Abstract
Nasopharyngeal carcinoma (NPC) is a common malignant tumor of the head and neck. Its pathogenesis is complicated and needs further investigation. The aim of this study was to investigate the expression and clinical significance of WWP1 in NPC. Bioinformatics approaches were used to evaluate the expression and functions of WWP1 in NPC. WWP1 protein expression was then detected by immunohistochemistry on a tissue microarray in an NPC cohort and its association with clinical features and prognosis was determined. In addition, WWP1 expression was knocked down in NPC cells using RNA interference, and their colony formation and invasion abilities were assessed. A total of 25 genes closely related to WWP1, which may be enriched in different pathways, were filtered out. WWP1 expression was significantly higher in NPC cells than in normal controls. High WWP1 expression was correlated with lymph node metastasis, tumor recurrence, clinical stage and poor prognosis. Knockdown of WWP1 resulted in attenuated proliferation and invasion of NPC cells. The results suggest that WWP1 may serve as a novel biomarker and prognostic factor for NPC and a potential therapeutic target worthy of further investigation:
Introduction
Nasopharyngeal carcinoma (NPC) is a malignant disease with remarkable geographic and racial distributions in the world. On the basis of the report of GLOBCAN 2018, 1 the incidence rate and mortality rate of NPC in the Western countries are decreasing after decades of cancer prevention and treatment. While in Southern China, the incidence rate is still very high possibly due to the high consumption of salted-preserved fish and the high prevalence of Epstein–Barr virus (EBV) infection. 2 More than 70% of NPC patients present neck lymph node metastasis at the time of diagnosis. 3 Although radiotherapy achieves favorable outcomes for patients at early stages, the occurrence of local recurrence and distant metastasis may lead to poor clinical outcomes. 4
Many factors such as genetic variation, viral infection, and environmental pollution exposure might contribute to the carcinogenesis of NPC, of which aberrant expression of genes may be a major one. For example, recent reports showed that Cyclin D1 is overexpressed in NPC tissues, which might predict good clinical outcomes. 5 Likewise, overexpression of INHBA may be associated with advanced clinical stages and poor prognosis in NPC patients. 6 As an oncogene in NPC, LAPTM4B knockdown inhibited the proliferation, migration, and invasion ability of NPC cells, and thus, LAPTM4B has been suggested to be a potential target for NPC therapy. 7 Hence, the aberrant expression of the genes might play crucial roles in the development of NPC.
WWP1 (WW domain-containing E3 ubiquitin protein ligase 1) is a multifunction protein that contains domains for ubiquitin transferring and substrate binding. As an intrinsic E3 ubiquitin ligase, WWP1 has been implicated as an oncogene, which is crucial in the maintenance and progression of various types of epithelial cancers. 8 Previous reports have shown that WWP1 was overexpressed in colon cancer, which predicted a poor prognosis. 9 Similarly, in hepatocellular carcinoma, high WWP1 expression might also have a correlation with poor clinical outcomes, and thus, WWP1 knockdown resulted in a decrease in cell proliferation, colony formation, and cell migration abilities. 10 However, to our knowledge, few studies have been devoted to the expression and possible roles of WWP1 in NPC.
In this study, we aimed to explore the expression of WWP1 in NPC tissues and further evaluate its clinical significance in this cancer.
Materials and Methods
Protein-Protein Interaction (PPI) Network Evaluation
To know the possible biological functions of WWP1, the analysis was performed in the STRING database. 11 The genes that might have a close relationship with WWP1 were filtered out using PPI network construction. Then, they were analyzed in the GATHER tool 12 including gene ontology (GO) function analysis, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis.
Transcription Factor Network Evaluation
Since a gene might interact with other genes, we predicted the target genes of WWP1 using the NetworkAnalyst tool. 13 A transcriptional factor database was selected. Then, GO analysis and pathway enrichment analysis were also performed to annotate the possible functions of WWP1 and its predicted target genes.
Evaluation of WWP1 mRNA Expression in NPC Tissues by Data Mining
Oncomine is a tool that includes the genes, pathways, and networks deregulated across 18,000 cancer gene expression microarrays, concerning the majority of cancer types and subtypes. 14 To evaluate the expression levels of WWP1 mRNA in both NPC tissues and the healthy controls, the Oncomine database 15 was used.
Cell Culture, qPCR Assay, and Western Blot Assay
Human NPC cell lines 5-8F, CNE2, HONE1, and human immortalized nasopharyngeal epithelial cell line (NP69) were obtained from Chongqing Aozhe Biotech company, and were maintained in RPMI-1640 medium (Gibco, USA) supplemented with 10% fetal bovine serum (FBS, Gibco). All cells were incubated at 37C with 5% CO2.
Total RNA was extracted from tissues or cells by TRIzol Reagent (Invitrogen, Thermo Fisher Scientific, USA) and then reversely transcribed to cDNA by PrimeScript RT reagent kit (TaKaRa, Japan) according to the manufacturer’s protocols. GAPDH mRNA was used as an internal control for WWP1 mRNA expression. The primer sequences were presented as follows:
WWP1: F: TGCTTCACCAAGGTCTGATACT, R: GCTGTTCCGAACCAGTTCTTTT.
GAPDH: F: ATTCCACCCATGGCAAATTC, R: GCATCGCCCCACTTGATT.
Each PCR amplification was performed in triplicate.
Western blot assay was conducted. In brief, cells were harvested, washed with ice-cold phosphate-buffered saline (PBS), and lysed with radioimmunoprecipitation assay (RIPA) buffer supplemented with protease inhibitor. Proteins were running on a 10% sodium dodecyl sulfate (SDS)–polyacrylamide gel electrophoresis and transferred to polyvinylidene difluoride membranes (Roche, Switzerland). Blots were then incubated in fresh blocking solution with an appropriate dilution of primary antibody at 4C for 24 hr. The sources of antibodies were as follows: WWP1 rabbit polyclonal, GAPDH rabbit polyclonal (BIOSS, China). The intensities of WWP1 were adjusted by the GAPDH band intensity.
NPC Cohort
An NPC tissue microarray (HNasN129Su01), containing 129 NPC tissues, was purchased from Shanghai Outdo Biotech Co., Ltd. The cohort contained 99 males and 30 females. Their ages ranged from 20 to 82 years (median: 47 years). The diagnoses were carried out between January 2010 and October 2011. The follow-up period was from May 2014 to March 2017 (Mean follow-up time was 75.19 months). All patients were clinicopathologically diagnosed as NPC and received no extra treatment before surgery. The clinical features of this chip were listed in Table 1. This study was approved by the ethics committee of Shanghai Outdo Biotech Co., Ltd. (SHYJS-CP-1810011). All samples were obtained after informed consent from the patients.
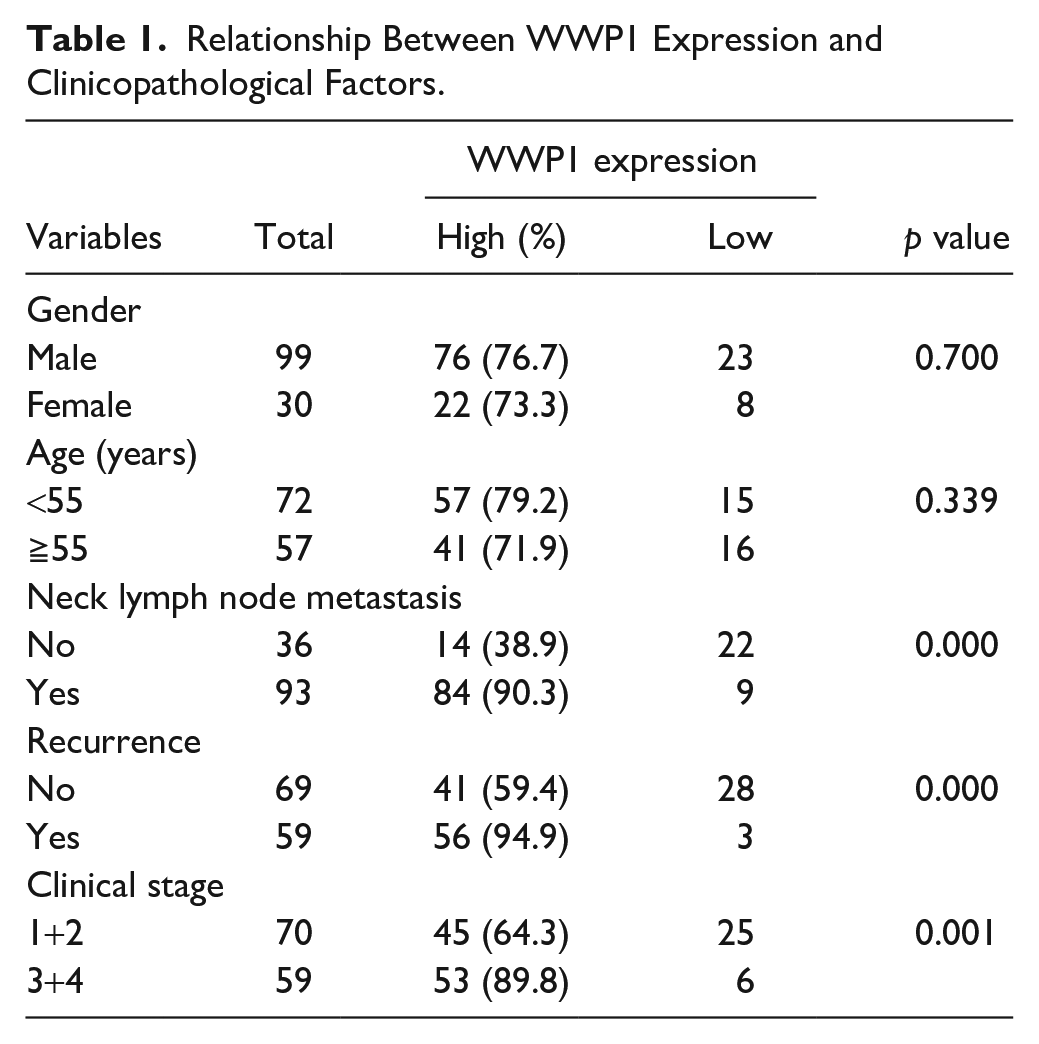
Relationship Between WWP1 Expression and Clinicopathological Factors.
Immunohistochemistry (IHC) Staining and Evaluation
The protein expressions of WWP1 were detected using the method of IHC. In brief, the slides were deparaffinized, rehydrated, and treated with 3% hydrogen peroxide for 20 min to inhibit endogenous peroxidase. The sections were rinsed with distilled water and saturated in phosphate-buffered saline for 5 min and then incubated with a 1:500 dilution of rabbit anti-polyclonal antibody (primary antibody; BIOSS) overnight at 4C. The staining was visualized using DAB solution and counterstained with hematoxylin. IHC staining was conducted in accordance with the manufacturer’s instructions.
The IHC stain results were identified by integrated scoring. The scoring method which combined the intensity and the percentage of positivity was previously described in the literature. 16 In brief, the staining intensities of WWP1 were scored from 0 to 3, where 0 = negative, 1 = weak, 2 = moderate, and 3 = strong. The percentages of positively stained cells were scored on scales from 0 to 4, in which 1 represents 0–25%, 2 represents 26–50%, 3 represents 51–75%, and 4 represents 76–100%.
The staining results were evaluated and scored independently by two observers without knowledge of the clinical information of the cases. When the samples were scored by the intensity and the percentage of positivity, a total score was obtained by multiplying these two parameters. Thus, the total score ranges from 0 to 12. Cut-off levels for the scoring were presented as follows: scores of ≥6 were classified as high expression, while those of <6 were classified as low expression.
Establishing Stable WWP1-Knockdown Cell Lines
The WWP1 shRNA with nonoverlapping sequences were synthesized by Jikai Gene (Shanghai, China) and packed into the lentiviral vector. The lentiviral shRNA (20 μl/ml medium) was added to 5-8F cells in a 6-well culture plate for 24 hr. Selection with puromycin (5 μg/ml) was performed on cells for 2 to 3 weeks to select WWP1-stable knockdown NPC cells. The cells transfected by WWP1 shRNA were named sh-WWP1-5-8F, and those transfected by the control sequences were named sh-NC-5-8F.
Colony Formation Assay
Cells were seeded into the 6-well plate at a density of 600 per well. The crystal violet was used to stain the cells following incubation at 37C in a 5% humidified atmosphere for about 10 days, and the colonies were counted to assess the clonogenic ability of the cells.
Transwell Migration and Invasion Assay
Transwell 24-well plates with 8-μm microporous filters (Corning, USA) were used to investigate the migration and invasion activity of the cancer cells. The cells (5-8F) were counted and resuspended in a serum-free medium. A total of 4×104 cells were seeded into the upper chambers, which were coated with (invasion) or without (migration) Matrigel (BD, USA). The bottom chambers were filled with 600 μl RPMI 1640 containing 20% FBS. For migration assay, the apparatus was incubated for 6 hr at 37C, whereas for 18 hr at 37C for invasion assay. After incubation, cells in the inner wall of the chambers were removed, and then cells in lower filters (outside the chambers) were washed with PBS twice and stained with Giemsa (Sigma-Aldrich, St. Louis, MO) for 10 min, respectively. At last, images were taken, and the number of the cells was counted.
Statistical Analysis
For continuous variables, differences between groups were analyzed with analysis of variances (ANOVA), a t-test, or a Wilcoxon Rank Sum Test according to the concrete types of the data. A chi-squared test was used to differentiate the rates of different groups. The Kaplan–Meier method was used to calculate the overall survival curves, and a log-rank test was used to determine differences in the survival rates. Cox multivariate regression survival analysis was conducted involving all the potential clinical factors. The analyses were performed by utilizing the MedCalc software (15.2.2; Mariakerke, Belgium). p<0.05 was considered to have statistical significance.
Results
Protein-Protein Interaction (PPI) Network Construction
In order to construct the PPI network, the gene WWP1 was assessed in the STRING with a confidence of 0.40. The network was established by a total of 26 genes including WWP1 (Fig. 1A). To learn the possible functions of the closely related genes of WWP1, the GO and pathway enrichment analyses were assessed. The data showed that these genes were enriched in a total of 55 GO terms and six pathways, as shown in Fig. 1B.
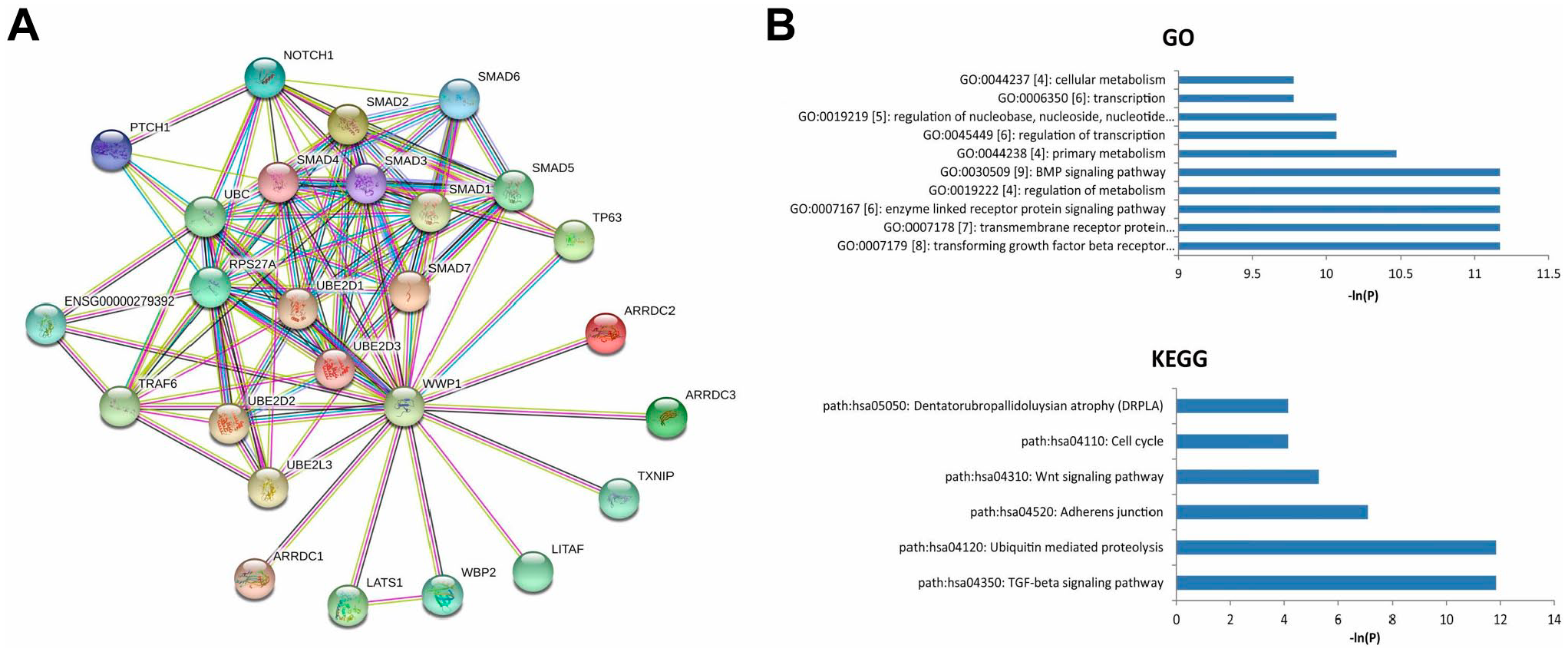
(A) PPI network for WWP1. (B) GO and KEGG enrichment analysis for the genes closely related to WWP1.
The top GO terms included GO:0007179 [8]: transforming growth factor-β (TGF-β) receptor signaling pathway, GO:0007178 [7]: transmembrane receptor protein serine/threonine kinase signaling pathway, GO:0007167 [6]: enzyme linked receptor protein signaling pathway, GO:0019222 [4]: regulation of metabolism, and GO:0030509 [9]: BMP signaling pathway.
Pathway enrichment analysis showed that these genes might be associated with several cancer-related pathways, such as path: hsa04350: TGF-β signaling pathway, path:hsa04120: Ubiquitin mediated proteolysis, and path:hsa04310: Wnt signaling pathway.
Expression of WWP1 mRNA and Protein in NPC Cases
Through the analysis in the Oncomine database, the expression levels of the WWP1 mRNA in NPC and the control tissues were evaluated. The data showed that the expression of WWP1 mRNA was significantly higher in NPC tissues than that in the control ones (p<0.05), as shown in Fig. 2A. Then, the samples of NPC were detected for WWP1 protein expression by IHC. Specific staining was mainly found in the nuclei and cytoplasm of cancer cells (Fig. 2B).
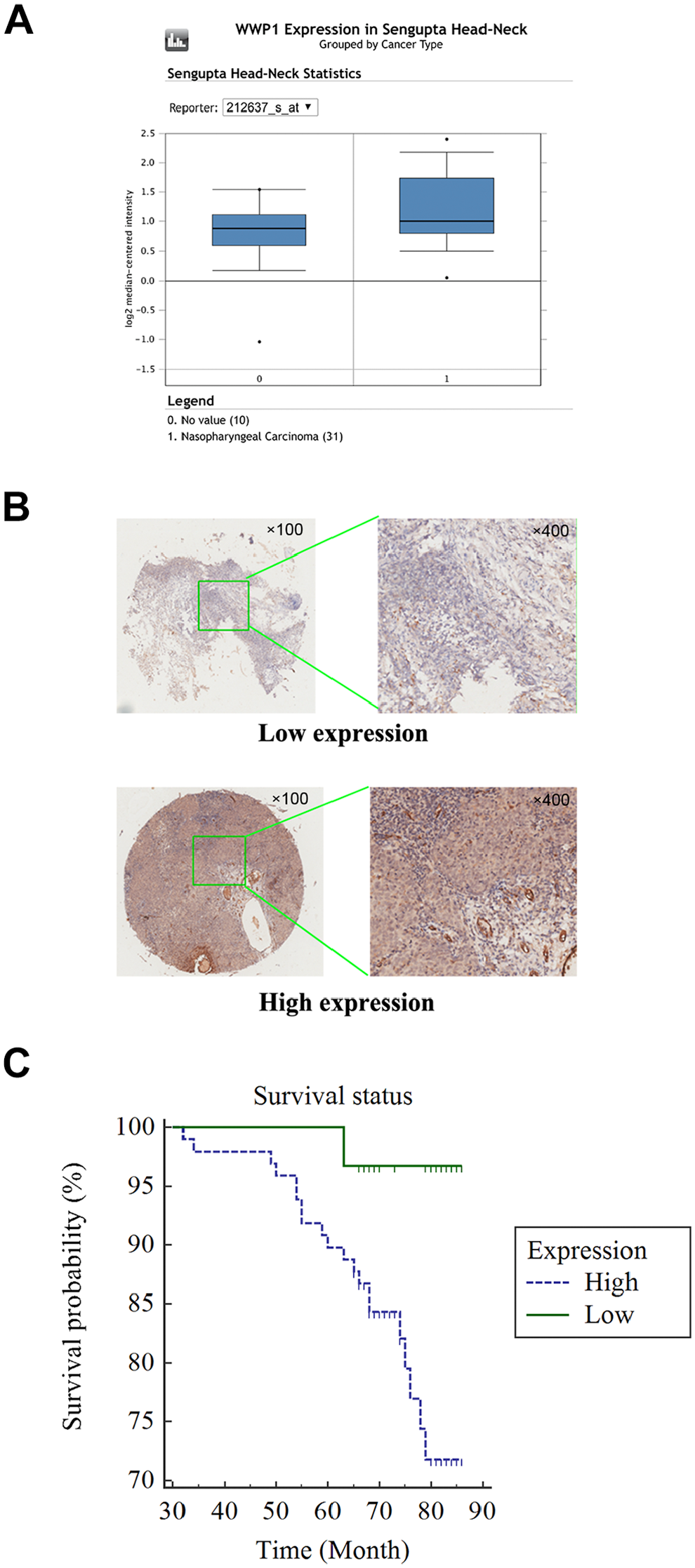
(A) The mRNA expression of WWP1 in NPC tissues and healthy controls (p<0.05, cancer vs control). (B) The protein expression of WWP1 in NPC samples was assessed by IHC on the basis of a tissue chip (100× and 400×). (C) A Kaplan–Meier curve indicated that patients with high expression of WWP1 have a shorter overall survival time than those patients with low WWP1 expression (p<0.05).
Relationship Between Clinicopathologic Features and WWP1 Expression
In this chip, five parameters could be assessed, including age, sex, neck lymph node metastasis, recurrence (local recurrence), and clinical stage.
The expressions of WWP1 in NPC samples were stratified into two groups (a high and a low group) according to the criteria mentioned.
As shown in Table 1, the relationship between WWP1 expression level and the clinicopathologic parameters was evaluated. No associations were shown between high WWP1 expression and age or sex, respectively (p>0.05). However, the data showed that high WWP1 expression might be correlated with the presence of neck lymph node metastasis (p<0.01), tumor recurrence (p<0.01), and advanced clinical stages (p<0.01), respectively.
Relationship of WWP1 Protein Expression With the Prognosis in NPC Cases
The prognostic value of WWP1 expression in NPC was further evaluated. The log-rank test showed that patients with high WWP1 expression had a shorter overall survival time than patients with low expression (p<0.05, Fig. 2C). Nevertheless, multivariate Cox regression analysis failed to show that the high expression of WWP1 protein may be an independent prognostic factor for NPC patients (Table 2).
Multivariate Analyses of WWP1 Protein Expressions and Other Clinical Markers Related to Overall Survival in NPC.
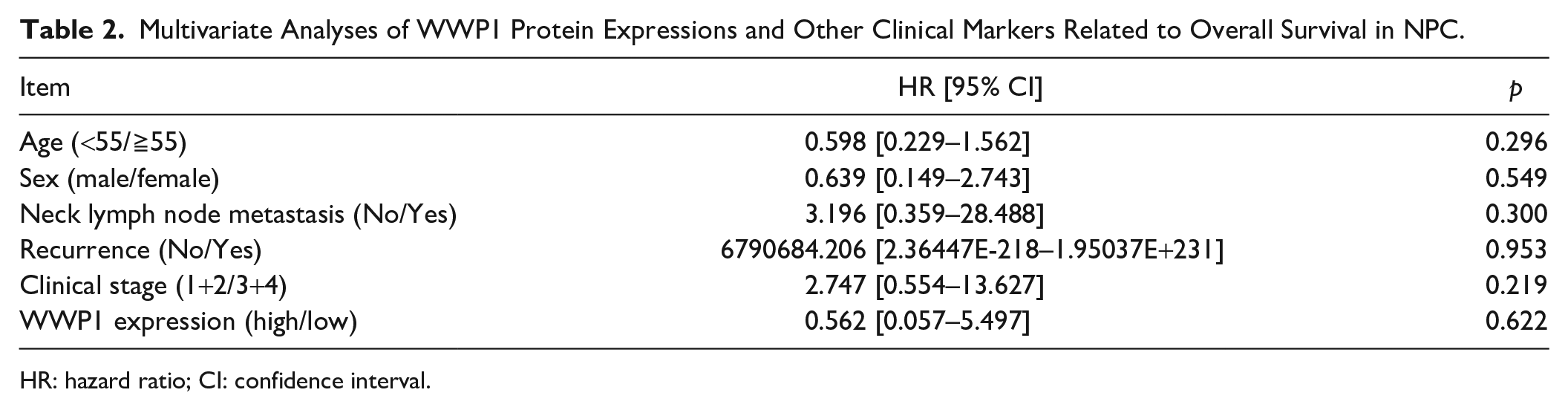
HR: hazard ratio; CI: confidence interval.
WWP1 Knockdown in NPC Cells Resulted in Attenuated Cell Proliferation and Weakened Cell Invasive Abilities
WWP1 mRNA was overexpressed in NPC cell lines relative to that in NP69 cells, particularly in 5-8F cells (p<0.05, Fig. 3A). Thus, 5-8F was selected in the following experiments. RNA interference technique was exploited in 5-8F cells to knockdown WWP1. As a result, WWP1 mRNA and protein were significantly downregulated in sh-WWP1-5-8F cells compared with those in sh-NC-5-8F cells, respectively (p<0.05, Fig. 3B and C). The number of colony formation of sh-WWP1-5-8F was significantly less than that of sh-NC-5-8F (p<0.05, Fig. 3D). The invasive capabilities of sh-WWP1-5-8F were markedly lower than that of sh-NC-5-8F (p<0.05, Fig. 3E).
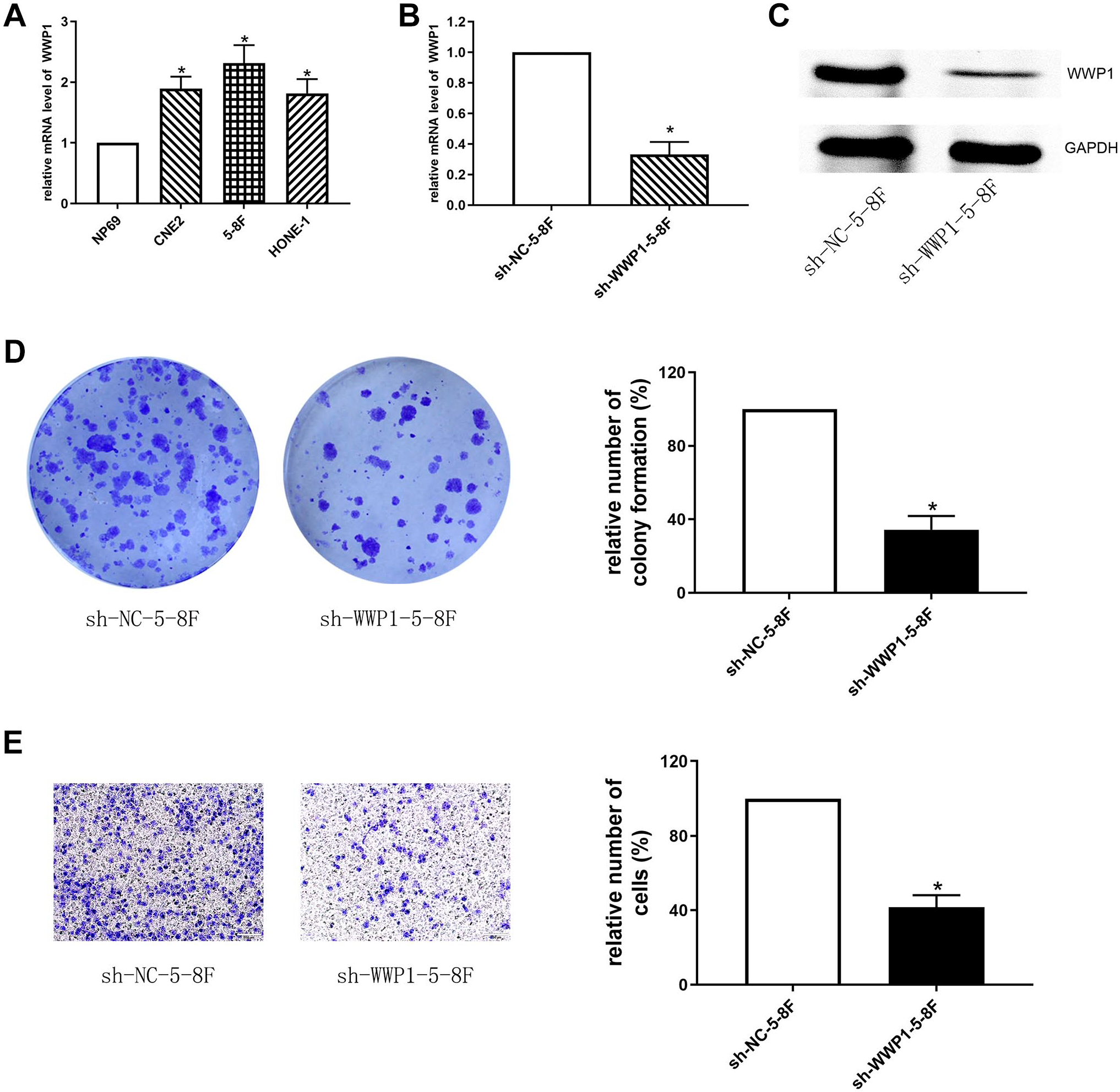
(A) The mRNA expression of WWP1 in NPC cell lines and NP69 cells (*p<0.05 vs NP69). (B-C) The expression of WWP1 mRNA (B) and protein (C) in sh-WWP1-5-8F and sh-NC-5-8F cells, respectively (*p<0.05). (D) The colony formation assay showed that the number of colony formations in sh-WWP1-5-8F was less than that in sh-NC-5-8F (*p<0.05). (E) The transwell migration and invasion assay showed that the invasive abilities of sh-WWP1-5-8F were weakened than those of sh-NC-5-8F (*p<0.05).
Discussion
We have explored the expression of WWP1 in an NPC cohort using a tissue chip. The results showed that high WWP1 expression may have an association with neck lymph node metastasis, tumor recurrence, and advanced clinical stages. The data confirmed that WWP1 may act as an oncogene in NPC.
Ubiquitination is a cellular process that involves adding one or more ubiquitins to a substrate, leading to protein degradation or transport. This process is carried out by ubiquitin-activating enzymes (E1), ubiquitin-conjugating enzymes (E2), and ubiquitin ligases (E3). 17 One of the significant ubiquitin ligases is WWP1, which interacts with PY motif-containing proteins, targeting them for ubiquitination and proteasomal degradation. WWP1 is involved in various physiological processes through ubiquitination or protein-protein interactions. Dysregulation of WWP1 expression is associated with various pathophysiological conditions, including tumors. 18
According to literature, WWP1 is highly expressed in various tumor tissues, such as colorectal cancer 9 and hepatocellular cancer. 10 It functions as an oncogene and is closely associated with malignant phenotypes such as tumor invasion, metastasis, stemness, and drug resistance. Silencing WWP1 through RNAi induces apoptosis in these cells and inhibits cancer cell malignancy. 19
In NPC, WWP1 was overexpressed in its cells and tissues relative to the normal cells, indicating that WWP1 might also act as an oncogene in the NPC development. Nevertheless, the molecular mechanisms of WWP1 in NPC development remain unknown. Using several bioinformatics analyses, we have learned that there are 25 genes that might be closely related to WWP1 in the biological processes by evidence. These genes were enriched in a number of GO biological process terms, such as the transforming growth factor-beta receptor signaling pathway, the transmembrane receptor protein serine/threonine kinase signaling pathway, and the regulation of metabolism. Moreover, pathway enrichment analysis showed that these genes may be enriched in several pathways such as the TGF-β signaling pathway, the Ubiquitin mediated proteolysis, and the Wnt signaling pathway. These data indicated that the genes might be involved in cancer progression through various signaling pathways, and the mechanisms by which WWP1 promotes cancer development are very complicated. For instance, evidence showed that WWP1 inactivates a tumor suppressor, PTEN, to raise cancer risk. 20 WWP1 facilitates the progression of lung cancer via enhancement of epidermal growth factor receptor (EGFR) recycling, stability, and downstream activity because WWP1 can induce Lys689 ubiquitination in the EGFR juxtamembrane region. 21 Moreover, pathways such as STAT3 and MMP-2 might be involved in WWP1-induced tumor development. 22 Hence, WWP1 might promote cancer lymph node metastasis and recurrence through complex molecular mechanisms.
The log-rank test showed that high WWP1 expression may predict an unfavorable prognosis. Nevertheless, multivariate Cox regression analysis failed to show WWP1 expression as an independent indicator for NPC. Since lymph node metastasis and tumor recurrence are signs of tumor deterioration, WWP1 might play critical roles in the progression of NPC. Given that overexpression of WWP1 might play a crucial role in cancer development, it may also be regarded as a target for treatment. For example, WWP1 could be targeted by miR-129-5p in laryngeal cancer research, which lowered the invasion abilities of cancer cells. 23 Using a compound from cruciferous vegetables, indole-3-carbinol, WWP1 could be inhibited and PTEN could, in turn, be reactivated in cancer cells. 24 Bortezomib, a proteasome inhibitor approved for the treatment of multiple myeloma, can prevent prostate cancer tumorigenesis and bone metastasis by inhibiting WWP1, Smurf1, and Smurf2 in prostate cancer. 25 Then, the malignant phenotype of tumor was subsequently weakened. The evidence indicated that WWP1 in NPC might also be regarded as a potential target for cancer treatment. However, most inhibitors affect both normal and cancerous cells, leading to undesirable side effects and toxicity. Coupled with the emergence of drug resistance, many inhibitors lose their efficacy over time, further limiting their therapeutic efficiency. Therefore, it is important to develop low-toxicity and efficient inhibitors in the future.
Notably, WWP1 may have both oncogenic and tumor-suppressive roles. For instance, in a glioma xenograft mouse model, WWP1 overexpression inhibited cell malignancy and tumor growth. 26 In addition, WWP1 inhibited breast cancer cell migration and bone metastasis, while its knockdown promoted bone metastasis of breast cancer cells. 27 Therefore, the functions of WWP1 are multifaceted, and the underlying mechanisms require further elucidation in future studies.
Several limitations might be involved in the present study. First, both the sample size and the number of clinical features for the NPC cohort are limited. Thus, any selection bias might exist in the results. Future studies enrolling large sample sizes with detailed phenotype information are needed. Second, besides bioinformatics methods, only IHC was used for the detection of WWP1. Although WWP1 was knockdown in NPC cells and the changes in the cell malignant phenotypes were observed, in-depth molecular mechanisms have not been explored. For IHC, Immunostaining was performed manually. However, more consistency in staining results would be achieved with an automated stainer. Thus, an automated stainer is needed to ensure stability and consistency of results in future studies. Moreover, a polyclonal antibody was used in this study. Since monoclonal antibodies produce more stable results than polyclonal antibodies. In future studies, we will select monoclonal antibodies whenever possible. Third, further in vivo studies may help us better understand the precise mechanisms of WWP1-related cancer development.
Conclusion
The data indicated that overexpression of WWP1 was related to neck lymph node metastasis, tumor recurrence, advanced clinical stages, and poor outcomes, respectively, in NPC patients, suggesting the possible prognostic value of WWP1 and raising the possibility of WWP1 as a target for NPC treatment.
Footnotes
Author Contributions
Conception and design (HC, XZ). Analysis and interpretation of data (CL, SH). Statistical analysis (JL, HZ). Manuscript editing and revising (HC, XZ).
Availability of Data and Materials
Data are available on reasonable request. The data used to support the findings of this study are available from the corresponding author on request.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was partly supported by the National Natural Science Foundation of China (51963006), the Guizhou Science and Technology Project (ZK2022-044), and the Cultivation project of Affiliated Hospital of Guizhou Medical University (gyfybsky-2021-60).
Ethical Approval
A commercial tissue microarray was used in this study, which was approved by the Institutional Review Board of Shanghai Outdo Biotech Co., Ltd. (SHYJS-CP-1810011).
