera by immunoblot analysis of transfected cells, antisera p1-N-1a and p1-N-1b were combined and affinity immunopurified using the peptide used for immunization coupled to a CNBr-Sepharose column. The resulting immunopurified antibodies were designated p1-N-1ab-p.
Immunoblot Analyses and Immunoprecipitation
Forty-eight hr after transfection, HEK293 cells were washed three times with PBS, followed by incubation in DMEM supplemented with 2% glutamate for 6 hr. Medium was removed, and cells were lyzed on ice in 100 μl standard lysis buffer (25 mM Tris-HCl, pH 8.0, 50 mM NaCl, 0.5% deoxycholic acid, 0.5% Triton X-100) and supplemented with 1% protease inhibitors (1 mM PMSF, 1 mg/ml pepstatin, 1 mg/ml aprotinin, 5 mg/ml antipain, 5 mg/ml leupeptin, 100 mg/ml trypsin-chymotrypsin inhibitor). To remove cellular fragments, medium samples were spun at 1000 × g for 5 min at 4C, the pellet discarded, and the cell culture medium spun at 10,000 × g for 15 min at 4C. The second supernatant was ethanol precipitated overnight at −20C, and pellets were resuspended in 100 μl of standard lysis buffer. All tissue samples were homogenized in 1 ml of standard lysis buffer with 1% protease inhibitors using a Precellys 24 homogenizer (peqlab Biotechnologie GmbH). Thirty μg of tissue lysates was subjected to immunoblot analyses.
Medium and lysate samples were boiled in standard 3-fold SDS-PAGE Laemmli buffer and separated by SDS-PAGE. Subsequent electroblotting onto nitrocellulose membranes was followed by blocking the membranes in TBS containing 0.2% Tween-20 and 5% non-fat milk for 90 min. The membranes were probed at 4C overnight with anti-pCLCA1 antibodies p1-N-1ab-p or p1-N-2a, the preimmune sera, or an irrelevant immuno-purified antibody (anti-horse ecr10). Specific and control antibodies were diluted 1:1000. Membranes were incubated with secondary horseradish peroxidase-conjugated anti-rabbit IgG for 1 hr, and protein labeling was developed using enhanced chemiluminescence (Thermo Fisher Scientific; Rockford, IL).
Immunoprecipitation of pCLCA1-transfected HEK293 cells was performed as described (Mundhenk et al. 2006). In brief, immunoprecipitation was performed using either the carboxy-terminal antibody or the N-terminal antibodies diluted 1:1000. Medium or cell lysates were incubated for 24 hr at 4C with the respective antibody and then with 20 μl protein A-Sepharose beads (Sigma-Aldrich; Munich, Germany). The immunoprecipitates were analyzed by immunoblotting with the carboxy-terminal antibody or the N-terminal antibodies. Unfortunately, both antibodies failed to detect any specific bandinthe immunoblot after immunoprecipitation (data not shown).
Analysis of pCLCA1 mRNA Tissue Expression Pattern
Total RNA from organs listed above was isolated using the Trizol method (Invitrogen) and purified using the RNeasy Mini Kit (Qiagen), including digestion with DNase I (Qiagen). Five hundred ng RNA was reverse transcribed as described above, and the cDNA served as template in the following PCR reactions: PCR amplification using the DreamTaq DNA Polymerase (Fermentas; St. Leon-Rot, Germany) included 34 cycles at 95C for 2 min, 95C for 30 sec, 61C for 30 sec, and 72C for 3 min, as well as a final extension step at 72C for 10 min. The primers used here were the same as for the cloning of pCLCA1, obtaining the pCLCA1 ORF (see above). To control for mRNA quality and efficacy of reverse transcription, a 68-bp product of the housekeeping gene EF-1a was RT-PCR amplified from each sample using primers 5'-CAAAAACGACCCACC-AATGG-3' (sense) and 5'-GGCCTGGATGGTTCAG-GATA-3' (antisense; Gruber and Levine 1997).
Immunohistochemistry
Tissues were either fixed in 4% neutral-buffered formaldehyde for 24 hr and embedded in paraffin or shock-frozen in liquid nitrogen and stored at −80C. Paraffin-embedded tissues were cut at 3-μm thickness, mounted on adhesive glass slides, and dewaxed in xylene, followed by rehydration in descending graded ethanol. Endogenous peroxidase was blocked by incubating the slides with 0.5% H2O2 in methanol for 30 min at room temperature. Antigen retrieval was performed using 15 min microwave heating (600 W) in 10 mM citric acid, pH 6.0, containing 0.05% Triton X-100. Frozen tissues were cut at 5-μm thickness, mounted on adhesive glass slides, and stored at −80C until further use. The cryostat sections were fixed in acetone for 10 min and dried for 20 min. Avidin-biotin blocking of the cryostat sections was performed according to the manufacturer's protocol (Dako North America, Inc., Carpinteria, CA). The slides were washed in PBS containing 0.05% Triton X-100 and blocked with PBS containing 2% BSA and 20% normal goat serum for 30 min. Anti-pCLCA1 antibodies, preimmune sera, or an irrelevant purified antibody (ecr10) were incubated overnight at 4C. As a further control, slides were incubated with antibodies that had been preincubated with 10 μg/ml of the corresponding pep-tides for 30 min. Antibody dilutions ranged from 1:500 to 1:40,000, with optimal results for 1:40,000 for both the purified and the non-purified antibodies. Sections were washed repeatedly in PBS containing 0.05% Triton X-100 and incubated for 1 hr with biotinylated secondary goat anti-rabbit antibody diluted 1:200. Color development was performed by incubating the slides with freshly prepared avidin-biotin-peroxidase complex (ABC) solution (Vectastain Elite ABC Kit; Vector Laboratories, Inc., Burlingame, CA), followed by repeated washes and exposure to diaminobenzidine tetrahydrochloride (Merck; Darmstadt, Germany). The sections were counterstained with hematoxylin, dehydrated in ascending graded ethanol, cleared in xylene, and coverslipped. Consecutive sections from each tissue sample were stained with hematoxylin and eosin for histological examination and with the periodic acid-Schiff (PAS) reaction to visualize mucins and mucin-producing cells. For immunofluorescent analyses, slides were incubated with the specific anti-pCLCA1 antibodies diluted 1:500 as described above, followed by incubation with an FITC-conjugated secondary anti-rabbit antibody (Dianova; Hamburg, Germany) diluted 1:200 in PBS for 1 hr.
Endoglycosidase Treatment
Cell lysates or cell culture medium was incubated with endo H or PNGase F (New England Biolabs; Frankfurt, Germany) prior to SDS-PAGE analysis with antibody p1-N-1ab-p. Briefly, an aliquot of each sample was incubated with 50 U/ml endo H or 25 U/ml PNGase F, respectively, for 1 hr at 37C. One sample was left untreated as negative control in each experiment.
Results
Structure of the Porcine CLCA Gene Locus
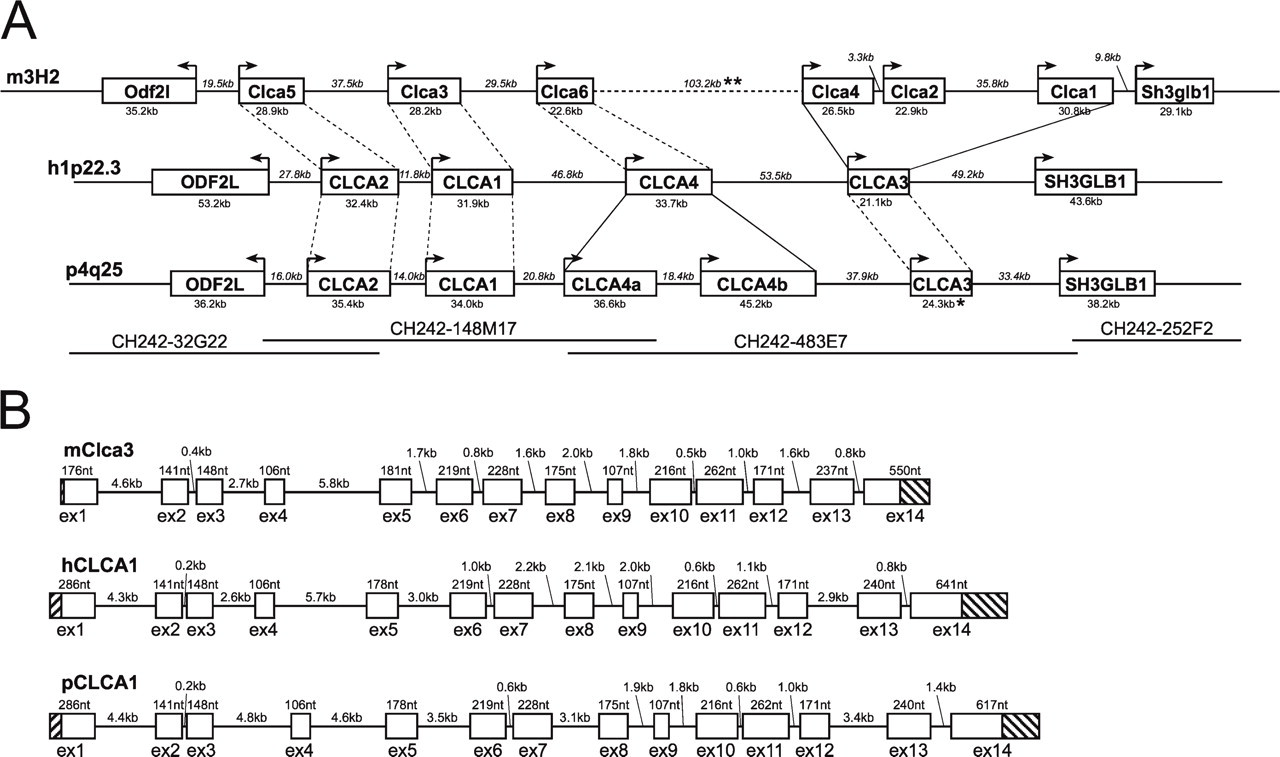
In CF patients, hCLCA1 has been described as a potential modulator of CF severity. A porcine ortholog of this gene has been described on the mRNA level (GenBank accession number NM_214148) and first functional analyses have been performed (Loewen et al. 2002a,b). However, inasmuch as both silencing and duplications of CLCA genes have been described in several species, it is unclear whether additional CLCA1 genes may exist in the porcine genome and whether the gene may be silenced in the pig. To test for the possibility of CLCA1 gene duplication or other variations in the porcine genome, we aimed at identifying all CLCA homologous genes in the pig. In mammals, up to six highly homologous CLCA genes have been described. Evaluating the localization of the published genes in human, chimpanzee, cow, horse, dog, mouse, and rat, we found a conserved and dense structure of a single locus containing all CLCA genes described and two flanking genes, “outer dense fiber of sperm tails 2-like” (ODF2L) and “SH3-domain GRB2-like endophilin B1” (SH3GLB1), in all species examined (data not shown). The porcine CLCA locus was identified by comparison of the human CLCA on chromosome 1p22.3 to porcine BAC end sequences that have been mapped to the porcine chromosome 4q25. Sequencing of the BACs CH242–32G22, CH242–148M17, CH242–483E7, and CH242–252F2 revealed a region of 315 kb between the flanking genes ODF2L and SH3GLB1. Comparison with published CLCA sequences from different species revealed the presence of five CLCA genes within this region. Exon 14 is missing from pCLCA3, which also contains numerous stop codons in the ORF and therefore probably represents a pseudogene (Figure 1A). Comparison of the exon-intron structure of CLCA1 genes in mouse, human, and pig showed a common structure with 14 exons. Different exon lengths were only seen in untranslated regions, as well as in exons 5, 13, and, more pronounced, exon 14 (Figure 1B). Because different exon lengths in the coding region represented codon triplets, the ORF of porcine CLCA1 is not affected by frameshift mutations or stop codons and encodes 915 amino acids. Thus, the predicted CLCA1 from the porcine CLCA locus encodes a potentially functional protein.
Phylogenetic studies and analysis of genetic distance distribution of the CLCA protein alignment definitely identified four branches of mammalian CLCA, with only one porcine gene classified to the human CLCA1 and murine CLCA3 (Figure 2; Klymiuk et al. 2008). Thus, there is clear evidence that only one CLCA1 copy exists in the porcine CLCA locus. This gene has a size of 34 kb and transcribes a predicted mRNA of 3094 nt in length.
Bioinformatics and Generation of Antibodies
To test the hypothesis that pCLCA1 is a fully secreted protein, similar to the human and murine orthologs hCLCA1 and mCLCA3 (Gibson et al. 2005; Mundhenk et al. 2006), we performed computational analyses on the pCLCA1 protein structure, including identification of hydrophobic domains and possible transmembrane regions. The Kyte-Doolittle, SOSUI, HMMtop, DAS, and PSORT II algorithms failed to predict any hydrophobic domains that could account for a transmembrane region, other than the canonical cleavable signal sequence at the N terminus of the protein (not shown). In contrast, TMPRED software suggested one potential transmembrane domain between aa 374 and 396. The NetNGlyc software for the prediction of N-linked glycosylation identified six consensus glycosylation sites at aa 503, 772, 806, 812, 838, and 893. Considering the conserved posttranslational cleavage site around aa 680 that has been found for all CLCA proteins characterized to date (Patel et al. 2008), only the predicted glycosylation site at aa 503 would be located within the N-terminal cleavage product, whereas the remaining five sites would be located within the carboxyterminal fragment (Figure 3).
According to the established model of universal CLCA protein processing (Patel et al. 2008), posttranslational cleavage of a primary pCLCA1 translation product into a larger N-terminal fragment and a smaller carboxy-terminal fragment was anticipated. Based on this model, three separate antibodies were generated and partially epitope-immunopurified, two directed against epitopes located in the predicted N-terminal fragment, designated p1-N-1a and p1-N-2a, and one directed against an epitope within the predicted carboxy-terminal fragment, designated p1-C-1a (Figure 3). Both antibodies directed against the predicted N-terminal cleavage product recognized an ~75-kDa and an ~120-kDa protein in immunoblots of lysates from pCLCA1-transfected HEK293 cells and a single 75-kDa protein in lysates from porcine trachea tissue (Figure 4). These observations are consistent with the models and protein sizes established for virtually all other CLCA proteins analyzed to date (Gruber et al. 2002), including a primary translation product of ~120 kDa that is detectable only in transfected and overexpressing cell culture systems and an N-terminal cleavage product of ~75 kDa that is the only detectable protein species in ex vivo tissue lysates (Leverkoehne and Gruber 2002; Anton et al. 2005; Bothe et al. 2008). In contrast, antibody p1-C-1a raised against the predicted carboxy-terminal fragment failed to detect any specific protein in transfected cells or tissue lysates (data not shown), although it yielded staining patterns identical to those of antibodies p1-N-1a and p1-N-2a in the immunohistochemical experiments (see below). The sequence of the pCLCA1 clone used for transfection of the cells was identical to the sequence obtained from the respective BAC clones, but was two amino acids shorter than the published sequence (GenBank accession number NP_999313), which might be due to allelic variations, as described for other CLCA homologs (Kamada et al. 2004; Anton et al. 2005).
(
Tissue and Cellular Distribution Patterns of pCLCA1
Previous studies suggested that the human hCLCA1 and its murine and equine orthologs, mCLCA3 alias gob-5 and eCLCA1, respectively, are similarly expressed by virtually all mucin-producing cells throughout the body (Gruber et al. 1998b; Leverkoehne and Gruber 2002; Anton et al. 2005). To explore whether this expression pattern is also conserved in the pig, we analyzed the distribution of pCLCA1 mRNA and protein in virtually all porcine organ systems by RT-PCR, immunoblot, and immunohistochemistry, respectively.
High expression of pCLCA1 mRNA was detected in the upper respiratory tract, including the nasal mucosa, trachea, and bronchi, with lower expression levels in the lung (Figure 5A). Strong expression was also found throughout the intestinal tract, with strongest expression in the colon. Prominent expression was also detected in the esophagus, but only a very faint PCR product was seen for the glandular part of the stomach. Conjunctival mucous membranes contained significant amounts of pCLCA1 mRNA, but all other structures of the eyes were negative. pCLCA1 mRNA was found neither in the heart, liver, pancreas, or skin (Figure 5A), nor in salivary gland, kidney, or gall bladder, or in any other organ tested (not shown).
Immunoblot analyses using the immunopurified antibody p1-N-1ab-p and the antibody p1-N-2a and tissue lysates as templates detected the 75-kDa pCLCA1 protein in the respiratory tract, again with stronger expression in the organs of the upper respiratory tract, throughout the intestinal tract, and in the glandular part of the stomach (Figure 5B), as well as in the conjunctival membranes (not shown). In contrast to the negative RT-PCR results, pCLCA1 protein was detectable at low levels in the pancreas (Figure 5B) and in the parotid gland (not shown). These differences between RT-PCR results and immunoblot results might be due to different numbers of excretory ducts and their goblet cells in whole-tissue lysates. No other organs contained pCLCA1 protein levels detectable by immunoblot analyses.
Immunohistochemical stainings and microscopical analyses of formalin-fixed tissue sections yielded virtually identical results for all animals included in this study. Moreover, all three antibodies generated either against the predicted N-terminal cleavage products (p1-N-1a and p1-N-2a) or against the predicted carboxy-terminal cleavage product (p1-C-1a) resulted in identical staining patterns in all tissues analyzed, indicating strict colocalization of both cleavage products.
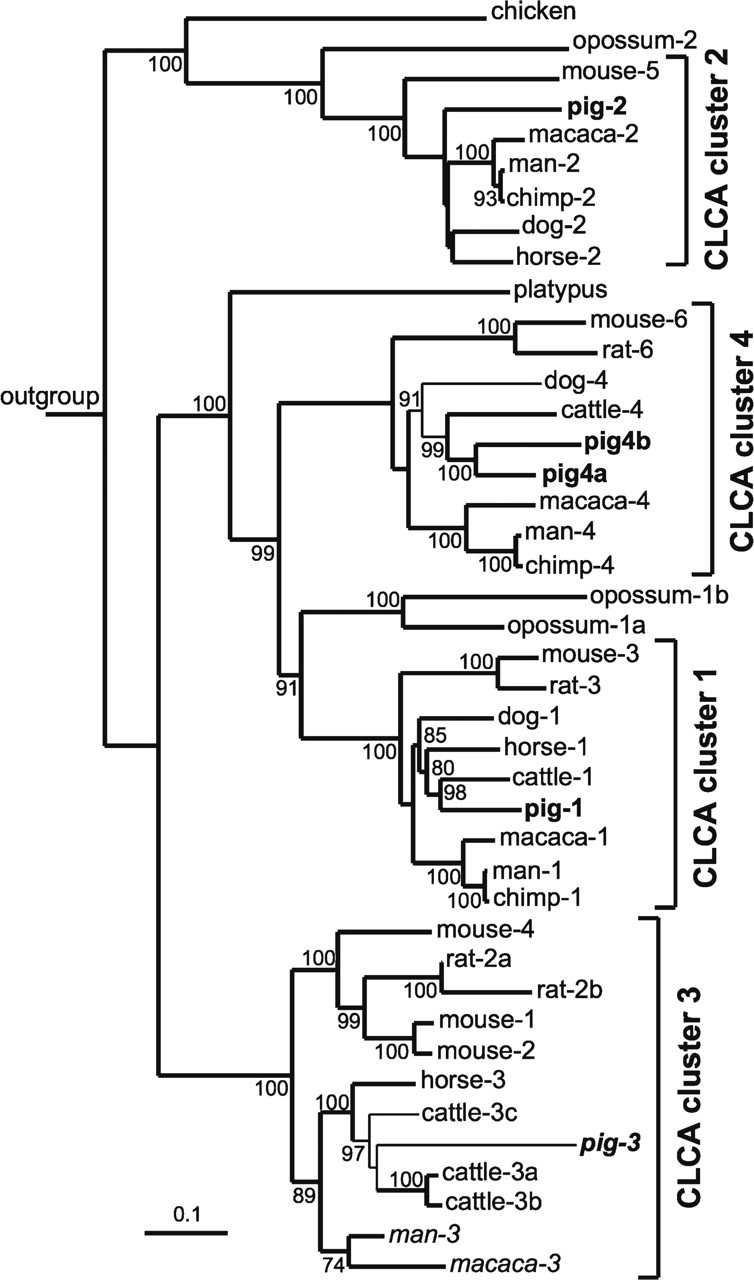
Phylogenetic tree of CLCA genes in different species. The tree was based on a single genetic distance tree, with the scale bar indicated. Branches occurring also in a most-parsimony tree are shown in bold, and branch nodes higher than 60% yielded from the consensus of 100 genetic distance trees are indicated. The alignment was based on amino acid sequences from the GenBank DNA database and represented CLCA genes in cow (XM_001252428, BTU36445, NM_181018, AF001263, NC_007301); chimpanzee (XM_ 001143250, XM_001142936, XM_524757); dog (XM_547299, XM_ 850237, XM_850235); human (NM_001285, NM_006536, NM_004921, NM_012128); horse (NM_001081799, XM_001495998, XM_001496218); chicken (XM_001234329); macaca (NM_001032912, XM_001109351, XM_001105446, XM_001109489); mouse (NM_ 009899, NM_030601, NM_017474, NM_139148, NM_178697, NM_207208); rat (NM_001013202, XM_217689, XM_001079170, NM_201419); opossum (XM_001365327, XM_001365401, XM_001362370); and platypus (XM_001511945), as well as the porcine sequences obtained from the analysis of the CLCA locus described above (in bold). The tree was outgrouped by BC091051 (Clca1 from frog), and the nomenclature of not-yet-classified sequences was based on the definition of human CLCA genes, whereas aberrant denotations in rodents were retained as commonly used. Amino acid sequences that do not obtain an open reading frame are shown in italics. CLCA sequences clustered by genetic distance distribution analysis are embraced by brackets and named clca cluster 1 to 4 according to the nomenclature of human CLCA family members. Based on the algorithm used (Klymiuk et al. 2008), the chicken, opossum, and platypus homologs were too distantly related to be included in the clusters.
Staining for the pCLCA1 protein was observed virtually exclusively in cytoplasmic mucin granules of goblet cells (Figure 6A) and other mucin-producing cells throughout the body, and was confirmed by parallel staining of serial tissue sections for mucins with the PAS reaction (Figure 6I, and inset in Figure 6B). In the respiratory tract, intraepithelial goblet cells of the nasal cavity, trachea, and bronchi, with a diameter of down to ~1 mm, were strongly labeled, with additional strong signals for the submucosal glands of the nose, trachea, and bronchi (Figure 6B-6D). Alveolar epithelial cells and other cells in the lungs were unstained. In the glandular part of the stomach, only a few of the deep cardiac glands stained positive, whereas parietal and chief cells were negative (Figure 6F). In the intestine, virtually all goblet cells were stained in the small and large intestinal crypts, with stronger staining in the more-apical crypt segments. Similar staining intensities were observed for the fewer goblet cells in the small intestinal villi (Figures 6G and 6J). In addition, the thin mucin layer covering the mucosal surface was also labeled in the respiratory and intestinal tracts in areas where it was not lost during sample processing (Figure 6H). Submucosal glands of the pharynx and esophagus also had strong expression of pCLCA1 (Figure 6E). Furthermore, the gall bladder, large excretory ducts of the parotid salivary glands, pancreas, and common bile duct had strongly stained intraepithelial goblet cells (Figures 6A and 6L). Mucin-producing and thus PAS-positive goblet cells of the conjunctival membranes, as well as the excretory ducts of the lacrimal gland, also had strong pCLCA1 expression, whereas all glands of the skin and hair follicles were negative. Interestingly, not all mucin-producing cells stained for the pCLCA1 protein, despite strong positivity in the PAS reaction, including Brunner's glands of the duodenum and mucous parts of the lacrimal gland (not shown).
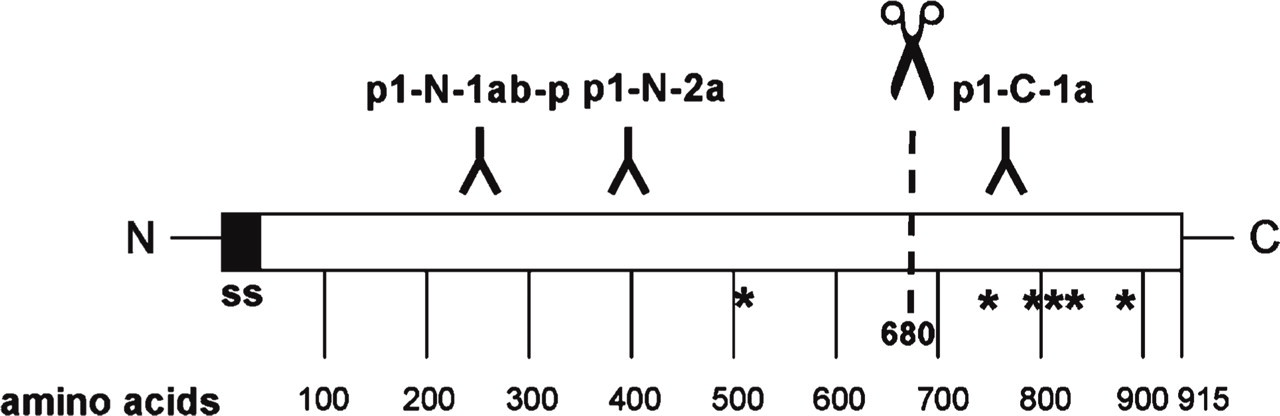
Predicted protein structure of pCLCA1 and location of peptides used for generation of antibodies. A cleavable signal sequence (ss), but no potential transmembrane domains, was predicted by computational sequence analyses (Kyte-Doolittle, SOSUI, HMMtop, DAS, and PSORT II). NetNGlyc predicted six potential asparagine-linked glycosylation sites (asterisks). According to the general model of CLCA protein processing, cleavage of the primary translation product is predicted around aa 680 (Patel et al. 2008). Two antibodies were generated against peptides from within the N-terminal segment and one antibody against a peptide corresponding to a sequence located in the predicted carboxy-terminal cleavage product.
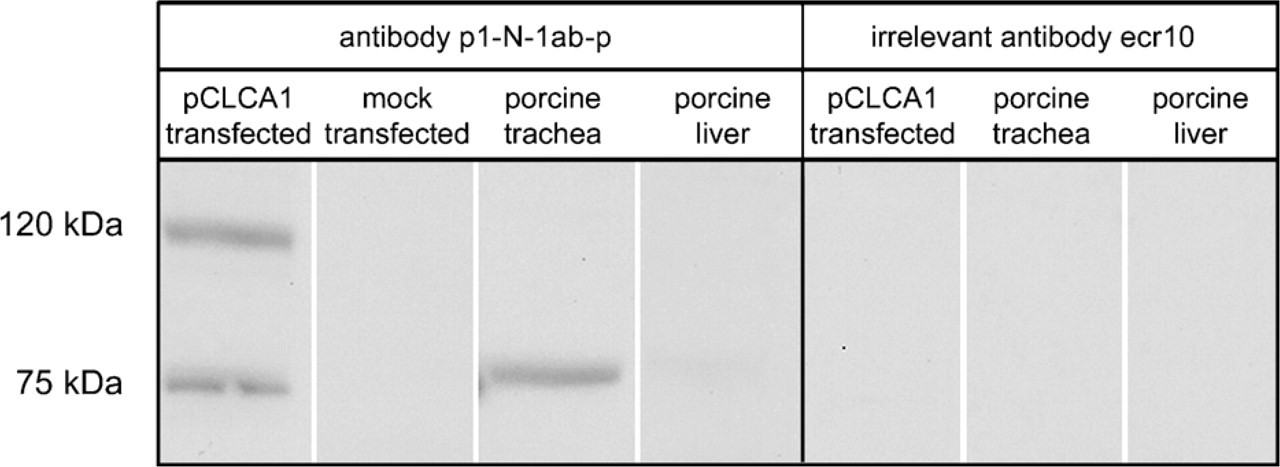
Detection of pCLCA1 in protein lysates from transfected cells and tissue lysates. Immunoblot analyses using anti N-terminal antibody p1-N-1ab-p (left panel) on lysates from pCLCA1-transfected HEK293 cells and tissue lysates from porcine trachea detected an ~75-kDa protein, consistent with the size of the predicted N-terminal cleavage product. The primary translation product of ~120 kDa in size was observed in overexpressing HEK293 cells only. No pCLCA1 protein was detected in the liver. Vector-alone-transfected HEK293 cells served as negative controls (mock transfected). Antibody p1-C-1a raised against an epitope within the predicted carboxy-terminal cleavage product failed to identify any proteins on the immunoblots (not shown), similar to an antibody directed against an equine CLCA protein (Anton et al. 2005), which was used as negative control (right panel).
Virtually identical staining patterns were observed when anti-pCLCA1 antibody binding on the tissues was visualized using immunofluorescence instead of enzymatic color development (not shown). The presence of pCLCA1 protein in mucin layers on mucosal membranes, however, was consistently shown with each of the three antibodies when the protein was detected on cryostat sections instead of paraffin-embedded tissues (Figure 6M), probably due to better mucin layer conservation in cryostat sections, in contrast to the loss of mucin layers during paraffin embedding of formalin-fixed tissues. Immunohistochemistry on cryostat sections of lung, large bronchi, esophagus, the glandular part of the stomach, small and large intestine, parotid gland, pancreas, and gall bladder identified strong expression of pCLCA1 protein in the mucus of all organs tested. The section of the colon is shown in Figure 6M. Staining with preimmune sera or the irrelevant antibody ecr10 as primary antibodies, or antibody p1-N-1ab-p that had been preincubated with the peptide used for immunization on paraffin-embedded sections or cryosections, failed to yield any staining in all tissues analyzed, regardless of the detection system (Figure 6N).
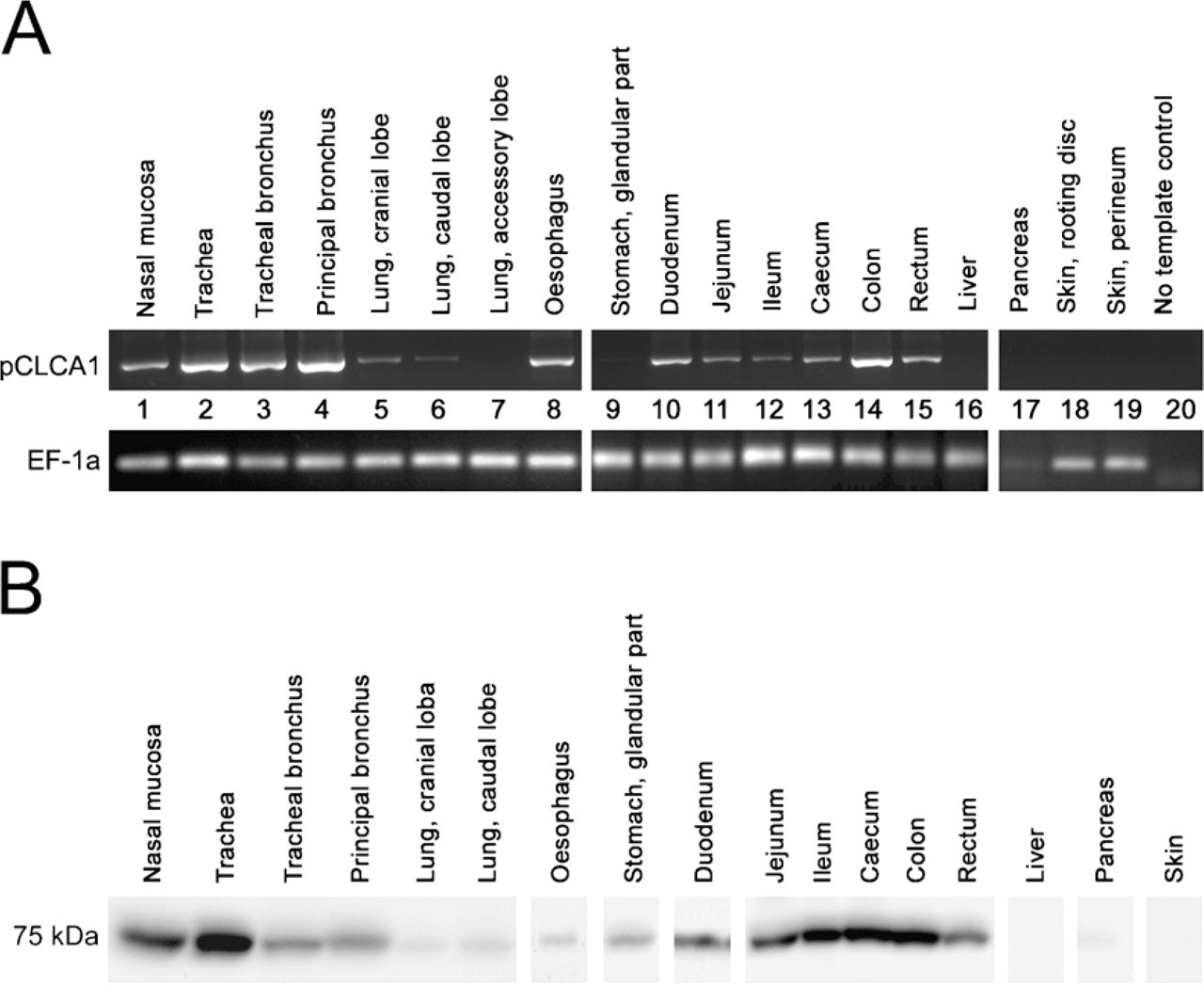
Tissue expression patterns of pCLCA1 mRNA and protein. (
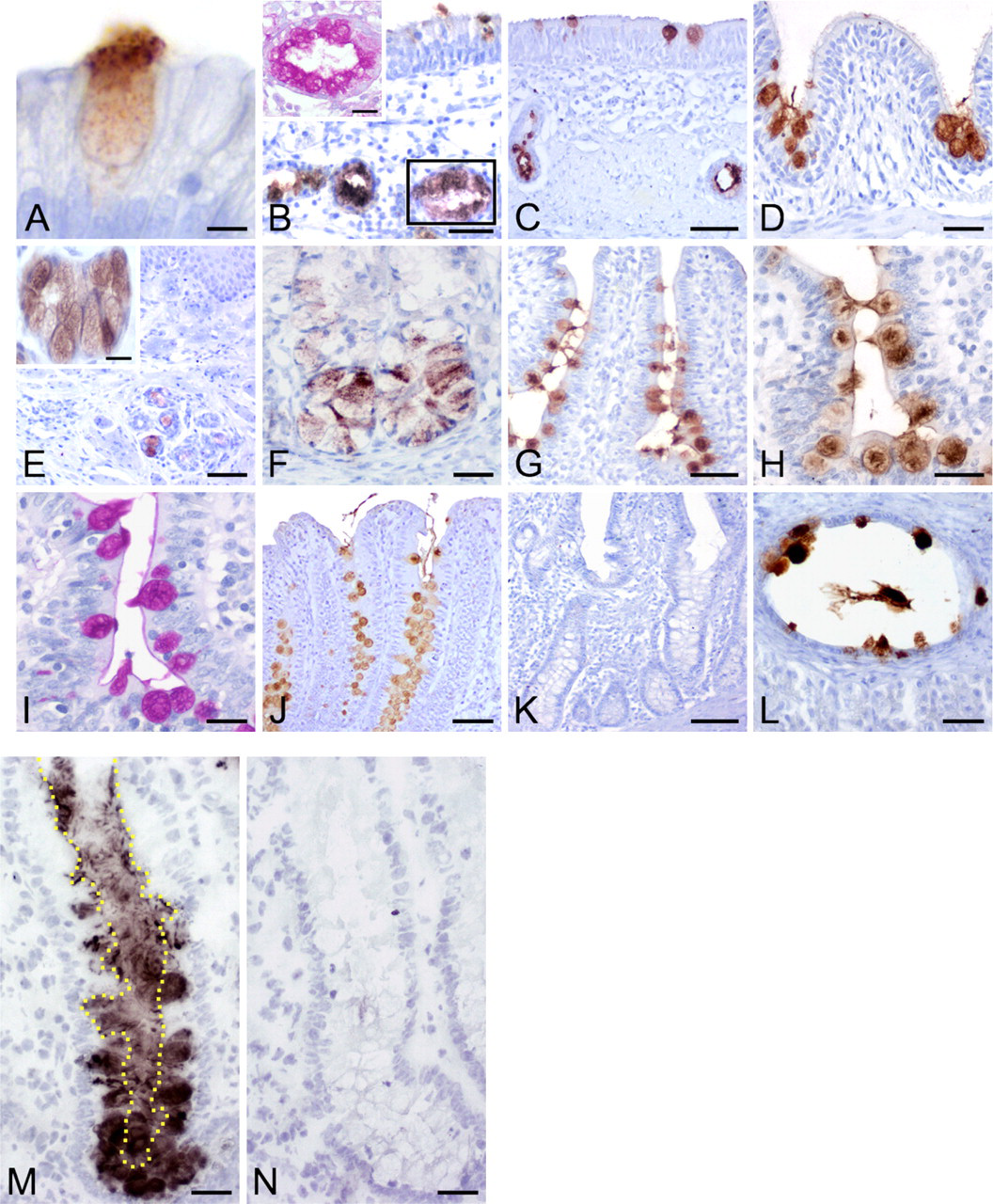
The pCLCA1 protein is expressed in goblet cell mucin granules. The pCLCA1 protein was detected immunohistochemically in cytoplasmic granules of goblet cells (
Protein Processing and Glycosylation Patterns
To challenge the hypothesis that pCLCA1 represents a soluble, secreted protein, the cell lysate and cell culture medium of transiently pCLCA1-transfected HEK293 cells were treated with endoglycosidases endo H and PNGase F, respectively, and immunoblotted using antibody p1-N-1ab-p (Figure 7). Both the primary translation product of ~120 kDa in size and the N-terminal cleavage product of ~75 kDa in size were detected in the cell lysate and were sensitive to endo H and PNGase F treatment, resulting in reductions in size by ~10 kDa and 3 kDa, respectively. Thus, both the primary translation product and the N-terminal cleavage products in the cell lysates were mannose-rich, immature glycosylation forms that have passed the endoplasmic reticulum but not yet the Golgi. In contrast, the two protein variants of ~130 kDa and ~80 kDa found in the cell culture medium were resistant to endo H but sensitive to PNGase F treatment, as indicated by a reduction in size from 130 kDa to 120 kDa and from 80 kDa to ~77 kDa, respectively. Thus, both the fully secreted 130-kDa and 80-kDa proteins were complex-glycosylated proteins processed in the Golgi (Figure 7, right panels).
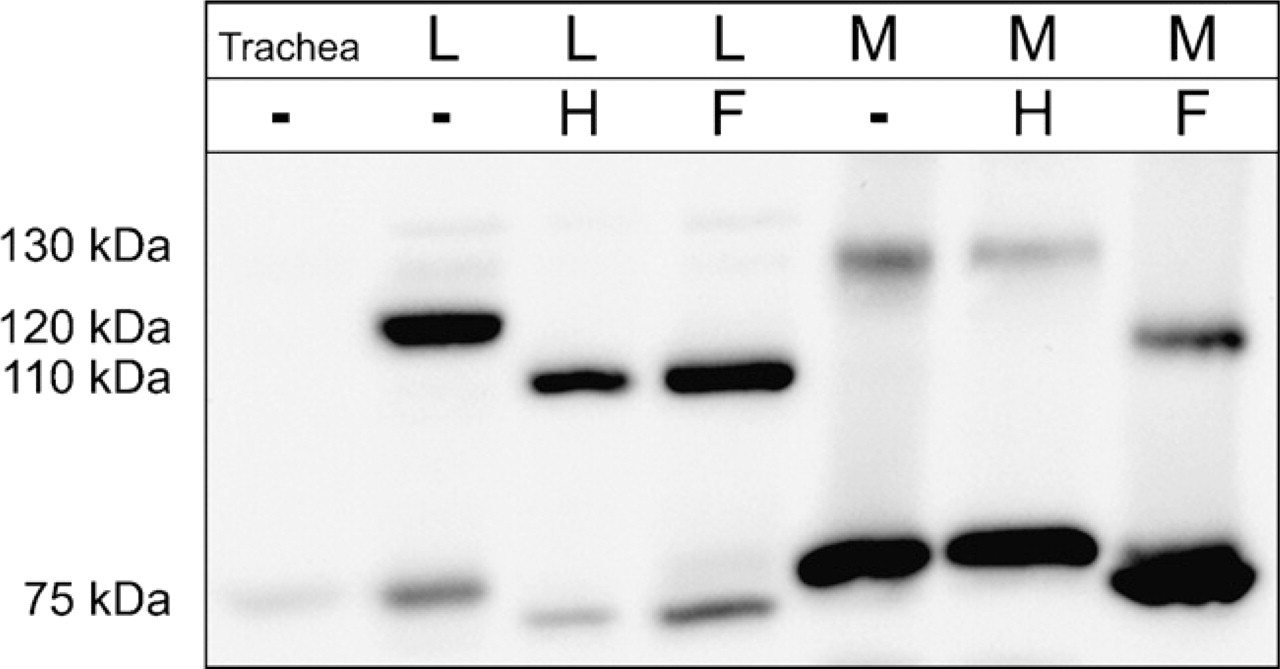
Cleavage and glycosylation of the pCLCA1 protein. Immunoblotting of the cell lysate (L) and culture medium (M) of transiently pCLCA1-transfected HEK293 cells after treatment with endo H (H) or PNGase F (F) identified the 120-kDa and the 75-kDa pCLCA1 proteins in the cell lysate as mannose-rich glycosylated forms. In contrast, the 130-kDa and 80-kDa products detected in the medium were only sensitive to treatment with PNGase F, as shown by reduction in size to ~120 kDa and 77 kDa, respectively, suggesting complex glycosylated proteins that had been processed in the Golgi. Overall expression in the trachea was weaker than in experimentally overexpressing cultured HEK293 cells.
Discussion
Despite their vast potential for the study of disease mechanisms and novel therapeutic interventions, the value of animal models of human diseases may be limited, owing to interspecies differences in anatomy, biochemical pathways, or genetic endowment. All mouse models of CF established to date, for example, display considerable differences from the human disease in terms of severity of the lesions, affected organs, and cellular pathways of anion conductance (Grubb and Boucher 1999; Scholte et al. 2004). The reasons for this are still poorly understood, but species-specific differences in genetic factors that modulate the disease are most probably involved (Collaco and Cutting 2008). Recent work has identified several such differences between human and murine CLCA genes and proteins that are thought to contribute to the different phenotypes of CF in human and mouse models (Ritzka et al. 2003; Bothe et al. 2008). These differences even give reason to speculate that they may contribute to the limited reproducibility of the human CF pathology in mice. First characterizations of a novel pig model of CF have supported expectations as to an improved model for mimicking the human functional and pathophysiological characteristics of the disease (Rogers et al. 2008b). Given the differences between several human and murine CLCA genes and proteins, however, we set out to characterize the porcine pCLCA1 and compare it with its human and murine orthologs in terms of gene structure, protein expression pattern, and cellular protein processing.
Previous work has identified considerable differences in the overall genomic CLCA locus as well as individual gene structures between humans and mice (Ritzka et al. 2003). Our detailed analyses of the genomic organization of porcine CLCA genes failed to identify any significant differences between the pCLCA1 gene and the genes encoding its human and murine orthologs, hCLCA1 and mCLCA3 alias gob-5 (Gruber et al. 1998a; Ritzka et al. 2003). The lack of two amino acids in both the cloned cDNA sequence and the genomic BAC clones at positions 525 and 526 when compared with the published mRNA sequence of pCLCA1 (GenBank accession number NM_214148; Loewen et al. 2004) is probably due to allelic variations that have previously been described for hCLCA1 and eCLCA1 in humans and horses, respectively (Kamada et al. 2004; Anton et al. 2005). Whether these allelic variations are accompanied by functional differences, as has been suggested for the allelic variants of hCLCA1 and hCLCA4, remains to be established (Ritzka et al. 2004).
Of note, the pCLCA1 gene and all other CLCA homologous genes in the pig are located in a region flanked by genes SH3GLB1 and ODF2L that contains no other known genes. Interestingly, the porcine pCLCA3 gene probably represents a pseudogene, similar to its human ortholog, hCLCA3 (Gruber and Pauli 1999), but unlike the corresponding murine orthologs mCLCA1, −2, and −4 (Leverkoehne et al. 2002). Although the overall genomic location and organization of the entire porcine CLCA gene locus was found to be similar to that of humans, we identified a gene duplication for the porcine ortholog of the human hCLCA4 and the murine mCLCA6 that appears to be unique to the pig. On the basis of sequence analyses, both pCLCA4 variants seem to code for fully expressed proteins of sizes similar to virtually all other CLCA proteins characterized to date (Patel et al. 2008). Whether this additional gene duplication in the pig results in a more complex CLCA protein function or regulation in the intestine, where hCLCA4 and the murine ortholog mCLCA6 are primarily expressed (Bothe et al. 2008), or whether the pCLCA4b variant may even be expressed in other cell types remains to be shown.
In addition to differences in genomic structure, certain orthologous CLCA proteins appear to be expressed in different cell types and subcellular structures between humans and mice, including hCLCA4 and mCLCA6 (Bothe et al. 2008). A previous study had localized the pCLCA1 mRNA by in situ hybridization to the crypt and villus epithelia of porcine ileum and to the surface epithelium and submucosal glands of the trachea (Gaspar et al. 2000). Our immunohistochemical detection of the pCLCA1 protein using specific antibodies included virtually all major organ systems. In addition to the cell types that had tested positive by in situ hybridization (Gaspar et al. 2000), we identified a much wider expression pattern in virtually all mucin-producing cells, including goblet cells. Importantly, the expressing cell types were found to be virtually identical to the cells that express the direct human, murine, and equine orthologs hCLCA1, mCLCA3, and eCLCA1 (Gruber et al. 1998b; Leverkoehne and Gruber 2002; Anton et al. 2005). The only exceptions were the large excretory ducts of the pancreas, the gall bladder, and the common bile duct, which have not been reported to express hCLCA1 or mCLCA3 in humans or mice, respectively, possibly due to either technical or study-related reasons or species-specific differences (Gruber et al. 1998b; Leverkoehne and Gruber 2002). Therefore, pCLCA1 can be assumed to share the as yet unknown functions of its orthologs hCLCA1 and mCLCA3 in mucin granules of goblet cells and as soluble constituents of the mucin layers on mucosal membranes. Despite strong PAS staining, no pCLCA1 signal was found in Brunner's glands. This might be due to the specialized function of these glands, which, in addition to glyco-proteins, secrete numerous additional factors, including bicarbonate, epidermal growth factor (EGF), bactericidal factors, proteinase inhibitors and surface-active lipids (Krause 2000). It is well accepted that EGF stimulates chloride secretion via a calcium-independent pathway (Carlos et al. 2007), raising the question of whether factors other than CLCA might be involved in chloride conductance within Brunner's glands. This hypothesis is supported by the observation that EGF is capable of inhibiting calcium-dependent chloride conductance, as shown in T84 human colonic epithelial cells (Uribe et al. 1996).
Future studies will have to address whether these CLCA family members act as soluble mediators of as-yet-unknown epithelial channel proteins or whether they serve other functions in the process of mucin secretion or as structural components of the mucous layer.
Also similar to hCLCA1, mCLCA3, and eCLCA1, the pCLCA1 protein data were found to be consistent with the general model of CLCA protein synthesis, cleavage, and processing, including shedding of both cleavage products through the secretory pathway (Patel et al. 2008). Unfortunately, the antibodies generated here against the carboxy-terminal cleavage product failed to identify their antigens in denaturing immuno-blots of the cell lysates or supernatant, despite strong reactivity in the immunohistochemical assays, which showed staining patterns identical to those of the antibodies directed against the larger N-terminal fragment. The reason for the lack of reactivity on immunoblots but strong and specific staining in immunohistochemistry cannot be explained at this point. However, it is generally accepted, and not an uncommon phenomenon, that some antibodies fail to detect their target proteins in immunoblot or immunoprecipitation, probably for conformational reasons, despite good results in the immunohistochemistry and vice versa.
Nevertheless, the absence of any potential transmembrane domain in both cleavage products and the fact that both the primary translation product and the N-terminal cleavage product were fully secreted by transfected cells with complex glycopatterns, together with the immunohistochemical detection of both cleavage products in the mucin layers, argue that both pCLCA1 cleavage products are secreted soluble proteins in the mucin layers of mucosal membranes. This model also fully complies with the models proposed for the human hCLCA1 (Gibson et al. 2005), the murine mCLCA3 (Mundhenk et al. 2006), and the equine eCLCA1 (Range et al. 2007). In fact, similar to the latter, first patch clamp studies on heterologously expressed pCLCA1 confirmed its role in mediating calcium-dependent transmembrane anion conductance. However, as a secreted protein, pCLCA1, like its human, murine and equine orthologs, cannot form an anion channel by itself. These data point toward an extracellular, indirect activation of the chloride conductance, possibly via an interaction with an as-yet-unknown chloride channel. Recent electrophysiological data on human hCLCA1 support the hypothesis that at least some CLCA proteins do not form chloride channels per se or enhance the trafficking or insertion of channels. Rather, hCLCA1 elevates the single-channel conductance of calcium-dependent chloride channels by lowering the energy barriers for ion translocation (Hamann et al. 2009). Unique electrophysiological features were observed for pCLCA1 that are at variance with observations on other CLCA homologs. For example, pCLCA1 is the first member of the CLCA family that is not inhibited by 4,4'-diisothio-cyanatostillbene-2,2'-disulfonic acid. Furthermore, pCLCA1 is the first CLCA protein to enhance cAMP-activated chloride conductance when expressed in the human colon carcinoma cell line, Caco-2, leading to the assumption that the CFTR channel may be involved (Loewen et al. 2002a,b). Further electrophysiological evidence of pCLCA1 pointed to a regulatory effect of this protein on other chloride channels rather than to an inherent chloride channel function (Loewen et al. 2004), supporting the hypothesis of an indirect activation of the chloride secretory pathway. Future side-by-side comparisons between pCLCA1 and other CLCA homologs will have to address whether these differences are real or due to different experimental settings.
In conclusion, pCLCA1 is expressed in virtually all CF-relevant tissues and mucosal membranes, and its genomic characteristics, tissue, and cellular expression patterns and protein processing are highly similar to those of its human ortholog, hCLCA1. Based on these results, there is as yet no reason to assume that species-specific differences of pCLCA1 may interfere with its characterization as a possible modifier of the CF phenotype in the pig and comparisons with human patients. However, the duplication of a putatively functional pCLCA4 variant that is absent from humans and mice has to be considered a pig-specific variation within the CLCA gene family so far. Future analyses will have to address the functional and biomedical significance of this duplication in normal and CF tissues in the pig.
Footnotes
Acknowledgements
This study was supported by the German Research Council (Deutsche Forschungsgemeinschaft MU 3015/1–1).
We thank Jana Enders for excellent technical support.
