Abstract
With recent advances in immunohistochemical (IHC) techniques, immunohistochemistry now plays a more important role in research, especially in mouse models where characterization of cellular patterns of protein expression has become critical. Even with these recent advances, a paucity of IHC quality antibodies for some proteins still exists. To address this, we have developed a novel IHC assay that utilizes a commercially available goat anti-DDDDK peptide polyclonal antibody on paraffin-embedded tissues from knock-in mice expressing proteins of interest tagged with a 3×FLAG epitope at physiologically relevant levels. Focusing on two 3×FLAG-tagged proteins for which specific antibodies were available, USP48 and RIPK3, we were able to validate our anti-DDDDK assay by comparing the IHC directed against the actual proteins to the anti-DDDDK IHC assay, which recognizes the FLAG epitope. We were also able to detect a third 3×FLAG-tagged protein, BAP1, for which quality reagents were not available. This universal IHC method will enable researchers to characterize the expression patterns of proteins of interest when specific antibodies are lacking.
Introduction
With the advent of personalized health care, immunohistochemistry (IHC) has come to play a critical role in research, especially when murine models are utilized to characterize biomarkers of interest. Since its inception by Dr Albert Coons in 1941 (Coons et al. 1941), IHC has witnessed many advances over the decades that have seen this procedure become a more efficient tool for many research applications. These advances include the development of mouse (Kohler and Milstein 1975) and rabbit monoclonal antibodies (Spieker-Polet et al. 1995), antigen retrieval techniques (Pileri et al. 1997; Shi et al. 1991; Tacha and Teixiera 2002), techniques for the amplification of weak signals (Toda et al. 1999), and the development of more efficient detection kits that now commonly use polymer technology (Sabattini et al. 1998). Consequently, IHC assays can be developed rapidly, especially when it pertains to the characterization of proteins of interest in murine tissues. Although IHC diagnostic assays are primarily developed for use in human tissues in combination with personalized drug therapy, early in their characterization potential biomarkers are usually investigated in murine disease models. Additionally, the characterization of the tissue-, cellular-, and disease-specific distributions of potential therapeutic targets is commonly initiated in murine models. Biomarkers can be characterized by western blotting, ELISA, FACS and a host of other techniques, but ultimately visual localization of proteins in the context of specific cellular and tissue morphological features requires IHC methodology.
Technological advances in IHC methodology have been accompanied by a dramatic increase in the availability of antibodies recognizing proteins from multiple species. The Human Protein Atlas Program (Uhlén et al. 2005) and search engines such as Biocompare, IHC World and Linscott’s Directory allow investigators to quickly identify IHC reagents for their research. Nonetheless, there can be paucity of well-characterized IHC reagents for novel biomarkers or therapeutic targets to be investigated in mice.
In order to circumvent limitations in the availability of mouse-specific antibodies for proteins of interest, we have developed a novel methodology involving knock-in mice with proteins of interest tagged with a 3×FLAG epitope that can be detected with a universal FLAG IHC assay. Using this knock-in approach, expression of the tagged protein remains under the control of the gene’s endogenous regulatory elements, preserving the expression of different splice variants at physiologically relevant levels. This approach is distinct from transgenic expression of an epitope-tagged mouse cDNA, which may not mimic the normal pattern of protein expression and often results in protein overexpression. Our FLAG IHC method uses a goat primary antibody, which eliminates the background noise associated with localizing mouse primary antibodies on mouse tissues. In this study, we confirm the validity of the FLAG IHC assay by performing comparisons to IHC assays specific to the proteins that we have tagged in knock-in mice: receptor-interacting serine/threonine-protein kinase 3 (RIPK3) and ubiquitin-specific peptidase 48 (USP48). Additionally, we highlight the utility of the IHC assay with detection of 3×FLAG-tagged mouse BRCA1-associated protein-1 (BAP1) in the absence of a quality BAP1 IHC antibody.
Materials & Methods
Mice
Ripk3−/−, Bap1−/−, and Bap1FLAG/FLAG mice have been described previously (Newton et al. 2004; Dey et al. 2012). The latter were backcrossed to C57BL/6N mice for >10 generations. Ripk3FLAG/FLAG mice were generated by genOway (Lyon, France) from gene-targeted C57BL/6 ES cells (Fig. 1A) and genotyped with primers (5’-GCA CAC TTC AGG CTC TTG CAG T-3’ and 5’-TTG CAT TCC CCA GGG AGA TC-3’) that amplified 306-bp wild-type (WT) and 372-bp knock-in (KI) DNA fragments. Usp48FLAG/FLAG and conditional Usp48fl/fl mice were generated from gene-targeted C2 C57BL/6 ES cells (Fig. 1B and 1C). Usp48fl/fl conditional knock-out (cKO) mice had exon 3 encoding catalytic Cys98 flanked by loxp sites and were crossed to the inducible general Cre deleter C57BL/6-Gt(ROSA)26Sortm9(Cre/ESR1)Arte (Taconic; Hudson, NY). The CreERT2 allele was maintained in a heterozygous state. CreERT2 was activated in mice by intraperitoneal injection of 160 mg/kg tamoxifen (Sigma-Aldrich, St. Louis, MO) dissolved in sunflower seed oil (Sigma-Aldrich) once a day for five consecutive days. Tissues were harvested one month after the final tamoxifen injection. Usp48FLAG genotyping primers (5’-GAA CAG AAA GCA TCA ATC AGC C-3’ and 5’-TGA CAT AAG GCA AGT GGG ACA-3’) amplified 420 bp WT and 520 bp KI DNA fragments. Usp48 genotyping primers (5’-AGC AGG GAA TCA CAC CGT ATG-3’, 5’-AGC AGT ACT GTC ATT TCC AGC C-3’, and 5’-TAA AGG GCG GCG CAT AAC-3’) amplified 483-bp WT, 517-bp cKO, and 375-bp KO DNA fragments. The neo selection cassette in the Ripk3FLAG/FLAG and Usp48FLAG/FLAG knock-in strains was flanked by loxP sites and removed by breeding to the general Cre deleter strain C57BL/6-Gt(ROSA)26Sortm16(cre)Arte (Taconic). The neo selection cassette in the Usp48fl/+ ES cells was flanked by frt sites and eliminated with adenovirus-expressing recombinase FlpO (Vector Biolabs; Philadelphia, PA). The Genentech animal care and use committee approved all protocols.
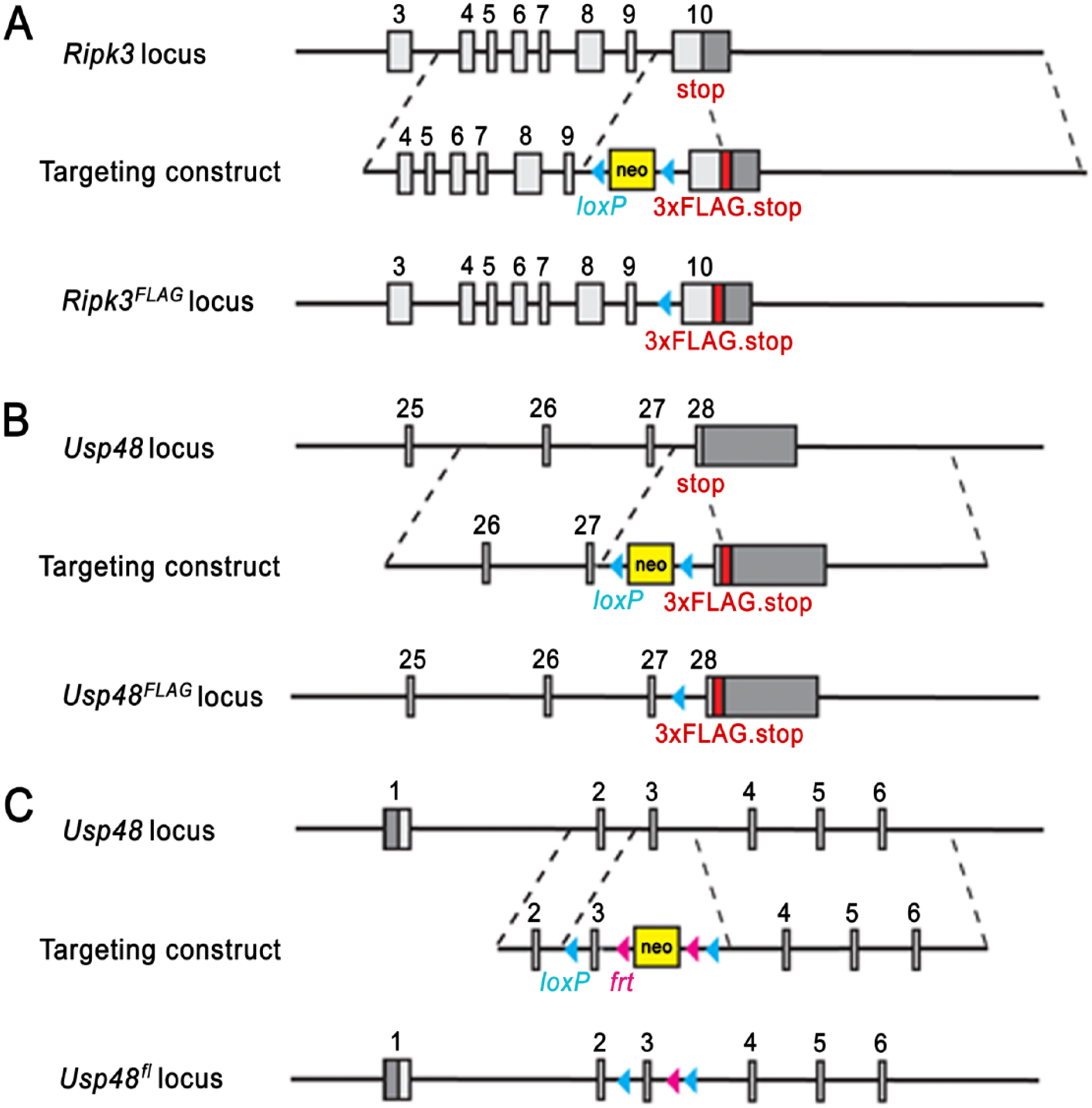
Ripk3 and Usp48 mutant mice: Targeting strategies for the generation of Ripk3FLAG/FLAG (A), Usp48FLAG/FLAG (B), and Usp48fl/fl (C) mice. Boxes represent exons with coding regions in light gray and non-coding regions in dark gray.
Western blot
For the detection of proteins on western blots, antibodies recognizing RIPK3 (rat monoclonal 1G6; Genentech, South San Francisco, CA), BAP1 (rabbit monoclonal 16B1, Genentech), β-actin (Cell Signaling Technology; Danvers, MA), HCF-1 (Clone 699A; Bethyl Labs, Montgomery, TX), or FLAG (Clone M2; Sigma-Aldrich) were utilized.
Immunohistochemistry
Formalin-fixed, paraffin-embedded serial murine tissue sections were cut at 4 µm, deparaffinized in xylenes and rehydrated through a graded series of alcohols. Sections were then pre-treated for antigen retrieval by incubation in a PT Module (Thermo Scientific; Waltham, MA) using fresh Target Retrieval (DAKO; Carpenteria, CA) for 20 min at 99°C followed by a 20-min cool down. After pre-treatment, all subsequent IHC steps were carried out at room temperature on a Thermo Scientific Autostainer 720 platform (Thermo Scientific).
After pretreatment with Target Retrieval, sections were blocked for endogenous peroxidase activity using KPL blocking solution (Kirkegaard and Perry Laboratories, Gaithersburg, MD) for 4 min and for endogenous avidin/biotin using an avidin/biotin blocking kit (Vector Labs, Burlingame, CA). Sections were subsequently blocked for non-specific binding sites for 30 min with either 10% donkey serum/3%BSA/PBS for FLAG IHC, 10% rabbit serum/3%BSA/PBS for RIPK3 IHC or proprietary TNB blocking buffer (Perkin Elmer; Waltham, MA) for USP48 IHC.
For FLAG IHC, sections were incubated for 60 min in 0.5 µg/ml goat anti-DDDDK polyclonal antibody (AbCam; Cambridge, MA) followed by incubation for 30 min with a biotinylated donkey anti-goat secondary antibody (Jackson ImmunoResearch; West Grove, PA). Sections were subsequently incubated for 30 min with Vector Elite ABC-HRP reagent (Vector Labs) followed by incubation for 5 min in a metal-enhanced DAB peroxidase substrate (Pierce Laboratories; Rockford, IL) for color development. For comparison of the RIPK3 IHC to its FLAG-tagged counterpart, adjacent sections were incubated for 60 min with 5 µg/ml rat anti-mouse RIPK3 monoclonal antibody (Clone 1G6; Genentech) followed by incubation for 30 min with a biotinylated rabbit anti-rat secondary antibody (Vector Labs). Sections were subsequently incubated for 30 min with Vector Elite ABC-HRP reagent (Vector Labs) followed by incubation for 5 min in a metal-enhanced DAB peroxidase substrate (Pierce Laboratories) for color development. For comparison of the USP48 IHC to its FLAG-tagged counterpart, adjacent sections were incubated for 60 min with 1 µg/ml rabbit anti-USP48 polyclonal antibody (Bethyl Labs) followed by peroxidase conjugated anti-rabbit Powervision (Leica Biosystems; Buffalo Grove, IL) for 30 min. Signal for USP48 required amplification by incubation of sections with biotinylated-TSA (Perkin Elmer) for 3 min followed by incubation in streptavidin-HRP complex (Perkin Elmer) for 30 min. Sections were then incubated for 5 min in a metal-enhanced DAB peroxidase substrate (Pierce Laboratories) for color development. All sections that underwent IHC were counterstained with Mayer’s Hematoxylin (Rowley Biochemical Institute; Danvers, MA), dehydrated, cleared with xylenes, and coverslipped. For negative controls, sections were incubated with the appropriate naïve isotype control antibody for each marker in place of the primary antibody.
The FLAG IHC assay for BAP1.3×FLAG was carried out in the same manner as for the RIPK3.3×FLAG and USP48.3×FLAG constructs minus the adjacent IHC for the direct detection of the endogenous protein, as commercial reagents were lacking for this marker. All imaging was performed using MetaMorph image analysis software on a BX51 Olympus microscope (Olympus Optical; Tokyo, Japan).
Results
To develop and validate an anti-FLAG IHC assay compatible with mouse tissues, we focused on two proteins, RIPK3 and USP48, for which validated IHC antibodies are available. Mouse RIPK3 is recognized by rat monoclonal antibody clone 1G6 (Genentech), and its kinase activity is essential for caspase-independent cell death, known as necroptosis (Cho et al. 2009; He et al. 2009; Newton et al. 2014). Mouse USP48 is recognized by a rabbit polyclonal antibody (Bethyl Labs) and is a member of the deubiquitinating enzyme family (Quesada et al. 2004; Altun et al. 2011). The expression patterns of RIPK3 and USP48 were evaluated in knock-in mice with a sequence encoding a 3×FLAG epitope tag inserted into the last coding exon ahead of the translation termination codon in order to produce C-terminally tagged RIPK3.3×FLAG or USP48.3×FLAG (Fig. 1A and 1B).
IHC with a goat anti-DDDDK antibody that recognizes the 3×FLAG epitope revealed cytoplasmic localization of RIPK3.3×FLAG in a host of adult Ripk3FLAG/FLAG mouse tissues (Fig. 2). Cytoplasmic labeling of RIPK3.3×FLAG was observed in small and large intestinal crypts, Kupffer cells, alveolar macrophages, pancreatic acinar cells, the thymic medulla, scattered lymphoid cells throughout the spleen and bone marrow, the cortex and para-cortex of the mesenteric lymph node, and in the endothelium and intra-vascular leukocytes of the heart. No labeling was observed in the kidney and skin. Interestingly, skin inflammation caused by caspase-8, Fas-associated death domain (FADD) or RIPK1 deficiency is dependent on RIPK3 (Bonnet et al. 2011; Weinlich et al. 2013; Rickard et al. 2014), suggesting that RIPK3 protein expression in skin is either below the detection threshold of our assay or might be increased in the absence of caspase-8, FADD or RIPK1. No labeling for RIPK3.3×FLAG was observed in wild-type tissues with the anti-DDDDK antibody, supporting the specificity of this antibody for FLAG-tagged proteins (Fig. 2). To validate the specificity of our in-house rat anti-RIPK3 antibody, 1G6, tissues from RIPK3-deficient mice were stained (Fig. 3; Newton et al. 2004). No labeling was observed in these mice. In addition, a similar pattern of expression was observed with the 1G6 antibody in wild-type tissues when compared with Ripk3FLAG/FLAG tissues stained with the anti-DDDDK antibody (Fig. 3). RIPK3 staining with 1G6 on adjacent sections of RIPK3.3×FLAG knock-in mouse tissues resulted in a similar staining pattern as seen with the anti-DDDDK antibody, further validating the specificity of our anti-FLAG IHC assay (Fig. 2). Of note, RIPK3 staining patterns in Ripk3FLAG/FLAG and wild-type tissues were comparable, indicating that fusion of a 3×FLAG tag to the C-terminus of RIPK3 did not alter the expression pattern or localization of the protein.
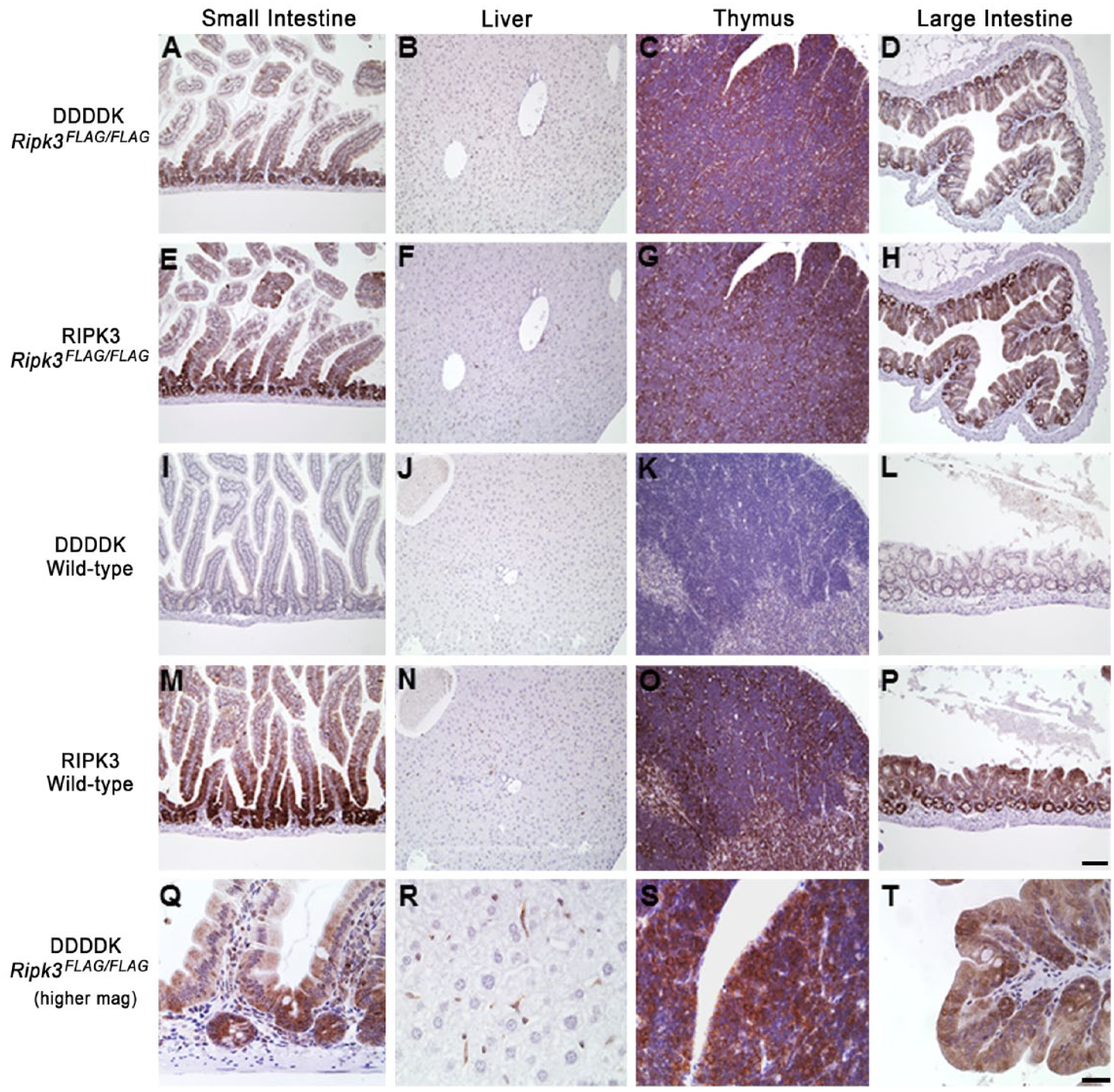
Tissues from 12-week-old Ripk3FLAG/FLAG mice stained with anti-DDDDK antibody (A–D, top panel) or anti-RIPK3 antibody in adjacent sections (E–H, second panel). RIPK3 is localized to the cytoplasm when utilizing both methods. No reactivity is observed in wild-type tissues with the anti-DDDDK antibody (I–L, third panel), and a similar pattern of expression is observed with the RIPK3 antibody (M–P, fourth panel) when comparing Ripk3FLAG/FLAG tissues to wild-type tissues. Higher magnification for anti-DDDDK IHC of Ripk3FLAG/FLAG tissues is shown in bottom panel (Q–T). Scale (A–P), 100 µm; (Q–T), 20 µm.
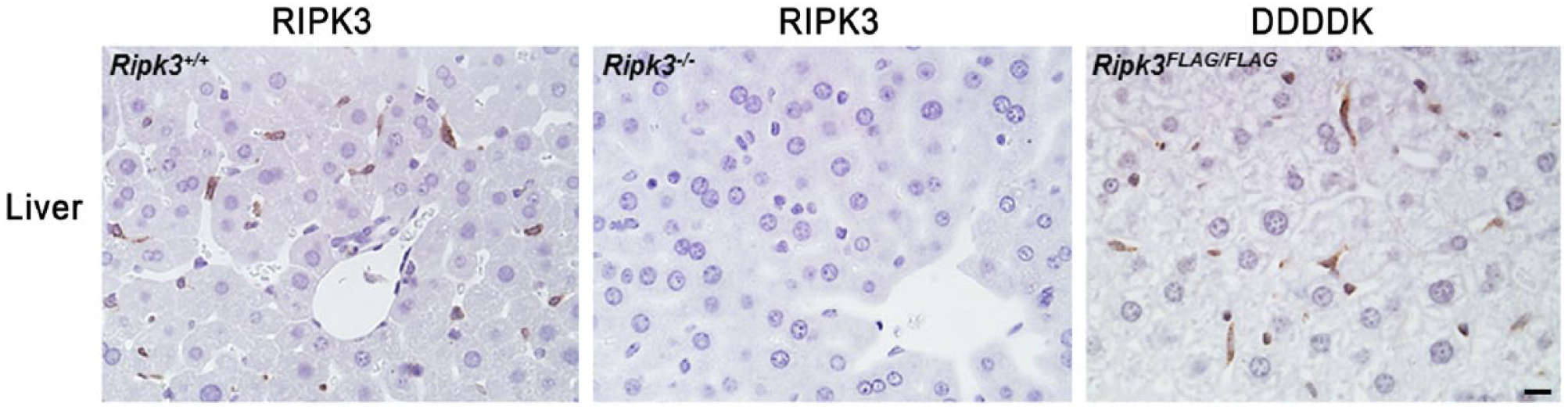
RIPK3 is detected with anti-RIPK3 antibody in Kupffer cells of the wild-type (Ripk3+/+) liver, but not in Ripk3−/− Kupffer cells. A similar pattern of expression is observed with the anti-DDDDK antibody when comparing Ripk3FLAG/FLAG tissues with wild-type tissues. Scale, 10 µm.
By western blotting, RIPK3.3×FLAG was detected in Ripk3FLAG/FLAG heart, lung, liver, thymus, and spleen, but was not detected in the bone marrow (Fig. 4). Very few positive cells were identified in the bone marrow using the FLAG IHC assay; therefore, it is suspected that RIPK3.3×FLAG levels in the entire bone marrow probably fell below the limit of detection by western blotting. Similarly, the weak signal in the liver by western blotting is likely due to the relative paucity of Kupffer cells in the liver as compared with the total cellular composition of the liver. RIPK3.3×FLAG was detected more readily by western blotting with the anti-FLAG antibodies than with a RIPK3 antibody, suggesting that the 3×FLAG tag can improve assay sensitivity. An additional lower molecular weight band was observed in the spleen and thymus of wild-type and Ripk3FLAG/FLAG mice, which is suspected to be a RIPK3 cleavage product or splice variant. The reference protein β-actin was not identified in the heart by western blotting because F-actin, but not β-actin, is expressed in the heart. The anti-DDDDK antibody was evaluated in adult Usp48FLAGFLAG mouse tissues that express USP48.3×FLAG and in embryos at 12.5, 15.5 and 18.5 days post coitum (d.p.c.). IHC detection of USP48.3×FLAG yielded a nuclear signal in a range of adult Usp48FLAG/FLAG mouse tissues (Fig. 5). Nuclear USP48.3×FLAG was observed in the cells of the small intestinal crypts as well as basolateral enterocytes with decreasing signal intensity as the cells differentiated and migrated up the villi. The large intestine and cecum exhibited a similar expression pattern. Nuclear USP48.3×FLAG was also observed in germ cells of the testes as well as ovarian follicles, throughout the gall bladder epithelium, and in the para-cortex of the mesenteric lymph node. No labeling was observed with anti-DDDDK antibodies in tissues from wild-type mice, except for some non-specific labeling of Leydig cells of the adult testes (Fig. 5). A commercially available anti-USP48 polyclonal antibody stained adjacent Usp48FLAG/FLAG tissue sections in a similar manner to the anti-DDDDK antibody in adult mouse tissues (Fig. 5). The specificity of the USP48 antibody was determined using tissues from Usp48fl/fl Rosa.CreERT2 mice that had been treated with tamoxifen to activate the ubiquitously expressed Cre recombinase and delete the Usp48 gene (Fig. 1C). Compared with wild-type tissues, USP48-deficient tissues exhibited negative staining for USP48 (Fig. 6).
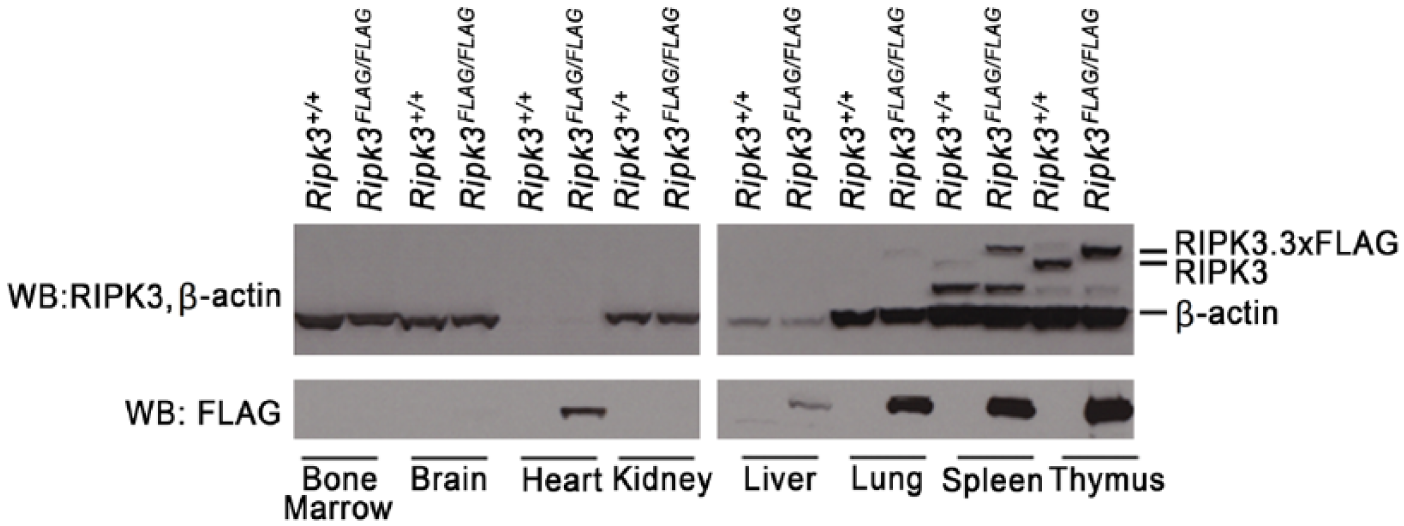
Western blots (WB) of wild-type (Ripk3+/+) and Ripk3−/− tissues. Note, F-actin, but not β-actin, is expressed in the heart. Additional bands identified in the spleen and thymus likely represent RIPK3 splice variants or cleavage products. The lighter β-actin band for the liver suggests that the samples may have been underloaded.
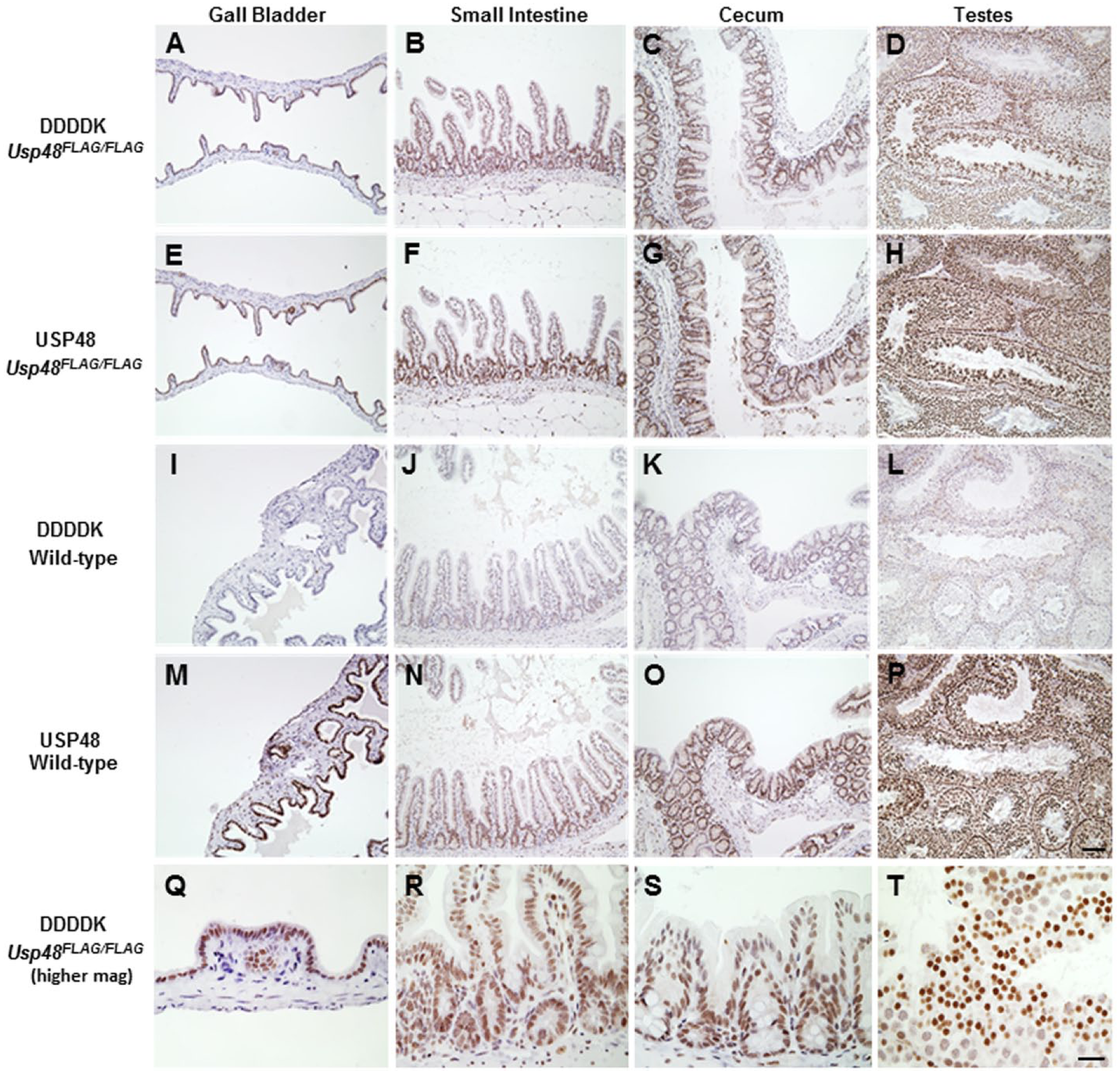
Tissues from 13-week-old Usp48FLAG/FLAG mice stained with anti-DDDDK antibody (A–D, top panel) or anti-USP48 antibody in adjacent sections (E–H, second panel). USP48 is localized to the nucleus when utilizing both methods. No reactivity in wild-type tissues is observed with the anti-DDDDK antibody (I–L, third panel), and a similar pattern of expression is observed with the commercially available USP48 antibody (M–P, fourth panel) when comparing Usp48FLAG/FLAG tissues to wild-type tissues. Higher magnification for anti-DDDDK IHC of USP48FLAG/FLAG tissues is shown in bottom panel (Q–T). Scale (A–P),100 µm; (Q–T), 20 µm.
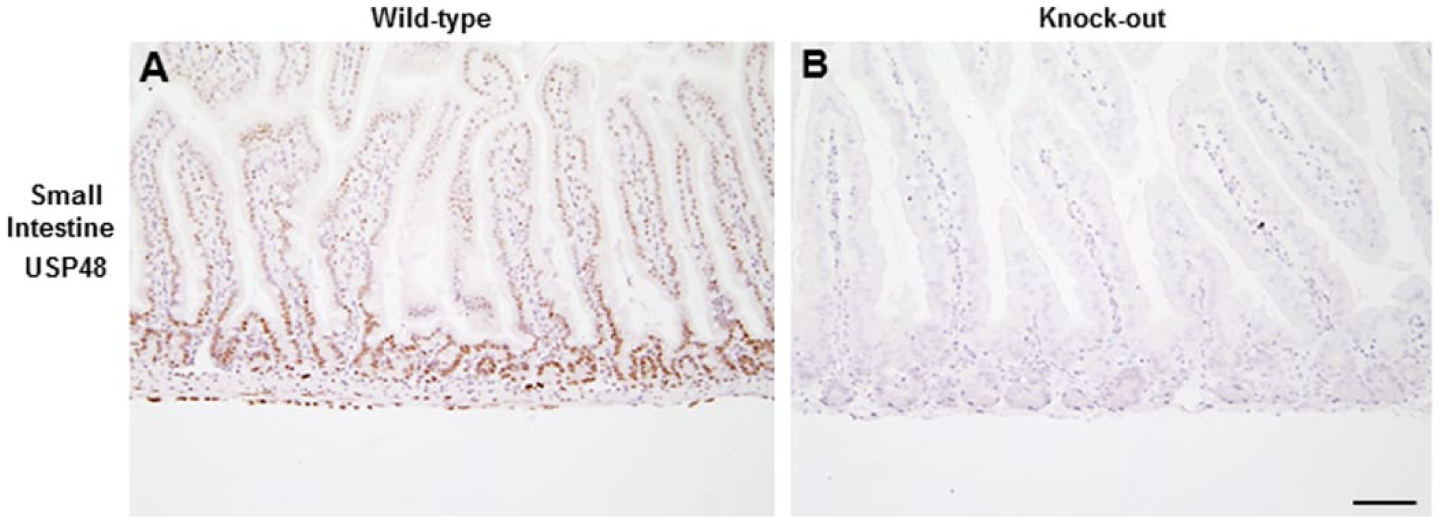
USP48 is stained by anti-USP48 antibody in cells of the intestinal crypts (A) but not in USP48-deficient crypts (B). Scale, 100 µm.
With the anti-DDDDK antibody, at 12.5 d.p.c. strong nuclear expression was observed in all tissues from Usp48FLAG/FLAG mice, whereas at 15.5 d.p.c., there remained moderate to weak expression in most tissues, although there was diminished immunoreactivity in the liver, brain and connective tissue (Fig. 7). By 18.5 d.p.c., USP48.3×FLAG continued to be detected in the ganglia, salivary gland, intestine, epidermis and thymus. There was also heterogeneous detection in the brain and connective tissues, and moderate to weak expression in the liver and lung. USP48 IHC results obtained with Usp48FLAG/FLAG embryos also correlated well at all time points evaluated, with strong nuclear expression of USP48 in all tissues at 12.5 d.p.c. and increasing differential expression at 15.5 and 18.5 d.p.c. (Fig. 7). Similar to what we observed for RIPK3, the addition of the C-terminal tag to USP48 did not alter the expression or distribution of USP48 (Figs. 5 and 7).
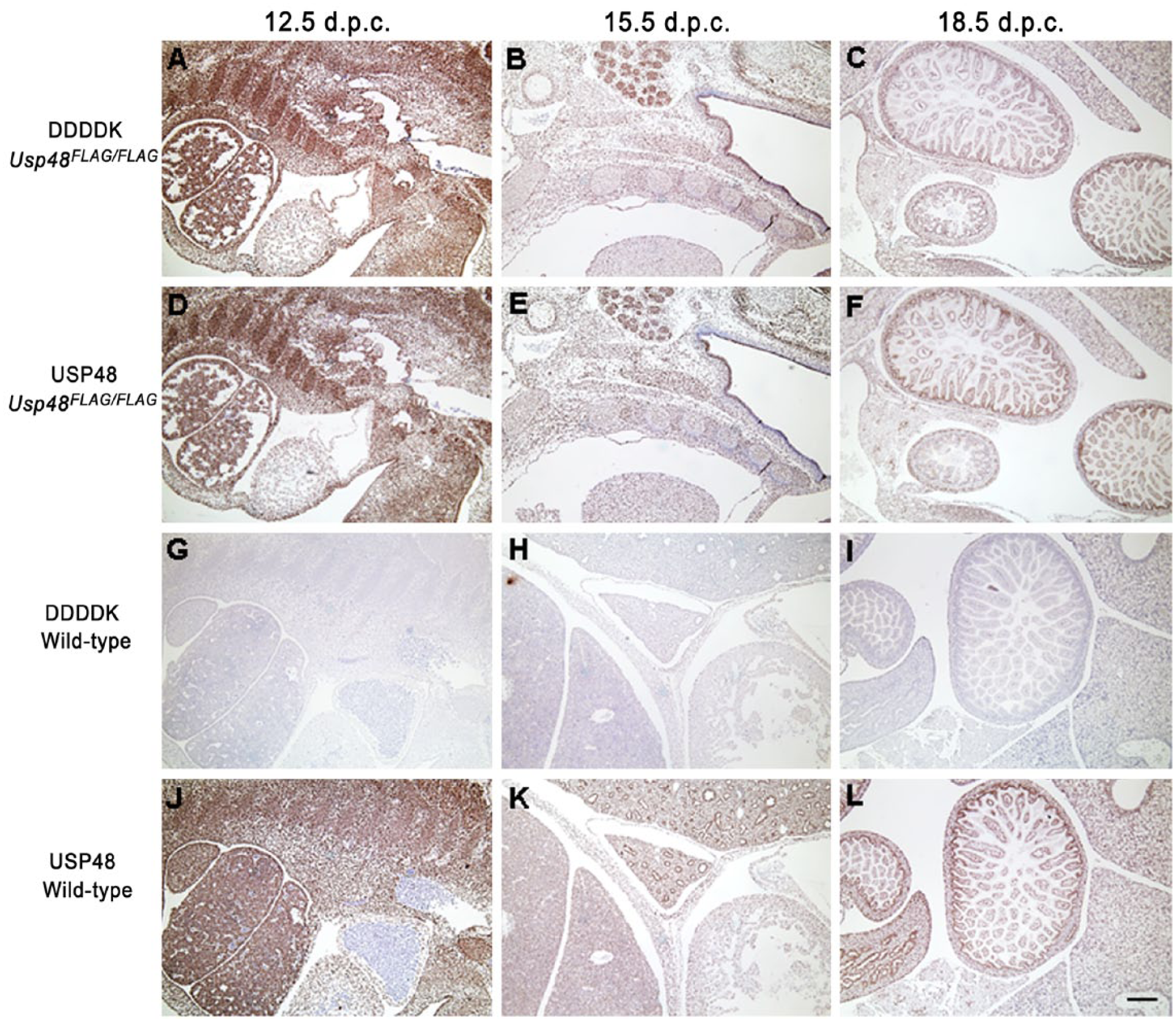
Usp48FLAG/FLAG embryos at different stages of development were stained with the anti-DDDDK antibody (A–C, top panel) or anti-USP48 antibody (D–F, second panel). USP48 is widespread in all tissues at 12.5 days post coitum (d.p.c.). Heterogeneous expression patterns are observed as the embryo matures to 15.5 and 18.5 d.p.c. No reactivity in wild-type embryos is observed with the anti-DDDDK antibody (G–I, third panel), and a similar pattern of expression is observed with the commercially available USP48 antibody (J–L, bottom panel) when comparing Usp48FLAG/FLAG embryos to wild-type embryos. Scale, 200 µm.
Next, we used our FLAG IHC assay to investigate the expression of the BAP1 tumor suppressor in Bap1FLAG/FLAG mice (Dey et al. 2012). BAP1 is a deubiquitinating enzyme for which quality IHC antibodies are unavailable. The anti-DDDDK antibody labeled nuclei in the bronchiolar epithelium as well as type II pneumocytes in the lung, although, staining observed in the pneumocytes was weaker when compared with that in the bronchiolar epithelium. Nuclear labeling was also observed in the gall bladder epithe-lium and Kupffer cells, and weaker nuclear labeling was identified in lymphocytes within the spleen, lymph node and thymus (Fig. 8). In these experiments, a weak background signal was also seen in control wild-type tissues, probably due to the non-specific binding of proteins containing sequences similar to the FLAG or 3×FLAG epitope. This background signal was not observed in the wild-type tissues used for the USP48.3×FLAG or RIPK3.3×FLAG IHC, but differences in tissue fixation and/or processing for the BAP1.3×FLAG tissues may have resulted in this mild background. By western blotting, BAP1 protein was detected in wild-type liver but not in Bap1−/− liver that was harvested three weeks post-Tamoxifen treatment, validating the FLAG IHC assay in the liver (Fig. 8).
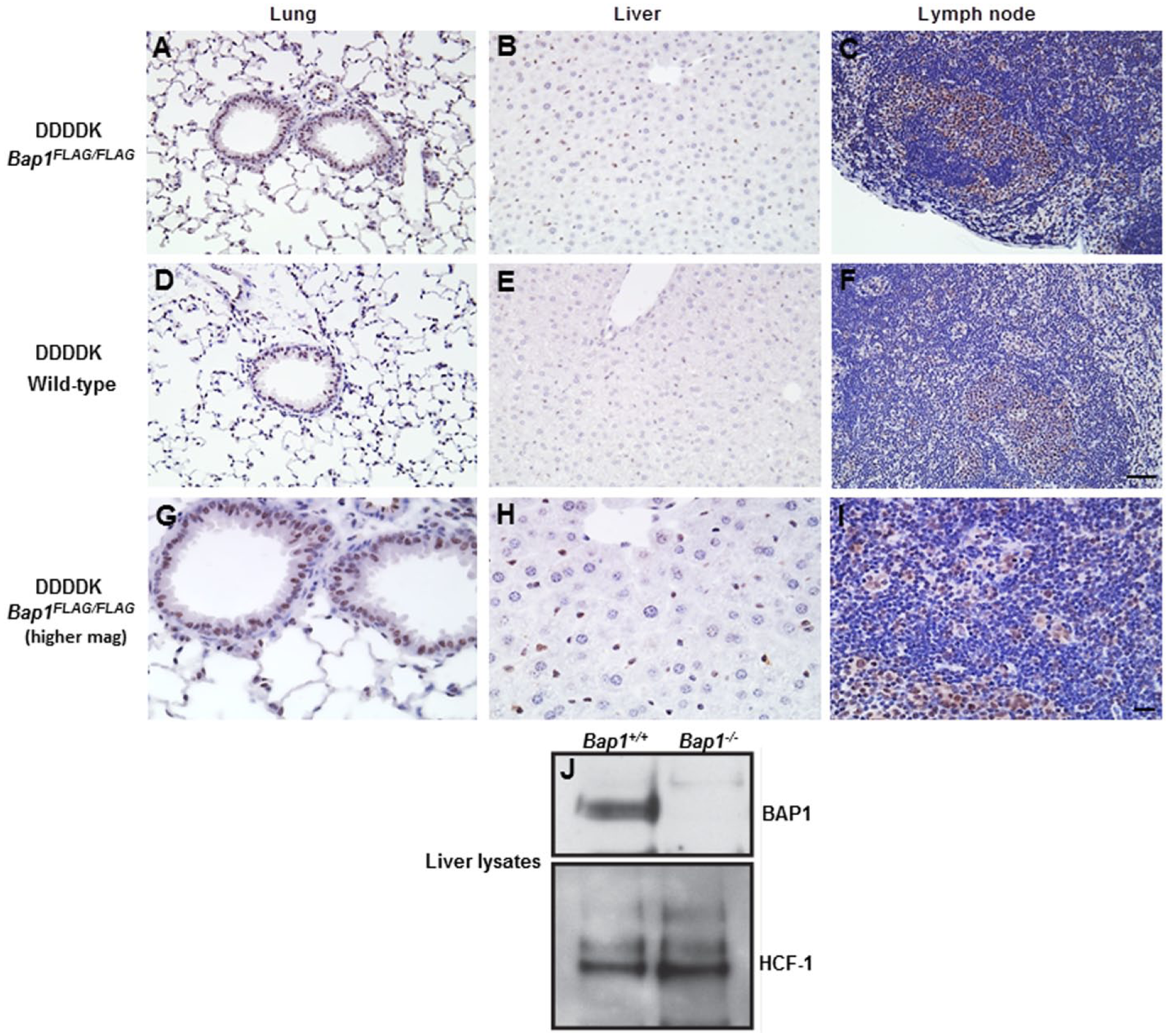
Tissues from 14-week-old Bap1FLAG/FLAG mice stained with anti-DDDDK antibody. Bap1 is localized to the nucleus (A–C, top panel). Background is observed in the corresponding wild-type tissues (D–F; middle panel), but at a much lower intensity. Higher magnification for anti-DDDDK IHC of Bap1FLAG/FLAG tissues is shown in bottom panel (G-I). Western blots of wild-type liver (Bap1+/+) confirms expression for BAP1 in the liver, with expression of BAP1 absent in Bap1−/− liver (Panel J). Scale (A–F), 100 µm; (G–I), 30 µm.
Overall, the labeling for BAP1.3×FLAG using the anti-DDDDK IHC assay was significantly weaker when compared with the signals for USP48.3×FLAG and RIPK3.3×FLAG. Immunoblotting with anti-FLAG antibody confirmed that BAP1 is less abundant than RIPK3 and USP48 in many tissues, including in the spleen (Fig. 9). The anti-DDDDK IHC that was performed on the knock-in spleens correlated well with the WB data (Fig. 9).
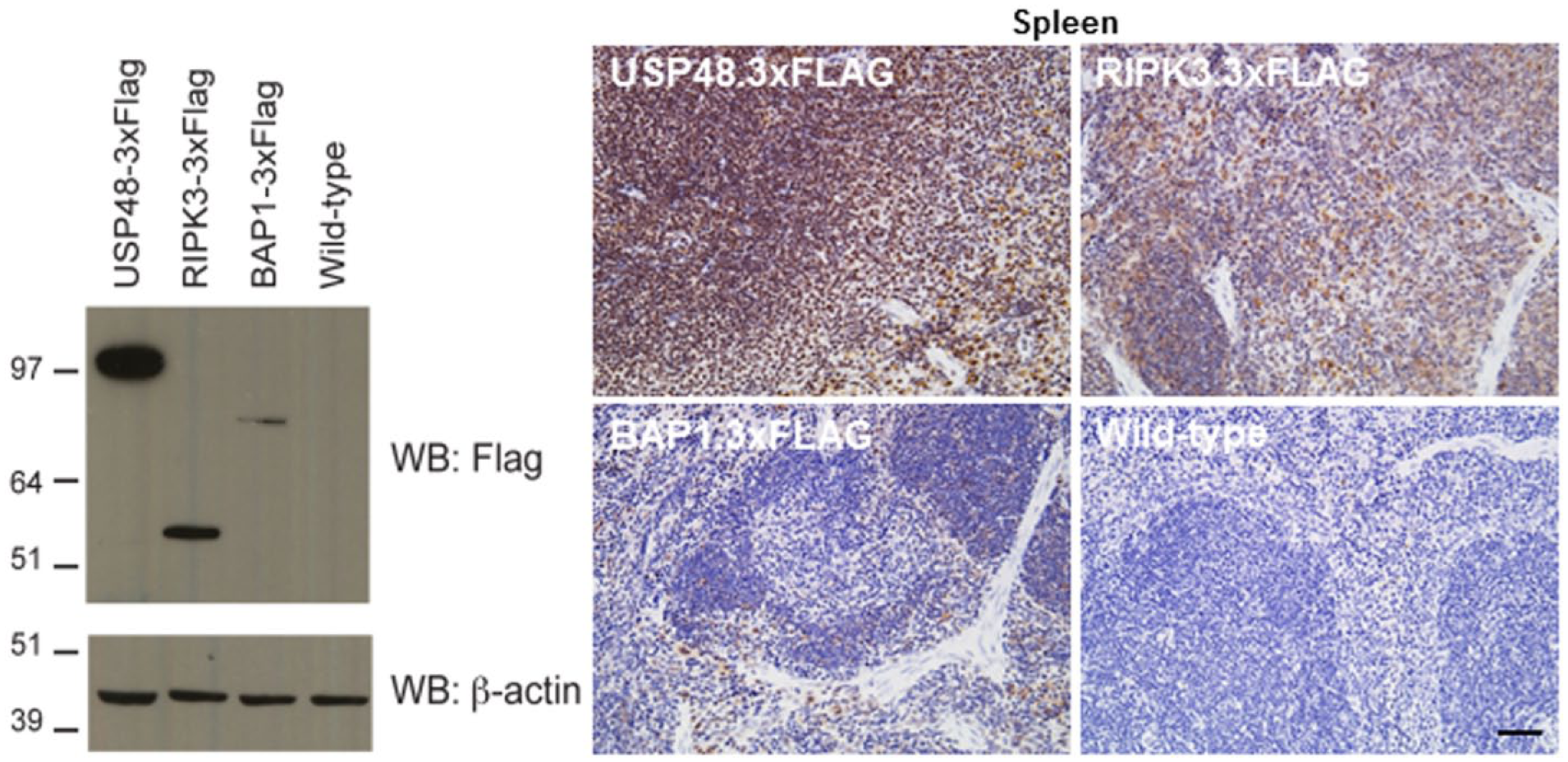
Immunoblotting of mouse USP48.3×FLAG (123 kDa), RIPK3.3×FLAG (56 kDa), and BAP1.3×FLAG (83 kDa) showing relative expression of USP48, RIPK3 and BAP1 in spleen (left panel). 3×FLAG IHC with anti-DDDK antibody showing correlation of expression levels with western blotting data using the same constructs (right panel). Scale, 100 µm.
Labeling with the anti-DDDDK antibody was observed in all adult mouse brain sections irrespective of genotype (Fig. 10). Multiple FLAG antibodies have been evaluated and our findings are consistent with the publication by Schäfer and Braun (1995), who clarified that this non-specific signal is likely due to cross-reactivity with proteins in the brain, such as β-phosphatases, that contain sequences very similar to the FLAG epitope. No such background was observed in the embryonic brain, which likely reflects differences in the proteome between the adult and fetal brains.
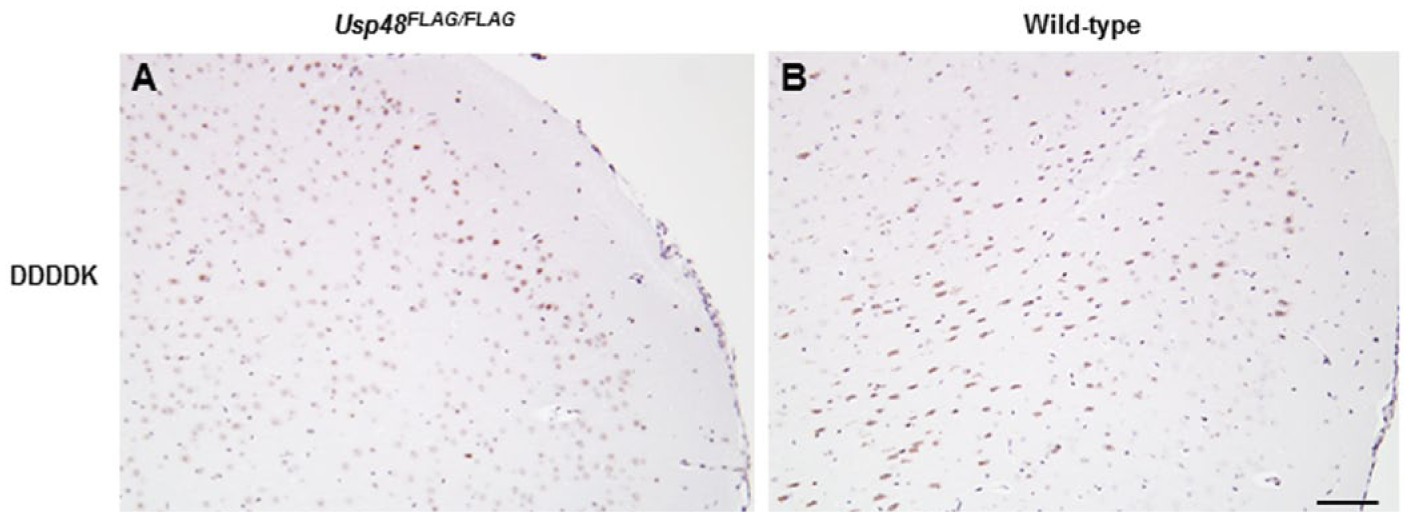
Brains from 3-week-old Usp48FLAG/FLAG (A) and wild-type (B) mice stained with anti-DDDDK antibody. Scale, 100 µm.
Discussion
When considering IHC as a method for the characterization of protein expression in murine tissues, the limiting factor is often a lack of commercially available antibodies. The novel method described in this study circumvents this issue by utilizing knock-in mice that express a 3×FLAG epitope-tagged protein that can be detected by IHC with a goat anti-DDDDK antibody. This method is also novel in that this is the first time a protein of interest has been triple tagged with the FLAG peptide, which we believe has increased the sensitivity of the assays utilized for this study. Up to now, only single tagged proteins have been utilized. This polyclonal antibody recognizes the FLAG epitope and, based on our results, specifically recognizes at least two 3×FLAG-tagged proteins (RIPK3 and USP48) in all mouse tissues examined, except in the adult brain. Therefore, it represents an extremely useful universal reagent for the detection and characterization of proteins in murine tissues.
Although the FLAG IHC method has been described in transgenic animal models, most often in fixed-frozen tissues that yield sub-optimal morphology (Campbell et al. 2012; Ruggiero et al. 2013; Seo et al. 2013), this method is novel in that it has been developed for paraffin-embedded knock-in mouse tissues. One major limitation of transgenic mice is that they often express the protein of interest at levels far in excess of the normal endogenous protein, leading to results not relevant to normal physiology. In contrast, knock-in mice should express the tagged protein at endogenous levels provided the tag does not compromise protein stability. This potential issue was not encountered with Ripk3FLAG/FLAG mice because levels of RIPK3.3×FLAG were comparable to wild-type RIPK3 in multiple mouse cell types by western blotting (Fig. 4). The addition of a 3×FLAG tag to BAP1 or USP48 did not grossly disrupt either protein because the Bap1FLAG/FLAG and Usp48FLAG/FLAG mice did not exhibit the embryonic lethality associated with loss of BAP1 or USP48 (Dey et al. 2012 and our unpublished data).
Another limitation of the FLAG IHC assay and its application to FLAG or 3×FLAG knock-in tissues appears to be non-specific binding of proteins containing sequences similar to the FLAG or 3×FLAG epitope, which appear to be particularly abundant in the adult brain. This issue may also prove problematic for very low abundance proteins in other tissues, because signal amplification increases the non-specific signals (unpublished data). Although the potential background staining that can accompany the FLAG IHC is a limitation of the assay, we believe the main caveat of the knock-in approach is that a tag can potentially compromise protein function; however, this issue was not encountered in either the Usp48FLAG/FLAG, Ripk3FLAG/FLAG or Bap1FLAG/FLAG mice. Nonetheless, we have established, using relatively well-expressed proteins (RIPK3 and USP48) and a less well-expressed protein (BAP1), that the IHC method can be used to detect a range of cytoplasmic and nuclear proteins. In addition to IHC, additional benefits can be realized with this knock-in approach, which include affinity purification of signaling complexes that can then be defined by mass spectrometry (Dey et al. 2012).
The alternative to making epitope-tagged knock-in mice is to generate quality antibodies that specifically recognize the protein of interest. Knock-in mouse and antibody generation each require a considerable investment of resources, but for antibody generation there is no guarantee that such efforts will yield the desired IHC reagent. In contrast, the path to generating a knock-in mouse is well defined. In addition, recent advances in genetic engineering using the CRISPR/Cas system (Yang et al. 2013) have the potential to further shorten time lines for the creation of epitope-tagged knock-in mice. One may also inquire if other tags may be better utilized, such as hemagglutinin (HA). The potential use for HA-tagged knock-in mice has been explored, and of the three HA-tagged knock-in mice that we have generated, two exhibited hypomorph or knock-out phenotypes, indicating that the addition of the HA tag was deleterious. The third HA-tagged knock-in mouse model made a functional protein, but the protein was expressed in such low abundance that we have never detected it using HA or protein-specific antibodies in any application. In comparison, we have observed a knock-out phenotype in one FLAG knock-in mouse (data not shown), so it is difficult to say that one tag will be consistently better than the other. In addition, one may also want to test the effects of tagging the N-terminus versus the C-terminus in cell-based overexpression systems prior to making the knock-in mice. This testing does, however, rely on some prior knowledge of the signaling pathways being investigated and, unfortunately, the effect of tagging an endogenous protein has not always been revealed in tests using overexpression systems. However, our current success with several FLAG knock-in strains suggests this is a viable approach for the detection of many unique proteins. We have not methodically evaluated whether homozygous FLAG/FLAG is critical or if FLAG/+ heterzygous mice could work, but one can predict that, for highly expressed genes, heterozygosity would suffice.
Although the FLAG IHC assay does not appear complex, there are some aspects of the method that should be emphasized. For example, although antigen retrieval buffers can be recycled, it was discovered during the development of this assay that the citrate retrieval buffer had to be freshly prepared or it compromised the detection of the DDDDK epitope. In addition, amplification reagents, such as TSA (Perkin Elmer, Waltham, MA) and HQ Amp (Ventana Medical Systems, Tucson, AZ) had to be excluded from this assay because they produced high background signal in wild-type tissues, most likely due to amplification of the non-specific signal from endogenous untagged proteins bearing FLAG-like epitopes.
In summary, we describe an IHC method that will enable researchers to characterize proteins in murine tissues in those instances when commercial reagents are lacking. This method is also universal in that a single protocol utilizing a commercially available antibody that recognizes the FLAG or 3×FLAG epitope can be used to characterize multiple markers of interest.
Footnotes
Acknowledgements
We thank Shari Lau (Genentech, Inc.) for IHC consultation/collaboration, Allie Maltzman and Mira Chaurushiya (Genentech Inc.) for generation of WB data, Reenal Pattni (Genentech, Inc.) for generation of the Usp48.3×FLAG KI construct, Merone Roose-Girma, Tim Soukup, Megan Schlatter, and Michele Bauer (Genentech, Inc.) for generation of previously unpublished mouse models and Charles Jones (Genentech, Inc.) for histology assistance.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: All funding was provided for by the
