Abstract
We reported previously that Nfic-deficient mice exhibit short and abnormal molar roots and severely deformed incisors. The objective of this study is to address the mechanisms responsible for these changes using morphological, IHC, and RT-PCR analysis. Nfic-deficient mice exhibited aberrant odontoblasts and abnormal dentin formation in molar roots and the labial crown analog of incisors. The most striking changes observed in these aberrant odontoblasts were the loss of intercellular junctions and the decreased expression of ZO-1 and occludin. As a result, they became dissociated, had a round shape, and lost their cellular polarity and arrangement as a sheet of cells. Furthermore, the dissociated odontoblasts became trapped in dentin-like mineralized tissue, resembling osteodentin in the overall morphology. These findings suggest that loss of the Nfic gene interferes with the formation of intercellular junctions that causes aberrant odontoblast differentiation and abnormal dentin formation. Collectively, these changes in odontoblasts contributed to development of molars with short and abnormal roots in Nfic-deficient mice.
Odontoblasts are tall and columnar in shape and are characterized by a highly polarized distribution of cellular organelles. They are joined and attached at their apical ends of cell bodies by terminal webs and associated fascia adherens junctions (Nishikawa and Kitamura 1987; Sasaki and Garant 1996). However, the regulatory mechanism for formation of the intercellular junctional complex in odontoblasts remains unclear.
The nuclear factor I (NFI) family of transcription-replication factors is encoded by four genes in mammals (Nfia, Nfib, Nfic, and Nfix) (Gronostajski 2000; Mukhopadhyay et al. 2001; Bachurski et al. 2003). The Nfi gene family encodes site-specific transcription factors essential for the development of a number of organ systems. NFI proteins seem to have unique cell type—specific transcriptional modulation properties. Nfia-deficient mice exhibit agenesis of the corpus callosum and other forebrain defects (das Neves et al. 1999; Shu et al. 2003), whereas Nfib-deficient mice possess unique defects in lung maturation and forebrain (Steele-Perkins et al. 2005), and Nfix-deficient mice showed defects in brain (Campbell et al. 2008) and spine (Driller et al. 2007). On the other hand, Nfic-deficient mice showed short molar roots and severe incisor defects (Steele-Perkins et al. 2003). Recently, we reported that the Nfic disturbance—induced short root formation is caused by abnormalities in odontoblasts and dentin formation but not in HERS (Park et al. 2007).
In this study, we investigated the possible mechanism by which disruption of Nfic interferes with the formation of intercellular junctions that causes dissociation of odontoblasts, leading to aberrant odontoblast differentiation and abnormal dentin formation.
Materials and Methods
Mice
Postnatal wild-type (WT) and Nfic-deficient mice [postnatal 7 (P7), 10, 14, and 18] were maintained at the specific pathogen free animal facility of the Dental Research Institute, Seoul National University. They were housed under controlled 12-hr day-night cycles and were provided with pelleted rodent diet and water. All animals were used for experiments according to the guidelines of the Dental Research Institute.
Tissue Preparation for Light and Electron Microscopy
The mice were anesthetized with isoflurane and cardiac-perfused with 2.5% glutaraldehyde in 0.1 M cacodylate buffer, pH 7.4. Mandibles were dissected, demineralized for 4 weeks in 10% EDTA (pH 7.4) at 4C, and processed for paraffin embedding. Tissues were postfixed in 1% osmium tetroxide for embedding in Epon mixture (Poly-science; Washington, PA). For transmission electron microscopic analysis, thin sections were cut using an ultramicrotome, stained with uranyl acetate and lead citrate, examined, and photographed using a transmission electron microscope (1200 EXII; JEOL, Tokyo, Japan).
Immunofluorescence Microscopy
Five-μm-thick paraffin sections were stained with hematoxylin and eosin (H&E). Subjacent sections selected for H&E staining were used for immunofluorescence microscopy. Deparaffinized sections were immersed in 0.6% H2O2/methanol for 20 min to quench the endogenous peroxidase activity. They were preincubated with 1% BSA in PBS for 30 min and incubated overnight at 4C with polyclonal rabbit anti-connexin-43 (Cx43), rabbit anti-ZO-1, or rabbit anti-occludin antibodies (Zymed Laboratories; South San Francisco, CA) at a 1:200 dilution. After washing, sections were incubated for 1 hr at room temperature with secondary antibody (anti-rabbit-IgG-FITC, 1:500; Invitrogen, Carlsbad, CA) and counterstained with 4′-6-diamidino-2-phenylindole (DAPI). Sections were examined by confocal laser scanning microscopy (LSM 5 PASCAL; Carl Zeiss, Jena, Germany).
MDPC-23 Cell Culture and Nfic Inactivation
MDPC-23 cells (a generous gift from Drs. C.T. Hanks and J.E. Nör, School of Dental Medicine, University of Michigan, Ann Arbor, MI) are spontaneously immortalized preodontoblastic cells that were originated from fetal mouse molar dental papilla (Hanks et al. 1998) and have the ability to differentiate into odontoblasts (Sun et al. 1998). They were cultured in DMEM (Invitrogen) supplemented with 10% FBS (Invitrogen) and antibiotics (penicillin-G, 100 U/ml; streptomycin, 100 μg/ml; Fungizone, 2.5 μg/ml; Invitrogen) at 5% CO2 in a 37C incubator. MDPC-23 cells were transiently transfected using Lipofectamine PULS reagent in Opti-MEM (Invitrogen) containing 2 μg of control vector (pLKO.1 control), RNAi NFIC plasmid (pLKO.1-NFIC shRNA), or pCMV-NFIC expression vector. Transfection was performed according to the manufacturer's instructions.
Primary Pulp Cell Culture
Under anesthesia, the mandibles were removed from 18-day-old WT and Nfic-deficient mice. After the incisors were dissected out, they were cut longitudinally into two pieces using a sterile razor blade. The pulp tissues were scraped carefully with a 27-gauge needle and placed on 60-mm culture dishes (Nunc; Rochester, NY). The explants were weighted down with a sterile cover glass and cultured in DMEM (Gibco BRL; Carlsbad, CA) supplemented with 100 IU/ml penicillin, 100 μg/ml streptomycin (Gibco BRL), and 10% FBS (Gibco BRL). The cells were treated with 50 μg/ml ascorbic acid and 10 mM β-glycerophosphate and cultured at 37C in a humidified atmosphere containing 5% CO2.
RT-PCR Analysis
Total RNA was extracted from MDPC-23 and primary pulp cells using TRIzol reagent according to the manufacturer's instructions (Invitrogen). Total RNA (2 μg) was reverse transcribed using 0.5 μg Oligo dT and 1 μl (50 U) Superscript III enzyme (Invitrogen) in a 20-μl reaction mixture at 50C for 1 hr The resulting mixture was amplified by PCR. One μl of reverse transcription product was subjected to PCR using the following cycling conditions: denaturation at 94C for 5 min, followed by 32 cycles for NFI-C (28 cycles for Cx43 and 35 cycles for ZO-1) each of denaturation at 94C for 30 sec, primer annealing at 55C for 30 sec, product extension at 72C for 45 sec, and final extension at 72C for 7 min. The primer sequences used are as follows: 5′-GAC CTG TAC CTG GCC TAC TTT G-3′ and 5′-TTT CCA CCA AAA ATG CAG GCT GG-3′ for NFI-C; 5′-TAC CAC GCC ACC ACC GGC-3′ and 5′-AAT CTC CAG GTC ATC AGG −3′ for Cx43; 5′-CAT AGA ATA GAC TCC CCT GG-3′ and 5′-GCT TGA GAC CTC ATA CCT GT-3′ for ZO-1; 5′-CAG CAG TAT GAA AGC GTG G-3′ and 5′-GCA CTC TTC TCC TGG TCC TG-3′ and 5′-GTT GAC CGT CCC TTC ACA GT-3′ for E-cadherin; 5′-GGA AGA AAA GGT TGG CAG AG-3′ for vimentin; 5′-AAG TTC GCA CTG GCA CCT AC-3′ and 5′-AAC CAA GAA GCC CTG GAG AC-3′ for α-tubulin; and 5′-ACC ACA GTC CAT GCC ATC AC-3′ and 5′-TCC ACC ACC CTG TTG CTG T-3′ for GAPDH. GAPDH was used as a control. PCR products were analyzed on a 1.2% agarose gel, stained with ethidium bromide, and visualized under UV light.
Western Blot Analysis
To prepare whole cell extracts, the cells were washed three times with PBS, scraped, and collected into a 1.5-ml tube. After centrifugation at 1000 × g for 5 min at 4C, supernatants were removed, and pellets were resuspended in lysis buffer (100 mM Tris, pH 7.4, 350 mM NaCl, 10% glycerol, 1% Nonidet P-40, 1 mM EDTA, 1 mM dithiothreitol, 10 μg/ml aprotinin, 10 μg/ml leupeptin, and 10 μg/ml pepstatin) and kept on ice for 15 min. Cell debris was removed by centrifugation at 16,000 × g for 15 min at 4C. Proteins (30 μg) were separated using 10% SDS-PAGE and transferred to a nitrocellulose membrane (Schleicher and Schuell Bioscience; Dassel, Germany). Membranes were blocked for 1 hr with 5% non-fat dry milk in PBS containing 0.1% Tween-20 (PBS-T), washed with PBS-T, and incubated overnight at 4C with rabbit anti-ZO-1 or rabbit anti-Cx43 antibodies (Zymed Laboratories; South San Francisco, CA) diluted 1:1000 in PBS-T. After washing, membranes were incubated for 1 hr at room temperature with horseradish peroxidase—conjugated anti-rabbit IgG (Santa Cruz Biotechnology; Santa Cruz, CA). Labeled protein bands were detected using an enhanced chemiluminescence (Amersham Biosciences; Amersham, UK).
Terminal Deoxynucleotidyl Transferase-mediated dUTP Nick End Labeling (TUNEL) Peroxidase (POD) Staining
Apoptotic cells were localized on 6-μm-thick paraffin sections using a terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling kit (TUNEL; In Situ Cell Death Detection Kit, POD) according to the manufacturer's instructions (Roche Biochemicals; Basel, Switzerland). Endogenous peroxidase was blocked by incubation for 10 min in 3% H2O2 before enzymatic labeling. Sections were washed in PBS and treated with a 50-μl TUNEL reaction mixture. The color reaction was achieved by incubation in PBS containing DAB after enzymatic labeling. Sections were counterstained with methyl green.
Results
Morphological Changes of Odontoblasts in Incisors From Nfic-deficient Mice
The incisor of a WT mouse showed circular dentin around the pulp in cross-section. The inner surface of dentin was lined with a single layer of tall columnar and well-organized odontoblasts (Figure 1A). In contrast, Nfic-deficient mice had an open area in the lingual root analog of the incisor as a result of lack of dentin formation, whereas the labial crown analog contained abnormal dentin in which aberrant odontoblasts were trapped (Figure 1B). Also, the surface of abnormal dentin was uneven and covered with polygonal, disorganized, and abnormal odontoblasts (Figure 1B). A longitudinal section of a molar from an Nfic-deficient mouse had a short and abnormal root compared with that of a WT mouse (Figures 1C and 1D).
In longitudinal sections of incisors from WT and Nfic-deficient mice, both showed neural crest—derived differentiating odontoblasts. Those in WT differentiated into normal odontoblasts that were elongated, highly polarized, and well organized as a sheet of cells (Figures 2A and 2B). However, those in Nfic-deficient incisors failed to differentiate into normal odontoblasts. These aberrant odontoblasts were round and dissociated and lost their polarity (Figures 2C and 2D). Electron microscopy of aberrant odontoblasts exhibited no intercellular junctional complex between them (Figure 2E). Many of them were trapped in osteodentin-like mineralized tissue, thereby resembling osteocytes.
Formation of Intercellular Junctions Between Odontoblasts in Nfic-deficient Mice
We studied the IHC localization of three major structural proteins of the junctional complexes, ZO-1, occludin, and Cx43, in WT and Nfic-deficient mice. Odontoblasts from WT mice showed strong ZO-1 expression in the apical region, whereas aberrant odontoblasts from Nfic-deficient mice showed no ZO-1 expression (Figures 3A and 3B). Occludin immunoreactivity was observed at the junctional complexes between odontoblasts from WT mice but not in aberrant odontoblasts from Nfic-deficient mice (Figures 3C and 3D). Cx43 immunoreactivity was observed in the junctional complexes between odontoblasts from WT mice and also in the abnormal odontoblasts from Nfic-deficient mice (Figures 3E and 3F).
To examine the effect of Nfic on intercellular junction formation, we performed RT-PCR analysis of E-cadherin, ZO-1, and Cx43 mRNA expression in normal and Nfic-inactivated MDPC-23 cells and primary pulp cells from WT and Nfic-deficient mice. ZO-1 mRNA was expressed strongly in normal MDPC-23 cells but weakly in the Nfic-inactivated MDPC-23 cells. Primary pulp cells from WT mice showed strong E-cadherin and ZO-1 mRNA expression, whereas those from Nfic-deficient mice exhibited a decreased level in their mRNA expression (Figure 4A). However, Cx43 expression remained unchanged in the cells after Nfic inactivation (Figure 4A). ZO-1 and Cx43 protein expression in normal and Nfic-inactivated MDPC-23 cells was confirmed by Western blot analysis. Cx43 protein expression was not changed in MDPC-23 cells after inactivation of the Nfic gene. However, ZO-1 protein expression was decreased in Nfic-inactivated MDPC-23 cells compared with normal MDPC-23 cells (Figure 4B).
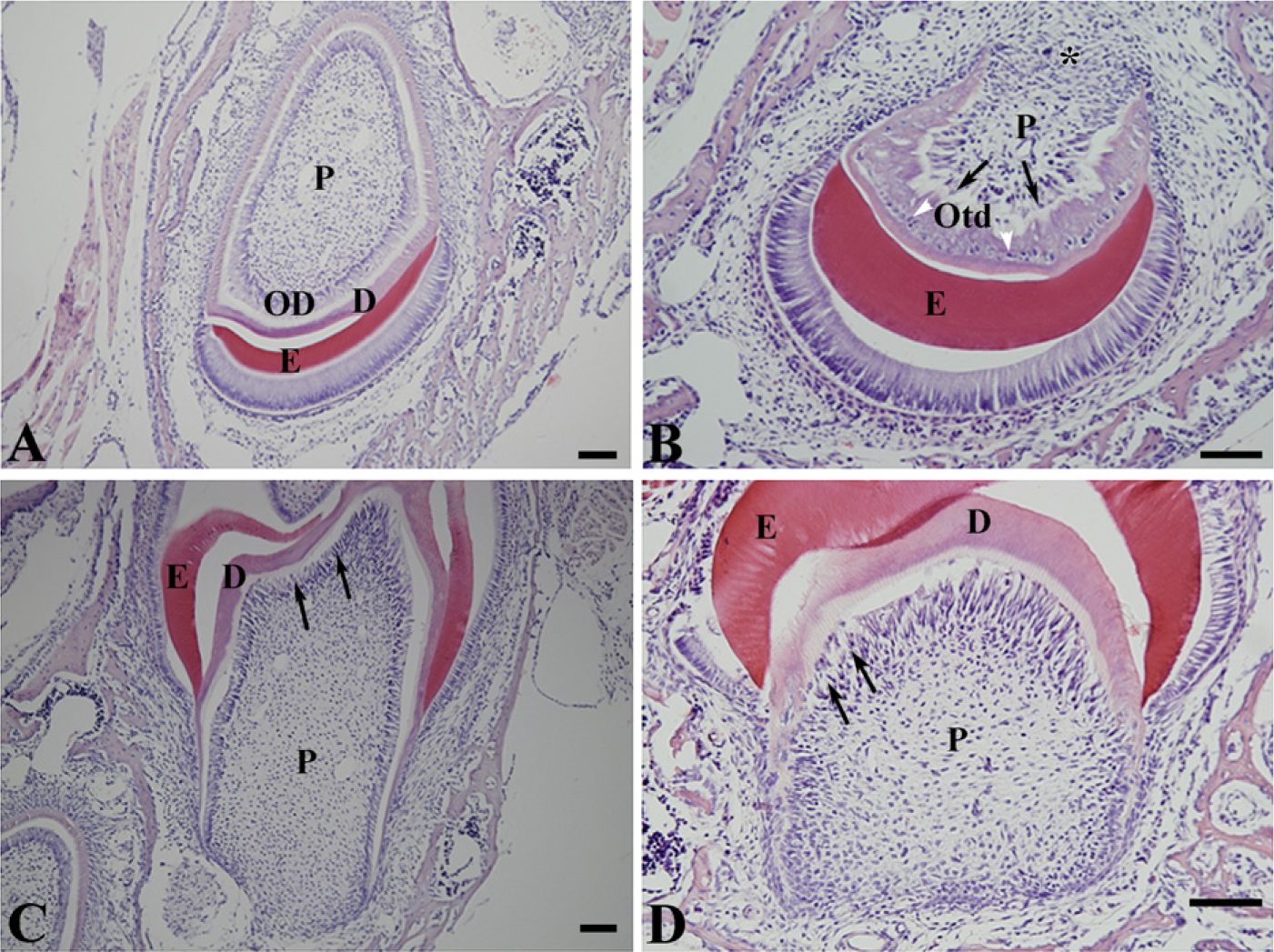
Light micrographs showing cross-section of incisors from P10 wildtype (WT) (
Odontoblast Apoptosis in Nfic-deficient Mice
TUNEL-positive odontoblasts were hardly observed in incisors from WT mice (Figure 5A). However, apoptotic cells were found in the subodontoblastic region and were more evident in the lingual region than the labial region of the pulp from Nfic-deficient mice (Figure 5B). Electron microscopic observations confirmed that many cells were apoptotic. The cells were characterized by condensed chromatin, swollen mitochondria, cellular inclusions, and vacuoles (Figure 5C).
Discussion
Here we showed that disruption of Nfic induced the formation of aberrant odontoblasts and abnormal dentin in both molar roots and the labial crown analog but not the lingual root analog of the incisors. Aberrant odontoblasts became round in shape, dissociated, and embedded in abnormal dentin. These changes in odontoblasts contributed to the development of molars with short roots and severely deformed incisors in Nfic-deficient mice (Steele-Perkins et al. 2003). We reported that, during odontoblast differentiation, Nfic mRNA is expressed specifically in odontoblasts but not in pre-odontoblasts (Steele-Perkins et al. 2003). Furthermore, in this study, we found that Nfic disruption did not interfere with the differentiation of dental papilla cells into preodontoblasts but disturbed the differentiation of odontoblasts from preodontoblasts. All these findings suggest that NFIC plays a key role in the terminal differentiation and function of odontoblasts.
One of the unique features of normal odontoblasts is the presence of terminal webs and associated fascia adherens junctions at the apical end of cell bodies (Nishikawa and Kitamura 1987; Sasaki and Garant 1996). Our finding that loss of Nfic caused odontoblast dissociation and entrapment in abnormal dentin provides direct evidence that intercellular junctions are responsible for alignment of odontoblasts as a single layer of cells, functioning as a unit (Linde and Goldberg 1993). Also, the junctional complex plays a crucial role in synchronized secretion of dentin matrix for formation of a uniformly even dentin surface and preventing their entrapment in the predentin/dentin during dentin formation (Linde and Goldberg 1993; Arana-Chavez and Katchburian 1997).
Intercellular junctions are formed during odontoblast differentiation from preodontoblasts (João and Arana-Chavez 2004) and differ morphologically from those found in highly polarized intestinal epithelial cells. Odontoblasts have the adhesion complex that consists of fascia adherens, fascia occludens, and gap junctions (Iguchi et al. 1984). The fascia adherens are composed of dense patches along the closely juxtaposed cell membrane of the neighboring odontoblasts and numerous actins inserted into the dense patches. The fascia occludens does not form a zona occludens as observed in epithelial cells. Numerous gap junctions are present between cell bodies of odontoblasts (Iguchi et al. 1984). However, little is known about the regulatory mechanism for intercellular junction formation in normal odontoblasts. ZO-1, a 225-kDa membrane protein, is a major structural protein that is present at the cytoplasmic surface of tight junctions and binds to claudins, occludin, ZO-2, ZO-3, and actin (Stevenson 1999; González-Mariscal et al. 2003). E-cadherin is a cell adhesion molecule that is connected to actin-based adherent junctions through catenins and thus is crucial for efficient cell—cell adhesion (Ozawa et al. 1990). Connexins are a family of transmembrane proteins that assemble to form vertebrate gap junctions, which are organized in hexametric assemblies in plasma membranes of adjacent cells to form intercellular channels (Kumar and Gilula 1996; Elenes et al. 2001). During odontoblast differentiation, both Cx43 and ZO-1 are expressed in differentiating odontoblasts (João and Arana-Chavez 2003), when NFIC is also expressed. Interestingly, aberrant odontoblasts in the incisors from Nfic-deficient mice showed the loss of intercellular junctions and decreased expression of ZO-1 and occludin. Furthermore, primary pulp cells from Nfic-deficient mice showed a decreased expression level of E-cadherin compared with that of WT pulp cells. These data provide the first evidence that Nfic is involved in intercellular junction formation by regulating the expression of junction components and that loss of Nfic results in the dissociation of odontoblasts.
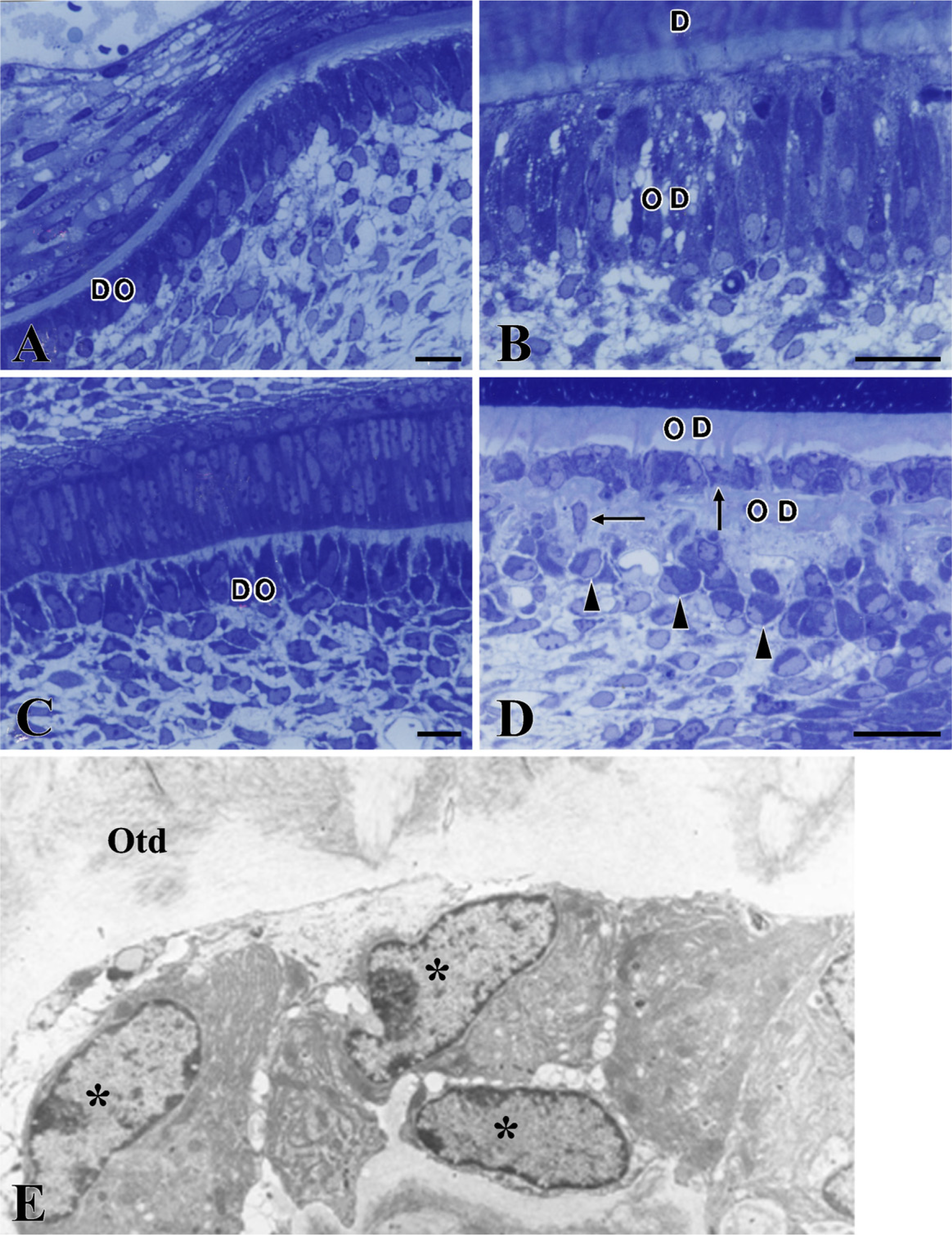
Light and electron micrographs showing differentiating odontoblasts and odontoblasts in longitudinal sections of mandibular incisors from P14 WT (
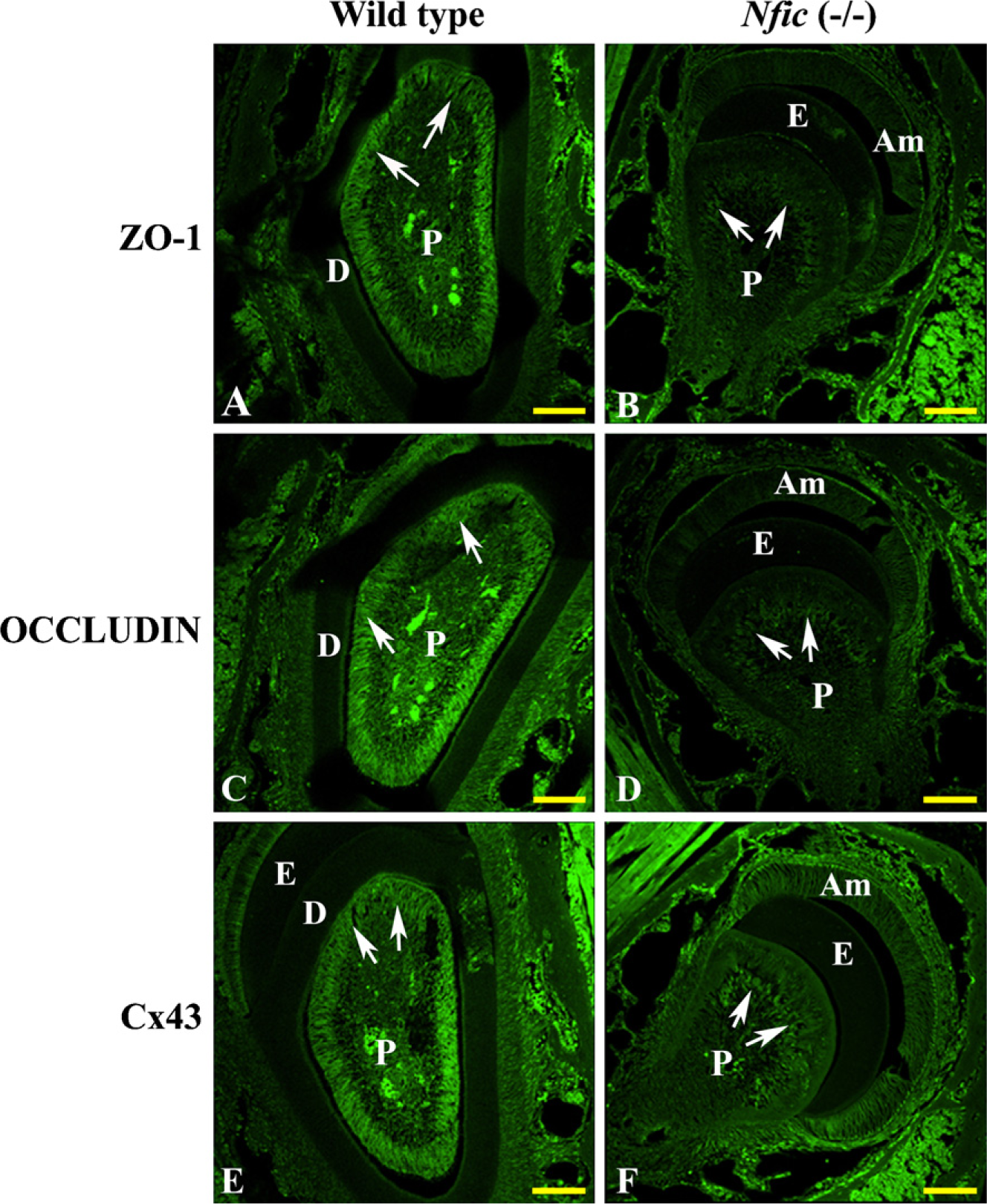
IHC localization of ZO-1 (
Although odontoblasts respond to external stimuli as a protective function, their ability to repair dentin is limited. When severe dentin damages from deep caries and trauma kill the original odontoblasts, pulp stem cells proliferate and differentiate into odontoblast-like cells that form reparative dentin (Smith et al. 1990). Reparative dentin contains abnormal odontoblasts with a round shape, no odontoblastic processes, and no polarity. With continuous deposition of matrix, they become trapped and form atubular abnormal dentin, also known as osteodentin, because it resembles bone in overall morphology. The mechanism for osteodentin formation during dentin repair is unknown. The morphological similarity between osteodentin and abnormal dentin found in Nfic-deficient mice led us to postulate that pulp stem cells may not be able to fully differentiate into normal odontoblasts during dentin repair and thus are formed by cells that do not express NFIC. In addition, transgenic mice that overexpress active transforming growth factor (TGF)-β1 predominantly in odontoblasts showed the same phenotypic changes as seen in Nfic-deficient mice (Thyagarajan et al. 2001). These transgenic animal studies strongly suggest a functional relationship between Nfic and TGF-β1—mediated signaling molecules such as Smads in odontoblast differentiation.
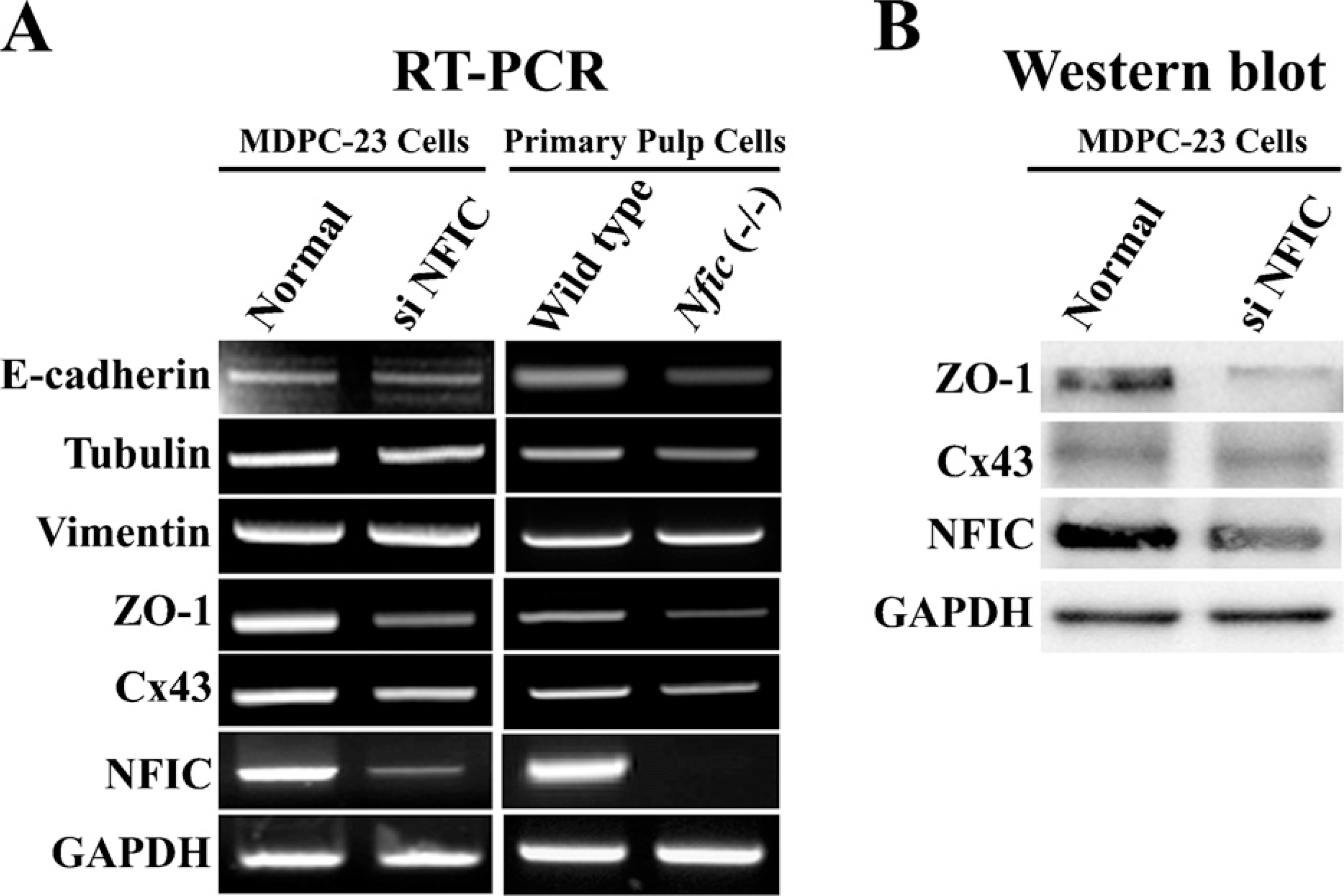
Expression of E-cadherin, tubulin, vimentin, ZO-1, and Cx43 mRNA and protein by RT-PCR (
Finally, TUNEL staining showed an increased number of apoptotic cells in incisors from Nfic-deficient mice. In addition, electron microscopy showed that some of the aberrant odontoblasts embedded in abnormal dentin undergo apoptosis. Thus, it seems most likely that development of molars with short and abnormal roots in the Nfic-deficient mice results from a combination of aberrant odontoblast formation and a reduced population of aberrant odontoblasts. However, the mechanism by which disruption of Nfic induces odontoblast apoptosis in a cell-specific manner remains unknown.
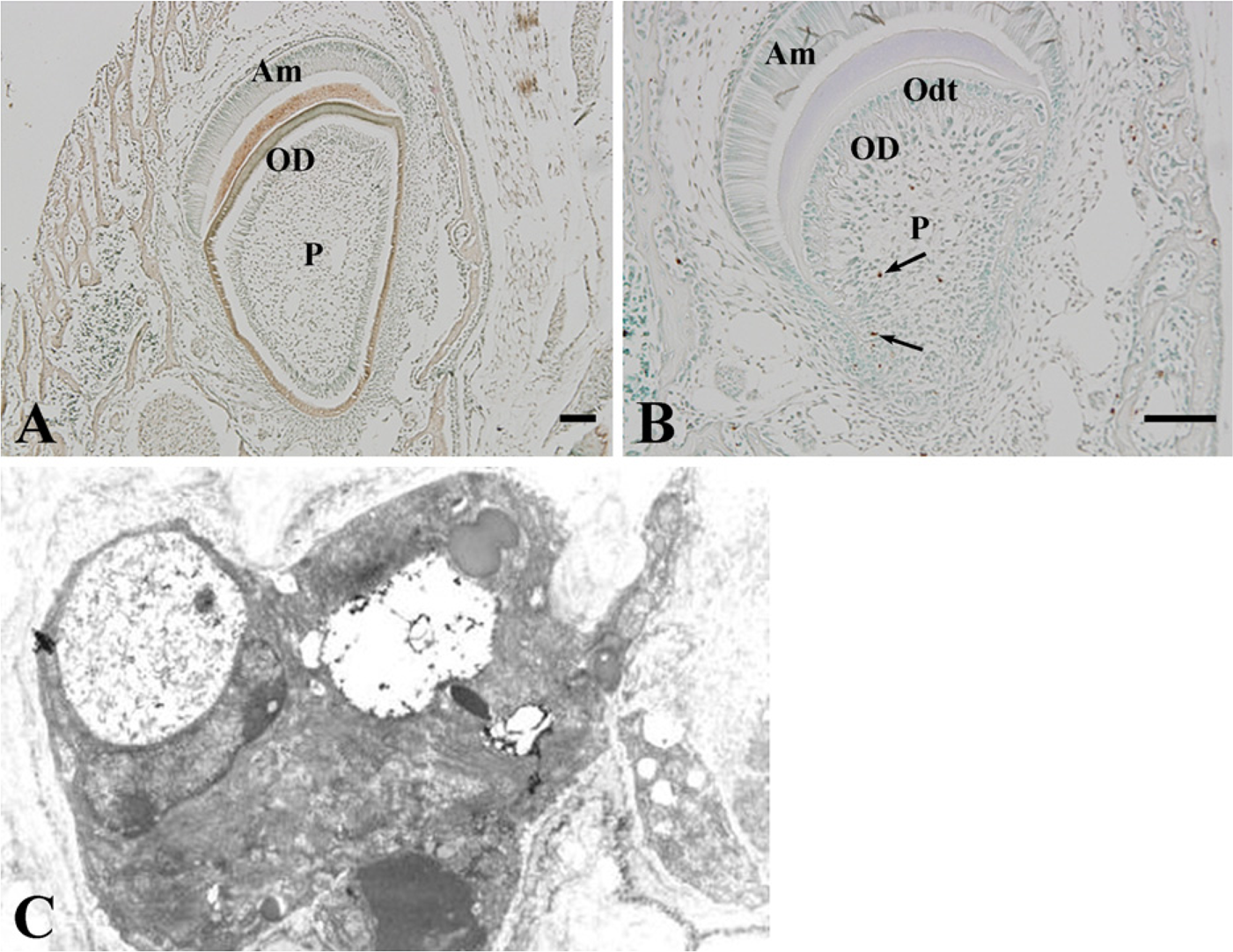
Terminal deoxynucleotidyl transferase—mediated dUTP nick end labeling (TUNEL) peroxidase (POD) staining of incisors from P10 WT (
In summary, the disruption of Nfic induces dissociation, phenotypic changes, and apoptosis of odontoblasts by interfering with the formation of intercellular junctions. The findings from this study help us to better understand the regulatory mechanisms for odontoblast phenotype, survival, and osteodentin formation. In addition, it will be interesting to know if the disruption of the Nfic gene is associated with patients with short roots.
Footnotes
Acknowledgements
This study was supported by RO1-2006-000-10581-0 from the Basic Research Program of The Korea Science and Engineering Foundation, a Korea Science and Engineering Foundation (KOSEF) grant funded by the Korean government (MOST; M10646010002-06N4601-00210), and a Korea Research Foundation grant funded by the Korean Government (MOEHRD, Basic Research Promotion Fund; KRF-2008-314-E00214). R.M.G. was supported by National Heart, Lung, and Blood Institute Grant 5R01HL080624.
