Abstract
Analysis of trisomy 8 cells and the chromosome-specific fluorescence in situ hybridization (FISH) signals on the ring-shaped nucleus of a neutrophil reveal that homologue chromosomes orient in diametrical opposition to each other. This positioning results in a separation of the two haploid sets of parental chromosomes organized as two exclusive groups. These two groups impart the nucleus a symmetry that fortifies immune protection by accelerating chemotaxis. The ring form of the nucleus is a legacy of the orientation of chromosomes as a rosette during metaphase and telophase stages. A dual control maintains this spatial order: (1) chromosomes are tethered to the centriole all through the cell cycle, and (2) during their circular orientation in telophase the chromosomes bind to each other with lamins, which reorganize the nuclear membrane of the daughter nuclei, generating an additional anchorage. Here, chromosomes serve as temporary packets to assure proper distribution of the nuclear DNA during mitosis. The remainder time of the cell cycle the chromosomes are chained together across the telomeres, allowing a continuous sequence of genes of the two genomes, maternal and paternal, thus facilitating easy reading of the gene sequence. Exceptions to these orders are either physiological and temporary, or pathological and disease causing.
Keywords
Significance Statements
PMN (polymorphonuclear) leukocytes with bilaterally symmetric and circular nuclei have functional significance.
The basic circular form of a nucleus attained by the circular order of chromosomes, oriented opposite to their homologs, facilitate scanning of the gene sequence.
Control and maintenance of expression of alleles of the two parental sets.
Introduction
About 20 years of our cytogenetic work for diagnosis and therapy control in malignant disease have given us the unique opportunity to observe fascinating phenomena of blood and bone marrow cells. Some of them, like genomic separation of the parental chromosomes, were described earlier in context of immunological aspects.1–3 Here we present some additional facets that reveal a structurally elegant organization of chromosomes in a defined linear order as Hilbert Curve. 4 This order allows a functional harmony to the rationale of gene expression summarized by the statement, “the activation of transcription factors takes place in a precise temporal and spatial sequence of their chromosomal order.”4–5
Materials and Methods
Thousands of blood and bone marrow specimens as smears, cell suspensions, or fresh tissues for short term culture were accessed in Munich or Jena, Germany, and in Phoenix, Arizona. The specimens were generally prepared using the standard cytogenetic procedures: In short, initial colchicine treatment and hypotonic shock followed by acetic alcohol fixation of the cells in suspension, dropping on slides and air-drying. Specimens received as smears were treated with acetic alcohol before in situ hybridization with chromosome-specific probes. Cytogenetic and molecular cytogenetic analyses were done using G-bands by trypsin using Giemsa (GTG) banding and molecular cytogenetic–based staining techniques applying fluorochrome labeled probes from Vysis/Abbott (Wiesbaden, Germany) as mentioned below. Details of materials and methods are provided elsewhere.1–3
Results
As outstanding phenomenon of the neutrophil nucleus, we find a symmetric (mirror symmetry or rotation symmetry of order two) and circular form. These features are very prominent in neutrophils and other polymorphonuclear (PMN) leukocytes seen in extra vascular tissues, especially in the focus of inflammation, where 75% or more leukocytes display this symmetry and circular form.
Being polymorphic, these circular nuclei develop segmentations, generally two to five segments. In well-fixed and stained specimens, another feature is remarkable: folding of this ring-shaped nucleus. First folding may create a pattern like the digit eight and may impart the nucleus a bilobed form. Further folding may result in a tri-, tetra- or penta-lobed nucleus. The connectives between the segments, initially as notches, may also be attenuated, almost to invisibility under a light microscope.
Cases With Trisomy 8
Cytogenetic evaluation of myelodysplastic syndrome (MDS), acute myelogenous leukemia (AML), and other leukemia, cases with trisomy 8 as the sole chromosome aberration revealed a unique orientation pattern. The three copies of chromosome 8 were found in two groups, one with two chromosomes closely associated, forming a duplex signal, while the other with one chromosome placed diametrically opposite. With fluorescence in situ hybridization (FISH) signals of the probes for centromere 8 and C-MYC or “C-MYC break apart” probes in 8q24 similar distribution patterns were registered in metaphase and interphase nuclei. The duplex FISH signals were generally opposite to the single signal of the normal homologue. Apparently, the two closely associated signals (“duplex”) represent the two members of chromosome 8 as the products of a nondisjunction.
Images of metaphase and interphase cells are captured with FISH signals representing this unique pattern (Fig. 1). Quantitative data on metaphases and interphase cells from seven patients are presented in Table 1, showing a high frequency of this orientation (39–63%).
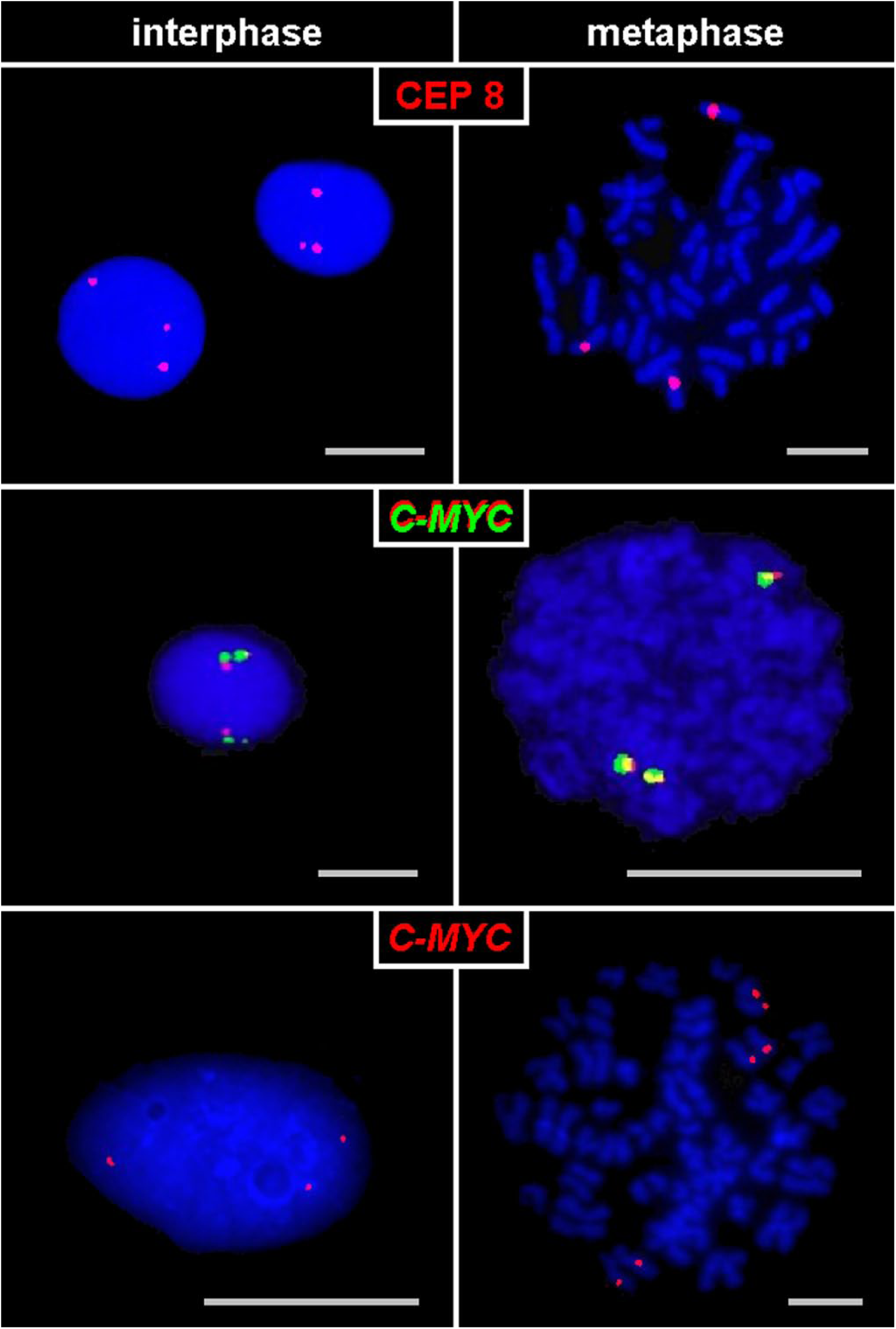
Cases of trisomy 8 hybridized with FISH probes for CEP 8, a C-MYC break apart probe (C-MYC in green and red) or a C-MYC probe (red): the duplex signals of the products of nondisjunction are localized diametrically opposite to their normal homologues both in metaphase and interphase. Abbreviations: FISH, fluorescence in situ hybridization; CEP, centromere.
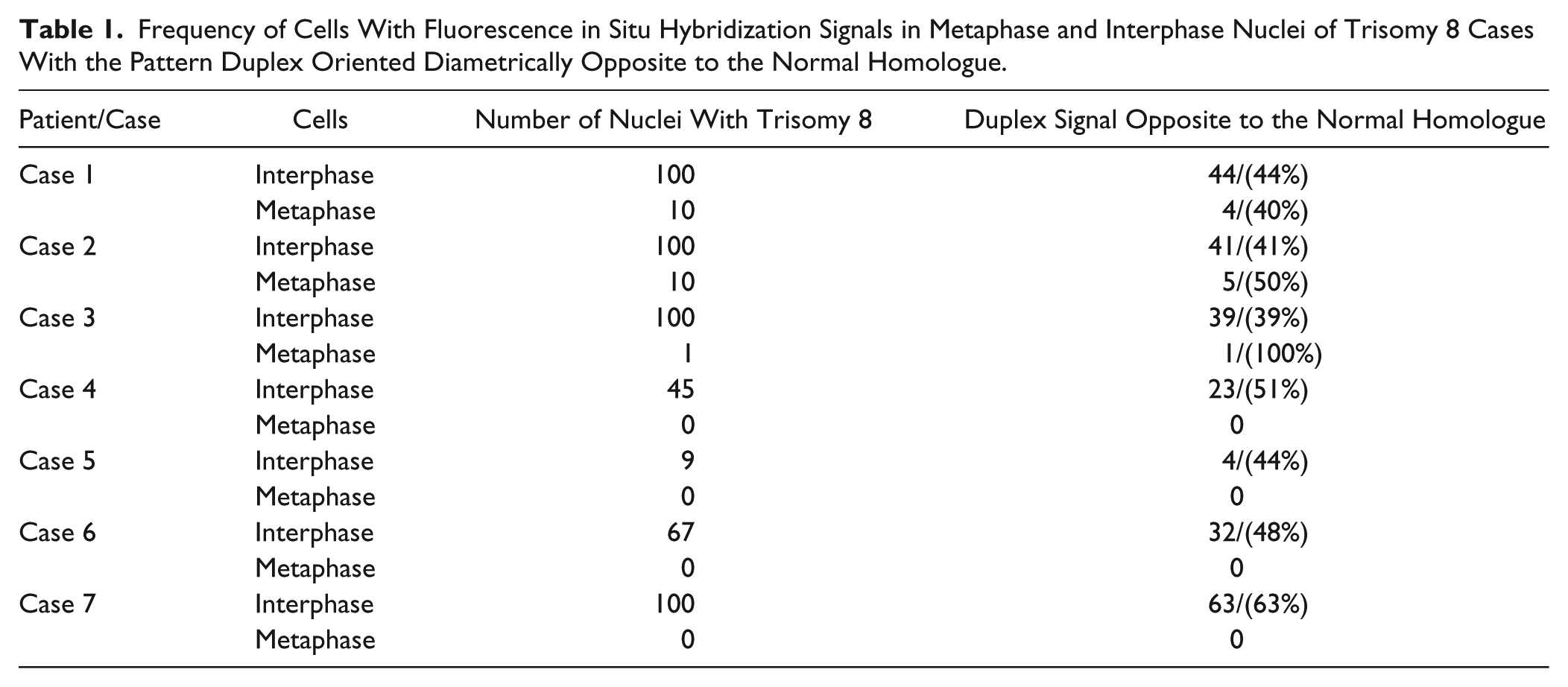
Frequency of Cells With Fluorescence in Situ Hybridization Signals in Metaphase and Interphase Nuclei of Trisomy 8 Cases With the Pattern Duplex Oriented Diametrically Opposite to the Normal Homologue.

Chromosomal abnormalities like polysomy, amplifications, or duplications also tend to localize diametrically opposite to their corresponding normal homologs (Fig. 2). This trend is also observed in cases of translocations (Fig. 3E).
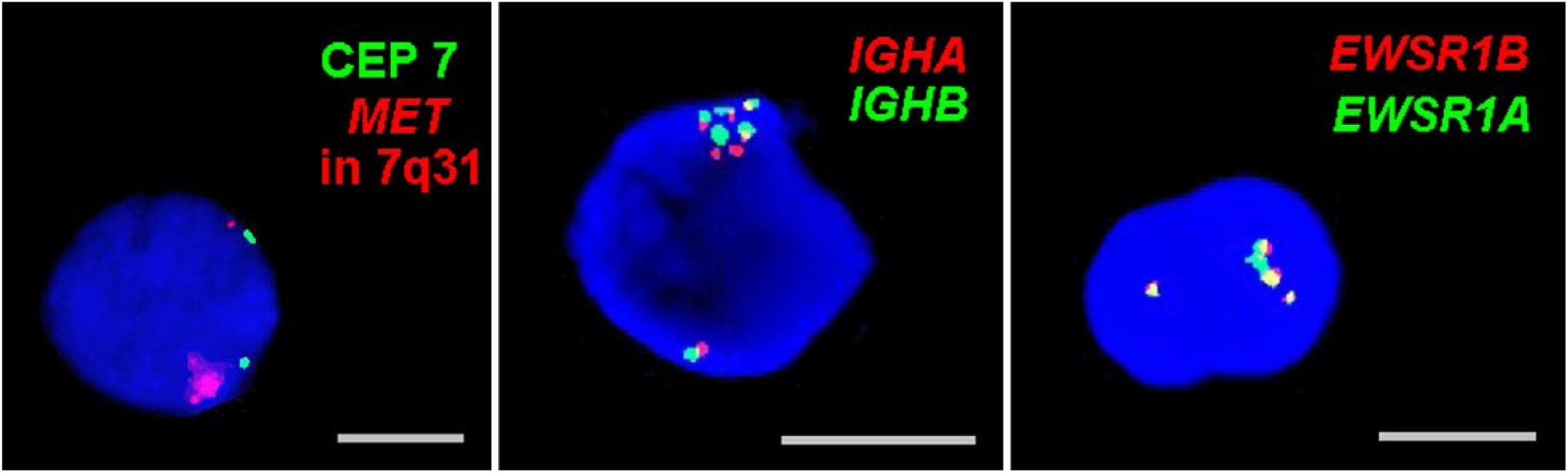
Cancer associated chromosome anomalies like amplifications, duplications, or polysomies occur in one of the two sets of chromosomes which orient opposite to the corresponding normal homologue. Here three such examples—one for a MET-amplification, and one each for a duplication of 14q23 (IGHA and IGHB) and of 22q12 (EWSR1A and EWSR1B). Abbreviation: CEP, centromere.
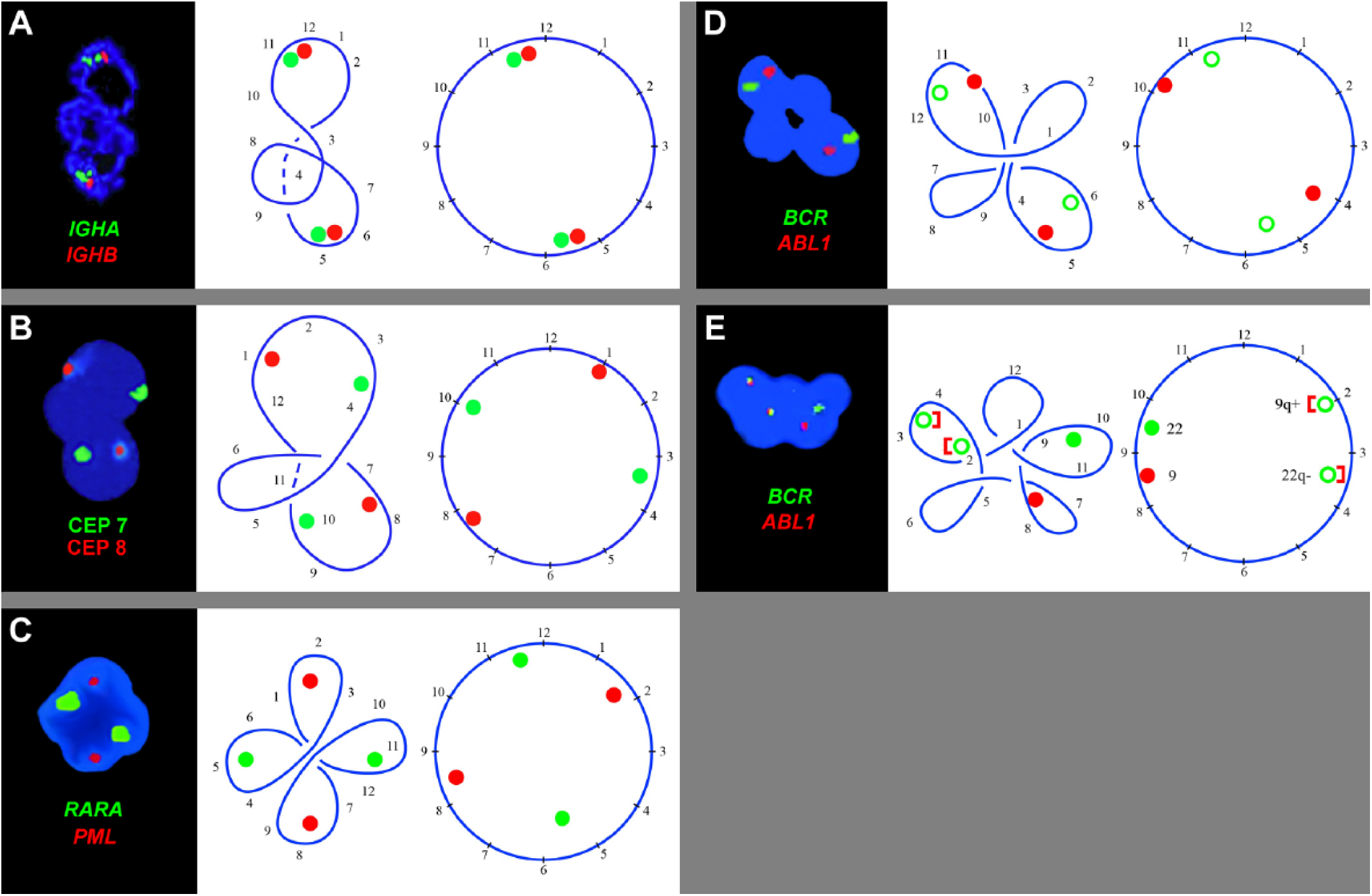
Five examples of annular neutrophil nucleus with diagramatic analysis showing the position of the chromosome-specific FISH signals. The numbers 1 to 12 are arbitrarily put on the circular nucleus. They guide to follow the foldings and loopings of the ring formed by the tandem association of the chromosomes at telophase. As probes were applied: (A) probes for IGHA and IGHB in 14q23. (B) centromeric probes for chromosomes 7 and 8. (C) probes for PML in 15q22 and of RARA in 17q21. (D–E) probes for ABL1 in 9q34 and of BCR in 22q11.2. Abbreviations: CEP, centromere.
Analysis of Neutrophils With Ring Shaped Nucleus
The ring-shaped nucleus of neutrophils and other PMN leukocytes with chromosome-specific FISH signals of one or more homologs, reveal the identical pattern: the homologs orient diametrically opposite each other (Fig. 3A–E). The circular, folded nuclei in Fig. 3 have two halves, apparently representing the two sets of parental chromosomes of a diploid cell.
The circular nucleus in Fig. 3A has apparently three loops. For analysis and follow up, we have inserted numbers on the basically circular nucleus as landmarks. These landmarks guide to untangle the loops or folding made by circular nucleus and reveal the two IGH (in 14q32) signals oriented as homologs opposite each other.
The nucleus in Fig. 3B has an extra loop and looks trilobed. When straightened following the landmark numbers 1 to 12, the FISH signals for chromosome 7 (red) and 8 (green) locate opposite to their homologs. The nucleus in Fig. 3C has four segments associated with four loops of the chromosomal ring. When opened following the given landmarks, the FISH signals of PML (in 15q22, red) and of RARA (in 17q21, green) localize diametrically opposite to their homologs. The nucleus in Fig. 3D has also four segments derived from four loops. When untangled, the FISH signals of ABL (in 9q34, red) and BCR (in 22q11.2, green) distribute opposite to their homologs. These two translocation probes, PML/RARA and BCR/ABL provide additional information. On a circular nucleus, the probes PML and RARA are far away from each other, while the probes ABL and BCR localize closer to each other (Figs. 3C, 3D, and 4).
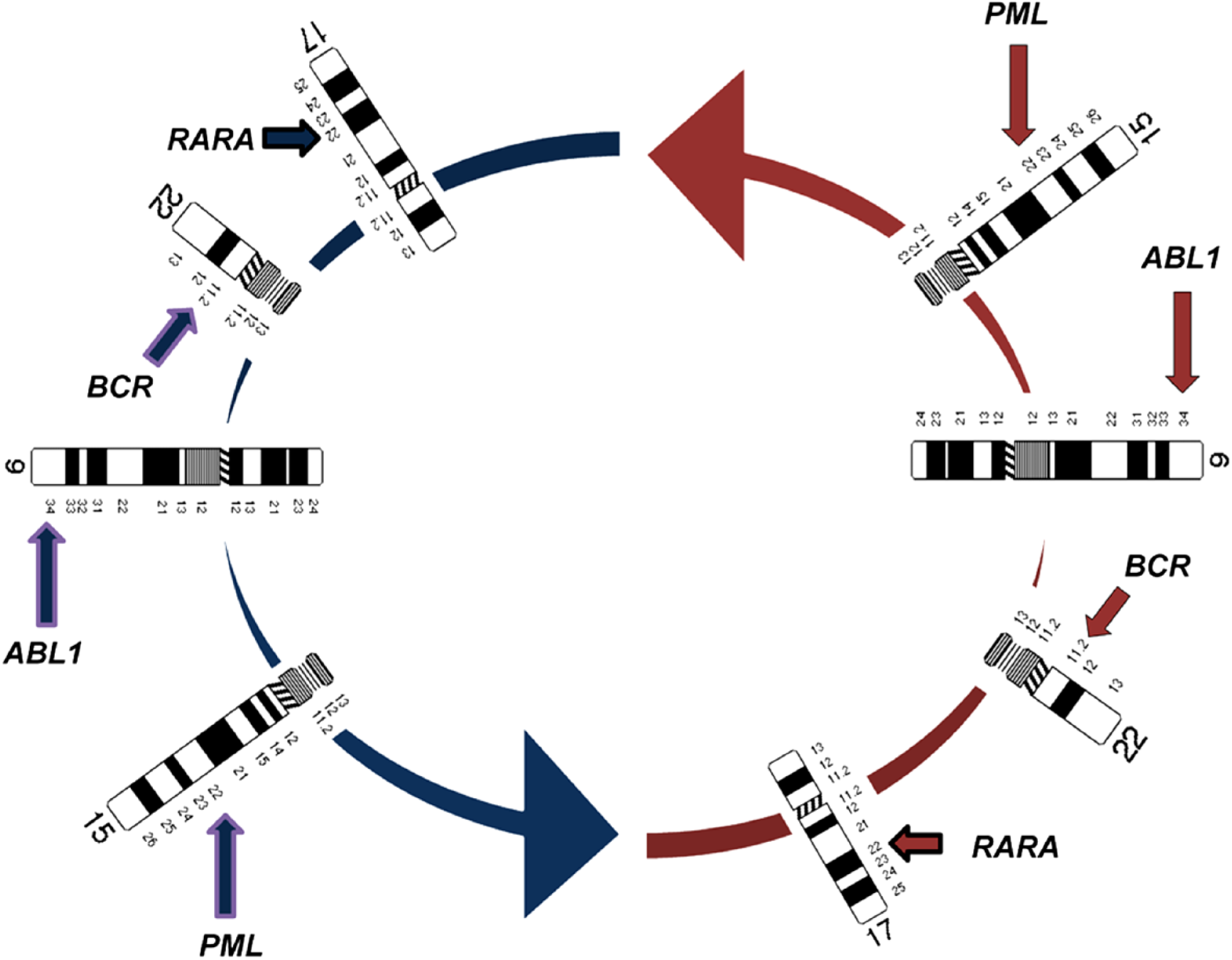
Relative positions of the chromosomes in a metaphase rosette or on a circular nucleus, as based on the results of the present study. Within each of the two parental genomes BCR and ABL are closer to each other than the PML and RARA.
The penta-lobed nucleus (Fig. 3E) of a patient with chronic myelogenous leukemia (CML) typical translocation displays two yellow fusion signals of 9q+ and 22q–, and one red and one green signal of ABL and BCR on the normal sister chromosomes (because the homologues generally orient opposite each other, the fusion signal opposite to ABL is designated 9q+ and the fusion signal opposite to BCR is 22q–). The domains delineated by landmarks 1 to 6 and 7 to 12 of this circular nucleus apparently represent the two parental halves of a symmetric nucleus.
Discussion
The nuclei of normal human neutrophils with two or more lobes may also share an annular shape 6 with several other species, like rats and mice. Ring-shaped nuclei of human granulocytes in acute erythroleukemia were seen earlier. 7 In patients with myelodysplastic crisis, we observed ring-like nuclei in about 80% of the myelocytes. 8 This phenomenon was also reported from other disorders of myelocytes.9–10 A note in Blood 11 supplies fascinating images of human neutrophils with circular nuclei. After elaborate studies of dividing human fibroblasts by cinematography and by staining fixed cells, Miles 12 reported that the telophase rosettes of chromosomes become coated with lamins and form a chain or ring. He also observed that this ring or chain of chromosomes folds to form a spheroid nucleus in fibroblasts.
The circular nucleus is a legacy of the chromosome rosette of the metaphase plate, seen also in the telophase. Each chromosome opposite to its homolog (Fig. 5B, C in Chaudhuri and Walther 2 ) brings forth a clear separation of the two parental sets of chromosomes. In this serial order, they get coated with lamins and form a circle of tandemly connected chromosomes.
Orientation of the Trisomy 8 Chromosomes
The distribution of homologous chromosomes opposite to each other is observed in both metaphase and interphase cells.1,2 Our data from trisomy 8 cases underscore this statement, by showing the adherence of trisomic chromosomes to this rule. Similar orientation pattern were observed also for trisomy 7, 9, and 21 by us and others. 13 A critical analysis of trisomy 21 using especially informative centromeric probes on translocation and free trisomy 21 provides a deeper insight. 14 This kind of orientation was also observed in many other types of cancer associated chromosome aberrations, such as polysomy, duplication, and amplification (Fig. 2), implicating involvement of only one set of the parental chromosomes. Moreover, certain reciprocal translocations like BCR/ABL are also observed to occur within one of the two parental genomes more frequently than by chance. 15 This provides a clue to disease etiology.
Three-dimensional analysis of interphase nuclei in mice and humans demonstrated that the average distance between the homologs is larger than that between the nonhomologs. 16 This fits well with the observation that the homologs lie diametrically opposite each other. Exceptions to this general trend, where the homologs fail to orient opposite to each other, arise temporarily in physiology or as a tissue specificity or may indeed be pathological.17,18 For example, NOR (Nucleolus Organizing Regions) bearing chromosomes congregate temporarily during the period of nucleolus formation, independently of their parental identity. 2 Certain domains of a chromosome may also move far from their spatial position for interactive association of genes. 19
Circular Nuclei of Neutrophils
The analysis of the neutrophil nuclei (Fig. 3A–E) reveals that they are predominantly circular and that the chromosomal homologs lie generally diametrically opposite each other. If this holds for each chromosome, we end up with two haploid sets of chromosomes separated in the two semicircles. These two halves are of equal weight both in quantity of DNA content and in chromosomal quality, each having a similar set of 23 chromosomes (22 plus X or Y) as we suggested earlier. 1
The Bilateral Symmetry of a Neutrophil Nucleus
The two sets of parental chromosomes in the two halves of a circular neutrophil nucleus possess a reciprocal serial order and therefore have a rotational symmetry of the order two. The process of folding and looping accompanied by local condensation and extension of chromatin connectives associated with segmentations impart the neutrophil nucleus the bilateral symmetry that may accelerate chemotaxis. 2 Eosinophils are bilobed with perfect bilateral symmetry and therefore faster than the neutrophils with their higher degree of lobulation. 20 Carvalho et al. 21 discussed generally the role of nuclear morphology in neutrophilic function. Our observation contributes concrete examples for this discussion in the areas of immunolgy 2 and genomic function.
Sequential Order of Chromosomes Within the Parental Sets
Our analysis of the two reciprocal translocations BCR/ABL, translocation t(9;22) in CML and PML/RARA, translocation t(15;17) in promyelocytic leukemia (PML) (Fig. 3C, D) indicates that chromosomes 9 and 22 within each parental set of chromosomes are closer to each other, than 15 and 17 (see also Fig. 4). This is in accordance with our finding based on karyotyped metaphases 9 that the translocation t(9;22) occurs predominantly within one parental genome as opposed to the translocation t(15;17), that may often involve one maternal and one paternal chromosome. This indicates a sequential chromosomal order within each parental set, each with a defined spatial position.
A Dual Control Maintains the Spatial Chromosome Order
This spatial order of chromosomes in the circular nucleus of a PMN cell as well as in a mononuclear cell like a fibroblast or other cells with spheroid nucleus is maintained by a dual control mechanism: (1) spindle fibers connect each chromosome of one of the two sets of chromosomes to one of the two centrioles of a diploid cell,2,11 and (2) the lamina associated domains of chromosome-specific heterochromatin make molecular contacts with nuclear lamina 22 assuring an anchorage of chromosomes to the nuclear membrane.
Gene Sequence in the Two Parental Sets of Chromosomes of a Diploid Cell
The analysis of the circular nuclei of neutrophils using chromosome-specific FISH signals presented here demonstrates that the entire nuclear structure determines the spatial relation of the chromosomes with respect to each other as well as with respect to the two parental genomes in a diploid nucleus. Ligation techniques, like Hi-C and others, to analyze the sequence of genes are of higher resolution and more precise, but require millions of cells and are not amenable to single cell level. Attempts of single cell Hi-C are quite limited. 23 The prospects and limitations of fine mapping of trait-associated genomic regions are much discussed. 24 The presented method of analysis of the circular nuclei may provide a broad cellular basis to the results obtained by ligation techniques. Genes are in a linear order on a chromosome. If the chromosomes become tandemly connected and attain a circular form, as we see in the circular nucleus (Fig. 3A–E), the abrupt end of the gene sequence at the telomeres will be overcome and give us a continuous, uninterrupted linear sequence of genes along the chromosomes of the two genomes in a defined order as Hilbert Curve 4 presenting the ideal condition for the scanning of the genes to maintain the process of gene activation envisaged by Kmita and Duboule. 5 This exclusive order of the two parental sets of chromosomes assures a platform to regulate the contest and concordance of the parental alleles in a coordinated fashion discussed earlier.2,25
Interphase Chromosomes
The aspects (1) homologs opposite each other, (2) separation of the parental sets of chromosomes, and (3) the chain of chromosomes tandemly connected forming a circle, compel us to reexamine the term chromosome. The conventional chromosomes are entities of mitosis when the total chromosomal DNA is packed in a number of parcels for the facility of division and distribution of the genetic information to the daughter nuclei. In the interphase, the phase between two mitoses, when the cells are actively fulfilling their function, we have a single chain of all chromosomes together. This logical form, without telomeric and other interferences, is the interphase chromosome as opposed to mitotic chromosomes,26,27 although the specific DNA sequences of the chromosomes are retained, 28 as expected in a Hilbert Curve.
Costello 29 described an identical linear order of chromosomes in the two parental sets in the metaphase of a protochordates. Habermann et al. 30 presented an optical section through a chicken fibroblast nucleus showing the mutually exclusive chromosome territories lying opposite to their homologs. Many organisms—plants, insects, and mammals including Homo sapiens—have the same basic organization of their chromosomes as has also been suggested in a critically analytical report by Misteli. 31 All in all it is highly probable that this is the basic principle of chromosomal organization: an important equipment with a high functional efficiency.
Footnotes
Competing Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
JPC and JUW planned the underlying experiments; JPC and TL drafted and wrote the manuscript; JPC, SK, TL and LS did the FISH-experiments. All authors provided to and agreed on the final version of the paper.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
