Abstract
Optic development involves sequential interactions between several different tissue types, including the overlying ectoderm, adjacent mesoderm, and neural crest mesenchyme and the neuroectoderm. In an ongoing expression screen, we identified that Tfap2β, Casq2, Penk, Zic1, and Zic3 are expressed in unique cell types in and around the developing eye. Tfap2β, Zic1, and Zic3 are transcription factors, Casq2 is a calcium binding protein and Penk is a neurotransmitter. Tfap2β, Zic1, and Zic3 have reported roles in brain and craniofacial development, while Casq2 and Penk have unknown roles. These five genes are expressed in the major tissue types in the eye, including the muscles, nerves, cornea, and sclera. Penk expression is found in the sclera and perichondrium. At E12.5 and E15.5, the extra-ocular muscles express Casq2, the entire neural retina expresses Zic1, and Zic3 is expressed in the optic disk and lip of the optic cup. The expression of Tfap2β expanded from corneal epithelium to the neural retina between E12.5 to E15.5. These genes are expressed in similar domains as Hedgehog (Gli1, and Ptch1) and the Wnt (Lef1) pathways. The expression patterns of these five genes warrant further study to determine their role in eye morphogenesis:
Introduction
Vertebrate eye development occurs as a series of highly coordinated induction events, with the lens vesicle, optic cup, optic stalk (OS), and precursor to the cornea all developing from the forebrain neuroectoderm.1,2 The onset of eye development occurs in tandem with the start of gastrulation, when the optic sulci are present in the anterior neural plate.1,2 The walls of the diencephalon then form the optic vesicles, which undergo coordinated invagination with the lens placode to form the lens vesicle and optic cup.1–3 The inner layer of the optic cup gives rise to the neural retina. 3 In contrast to the inner layer, the outer layer of the optic cup matures into the retinal pigmented epithelium (RPE) and OS. Elsewhere in the developing eye, the lens vesicle gives rise to the lens, whereas its overlying ectoderm matures into the cornea. In contrast, the iris, sclera, and ciliary body mature from the neural crest mesenchyme.1,4
In an ongoing RNASeq screen in the laboratory focusing on skull morphogenesis, we determined that the expression of five genes (Tfap2β, Casq2, Penk, Zic1, and Zic3) were also expressed in the optic region. Transcription factor Activating Enhancer-Binding Protein 2 Beta (Tfap2β), an alternative splice form of Tfap2, is mainly expressed in epithelial and neural crest cell lineages during craniofacial and skeletal development.5,6 Conditional deletion of Tfap2β in mouse cranial neural crest results in postnatal ocular defects typified by opacity. 7 The Calsequestrin (Casq2) is a Ca2+-binding protein in myocardial sarcoplasmic reticulum; Casq2 variants lead to abnormal muscle development and cardiac function. 8 Casq2 expression is initially found in cardiac muscle, but later its expression domains include the myotome and cardiac muscle. 9 Proenkephalin (Penk) gene encodes small endogenous opioid peptides, which are expressed in various tissues, including the nasal region, and Penk plays a critical role in cell proliferation and differentiation.10–13 Zic1 and Zic3 belong to zinc finger protein of the cerebellum family of transcription factors and are widely expressed, but spatially restricted, from gastrulation through organogenesis.14–16 Zic1 and Zic3 are expressed in peripheral margin of the optic cup.15,16
Several signaling pathways are involved in the eye development. Gli1 is expressed in the RPE, POM, and ventral eye cup. 17 Canonical Wnt/β-catenin pathway is active throughout retinal development. 18 Interestingly, in Tfap2β null mice, the expression of DKK1 is reduced in corneal epithelium, implying that Tfap2β may be involved in the Wnt signaling pathway. 18
While the role of Tfap2β, Casq2, Penk, Zic1, and Zic3 are well documented in other organs, their specific roles in the developing eye are less apparent. In this study, we describe the complimentary, expression patterns of Tfap2β, Casq2, Penk, Zic1, and Zic3 in the early stages of eye development. We also propose a pattern of expression for Zic1, and Zic3 that is similar, but more restricted then those previously documented in the eye.15,16 Our expression data provide new genes that are expressed in discrete optic regions that will aid in understanding vertebrate eye development.
Materials and Methods
Animals
Embryos were collected from timed pregnant CD-1 females purchased from Charles River (E12.5 and E15.5) or C57/Bl6J females (E10.5 and E16.5). The pregnant dams were euthanized by CO2, and the embryos were collected via C-section on the appropriate day of pregnancy. We confirmed the stage of the embryos with morphology. All animal husbandry procedures and protocols were approved by and performed under the oversight of the University of Pittsburgh Institutional Animal Care and Use Committee.
Antisense RNA Probe Preparation
We homogenized E15.5 skulls and extracted the RNA using the Trizol reagent (Invitrogen, Waltham, MA) according to standard procedures. SuperScript Vilo (Invitrogen) was used to reverse transcribe the RNA into cDNA. Using the cDNA template, we amplified fragments of Casq2, Penk, Zic1, and Zic3 using the primers in Table 1. 13 The fragments were cloned into the dual-promoter TOPO-TA cloning kit (Thermo Fisher, Waltham, MA) and purified using the Plasmid Midi Kit (QIAGEN, Germantown, MD). To ensure that we cloned the appropriate genes, we sequenced all plasmids to verify sequence identity and orientation of each fragment prior to probe synthesis. Dr. Trevor Williams kindly provided the Tfap2β clone. 19 All antisense digoxigenin-labeled RNA probes were transcribed from linearized clones using the appropriate RNA polymerase.
Primers Used to Clone and Generate Antisense in situ Probes for the Genes.

Amplicon size (bp).
Wan et al. 13
Histology and Section in situ Hybridization
Embryos were fixed in 4% paraformaldehyde at 4C overnight and embedded and infiltrated in paraffin using standard protocols. The embryos were cut in 8-µm sections and adjacent sections were placed on TESPA (triethoxysilane)-coated Superfrost Plus slides. Sections for in situ hybridization were selected based on the histology in the neutral fast red stained adjacent sections. For nuclear fast red staining, the slides were deparaffinized, rehydrated, and stained in 0.1% nuclear fast red (Sigma, St. Louis, MO). Following staining, the slides were dehydrated, mounted in Richard-Allen Scientific Mounting Media (Fisher) and coverslipped. The section in situ hybridization were performed according to standard protocols and detected with BM Purple. 13 The development reaction was stopped after appropriate signal had developed and the slides were counterstained with eosin, dehydrated, mounted in Richard-Allen Scientific Mounting Media (Fisher), and coverslipped. We observed very little variation in the expression domains within the three biological replicates that we analyzed; the images are representative of the three individuals for each stage and probe.
Microscopy
Histological and section in situ images were collected on a Zeiss Axioskop A1 microscope with the MrC3 camera and Zen imaging software. Image composites of the entire optic region at E12.5 and E15.5 were stitched together using the automatic settings in Zen software. The figures were assembled in Adobe Photoshop.
Results
Tfap2β, Casq2, Penk, and Zic1 are Expressed in Distinct Spatial Domains at E10.5
At E10.5, the optic cup has been induced and the lens placode is being induced and the neural crest cells are being excluded between the corneal epithelium and the optic cup. 20 We observed the expression of Tfap2β expressed in the superficial and corneal epithelium as well as the neural crest cells adjacent to the eye (Fig. 1B). Casq2 expression is found peppered through the neuroepithelium, as well as in the mesenchyme that will be invaded by the eyelid epithelia bud. At E10.5, we observed little expression of Penk (Fig. 1C and D). Both Zic1 and Zic3 are robustly expressed in the OS and cup (Fig. 1E and F).
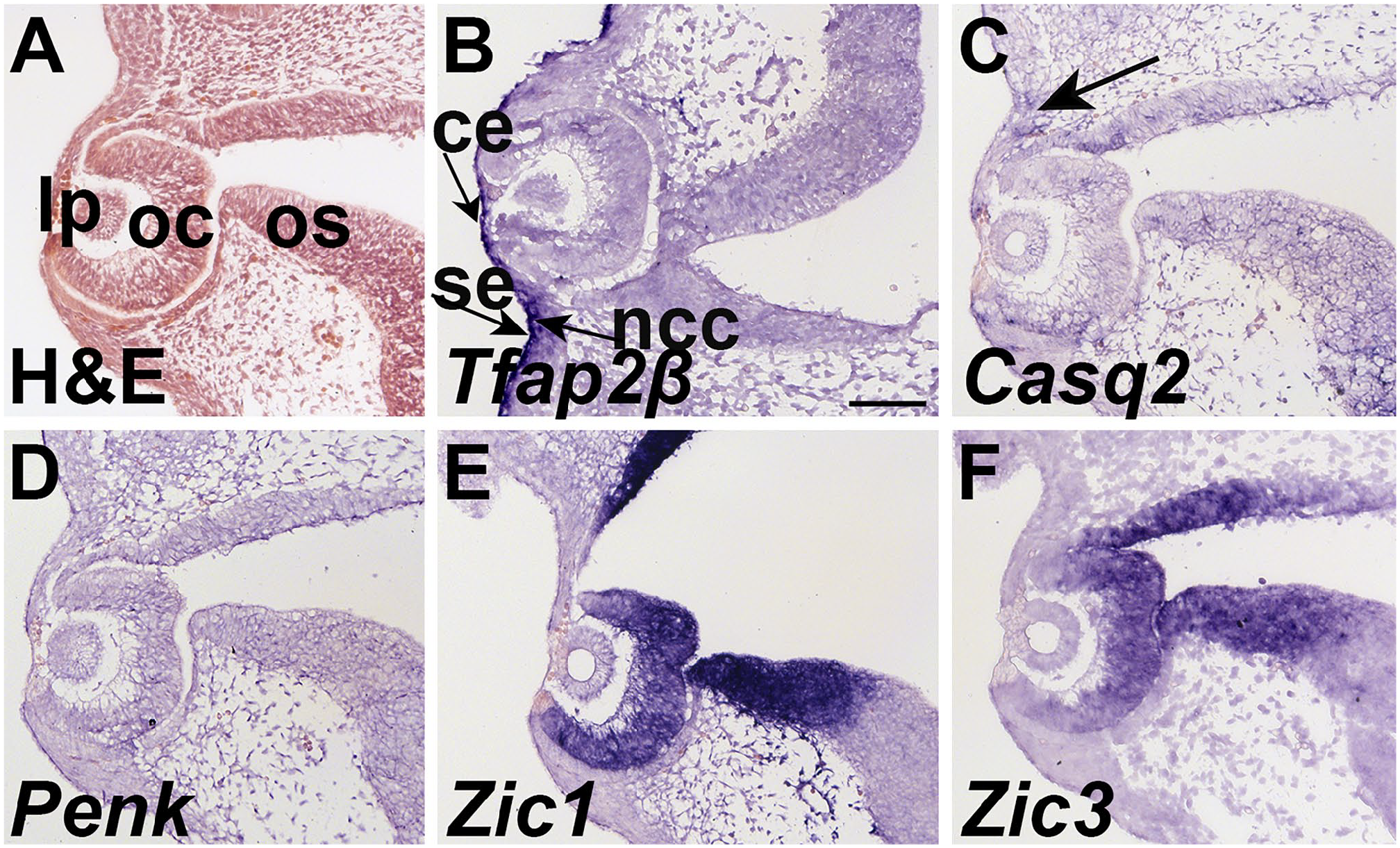
Tfap2β, Casq2, Penk, and Zic1 are expressed in and surrounding the eye at E10.5. Transverse sections through the eye region at E10.5 hybridized to (B) Tfap2β, (C) Casq2, (D) Penk, (E) Zic1, and (F) Zic3. (A) H&E staining shows the histology of the ocular region. (B) Tfap2β is expressed in the mesenchyme and epithelium. Tfap2β is expressed in the eyelid epithelium and is also present in the condensed mesenchyme of the future eyelid. Tfap2β expression is found in the superficial and corneal epithelium. (C) Casq2 is expressed in a subset of the mesenchyme near the lid (black arrow). Casq2 expression is also peppered through the optic stock and cup. (D) Penk expression is absent from the ocular region at E10.5. (E) Zic1 is expressed in the optic cup, and the lateral edge of the optic stock. (F) Zic3 is also expressed in the optic cup and stalk. Scale bar in A = 100 μm and applies to all. Abbreviations: lp, lens placode; oc, optic cup; os, optic stalk; ce, corneal epithelium; se, superficial epithelium; ncc, neural crest cells.
Tfap2β, Casq2, Penk, and Zic1 are Expressed in Distinct Spatial Domains at E12.5
At E12.5, when most of the tissues of the eye are present. Tfap2β is expressed in the corneal epithelium and mesenchyme. The tissue in the hyaline cavity expresses Tfap2β, but it is not expressed in the optic cup of the retina (Fig. 2B). Tfap2β is expressed in the epithelium of the eyelids and is absent from the base of the groove (Fig. 2B, arrowhead). Casq2 expression was detected in the extrinsic ocular muscles (Fig. 2C). Penk is expressed in periocular mesenchyme as well as frontal bone primordium (Fig. 2D) but is absent from the globe and associated tissues. Zic1 is expressed in the neural retina and optic nerve (Fig. 2E). Zic3 is not expressed in tissues surrounding the eye at E12.5 (Fig. 2F).
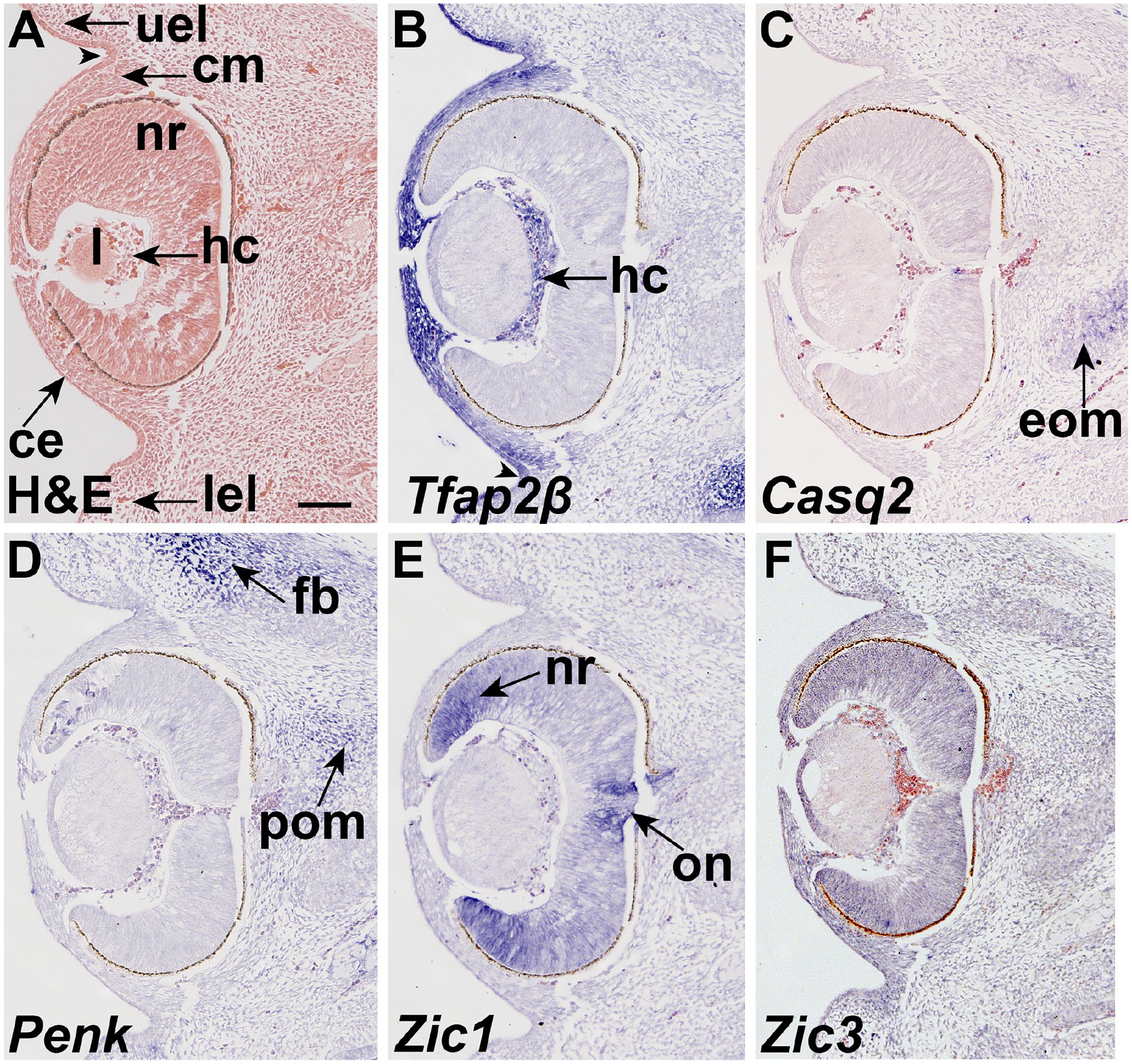
Tfap2β, Casq2, Penk, and Zic1 are expressed in and surrounding the eye at E12.5. Coronal sections of an E12.5 embryo hybridized to antisense probes to (B) Tfap2β, (C) Casq2, (D) Penk, (E) Zic1, and (F) Zic3. (A) Nuclear fast red staining shows the histology of the ocular region. (B) Tfap2β is expressed in the mesenchyme and epithelium. Tfap2β is expressed in the eyelid epithelium and is absent from the base of the groove (arrowhead). Tfap2β expression is found in the corneal epithelium and mesenchyme, as well as tissues within the hyaloid cavity. (C) Casq2 is expressed in the extrinsic ocular muscles (black arrow). (D) Penk is expressed in periocular mesenchyme (arrow) as well as frontal bone primordium (arrow). (E) Zic1 is expressed in the neural retina (black arrow), and the optic disk surrounding the optic nerve (arrow). (F) Zic3 is expressed in neural retina (nr) at E12.5. Scale bar in A = 100 μm and applies to all. Abbreviations: ce, corneal epithelium; cm, corneal mesenchyme; eom, extrinsic ocular muscles; fb, frontal bone primordia; hc, hyaline cavity; l, lens; lel, lower eye lid bud; nr, neural retina; on, optic nerve; pom, peri-ocular mesenchyme; uel, upper eye lid bud.
Wnt and Hedgehog Signaling Effectors Have Restricted Expression Patterns Throughout E10.5–E12.5
Since we observed that Tfap2β, Casq2, Penk, and Zic1 genes expressed in discrete anatomical regions of the developing eye, we next determined if their expression domains overlapped with signaling pathways that are required for eye development. Lef1 is a transcriptional target of Wnt signal pathway, and we used its expression domain to define where active Wnt signaling is occurring. At E10.5, Lef1 is mainly expressed in the upper eye lid (uel) and os (Fig. 3A). The expression of Lef1 is maintained in the eyelid bud mesenchyme at E12.5. The expression of Lef1 in OS develop into the corneal and periocular mesenchyme (Fig. 3D).
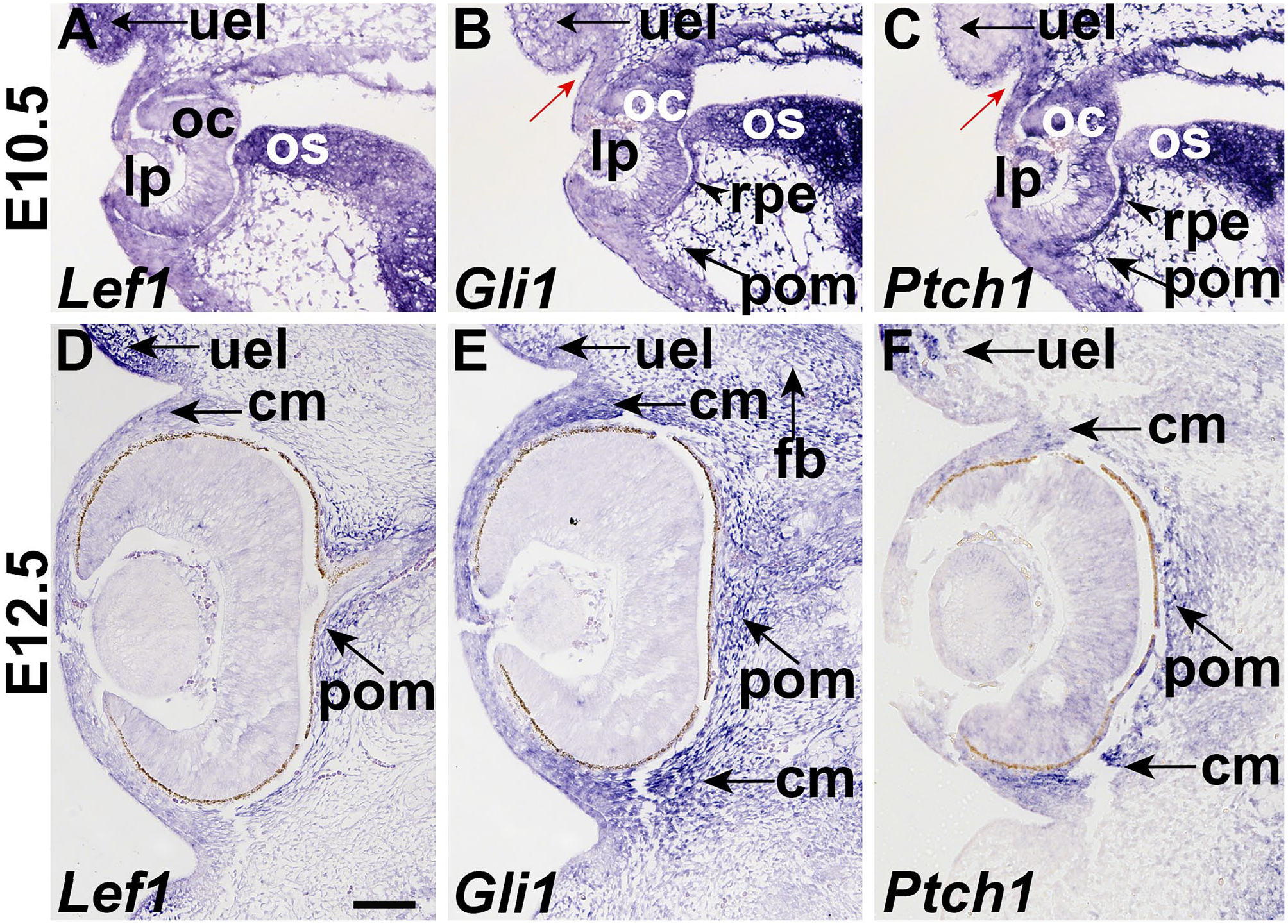
Hh and Wnt signaling is found in the periocular mesenchyme. Section in situ hybridization of (A, D) Lef1, (B, E) Gli1, and (C, F) Ptch1 in coronal sections of E10.5 (A-C) and E12.5 (D-F) skull. Refer to Figs. 1 and 2 for histology of adjacent sections. (A) Lef1 is expressed in the upper eye lid (uel), lens placode (lp), optic cup (oc), and optic stalk (os). (B) Gli1 is expressed in the invaginating eyelid epithelium (blue arrow), and the condensed mesenchyme adjacent to it. In addition, Gli1 expression is found in the optic cup (oc), optic stalk (os), retinal pigmented epithelium (rpe), and periocular mesenchyme (pom). (C) Ptch1 is expressed in early eyelid (blue arrow) and is excluded from the adjacent condensed mesenchyme. Ptch1 expression is found in the optic cup, optic stalk, pigmented retinal epithelium and periocular mesenchyme (arrow). (D) Lef1 was expressed in the eyelid bud mesenchyme (arrow), and periocular mesenchyme adjacent to the optic nerve (arrow). (E) Gli1 is expressed in the eyelid (arrow), corneal and periocular mesenchyme as well as the frontal bone. (F) Ptch1 is expressed in the eyelid, periocular and corneal mesenchyme (arrow). Scale bars in A = 100 μm and applies to all. Abbreviations: cm, corneal mesenchyme; fb, frontal bone primordia; lp, lens placode; oc, optic cup; os, optic stalk; pom, peri-ocular mesenchyme; rpe, pigmented retinal epithelium; uel, upper eye lid bud.
To identify where active Hedgehog (Hh) signaling is occurring we assayed the expression of Gli1, a transcription factor that is a target and activator of the pathway, and Ptch1, a Hh co-receptor, that is a target and inhibitor of the pathway. At E10.5, Gli1 and Ptch1 are broadly expressed in lens placode (lp), optic cup (oc), os, rpe, and periocular mesenchyme (pom). Gli1 is expressed in mesenchyme of upper eye lid (uel), while Ptch1 is in surface mesenchyme of upper eye lid (uel). At E12.5, Gli1 is broadly expressed in the eyelid bud mesenchyme, corneal mesenchyme, and periocular mesenchyme as well as the frontal bone primordium (Fig. 3E). Ptch1 is expressed in a few cells in the lens, the early eyelid bud and periocular mesenchyme (Fig. 3F).
Tfap2β, Casq2, Penk, Zic1, and Zic3 Have Discrete Expression Patterns at E15.5
At E15.5, Tfap2β was still expressed in the epithelium surrounding the eyelid, and bulbar conjunctiva mesenchyme adjacent to the corneal epithelium. Tfap2β is expressed in innermost layer of the outer nuclear layer of the retina, and a few cells in the outermost layer of the neural retina (Figs. 4B and 5B). Tfap2β is expressed in the corneal endothelium at E15.5 (Figs. 4B and 5B). Casq2 expression is continues to be expressed in the extrinsic ocular muscles (Figs. 4C and 5C). Casq2 is also expressed in the skeletal muscle surrounding the femur (Fig. 5D). Penk is detected in the sclera adjacent to the pigmented retinal epithelium as well as the perichondrium of the nasal cavity and sparsely through the bulbar conjunctiva (Figs. 4D and 5E; Wan et al. 13 ). Zic1 expression is detected in the outer nuclear layer of the retina, especially in the optic disk and the lip of the optic cup (Figs. 4E and 5F). We detected Zic3 expression at E15.5, only expressed in the optic nerve and lip of the optic cup (Figs. 4F and 5G).
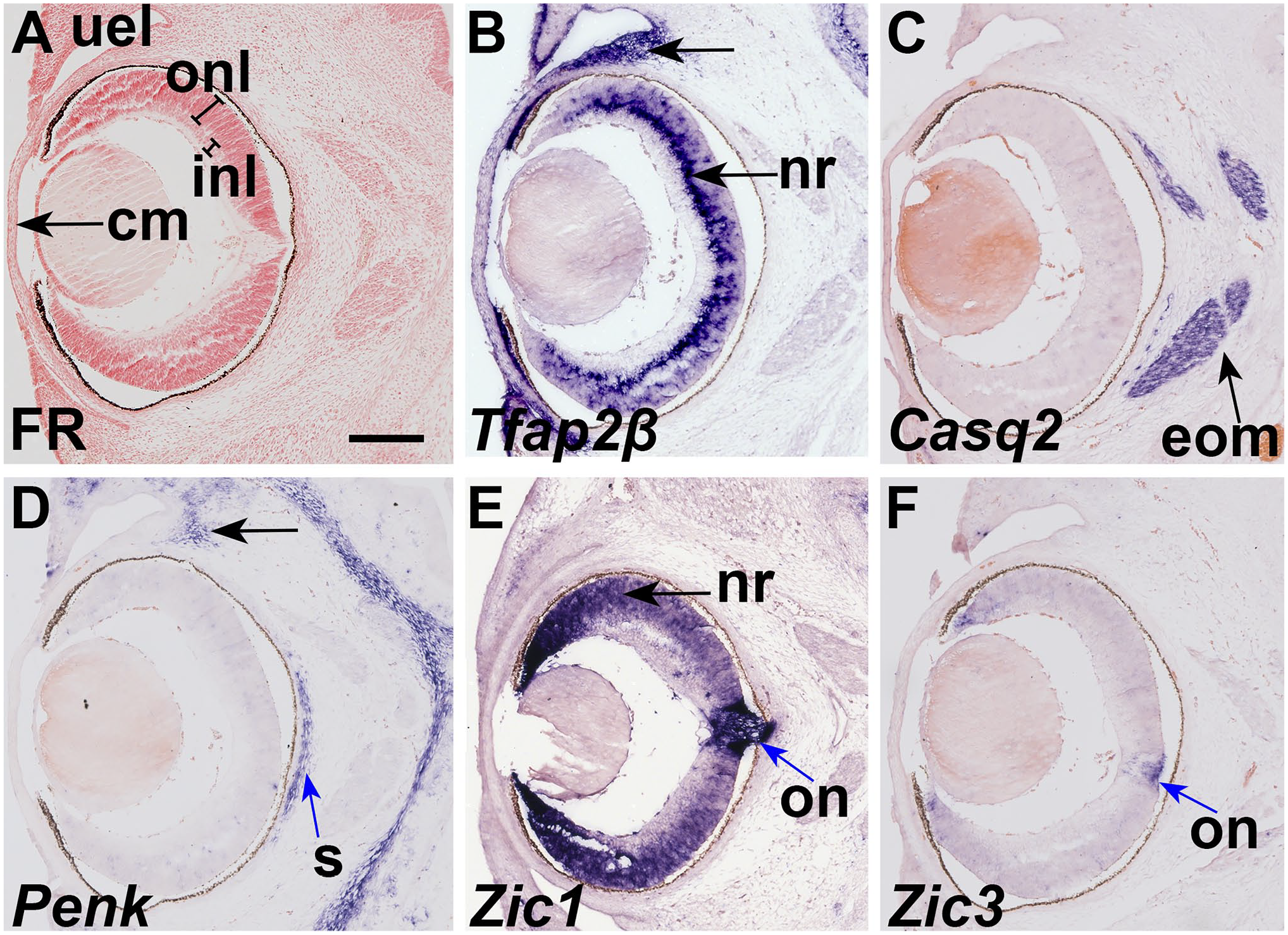
Tfap2β, Zic1, Zic3, Penk, and Casq2 expression define the neuronal, skeletogenic, and muscular cells in the E15.5 optic region. In situ hybridization of E15.5 coronal sections with probes to (B) Tfap2β, (C) Casq2, (D) Penk, (E) Zic1, and (F) Zic3. (A) Nuclear fast red staining of an adjacent section. (B) Tfap2β is expressed in the epithelium of the early eyelid (arrow), innermost layer of the outer layer of the neural retina (arrow), and the bulbar conjunctiva (arrow). (C) Casq2 is expressed in the extrinsic ocular muscles (arrow). (D) Penk is expressed in the sclera (blue arrow) and bulbar conjunctiva (black arrow). (E) Zic1 is expressed in the neural retina (arrow), strong in the optic nerve (blue arrow) and optic disk. (F) Zic3 is expressed in the optic nerve (blue arrow). Scale bars = 100 μm. Abbreviations: cm, corneal mesenchyme; eom, extrinsic ocular muscles; inl, inner nuclear layer; on, optic nerve; onl, outer nuclear layer; nr, neural retina; s, sclera; uel, upper eye lid bud.
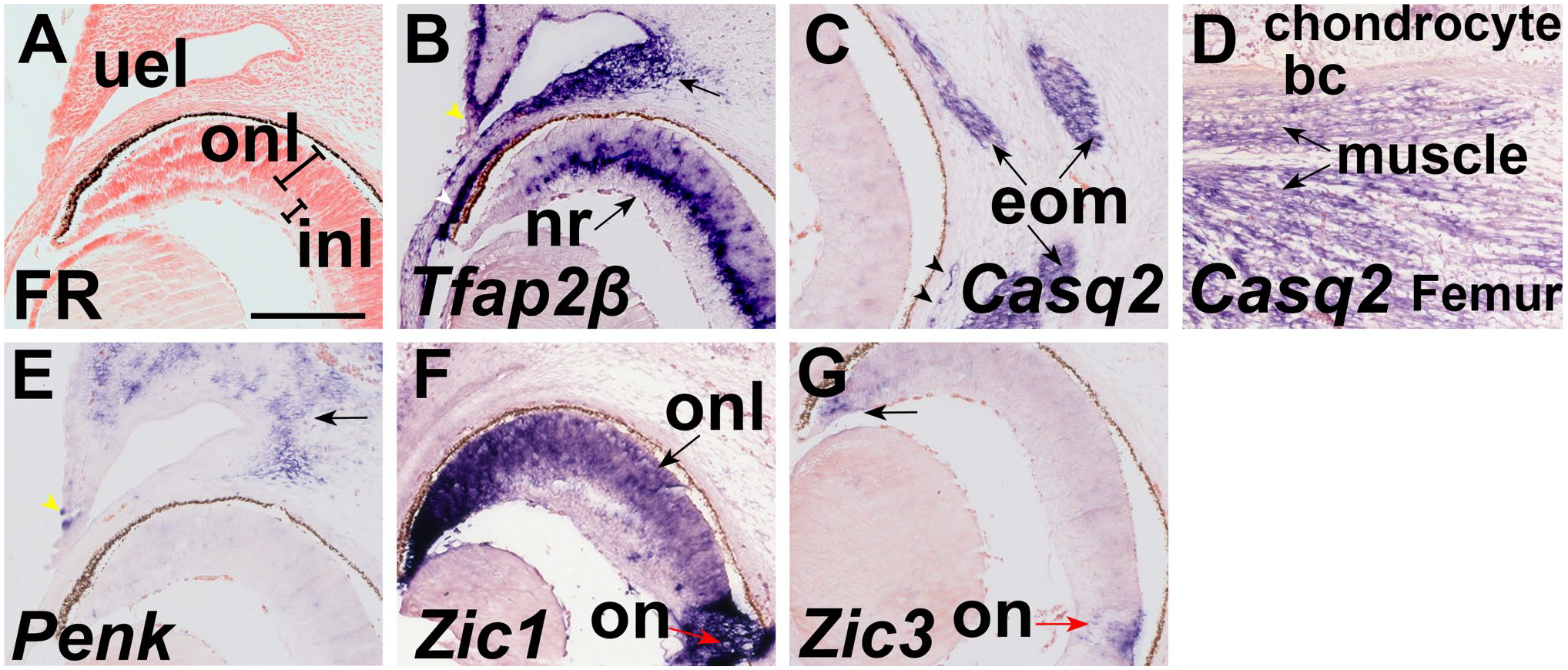
High power images of Tfap2β, Zic1, Zic3, Penk, and Casq2 expression at E15.5 (Β) Tfap2β, (C) Casq2, (D) Penk, (E) Zic1, and (F) Zic3 in coronal sections of E15.5. (A) Nuclear fast red staining. (B) Tfap2β is the ectoderm adjacent to the periderm cells in the eyelid bud (white arrowhead). Tfap2β is expressed in the innermost layer of the outer nuclear layer of the neural retina, and some cells in the outermost layer (arrow). Tfap2β is expressed in the corneal endothelium (white arrowhead). (C) Casq2 is expressed in the extrinsic ocular muscles (arrow) and vessels (black arrowheads). (D) Penk is expressed in the mesenchyme of the eyelid. The distal eyelid ectoderm also expresses Penk. (E) Zic1 is expressed in the neural retina (arrow) and has more intense expression in the optic nerve. (F) Zic3 is expressed in the optic nerve (red arrow). Scale bar in A = 200 μm and applies to all. Abbreviations: bc, bone collar; eom, extrinsic ocular muscles; inl, inner nuclear layer; on, optic nerve; onl, outer nuclear layer; nr, neural retina; uel, upper eye lid bud.
Between E15.5 and E17.5, the eyelid buds extend over the cornea and fuse together at the midline.21,22 This process is accomplished by the extension and fusion of the periderm at the leading edge of the eyelid bud.21,22 Penk and Tfap2β is found in the epithelium underlying the periderm (yellow arrowhead, Fig. 5B and D).
Discussion
In this study, we have performed an in situ hybridization expression analysis of Tfap2β, Casq2, Penk, Zic1, and Zic3 in the mouse eye at E12.5 and E15.5. We found that these five genes have a complimentary expression pattern to each other, suggesting that their functions may be coordinated in normal eye development. Tfap2β expression is found in the outer nuclear layer of the neural retina. In line with its function in cardiac muscle Casq2 expression is found in the external ocular muscles from E12.5 to E15.5. Zic1 was consistently expressed in the outer nuclear layer of the neural retina, including the lip of the optic cup between E12.5 and E15.5. Both Zic3 and Penk expand their region of expression between E12.5 and E15.5. The expression patterns here suggest that these genes may play a role in the eye development in a stage-dependent manner.
We chose to study the location of signaling pathways using the expression of Lef1 for Wnt/β-catenin signaling, because it is an epithelial transcription factor that is the target of activated Wnt signaling. The expression domains of Lef1, Gli1, and Ptch1 at E10.5 suggests that the eyelid morphogenesis may share similarities to organs that develop as the result of epithelial and mesenchymal interactions Similar to the tooth bud, we observed a central region of Hh signaling that is found in an invaginating epithelium, adjacent to the Lef1 domain. This epithelium is adjacent to a highly condensed mesenchyme. More work is needed to confirm if other morphogenetic and signaling processes are shared between the eyelid and tooth bud.
Our expression data show that Tfap2β, Zic1, and Zic3 have complimentary and overlapping expression domains within the neural retina. Tfap2β is expressed in the outer layer of neural retina and is excluded from the lip of the optic cup. Zic1 is found in the entire neural retina and the enriched in the lip of the optic cup, while Zic3 is only found in the lip of the optic cup. These complementary and overlapping expression patterns suggest that these transcription factors cooperate in the development of the neural retina. These data support the hypothesis that the domains of Zic1, Zic3, and Tfap2β define the medial-lateral axis of the neural retina.
Tfap2β (Figs. 1B and 2B) and Lef1 expression (Fig. 3A and D) partially overlap in and the developing eyelid mesenchyme and corneal epithelium at both E10.5 and E12.5. In Tfap2β-null mice, the expression of DKK1 is reduced in corneal epithelium. 19 DKK1 is an inhibitor of Wnt signaling, implying that Wnt signaling is increased in the developing cornea in Tfap2β-null mice. 19 Based on our expression data, we would predict that Lef1 expression would be increased in the Tfap2β-null animals and suggest that Tfap2β may function in a negative feedback loop to reduce Wnt signaling in corneal development.
Importantly, we observed overlapping expression domains of Zic1 and Zic3 with Gli1 and Ptch1 in optic cups and OS at E10.5, while the expression of Zic1 and Zic3 are not overlapped with Gli1 and Ptch1 at E12.5. Zic proteins has been reported to interact with Gli proteins, 23 and acts upstream of Sonic Hh target genes 24 to regulate forebrain development. These observations imply that Zic proteins regulate the early eye development through Hh signal.
Taken together, we have shown that Tfap2β, Casq2, Penk, Zic1, and Zic3 make a panel of five genes that are expressed in distinct, complementary, and overlapping tissues that will facilitate the study of eye morphogenesis.
Footnotes
Acknowledgements
The authors are grateful to the Dean’s Summer Scholar Program at the University of Pittsburgh’s School of Dental Medicine for supporting Casey White in this project. Nadine Robert is a LASPAU Fulbright scholar.
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
YW performed the experiments, analyzed data, and drafted the manuscript. CW analyzed data and drafted the manuscript. NR analyzed data. MBR analyzed data. HLS-R conceived and managed the project, analyzed data, and wrote the paper. All authors have read and approved the final manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by P30-NIH-DE020740, and start-up funds from the University of Pittsburgh School of Dental Medicine (HLS-R).
