Abstract
The biosynthesis of heparan sulfate (HS) proteoglycans occurs in the Golgi compartment of cells and will determine the sulfation pattern of HS chains, which in turn will have a large impact on the biological activity of the proteoglycans. Earlier studies in mice have demonstrated the importance of HS for embryonic development. In this review, the enzymes participating in zebrafish HS biosynthesis, along with a description of enzyme mutants available for functional studies, are presented. The consequences of the zebrafish genome duplication and maternal transcript contribution are briefly discussed as are the possibilities of CRISPR/Cas9 methodologies to use the zebrafish model system for studies of biosynthesis as well as proteoglycan biology.
Introduction
The diverse family of proteoglycans contains proteins substituted with one to more than hundred sulfated glycosaminoglycan chains, linear polysaccharides built up of repeating disaccharide units.1,2 The glycosaminoglycans found in proteoglycans include chondroitin sulfate (CS), dermatan sulfate (DS), keratan sulfate, heparan sulfate (HS), and heparin. Interestingly, although the majority of vertebrate proteins are glycoproteins, less than 100 proteins have been identified as proteoglycan core proteins. 3 The HS proteoglycans, which is the subject of this review (see Fig. 1), are present on the cell surface of most, if not all, vertebrate cells. In addition, HS proteoglycans are found in the extracellular matrix, particularly abundant in basement membranes.4,5 Heparin, with the same sugar backbone as HS but with a higher degree of sulfation, is found in mast cell granules. 6
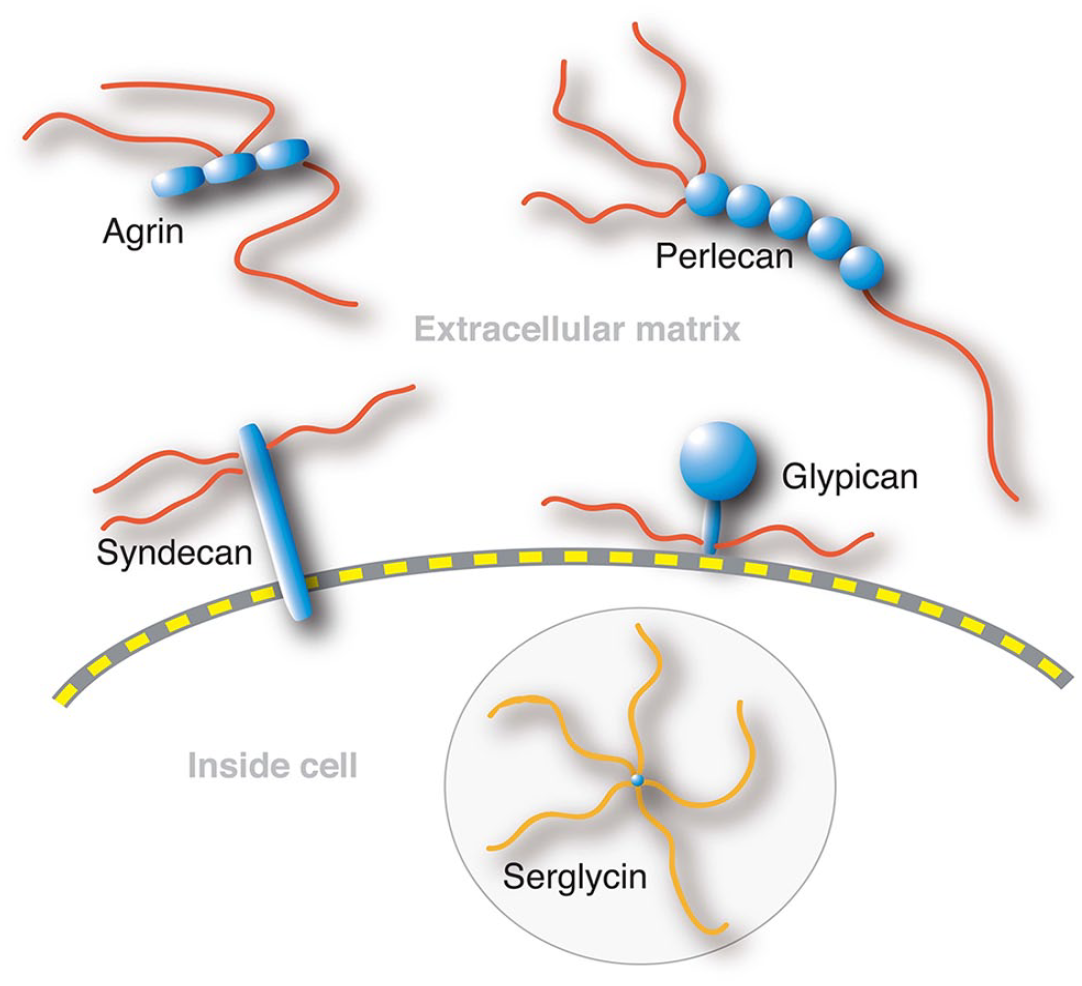
Heparan sulfate proteoglycans. Syndecans and glypicans are cell surface proteoglycans, whereas agrin and perlecan are present in the basement membranes. Serglycin with heparin chains is stored in mast cell granules.
The biological functions of HS proteoglycans rely, to a large extent, on interactions with proteins, where HS binding will modulate their functions. 7 The sulfation pattern of the HS chain will determine affinity for the protein ligands and is thus essential for function. The nature of the core protein and its level of expression will also affect HS function by influencing, for example, the localization of the HS chains (cell surface vs extracellular matrix) and the concentration of HS in these locations. During embryonic development, the abilities of heparan sulfate proteoglycans to act as coreceptors and to participate in the generation and maintenance of morphogen gradients are essential. Mouse embryos lacking HS die early during development, that is, at the time of gastrulation. 8
The sulfation pattern of HS is determined during biosynthesis in the Golgi compartment 9 ; however, extracellular editing by SULF 6-O-sulfatases will also change the HS structure after biosynthesis. 10 These enzymes, two in mammals and three in zebrafish (Sulf1, Sulf2a, Sulf2b), 11 will act on highly sulfated regions of the HS chain, removing 6-O-sulfate groups and thereby altering the affinity for several growth factors. 10
Zebrafish belongs to the ray-finned fishes (infraclass Teleostei) and has several advantages as a model organism. However, due to a whole-genome duplication in teleost fish, the number of genes encoding biosynthesis enzymes and proteoglycan core proteins are larger than in other vertebrates. 12 Although the expression of one of the duplicated genes may be lost (non-functionalization), both copies may be retained with similar or novel functions. The increased number of genes important for HS biosynthesis complicates studies in zebrafish. However, some of these genes have been non-functionalized (see Fig. 2 and below). Most of what we know about HS and heparin biosynthesis has been learnt in studies of rats and mice, in vivo and in cell cultures and also in cell-free systems with recombinant enzymes.9,13–15 However, there is no reason to believe that the basic mechanisms would be different in zebrafish, in particular because the structural features of zebrafish HS are similar to those of mammalian HS.16,17
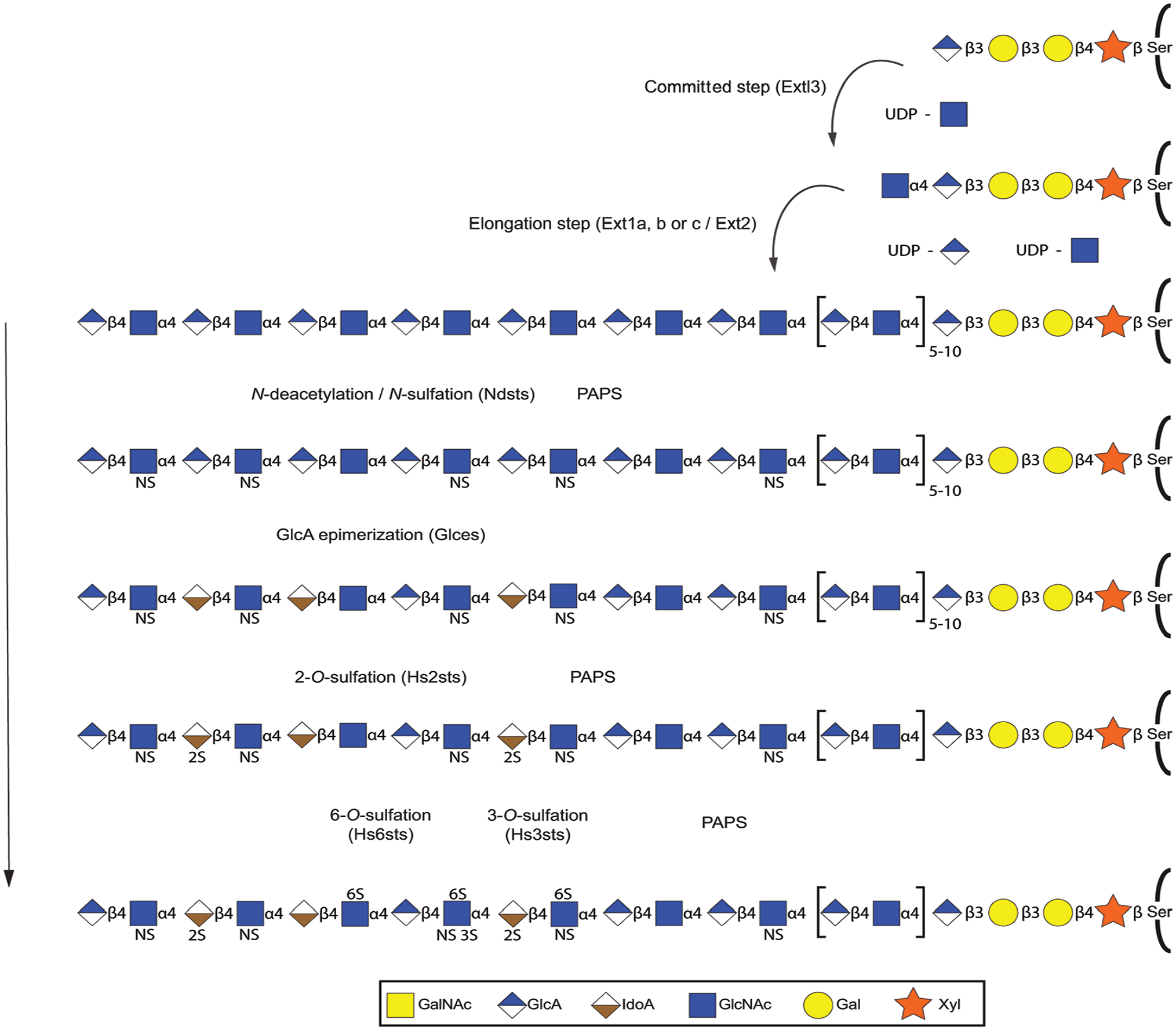
Polymerization and modification reactions during HS biosynthesis. After synthesis of the linkage region (see Fig. 3), the first HS-specific glycosyltransferase Ext13 adds a GlcNAc residue, followed by alternating additions of GlcA and GlcNAc by the HS copolymerase composed of one Ext1 (a, b, or c) and one Ext2 subunit. Note the dependence of UDP-sugars for these steps and the requirement for the sulfate donor PAPS for the following sulfation reactions. As the chain is growing, one or more of the bifunctional Ndst enzymes replace N-acetyl groups of selected GlcNAc residues with N-sulfate groups. Then follows C5-epimerization of GlcA residues into IdoA by Glce (a or b). This reaction occurs close to newly N-sulfated glucosamine residues, and the IdoA residues are the preferred substrate of the HS 2-O-sulfotransferases (1a or 1b). Then follows 6-O-sulfation by Hs6st enzymes (1a or 1b, 2, 3a or 3b) and 3-O-sulfation by one or more of the nine 3-O-sulfotransferases. Abbreviations: HS, heparan sulfate; GlcNAc, N-acetylglucosamine; GlcA, glucuronic acid; PAPS, 3′-phosphosadenosine-5′-phosphosulfate; IdoA, iduronic acid; NS, N-sulfate group; UDP, uridine diphosphate.
It Starts With a Core Protein
HS is synthesized in the Golgi compartment, where the growing HS chains are attached to serine residues in selected proteins. The major cell surface HS proteoglycans are syndecans and glypicans (see Fig. 1). Although the 6 mammalian glypican genes are paralleled by 10 zebrafish genes, 18 only three zebrafish syndecans have been identified compared with four in mammals. Among the extracellular matrix proteoglycans, single genes for perlecan and agrin are present in both mammalian and zebrafish genomes,19,20 whereas the two copies of the duplicated zebrafish collagen18a1 genes are expressed. 21 Serglycin, the core protein which in mammals carries heparin chains, is encoded by single genes both in zebrafish 22 and in mammals. The expression of core proteins will influence the location and amounts of HS and thereby affect the functional properties of the proteoglycans. Although the structure of HS primarily seems to depend on the type of cell responsible for biosynthesis, it has also been reported that the core protein may influence the outcome of biosynthesis.23,24
. . . and Biosynthesis of a Linkage Region
One or more serine residues in the core protein will serve as acceptor for the first sugar of the linkage region, a xylose, added by a xylosyltransferase (see Table 1 for enzymes participating in HS biosynthesis). Although the serine residue almost always is followed by a glycine, a consensus sequence for GAG attachment has so far not been possible to define. After xylosylation, two galactose residues, one at a time, will be transferred by two different galactosyltransferases. The fourth sugar, a glucuronic acid (GlcA), completes the linkage region. The same tetrasaccharide also functions as a linkage region in the biosynthesis of the glycosaminoglycans CS and DS. The galactose-xylose disaccharide residue can be transiently phosphorylated by Fam20b kinase, and the galactose units, at least in CS/DS GAG chains, may also be sulfated. 25 Although the role of sulfation is unclear, the transient phosphorylation of xylose appears to be essential for both HS and CS/DS chain elongation.26,27 A phosphatase removing the phosphate group on xylose has also been identified. 28 Zebrafish mutants for several of the enzymes taking part in the biosynthesis of the linkage region are available (see Table 2 and Fig. 3). Their phenotypes will reflect altered biosynthesis of both HS and CS/DS.
Genes Encoding Human and Zebrafish Heparan Sulfate Biosynthesis Enzymes.
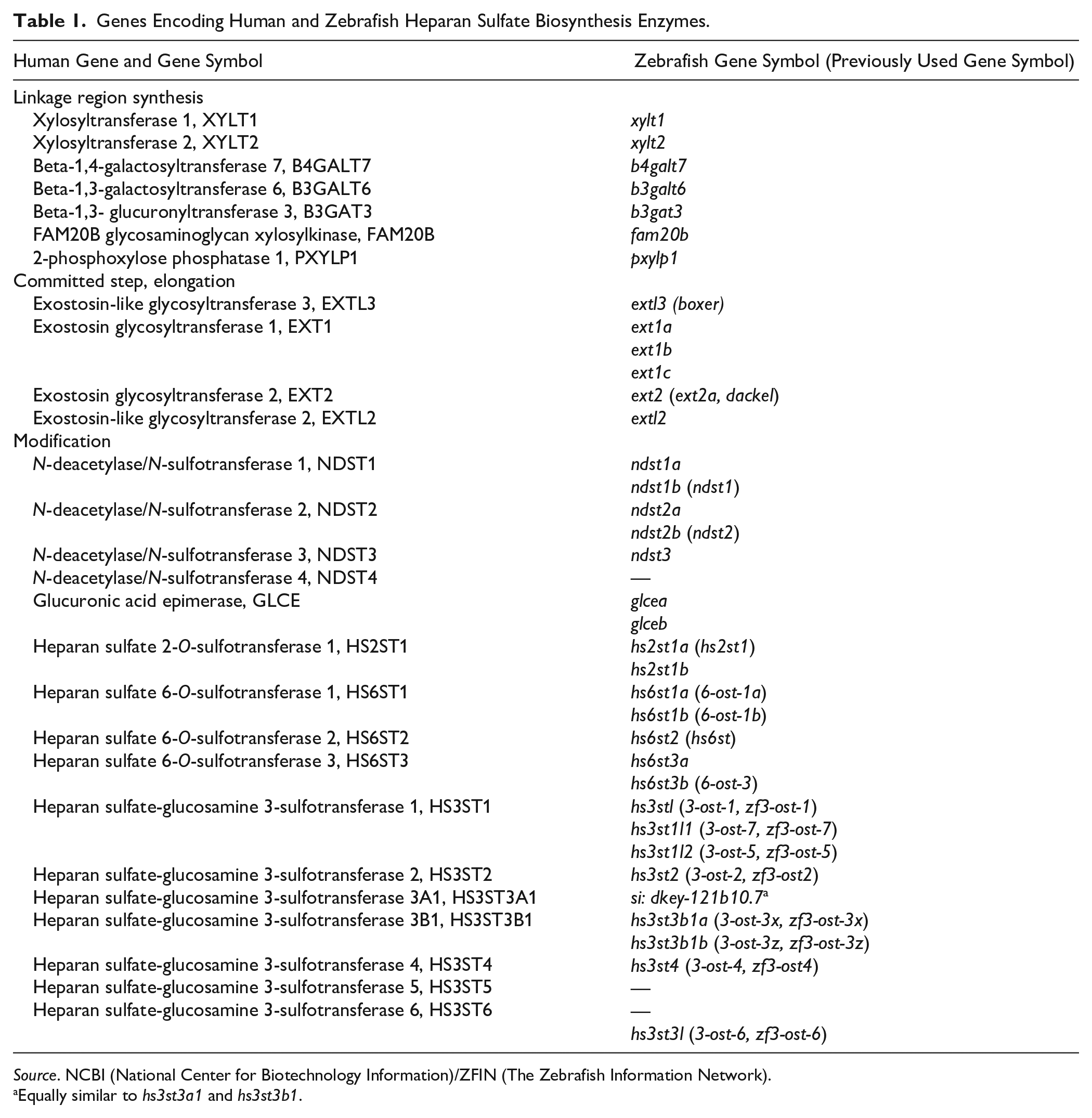

Source. NCBI (National Center for Biotechnology Information)/ZFIN (The Zebrafish Information Network).
Equally similar to hs3st3a1 and hs3st3b1.
Glycosaminoglycan Mutants.

ONIM indicates Online Mendelian Inheritance in Man; CNS, central nervous system; UDP, uridine diphosphate.
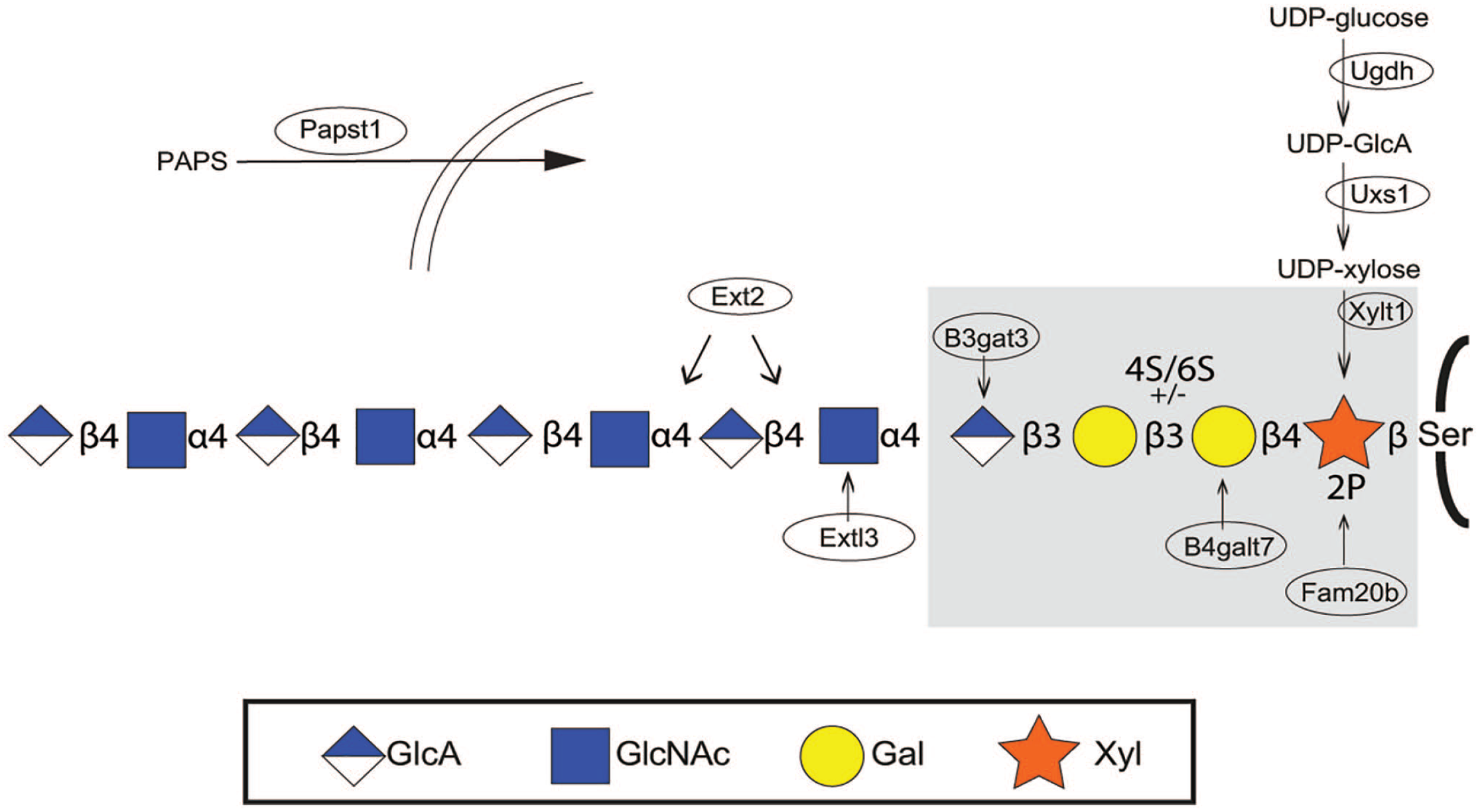
Glycosaminoglycan mutants. Most of the zebrafish glycosaminoglycan mutants affect biosynthesis of the common HS and CS/DS linkage region (shaded in the Figure). XYLT1 is one of the two xylosyltransferases adding xylose to selected serine residues in the core protein. Phosphorylation of xylose is carried out by Fam20b, for which a mutant is available. This is also the case for the enzymes responsible for the synthesis of UDP-xylose (Uxs1) and its precursor UDP-GlcA (Ugdh). Lack of UDP-GlcA will also affect polymerization of the glycosaminoglycan chains (see Fig. 2). Mutants are also available for two of the three enzymes completing the linkage region, B4galt7 adding the first galactose unit, and B3gat3 adding the fourth sugar, a GlcA. The two HS-specific enzymes for which mutants are available are Ext13 and Ext2, affecting HS polymerization (see Fig. 1). The ninth mutant affects transport by Papst1 of the sulfate donor PAPS into the Golgi compartment. Enzymes for which mutants have been isolated are placed within ellipses. Abbreviations: HS, heparan sulfate; CS, chondroitin sulfate; DS, dermatan sulfate; GlcA, glucuronic acid; PAPS, 3′-phosphosadenosine-5′-phosphosulfate; GlcNAc, N-acetylglucosamine; UDP, uridine diphosphate glucose.
. . . Followed by a Committed Step
Although most GAG attachment sites are specific for HS or CS/DS, respectively, some are not; an example is serglycin which in many cells is a CS proteoglycan, but in connective tissue–type mast cells it is substituted with heparin chains 6 ; another example is perlecan where the C-terminal GAG attachment site can be substituted with either HS or CS. 50 However, transfer of an N-acetylglucosamine (GlcNAc) residue to the tetrasaccharide linkage region by the exostosin-like glycosyl transferase 3 (Extl3) will commit the biosynthesis to HS (or heparin). Accordingly, HS amounts are reduced in extl3 mutant zebrafish larvae,16,17 discussed further below.
Elongation and Modification Occur Together
The HS chain is polymerized through alternating transfer of GlcA and GlcNAc by the HS copolymerase composed of exostosin glycosyltransferase 1 and 2 (Ext1/Ext2). At the same time as the chain is growing, it is modified, the first modification being N-deacetylation followed by N-sulfation of GlcNAc residues. Most subsequent modifications occur in N-sulfated regions of the GAG chain and starts with epimerization of GlcA into iduronic acid (IdoA), followed by 2-O-sulfation of IdoA. After epimerization and 2-O-sulfation follow 6-O-sulfation and 3-O-sulfation of GlcNAc and its N-sulfated counterpart (see Fig. 2). All sulfation reactions rely on the presence of the universal sulfate donor 3′-phosphosadenosine-5′-phosphosulfate (PAPS).
Multiple Zebrafish Enzymes
Although the zebrafish genome contains a single ext2 gene, three ext1 orthologs have been identified (see Table 1). As efficient polymerization of HS requires both Ext1 and Ext2, the single Ext2 would be expected to be necessary for HS biosynthesis. Interestingly, an ext2 mutant is available with multiple severe phenotypes (see below, Table 2 and Fig. 4). In mice, gene knockouts of Ext2 and Ext1 are lethal.8,48 Maternal mRNA contribution is most likely the reason why the phenotype is milder in the zebrafish mutants. In addition to Ext1, Ext2, and Extl3, a fourth glycosyltransferase, Extl2, may also affect HS biosynthesis. Mice lacking this glycosyl transferase are viable and fertile, 51 but this enzyme has not been studied in zebrafish.
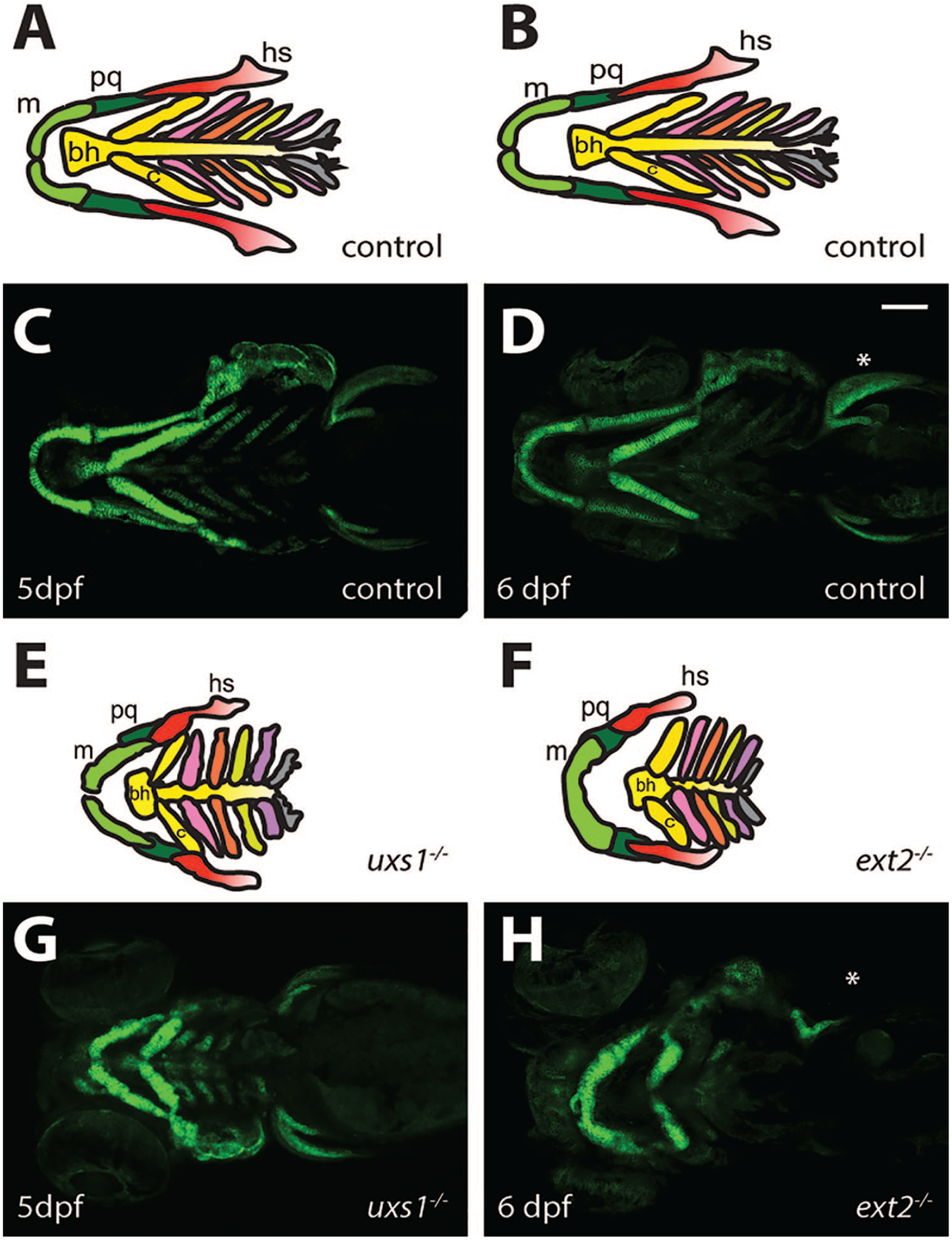
Pharyngeal cartilage morphology. Many zebrafish proteoglycan mutants develop malformations in the pharyngeal cartilage. Schematics of control larvae at 5 (A) and 6 (B) dpf give an overview of the organization of the pharyngeal cartilage elements seen in the ventral views of Tg(−1.7col2a1a:EGFP-CAAX) control larvae (C, D). uxs1−/− (E, G) and ext2−/− (F, H) larvae have disrupted intercalation of chondrocytes, resulting in abnormal-shaped pharyngeal cartilage elements. In addition, ext2−/ larvae lack pectoral fins (*).Scale bar, 100 µm. Abbreviations: m, Meckel’s cartilage; pq, paloquadrate; hs, hyosymplectic; bh, basohyal; c, ceratohyal; dpf, days post fertilization.
Bifunctional Ndst (N-deactylase/N-sulfotransferase) enzymes, five in zebrafish 52 (see Table 1), will form N-sulfated domains on the growing HS chain. Of the five, two are Ndst1 orthologs and two are Ndst2 orthologs. The fifth zebrafish ndst (ndst3) is most likely an ortholog of the Ndst gene that in mammals through a local gene duplication gave rise to Ndst3 and Ndst4. 52 All ndst transcripts are maternally contributed and widely expressed, the spinal cord being an exception; here, only ndst3 transcript could be detected. 52
The single mammalian C5-epimerase (Glce) and 2-O-sulfotransferase each have two orthologs in zebrafish. 53 Five zebrafish 6-O-sulfotransferase and nine 3-O-sulfotransferase genes have been identified in the zebrafish genome compared with three and seven in mammals.54,55 Although the distribution of the modification enzymes (sulfotransferases and epimerases) has been studied on the transcript level,52–55 little is known about the contribution of the different isoforms to HS synthesis or their enzymatic characteristics. Some HS biosynthesis enzymes, for example, Ext2 and Extl3, are ubiquitously expressed in most tissues during zebrafish embryo development, 16 whereas others, such as members of the 3-O-sulfotransferase family, show a distinct spatial and temporal expression. 55 Enzymes with such regulated expression could possibly have instructive functions for HS, changing the affinities of the polysaccharide for binding proteins.
Glycosaminoglycan Biosynthesis Mutants
Forward genetic screens where mutagens have been used to randomly introduce mutations in the zebrafish genome have been one way of linking specific genes to their function. In this manner, nine genetic mutants with defective biosynthesis of glycosaminoglycans have been identified (Fig. 3 and Table 2; see Habicher et al. 56 ). Slc35b2 is one of the two transporters of the sulfate donor PAPS from the cytoplasm into the Golgi compartment, affecting all sulfation reactions in this organelle; uridine diphosphate glucose (UDP)-glucose dehydrogenase (Ugdh) converts UDP-glucose into UDP-GlcA, needed for the biosynthesis of sulfated glycosaminoglycans and hyaluronan, possibly affecting also detoxification reactions in the liver; UDP-glucuronate decarboxylase 1 (Uxs1) makes UDP-xylose from UDP-GlcA necessary for the biosynthesis of the shared HS and CS/DS linkage region. Synthesis of the linkage region will also be affected by the four mutants fam20b, xylt1, b4galt7, and b3gat3. Cartilage defects and in particular abnormal jaw and pharyngeal cartilage structures (see Fig. 4) are phenotypes strikingly consistent in all identified mutants affecting both HS and CS/DS biosynthesis (Table 2).
Finally, ext2 and extl3 mutations will directly influence HS biosynthesis only. These two mutants show reduced pectoral fins, 57 deformed pharyngeal cartilage 17 (Fig. 4), defective sorting of axons in the optical tract, 16 and defective thymopoiesis. 58 Interestingly, although the defects are qualitatively similar, the phenotypes are more severe in the ext2 mutant, possibly caused by lower amounts of maternal ext2 mRNA in the early embryonic stage. When examined 6 days post fertilization (dpf), HS levels were reduced in both mutants; the reduction was most pronounced in the ext2 mutant which contained less than 20% of the control levels. 17 The low levels of HS synthesized were highly modified in both mutants, possibly reflecting a higher ratio of HS modification enzymes relative to the available HS polysaccharide substrate. Interestingly, extl3 mutants contained elevated levels of CS, 17 tentatively as a result of a compensatory mechanism to replace the lost HS. This phenomenon has also been documented in HS-deficient CHO-cells and in embryonic stem cells with altered HS production, and also in vivo in cartilage of mouse embryos with defective HS biosynthesis.8,59–61 A similar increase in CS amounts was not seen in the ext2 mutant, and it was speculated that the “committed step,” glycosyltransferases adding GlcNAc for HS (Extl3) or GalNAc for CS biosynthesis, could be critical in determining HS and CS/DS amounts. The more severe cartilage phenotype of the ext2 than the extl3 mutant could be another explanation.
Future Perspective
Much is yet to be known about HS biosynthesis, and the zebrafish model system will be a powerful tool in these studies. In addition to studies of mutants, morpholino knockdown experiments blocking either translation or splicing of specific genes have previously been used to give valuable information about gene function. However, usage of morpholinos has been linked to unspecific upregulation of the p53 pathway, leading to increased levels of apoptotic neural death and defects within craniofacial cartilage. 62 The more recent CRISPR/Cas9 technology has now provided the genome engineering field with an efficient tool for generating targeted mutations. 63 The method is based on the usage of a guide RNA (gRNA) and the Cas9 endonuclease complex introducing a double-stranded break at the site of gRNA base pairing to the genomic DNA. Non-homologous end joining (NHEJ), a genomic DNA repair system prone to mistakes, frequently causes frameshift mutations, leading to loss-of-function of the targeted genes. The CRISPR/Cas9 technologies further open up for virtually endless possibilities for studies of “humanized” transgenic zebrafish (i.e., where mutations similar to human patients are introduced) and to tag and exogenously express enzymes to monitor phenotypic changes. In addition, the technique is increasingly used for specific labeling of distinct cell types by the expression of fluorescent proteins and genetic fingerprints, which significantly increases our ability to detect phenotypic changes of HS mutants.
Comparisons of CRISPR/Cas9 knockout phenotypes with those obtained using morpholino knockdown approaches can now be applied to confirm (or modify) the conclusions drawn from morpholino studies. Some of the diverging results between knockouts and knockdowns may tentatively also be explained by nonsense-mediated decay of mRNAs containing CRISPR/Cas9-generated premature stop codons, resulting in selective upregulation of genes with similar sequence.64,65 In this case, gene knockdown with morpholinos where no genetic compensation occurs may result in a more severe phenotype than knockout of the same gene.
Gene knockouts of the HS biosynthesis enzymes one by one or in combination would make it possible to generate a “living library” of fish or larva with different HS (or CS) structures for studies of both biosynthesis, gene function and pathology. In combination with single cell analysis techniques and advanced imaging techniques now available, there will be vast possibilities to increase our knowledge in the proteoglycan area.
Footnotes
Author Contributions
All authors contributed to the literature review and manuscript preparation.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research work performed by the authors was supported by The Swedish Cancer Society, SciLifeLab, and Stiftelsen för Proteoglykanforskning at Uppsala University.
