Abstract
Sorafenib represents the first effective targeted therapy for advanced stage hepatocellular carcinoma (HCC); however, adequate patient stratification regarding sorafenib-responsiveness is still missing. Our aim was to analyse the association between the pretreatment microRNA profile of HCC and patient survival under sorafenib treatment. Total RNA was extracted from diagnostic fine-needle aspiration biopsy (FNAB) cytological smears of 20 advanced stage HCC patients collected between June 2008 and July 2012. All patients underwent sorafenib administration after FNA. Clinicopathological and survival data were recorded. Fourteen frequently deregulated miRNAs in HCC (miR-17-5p, miR-18a, miR-21, miR-34a, miR-122, miR-195, miR-210, miR-214, miR-221, miR-222, miR-223, miR-224, miR-140, miR-328) were tested by qRT-PCR. NormFinder software was used to select proper miR (mir-140) as a reference. Satisfactory amount of total RNA was obtained from all the considered samples (mean 10.8 ± 9.3 µg, range 0.2–32.2 µg). Among the analysed miRNAs, high miR-214 expression was associated with smaller tumor size (p=0.019), whereas high miR-17-5p expression correlated with better Eastern Cooperative Oncology Group performance status (p=0.003). The survival analysis revealed that high miR-224 expression was associated with increased progression-free and overall survival (PFS p=0.029; OS p=0.012). Pretreatment microRNA profiling, especially miR-224 expression, might serve as an ancillary tool for the better assessment of expected survival rates for patients under sorafenib treatment.
Introduction
Hepatocellular carcinoma (HCC) is one of the most common malignancies worldwide with high mortality rates and growing incidence in the western world (Jemal et al. 2011). Although potentially curative surgical treatments are available, only a minority of patients with localized small tumors are eligible for these therapeutic options. Unfortunately, a great percentage of patients are diagnosed at an advanced stage of disease (Pinter et al. 2011), for whom sorafenib, a multi-kinase inhibitor, is the first effective systemic molecular-targeted treatment option (Lachenmayer et al. 2010; Llovet et al. 2008).
Sorafenib has been shown to significantly prolong overall survival in patients with advanced HCC (Llovet et al. 2008). However, there are few validated and reliable molecular markers with which to predict patient outcome under sorafenib treatment. Several markers that could potentially stratify HCC patients according to sorafenib responsiveness were investigated recently: PDGFR-β and c-Met were suggested as predictors of prognosis and therapeutic effectiveness in sorafenib-treated HCC patients (Chu et al. 2013), whereas Mcl-1 and pERK expressions were found to be associated with overall survival (Personeni et al. 2013). The activation of c-Jun N-terminal kinase in HCC patients correlated inversely with therapeutic response to sorafenib (Hagiwara et al. 2012) and further factors, including serum cholinesterase levels (Takeda et al. 2013), plasma VEGF concentration changes (Tsuchiya et al. 2014) and VEGF and VEGFR gene polymorphisms (Scartozzi et al. 2014) were also presented as potential clinical predictors of sorafenib sensitivity. Given the high costs and relatively high percentage of “non-responders” to sorafenib [disease control rates have been indicated at 43% (Llovet et al. 2008) and 53% (Cheng et al. 2009)], the identification of potential predictive biomarkers represents a challenge for research and is of great interest.
MicroRNAs (miRNAs, miRs) are small (19-25 nt), non-coding, posttranscriptional regulators of gene expression commonly dysregulated in tumors. They target both tumor suppressor and oncogenic pathways (Gramantieri et al. 2008; Lujambio and Lowe 2012) and their altered expression carries great diagnostic and prognostic potential (Calin et al. 2005; Lu et al. 2005; Minguez and Lachenmayer 2011). In addition, miRNAs are highly stable and are reliably detected in archival clinical samples and cytology specimens, and are therefore considered to be ideal biomarker candidates (Fassina et al. 2012). Several in vitro data suggest that miRs can modulate the sensitivity of HCC tumor cells to therapy; for example, miR-122 sensitizes HCC cells to adriamycin, vincristin (Xu et al. 2011) and sorafenib (Bai et al. 2009); miR-34a induces sensitivity to sorafenib in human HCC cells (Yang et al. 2013); and miR-21 induces resistance to interferon-α/5-fluorouracil in HCC cells (Tomimaru et al. 2010). Moreover, low miR-21 expression has been associated with better clinical response and overall survival and was suggested as a potential predictor for the clinical response to IFN-α/5-FU combination therapy (Tomimaru et al. 2010).
HCCs at advanced stages are often associated with a significantly elevated risk of bleeding; therefore, fine-needle aspiration biopsy (FNAB) is favored over conventional liver biopsy (Fassina et al. 2010). There are few data of miRNA expression in FNAB samples. Simonato et al. (2013) recently demonstrated that archival cytology smears are suitable for obtaining high-quality RNA to be used in microRNA expression profiling. To our knowledge, there is no report to date on miRNA expression of liver FNAB samples, particularly for sorafenib-treated HCCs. Given the great predictive potential of miRNAs, our aim was to investigate the pretreatment miRNA expression of diagnostic, ultrasound-guided FNAB samples of sorafenib-treated, advanced stage HCC patients and to correlate the obtained results with clinical data. We analysed a panel of 14 miRNAs that are frequently dysregulated in HCCs (Karakatsanis et al. 2013; Varnholt et al. 2008). This is the first study to reveal a significant association between pretreatment miRNA expression of FNAB samples and the outcome of sorafenib-treated HCC patients.
Materials & Methods
Patients and Samples
Twenty HCC FNAB samples from the archives of the 2nd Department of Pathology, Semmelweis University, Budapest, were analysed in this study. In Hungary, sorafenib treatment is applicable only after pathological morphological confirmation of HCC diagnosis: either histology or FNAB sampling is mandatory in order to treat the patients. Diagnoses of HCC were established from FNAB samples by two expert cytopathologists (BJ and ES) between June 2008 and July 2012. Patients were subsequently treated with sorafenib at the Department of Oncology, United Saint Stephen and Saint Laszlo Hospital and Outpatient Clinics, Budapest.
All patients in this study met the following criteria: no prior surgery, loco-regional or systemic treatment, no subsequent loco-regional treatment after the end of sorafenib therapy, Child-Pugh class A or B, Eastern Cooperative Oncology Group (ECOG) performance status ≤1, more than 3 months survival after diagnosis, available follow-up data (follow-up deadline June 2013). Overall survival (OS) was defined as the time period from the start of sorafenib therapy until the date of death or end of follow-up. Progression-free survival (PFS) was defined as the time period between sorafenib initiation and progression or end of follow-up. Data and material were obtained with permission from the Regional Ethical Committee of Semmelweis University (# 137/2008) following the ethical guidelines of the 1975 Declaration of Helsinki and with informed consent of the patients.
The presence of >2000 well-preserved and well-visualized HCC cells (as observed in 10 fields at 4× magnification) was appointed as minimum accepted cellularity. Only slides with a proportion of tumor cells >50% and without massive necrotic debris were selected (Fig. 1).
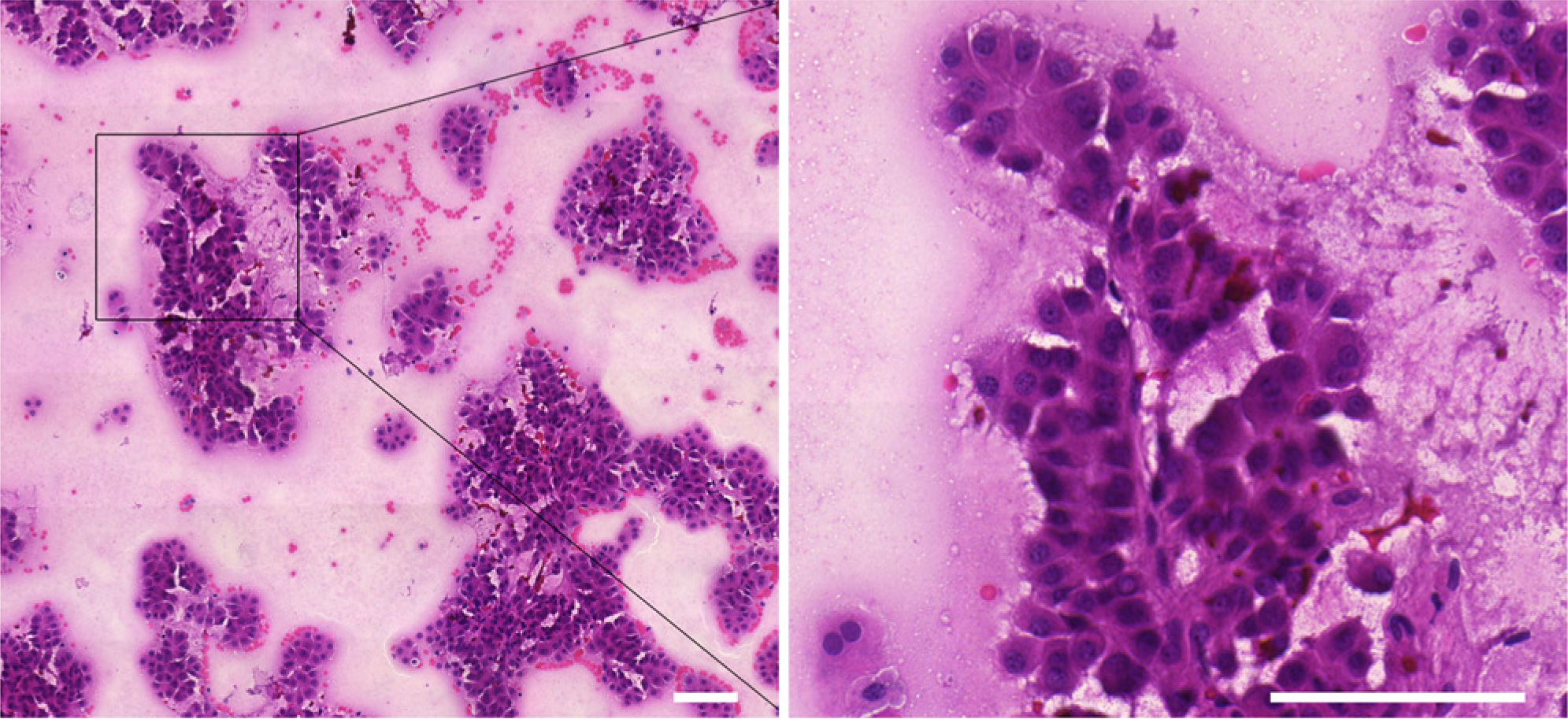
Representative image of a hepatocellular carcinoma sample obtained by fine-needle aspiration biopsy. Irregularly shaped tumor aggregates with transgressing endothelium can be seen. The image on the right shows a magnified view of the image on the left. Scale bars 100 µm.
Real-time RT-PCR Analysis
RNA Isolation
Total RNA was isolated from archival FNAB slides (fixed by Cytofix spray) using RNeasy FFPE Kit (Qiagen; Hilden, Germany). Briefly, slides were immersed in xylene for 72 hr to remove coverslips. Cells were scraped off the slides with sterile disposable knives and transferred into 1.5 ml tubes where proteinase K digestion was performed for 12 hr. Further procedures were carried out according to the manufacturer’s instructions, with modification for co-purification of miRNA (Doleshal et al. 2008). Traces of genomic DNA were removed using Turbo DNase digestion (Ambion; Austin, TX) following the supplier’s protocol. RNA concentration and quality were measured using a NanoDrop ND-1000 Spectrophotometer (NanoDrop Technologies Inc.; Wilmington, DE).
Reverse Transcription (RT) and Real-Time PCR (qPCR)
The following TaqMan MicroRNA Assays (Applied Biosystems; Foster City, CA) were used to determine the levels of the selected 14 miRNAs: miR-17-5p (ID:000393), miR-18a (ID:002422), miR-21 (ID:000397), miR-34a (ID:000426), miR-122 (ID:002245), miR-140 (ID:000462), miR-195 (ID:000494), miR-210 (ID:000512), miR-214 (ID:002306), miR-221 (ID:000524), miR-222 (ID:002276), miR-223 (ID:002295), miR-224 (ID:002099), miR-328 (ID:000543) and U6 (ID:001973). RT and qPCR were performed according to manufacturer’s instructions. Briefly, RT was carried out using TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems) in a final volume of 7.5 µl containing 10 ng total RNA. qPCR was performed using TaqMan Universal PCR Master Mix No AmpErase UNG (Applied Biosystems) in a final volume of 10 µl containing 0.65 µl RT product using an ABI Prism 7000 Sequence Detection System (Applied Biosystems). Reactions were run in triplicate, including no-template controls.
Relative quantification of target miRNA expression was performed using 2Cq(ref)-Cq(target) method, with miR-140 as the reference. From the candidate reference small RNAs (U6, miR-328, miR-140) to be analysed, miR-140 proved to be the most stable miRNA in our sample set, as defined by the NormFinder application (Andersen et al. 2004). Additionally, miR-140 was also used as reference in a previous study on HCCs (Varnholt et al. 2008).
Statistical Analysis
Statistical analyses were carried out using STATISTICA software v11 (StatSoft; Tulsa, OK). Patients were dichotomized into “high miR expression” and “low miR expression” groups according to the median relative expression of a given miR in order to obviate the effects of extreme values. Associations between miR expression and clinicopathological characteristics were examined using Fisher’s exact test. Survival curves were computed using the Kaplan-Meier method and differences were assessed using the log-rank test. Multivariate survival analyses were performed using the Cox proportional hazards model. P values ≤ 0.05 were considered statistically significant.
Results
Patient Characteristics and FNAB miR Expression
A total of twenty sorafenib-treated patients were included in the present analysis. The median age at the time of diagnostic aspiration was 68 years (range, 52–82 years), with a male:female ratio of 16:4. Patients were mainly Child-Pugh class A (16/20), with ECOG performance status 0 (13/20). The etiology of the underlying liver disease was predominantly viral (11/20), and tumor sizes ranged from 1.5 to 20 cm (median, 5 cm). Tumors were mostly multinodular (14/20). Elevated serum alpha-fetoprotein (AFP) levels (≥10 ng/ml) were present in 13/20 patients (Table 1). Of the 20 patients, 15 died during the follow-up period, with 5 patients still alive at the end of the study. The median follow-up time was 33.6 months, and the median OS and PFS for all patients was 6.2 months (95% CI 2.7–9.3 months) and 5 months (95% CI 3.5–6.5 months), respectively.
Baseline Clinicopathological Characteristics of Patients (n=20).
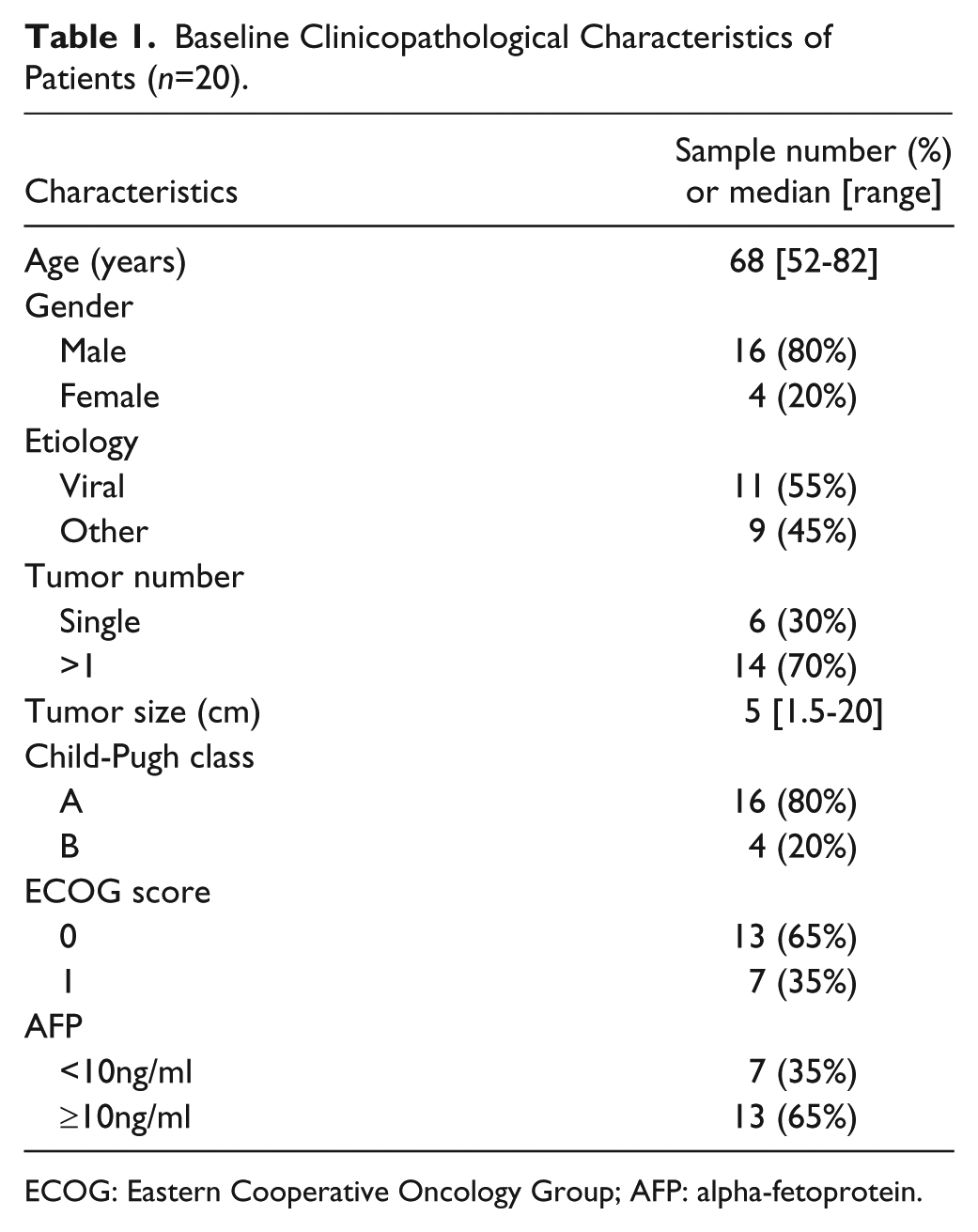
ECOG: Eastern Cooperative Oncology Group; AFP: alpha-fetoprotein.
A satisfactory amount of total RNA was obtained from all considered samples. The mean ± SD RNA yield was 10.8 ± 9.3 µg (range, 0.2–32.2 µg) with an OD260/280 ratio of 2.0 ± 0.1. The normalized relative expression values of miRs are shown in Table 2.
miRNA Expression in Hepatocellular Carcinoma Fine-needle Aspiration Biopsy Samples Relative to miR-140.
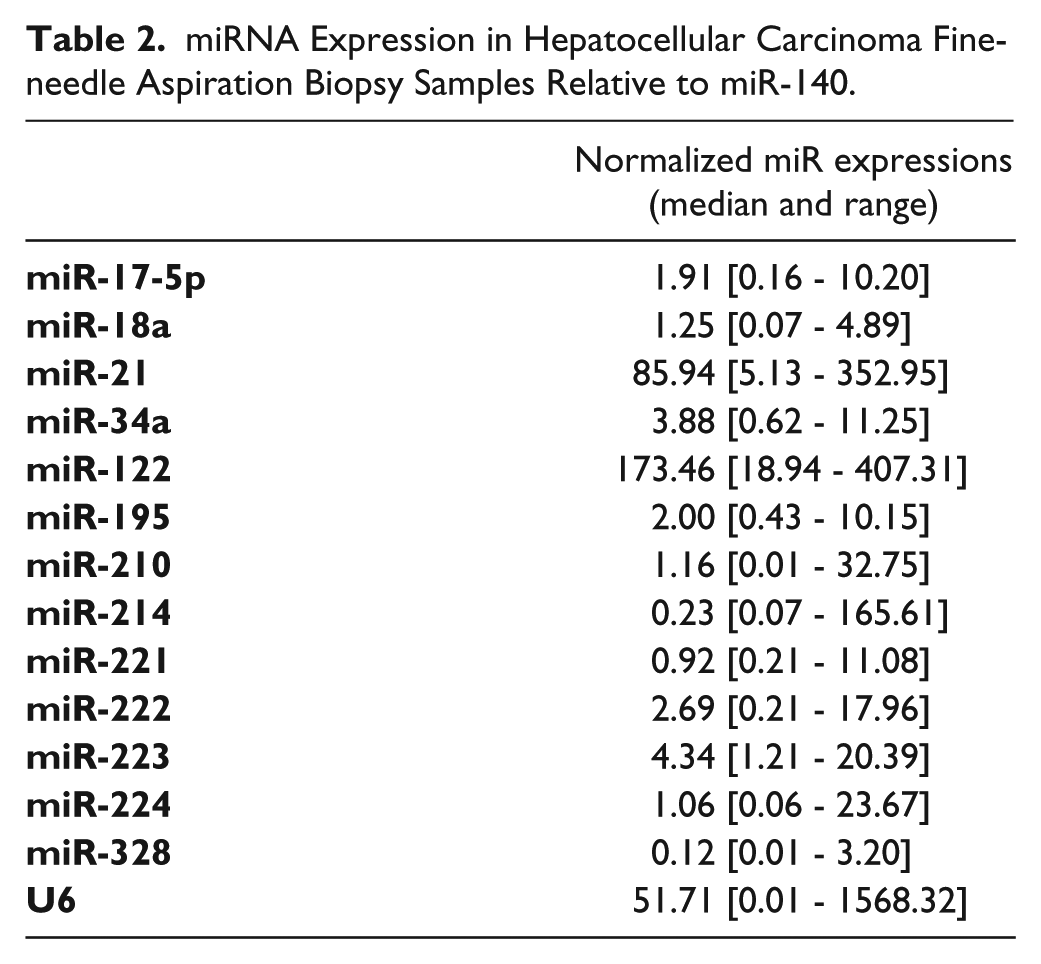
Relationship between Pretreatment miR Expression and Clinicopathological Features
Relationships between miR expression and clinicopathological features, such as age (<68 years / ≥68 years), gender (male/female), etiology (viral / other), tumor number (single / multinodular), tumor size (≤5cm / >5cm), Child-Pughstatus (A / B), ECOG status (0 / 1) and AFP levels(<10 ng/ml / ≥10 ng/ml), were analysed to identify possible associations. Median miR expression values were used to divide patients into high and low expression groups.
High miR-214 expression was more frequent in patients with smaller tumors (≤5 cm) than in larger ones (>5 cm) (80% vs 20%, p=0.019). High miR-17-5p expression was more frequent in patients with better performance status than in patients with an ECOG score of 1 (77% vs 0%, p=0.003). All other analysed miRs did not reveal significant correlations with patient characteristics (for detailed 2×2 contingency tables, see Supplementary Table S1).
Relationship between Pretreatment miR Expression and Outcome under Sorafenib Treatment
To analyse possible correlations between miR expression and clinical outcome under sorafenib treatment, Kaplan-Meier analyses and log-rank tests were performed on “high miR expression” and “low miR expression” patient groups. We found that both PFS and OS rates were better in the “high miR-224” group than in the “low miR-224” group (PFS, p=0.029; OS, p=0.012) (Figs. 2 and 3). The other analysed miRs did not show any significant associations with either PFS or OS of sorafenib-treated patients in our sample set (Table 3).
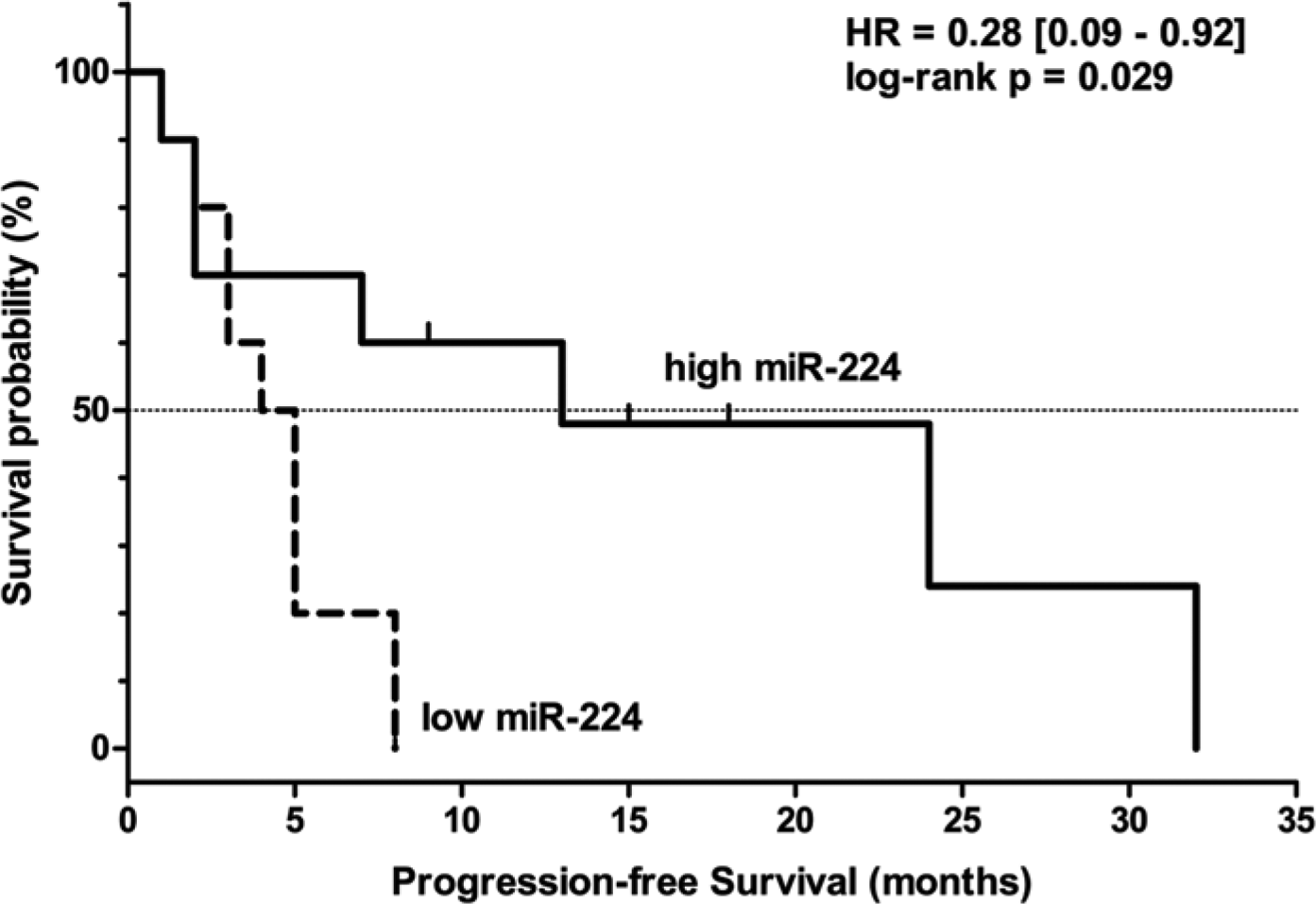
Kaplan-Meier curve showing progression-free survival of sorafenib-treated HCC patients according to high and low pretreatment miR-224 expression levels.
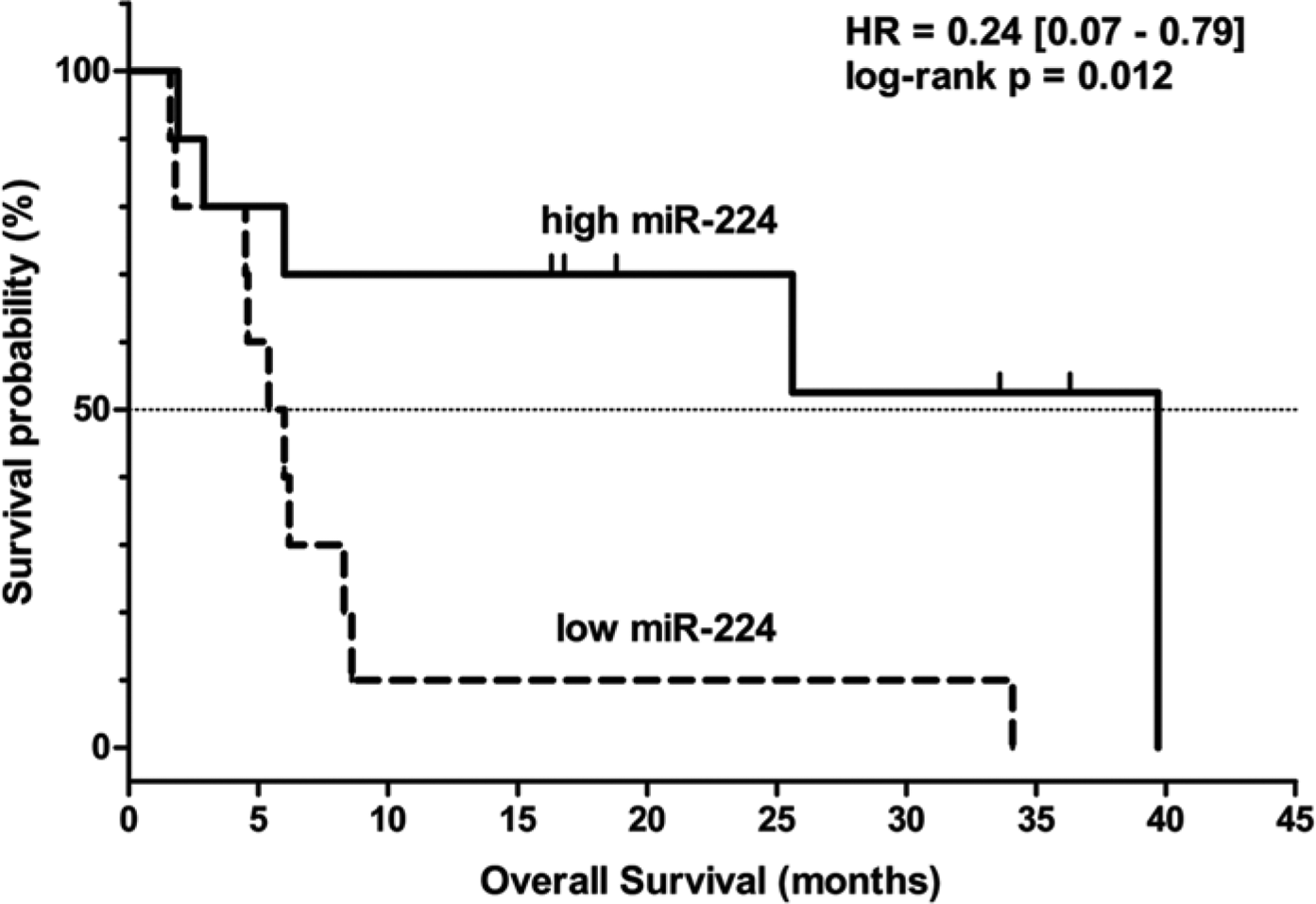
Kaplan-Meier curve showing overall survival of sorafenib-treated HCC patients according to high and low pretreatment miR-224 expression levels.
Results of Log-rank Statistics with Hazard Ratios of Univariate Cox Regression.
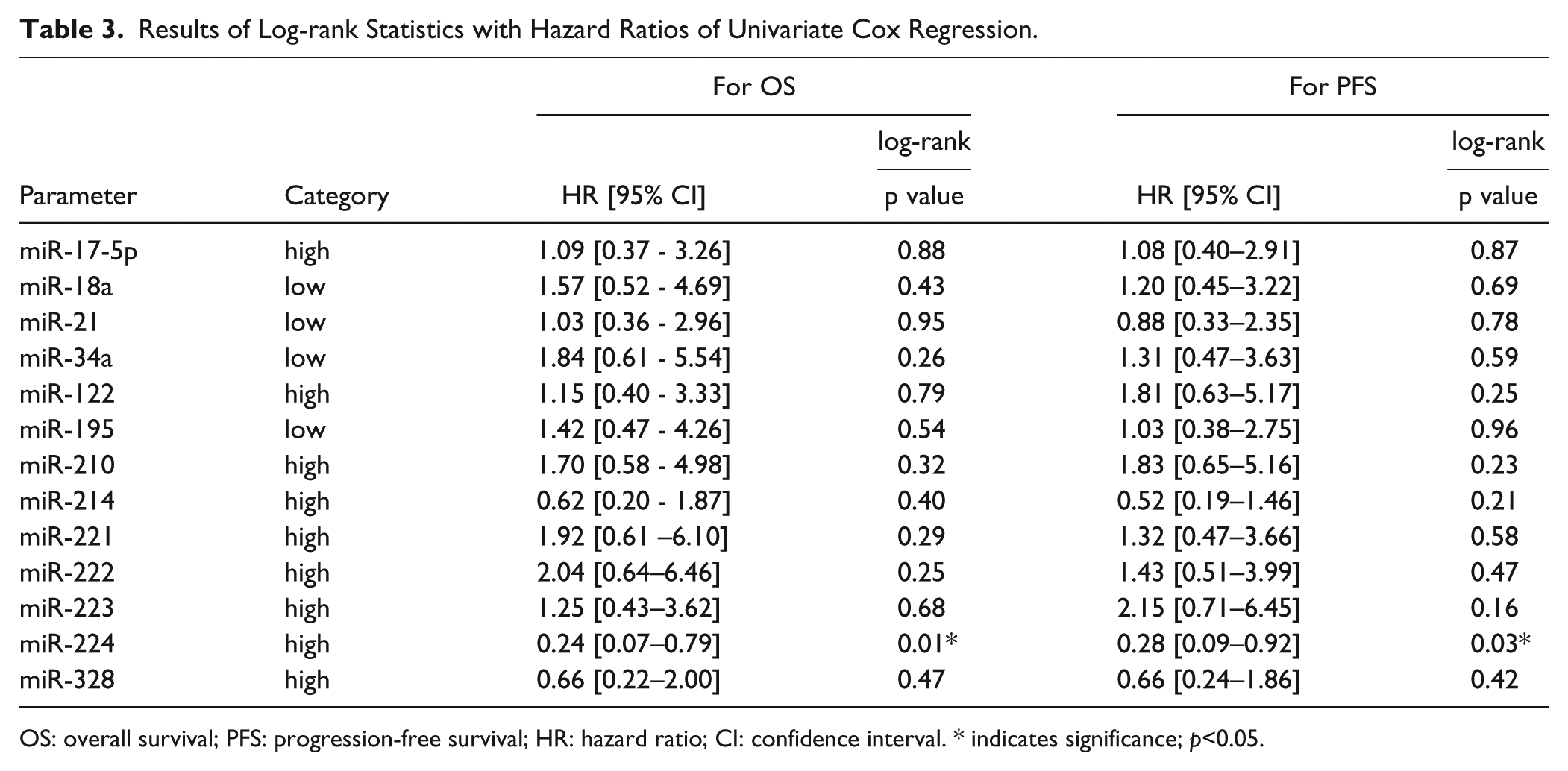
OS: overall survival; PFS: progression-free survival; HR: hazard ratio; CI: confidence interval. * indicates significance; p<0.05.
An analysis of the impact of clinicopathological characteristics on OS and PFS showed that none of the investigated parameters had a significant association with OS and only tumor size showed a marginal correlation with PFS (log-rank p=0.08; data not shown).
Although the parameters did not prove to be statistically significant, as potentially relevant prognostic factors they were forced into the Cox multivariate analysis in order to study whether miR-224 is an independent predictor of survival. Clinicopathological parameters, such as gender, age, tumor number, tumor size, Child-Pugh score, ECOG status and AFP level, were analysed. Based on this analysis, dichotomized miR-224 expression was found to be an independent prognostic factor for both OS and PFS (high vs low, OS HR=0.11 [95% CI 0.013–0.913] p=0.04, PFS HR=0.125 [95% CI 0.015–1.034] p=0.05) (Supplementary Table S2).
Discussion
Advanced HCC remains a devastating disease, with dismal prognosis. The introduction of sorafenib (Nexavar®, Bayer-Schering Pharma)—the first approved targeted therapeutic agent for HCC—represents a milestone in the management of this cancer type. Sorafenib is a potent multi-kinase inhibitor targeting proliferative and angiogenic signaling pathways, among others Raf/MEK/ERK signaling, vascular endothelial growth factor receptors 2 and -3 (VEGFR-2,-3) and platelet-derived growth factor receptor-β (PDGFR-β) (Wilhelm et al. 2004). Although sorafenib was proved to significantly prolong OS, it provides only limited clinical benefit due to the heterogeneity of the treated patients (Lachenmayer et al. 2010). Moreover, predicting the outcome of sorafenib-treated patients is still an unmet medical need because there is no consensus regarding the utility of such potential clinical or molecular markers (Baek et al. 2011, Pinteret al. 2011).
MicroRNAs are key epigenetic regulators of gene expression, interfering with both tumorigenic and tumor suppressor pathways and thus playing a significant role in carcinogenesis. In addition, alterations in the expression of miRNAs are thought to bear strong prognostic and predictive potential in malignant diseases such as HCC, among others. Their high stability and effective retrieval from archival clinical samples make miRNAs ideal biomarkers for diagnosis, prognosis and therapy responsiveness (Giordano and Columbano 2013). Even small and delicate samples, such as FNABs, are suitable for proper miRNA analysis, and this is supported by several reports on aspirated samples from the thyroid gland (Kitano et al. 2012), pancreas (Ali et al. 2012) and lung (Petriella et al. 2013).
In light of these studies, our aim was to investigate miRNA profiles of FNAB samples taken from advanced HCC patients and to correlate the obtained results with the prognostic data. Because the diagnostic FNABs were taken from pretreatment patients who had not received any prior surgical, loco-regional or systemic treatment, one could assume that the observed miRNA expression profiles truly reflect the unaltered molecular background of the tumors.
Our analysis revealed associations between miRNA expression and clinicopathological features of the patients. High miR-214 expression was associated with smaller tumor size, which is in line with previous data, as miR-214 is a known tumor suppressor miRNA shown to suppress the growth of HCC cells in vitro (Wang et al. 2012). Further, high miR-17-5p expression was associated with better ECOG performance status in the present study, which appears to contradict the findings of others, as elevated miR-17-5p expression has been previously correlated with worse prognosis and shortened survival times (Chen et al. 2012). However, we did not find a significant association between miR-17-5p and the outcome in patients treated with sorafenib.
When further assessing the prognostic significance of miRNAs, we found no significant correlations between other analysed miRs and prognosis, with the exception of miR-224. In vitro studies have previously proposed that certain miRNAs sensitize HCC cells to sorafenib: miR-122 restoration in HCC cells has been shown to significantly reduce clonogenic survival and growth upon sorafenib treatment (Bai et al. 2009), whereas miR-34a was shown to potentiate sorafenib-induced toxicity and apoptosis (Yang et al. 2013). In our sample set, neither of these miRs correlated with patient survival.
Intriguingly, pretreatment miR-224 expression proved to be associated with OS and PFS under sorafenib treatment. miR-224 is frequently upregulated and thus known as an oncomir in HCC, targeting cell proliferation, migration, invasion and apoptosis (Zhang et al. 2013). Upregulation of miR-224 has been described to confer poor prognosis to cervical cancer (Shen et al. 2013) and glioma (Lu et al. 2013) patients. In HCC patients, miR-224 expression did not show a correlation with survival; however, when miR-224 upregulation and simultaneous SMAD4 downregulation were taken into account together, they were associated with poorer survival of HCC patients (Wang et al. 2013). Although this might appear contradictory to our results, the high miR-224 phenotype that we found might also be associated with a better responsiveness to sorafenib and thus with better clinical outcome upon treatment. Moreover, in colon cancer cells, miR-224 confers sensitivity to methotrexate (Mencia et al. 2011) and, in HCC cell lines, celastrol exerts its antimetastatic effect through the down-regulation of miR-224 (Li et al. 2013). Curiously, a bipartite role has been described for miR-224 in HCC cell cycle regulation because it has the potential to promote both cell proliferation and apoptotis (Wang et al. 2008; Zhang et al. 2013). Thus, on the one hand, high miR-224 might promote tumor progression, whereas, on the other hand, it might have the potential to render HCC cells susceptible to (drug-induced) apoptosis.
Among the several putative miR-224 targets are PDGFR-α and PDGFR-β (Dweep et al. 2011; Murakami et al. 2006). Interestingly, the expression of both molecules was shown to predict poor prognosis (Chen et al. 2011) and shorter OS in patients with resected HCC (Patel et al. 2011). Moreover, PDGFR-β is a known target of sorafenib (Wilhelm et al. 2004). Intriguingly, in a previous study with an experimental design similar to ours, low PDGFR-β expression of biopsy specimens from sorafenib-treated patients was shown to be associated with longer OS (Chu et al. 2013). This is in line with our results, where we also found significantly prolonged OS of patients with high expression of miR-224, the potential negative regulator of PDGFR-β.
The limitations of this study are the relatively small sample size, the lack of non-tumorous control and the retrospective nature of the analysis. Larger cohorts and prospective studies would be needed to validate the utility of this approach. Nonetheless, this study supports the potential use of miRNAs as biomarkers and predictors of clinical response to therapy. miRNAs have already been suggested by other authors to influence or predict sorafenib sensitivity (Xia et al. 2013; Zhou et al. 2011); therefore, these miRNAs should also be tested on clinical samples. Our approach is a promising one for the further identification of miRNAs that can predict patient responsiveness to therapy. To the best of our knowledge, our study is the first to evaluate the miRNA profiles of FNAB samples harvested from HCC patients. Pretreatment microRNA profiling, particularly for miR-224 expression, might serve as an ancillary tool for the better assessment of expected survival of patients under sorafenib treatment.
Footnotes
Acknowledgements
The authors would like to express their gratitude to Mrs. Csilla Horváth for her precious assistance in PCR experiments.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by grants from the
