Abstract
The thoracic diaphragm is a unique skeletal muscle composed of costal, crural, and central tendon domains. Although commonly described in medical textbooks, newer insights into the diaphragm cell composition are scarce. Here, using reporter mice, combined with gene expression analyses of whole tissues and primary cultures, we compared the diaphragm domains and their myogenic progenitors (i.e., Pax3/7 satellite cells). The outcomes of these analyses underscore the similarities between the myogenic aspects of the costal and crural domains. Expression levels of all myogenic genes examined (except Pax3) were strongly affected in mdx (dystrophin-null) mice and accompanied by an increase in fibrosis- and adiposity-related gene expression. Cell culture studies further indicated the presence of a non-myogenic Pax3-expressing population, potentially related to vascular mural cells. We additionally investigated the diaphragm vasculature. XLacZ4 and Sca1-GFP transgenes allowed a fine definition of the arterial and microvasculature network based on reporter expression in mural cells and capillary endothelium, respectively. We also provide insights into the organization of the diaphragm venous system, especially apparent in the central tendon and exhibiting arcades lined with fat-containing cells. The novel information in this “contemporary atlas” can be further explored in the context of diaphragm pathology and genetic disorders.
The thoracic diaphragm (diaphragm, as commonly referred to) is a complex skeletal muscle organ essential for supporting respiration in mammals (De Troyer and Estenne 1988; Rowley et al. 2005). The diaphragm also exerts nonrespiratory functions, including promoting venous return, lymphatic sieving, and gastroesophageal reflux restriction (De Troyer et al. 1981; Martin et al. 1983; Mittal 1993; Abu-Hijleh et al. 1995; Loukas et al. 2005). This diaphragm multitasking is supported by a unique anatomy. The diaphragm is divided into costal and crural muscle domains with an interceding central tendinous domain (Blair 1923; Anraku and Shargall 2009). The costal (ventral) domain is the larger component of the diaphragm and is arrayed radially from the central tendon to the ribs. The crural (posterior) domain tacks the diaphragm to the dorsal wall and the spinal column, and it wraps and provides apertures for the esophagus and the aorta. The central tendon, primarily a supportive connective tissue structure, provides passage of the inferior vena cava.
Whether the costal and crural domains represent two distinct muscles, possibly of different embryological origins, has been a subject of debate (Pickering and Jones 2002; Babiuk et al. 2003). While the crural and costal muscles act in synchrony throughout respiration, the activities of these two muscular regions can diverge during certain events, such as swallowing and emesis (Pickering and Jones 2002). There is also evidence that the crural muscle domain can act independently to control esophageal reflexes, but this is not a common feature for all mammals (De Troyer et al. 1981; Mittal 1993; Pickering and Jones 2002). Moreover, observed variations between these two muscle domains based on metabolic activity were not found to be necessarily consistent for all mammals, and conclusions appeared to vary depending on the specific metabolic activity tested (Powers et al. 1990; Hodge et al. 1997). The diaphragm muscle has been analyzed extensively in studies associated with the mdx (dystrophin-null) mouse, a murine model of Duchenne muscular dystrophy (Faulkner et al. 2008; Mizunoya et al. 2011). Compared to other muscles in the mdx mouse, the diaphragm shows earlier signs of the pathology with measurable muscle degeneration and fibrosis (Stedman et al. 1991; Ishizaki et al. 2008). It was reported that both costal and crural domains displayed pathological features in the mdx diaphragm, but there is only limited information specific to muscle domain comparison (Anderson et al. 1998).
The present study has been driven initially by our interest in comparing satellite cells across different muscles in the mouse. Satellite cells, myogenic stem cells situated between the myofiber basal lamina and plasma membrane, are responsible for homeostasis and repair of myofibers in adult muscles (Hawke and Garry 2001; Yablonka-Reuveni 2011). Satellite cells and their proliferating progeny commonly express the paired box transcription factor, Pax7, across all muscle groups and species examined to date (reviewed in Yablonka-Reuveni 2011). Differently, Pax3 is only expressed by a subset of satellite cells in some muscles yet prevalent in diaphragm satellite cells (Relaix et al. 2006; Day et al. 2007; Yablonka-Reuveni 2011; Boutet et al. 2012). In the present study, we were particularly interested to compare Pax3 and Pax7 expression in the two different muscle domains of the diaphragm, an aspect that has not been emphasized in the aforementioned publications.
We were additionally interested in getting insights into the diaphragm vasculature with emphasis on comparing vascular structures and density across the three diaphragm domains. The crucial roles of the diaphragm are sustained by an extensive arterial, capillary and venous network, adapted to maintain adequate blood flow and fulfill the high demand for gas exchange during the continuous diaphragmatic functioning. Nevertheless, there is only limited information about the mouse diaphragm vasculature, with detection based primarily on electron microscopy or injected ink tracing (Simionescu et al. 1978a, 1978b). Apart from human studies, morphological details available on diaphragmatic vasculature mostly rely on non-murine mammals including bat, dog, sheep, and rat (Beck and Baxter 1960; Martin et al. 1983; Comtois et al. 1987; Boczkowski et al. 1990).
Here, we used a series of skeletal muscle reporter mice to analyze the costal and crural muscle domains and distinguish the structural transition at the myotendinous junction. This microanatomy analysis was accompanied with cell culture studies to compare the domains for myogenic progenitor performance and gene expression. Diaphragm domains were also compared between wildtype and mdx mice. Furthermore, we investigated the vascular structures in the diaphragm using transgenic mice with specific reporter expression in capillary endothelium and in mural cells of blood vessels (i.e., smooth muscle cells of larger vessels and capillary pericytes; Armulik et al. 2011). Surprisingly, our comparative cell culture studies led us to identify a non-myogenic Pax3-expressing cell population that appears to share characteristics of vascular mural cells. Collectively, this study provides new insights into the mouse diaphragm domain organization, vasculature, and myogenic/non-myogenic progenitors.
Materials and Methods
Animals
Female mice (from colonies we have maintained at the University of Washington) were used throughout this study. Animal care and experimental procedures were approved by the Institutional Animal Care and Use Committee at the University of Washington. Myf5nLacZ, knockin heterozygous (Tajbakhsh et al. 1996; Beauchamp et al. 2000), and the heterozygous transgenes MLC3F-nLacZ (Kelly et al. 1995; Beauchamp et al. 2000), XLacZ4 (Tidhar et al. 2001), nestin-GFP (Mignone et al. 2004), and Sca1-GFP (Ma et al. 2002) were all in a C57BL/6 strain background and used in our previous studies (Day et al. 2007; Kirillova et al. 2007). Double reporters nestin-GFP/Myf5nLacZ and Sca1-GFP/XLacZ4 mice were generated by crossing heterozygous parental mice. α7integrinLacZ, knockin, C57BL/6 background was used as a heterozygous or homozygous (the latter null produced a stronger signal); this mouse was developed and kindly provided by Dean Burkin, University of Nevada (Flintoff-Dye et al. 2005). MyoDiCre, knockin, heterozygous, was developed and kindly provided by David Goldhamer, University of Connecticut (Kanisicak et al. 2009), and was crossed to C57BL/6 background. Pax3Cre (Pax3tm1(cre)Joe/J), knockin, heterozygous, from Jackson Laboratory (Engleka et al. 2005), was crossed to C57BL/6 background. Males MyoDiCre or Pax3Cre were each bred with the knockin reporter female Gt(ROSA) 26Sortm4(ACTB-tdTomato,-EGFP)Luo/J from Jackson Laboratory, referred in the present study as Rosa26mT/flox/mG. The Rosa26mT/flox/mG mouse harbors ubiquitous expression of the fluorescent reporter Tomato in the cytoplasmic membrane, but upon Cre-mediated recombination, the targeted cell lineage switches to express GFP (Muzumdar et al. 2007). We also generated triple heterozygous Pax3Cre × Rosa26mT/flox/mG × XLacZ4 mice. The dystrophin-null mdx4cv (C57BL/6 background) was kindly provided by Jeffery Chamberlain (University of Washington) (Chapman et al. 1989; Im et al. 1996). Control wildtype mice were C57BL/6 bred in parallel with the aforementioned collections. Mice were typically 6 months old, except for wildtype-mdx comparisons where mice were 12–14 months old.
Diaphragm Isolation
Unless otherwise noted, whole diaphragms were isolated free of intercostal muscles and were carefully cleaned of extraneous connective tissue and fat, as well as associated diaphragm tissues (i.e., aorta, esophagus, and vena cava) with great care taken to avoid damaging vasculature or other structures intrinsic to the diaphragm.
Histology and Fluorescent Imaging of Whole Diaphragm
Freshly isolated whole diaphragms were imaged immediately upon harvesting using phase/fluorescent microscopy or first fixed for 10 min in 2% paraformaldehyde solution fortified with 1% sucrose then stained with X-gal (followed by Oil-red-O in some instances) as previously detailed (Koopman et al. 2001; Day et al. 2007). A Nikon Stereoscopic Zoom Microscope (SMZ1500, with fluorescent capacity) equipped with a Digital Sight DS-Ri1 camera was used for analysis. Alternatively, higher-resolution images were taken with an inverted fluorescent microscope (Nikon Eclipse, TE2000-S). This microscope was also used for the cell culture studies detailed below.
Primary Myogenic Cultures
The cleaned diaphragm was subsectioned into costal, crural, central tendon and a narrow strip of transitional zone at the muscle-tendon borders of both the costal and crural domains. The different subsections were digested for 1 hr in 0.1% pronase and cells were released and cultured according to our routine procedure (Shefer et al. 2006; Danoviz and Yablonka-Reuveni 2012). Cells were seeded in 35-mm tissue culture plates coated with 2% gelatin, and cultures were maintained in DMEM-based medium containing 20% fetal calf serum, 10% horse serum, 1% chicken embryo extracts; this medium promotes myoblast proliferation, spontaneous differentiation, and fusion into myotubes (Shefer et al. 2006; Danoviz and Yablonka-Reuveni 2012). The seeding density for muscle domain cultures was 1–2 × 105 cells per plate, while transitional zone and central tendon cultures were initiated with all of the recovered cells, as numbers were too low for accurate cell counting.
Single Myofiber Isolation and Culture
Intact myofibers were released from whole diaphragm and extensor digitorum longus (EDL) muscle by collagenase digestion, following the same procedure as previously published for EDL myofibers (Shefer et al. 2006; Day et al. 2010). Our detailed protocol for myofiber isolation from the diaphragm, along with an update of EDL myofiber isolation, is included in a forthcoming publication (Keire et al. 2013). Muscles used for single myofiber isolation were typically from nestin-GFP transgenic mice (or double reporter nestin-GFP/Myf5nLacZ mice). As described for EDL myofibers (Day et al. 2007; Day et al. 2010), satellite cells can be detected in isolated myofibers from diaphragms of these reporter mice based on direct detection of GFP expression, coinciding with the expression of the satellite cell marker Pax7 and Myf5-driven β-gal (Supplemental Fig. S1, panels A, B). Isolated myofibers were cultured in individual wells of 24-well trays coated with Matrigel (Shefer et al. 2006; Day et al. 2010). Culture medium was the same as detailed above. In this culture system, satellite cells migrate from the myofiber and establish a myogenic culture with proliferating and differentiating progeny, similar to a primary culture. Notably, compared to myogenic cultures established on gelatin, the Matrigel substrate sustains proliferation for an extended period of time.
The use of the nestin-GFP reporter has allowed us to determine satellite cell number and, importantly, myofiber purity upon myofiber culturing. For example, capillary endothelial cells also express nestin-GFP (Day et al. 2007), and the detection of additional GFP+ cells on the myofiber surface indicates the presence of capillary fragments, which would contribute non-myogenic cells to the resulting myofiber culture. Even with the more extensive myofiber rinsing included in our updated isolation protocol (Day et al. 2010; Keire et al. 2013), residual interstitial cells can still be occasionally co-isolated with myofibers, resulting in a “mixed progeny” culture. The extent of myogenic purity of the cultures can be easily assessed by cell morphology. Morphology-based cell identity can be confirmed by X-gal staining of cultures prepared from Myf5nLacZ mice (Supplemental Fig. S1, panels C–F).
Immunofluorescence
All immunofluorescence details (including fixation of myofibers and cultured cells in 2% paraformaldehyde, primary and secondary antibodies, microscopy, and imaging) followed our routine protocols (Shefer et al. 2006; Day et al. 2010; Danoviz and Yablonka-Reuveni 2012).
RNA Isolation and Semi-quantitative RT-PCR
RNA was isolated from whole diaphragms, diaphragm subdomains, control tissues, and cell cultures following our previously published protocols (Day et al. 2007) except that samples were first extracted with Trizol / phenol-chloroform, and the resulting aqueous layer was then processed for RNA purification using Qiagen RNeasy mini columns (Qiagen, Valencia, CA). When diaphragm domains were harvested, two littermate mice were typically used in parallel; the costal and crural muscles were processed and analyzed individually per each mouse, but the central tendon was pooled from two mice due to the limited amount of tissue recovered from this domain.
Conditions for cDNA synthesis (using iScript, BioRad, Hercules, CA), PCR (using HotStarTaq DNA polymerase, Qiagen), agarose gel product separation and visualization (using SYBR Green, Life Technologies, Gaithersburg, MD) were similar to those previously published (Day et al. 2007), except that in the present study, cDNA was generated from 500 ng total RNA per reaction. The number of PCR amplification cycles was 12 for 18S and 26–35 for all other genes (specific details are provided later in the figures). PCR reactions for 18S were also performed with cDNA diluted at 1:100 to 1:1000, to permit log-linear amplification within the same cycle range as the others genes. Primer sets for 18S (Day et al. 2007), adipsin (Li et al. 2008), myogenin (Kwiatkowski et al. 2008), Pax3 (Graves and Yablonka-Reuveni 2000), and Pax7 (Day et al. 2007) resulted in products of 323, 465, 551, 497, and 466 bp, respectively. Forward/reverse primer sets (and product size) for the additional genes analyzed were as follows:
ADAM12, TAAAACGTACAGCTTAGAGCC/TCAAC GTGATTGGCGATCTC (300 bp);
Col1-α1, AGGCCACGCATGAGCCGAAG/AGCACT CGCCCTCCCGTCTT (316 bp);
α-klotho, CCCTGGCCGACCATTTCAG/GCAGAAG AGACGAGAGGTTG (417 bp);
β-klotho, GACGAGGGCTGTTTTATGTG/TCCTTCGCTCACCACACAC (467 bp);
MRF4, TCGTCGGAAAGCAGCTACCCT/CTGGGG AGTTTGCGTTCCTCT (426 bp);
Myostatin, ACGCTACCACGGAAACAATC/CTGGTC CTGGGAAGGTTACA (399 bp);
TBP, TCCGCAGTGCCCAGCATCAC/TCTGGCCTG AGGCACCCTGT (407 bp);
Tcf21, ACATTCACCCAGTCAACCTG/CGGTAAGTA CAGTTGGTCTCCT (301 bp);
TGF-β1, AACAATTCCTGGCGTTACCTTGG/GAGC AGGAAGGGTCGGTTCAT (252 bp).
Expression of 18S and TBP (TATA box binding protein) housekeeping control genes served as quality and loading controls.
Results and Discussion
Distinguishing Mouse Diaphragm Domains by Myogenic-Reporter Expression
Figure 1 depicts an abdominal view of an isolated mouse diaphragm cleaned from external connective tissue and fat and separated from associated tissues (i.e., esophagus, aorta and its surrounding fat, vena cava and its surrounding fat). The diaphragm image and accompanying schematic show the costal, crural, and central tendon domains, as well as the localization of the different diaphragmatic apertures. Note that in this and all subsequent figures, left or right location of the inferior vena cava reflects abdominal or thoracic view of the diaphragm, respectively.
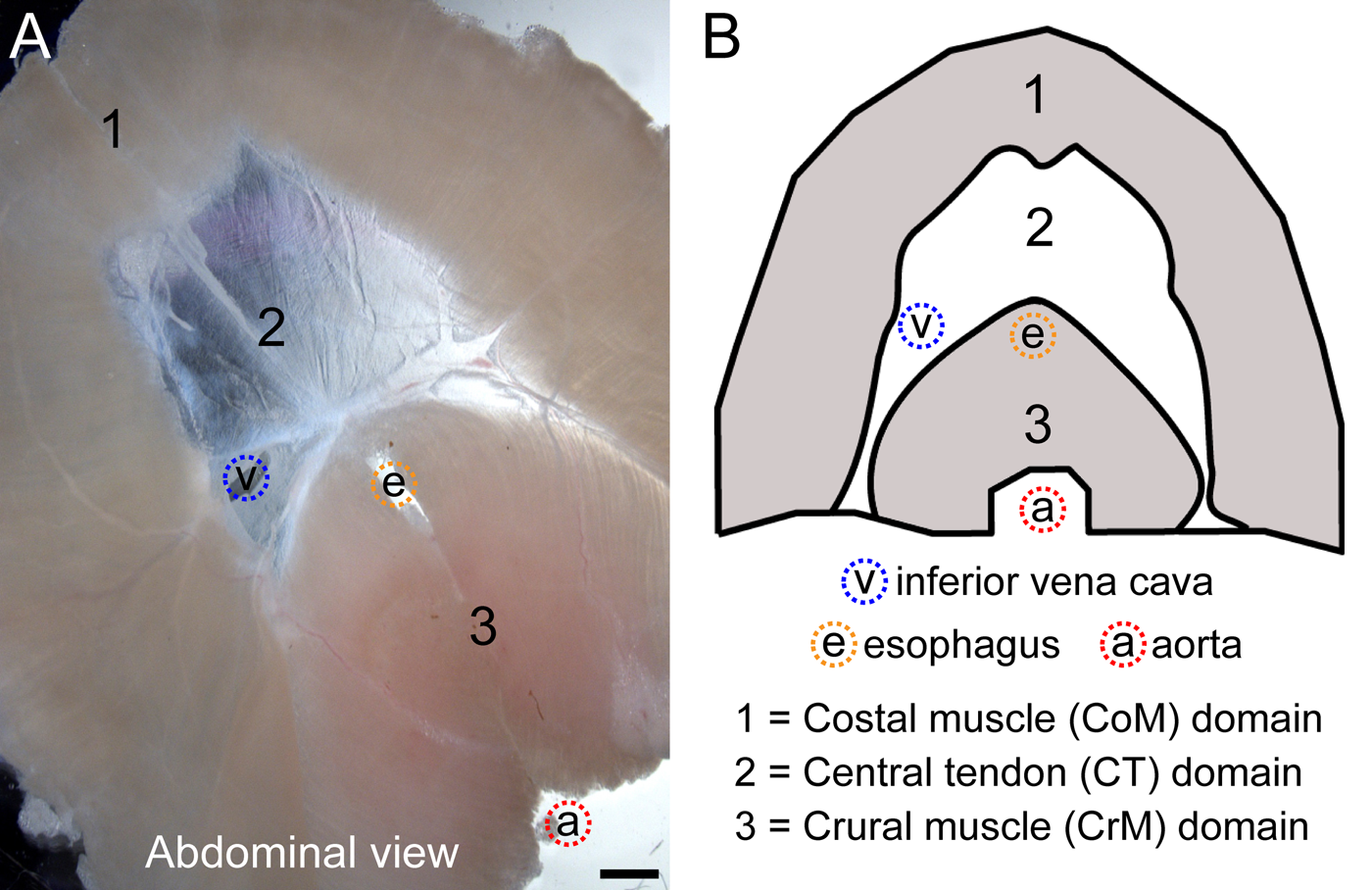
Abdominal view of isolated mouse diaphragm (A) along with a schematic view of the diaphragm muscle and tendon domains (B). Here and in all subsequent figures: (i) blue, red, and orange dotted circles indicate the apertures for the inferior vena cava (v), aorta (a), and esophagus (e), respectively; (ii) left or right location of the inferior vena cava in diaphragm images reflects abdominal or thoracic view of the diaphragm, respectively. An array of channel-like structures can be seen within the central tendon, which is part of the venous drainage system of the diaphragm and is discussed later. Scale bar, 5 mm (A).
We further analyzed whole diaphragms using a series of muscle-related reporter mice (Fig. 2). Figure 2A demonstrates a uniform MyoDiCre-driven GFP expression in both muscle domains and no expression in the central tendon. This pattern is in accordance with endogenous expression of the myogenic-specific transcription factor MyoD. However, the detection of MyoDiCre-driven reporter in adult muscles is in fact reflective of ancestral expression of MyoD in the myogenic progenitors during muscle development (Kanisicak et al. 2009). The cross between MyoDiCre and Rosa26mT/flox/mG also permits demonstration of the presence of non-myogenic interstitial cells in the muscle domains, which fluoresce red in the absence of MyoD-driven Cre expression (Fig. 2A′). This red fluorescence along with the DAPI nuclear staining additionally depicts the cellular (and non-myogenic) nature of the central tendon (Fig. 2A′′, B). Without such “counterstaining”, the central tendon, which is made of a thin layer of connective tissue, appears almost void of cells compared with the thicker muscle domains. We also verified, using freshly isolated myofibers, myofiber cultures, and standard primary cultures, that GFP expression is distinctively only within myofibers and myogenic progeny, while Tomato+ cells are always within the non-myogenic populations (data not shown).
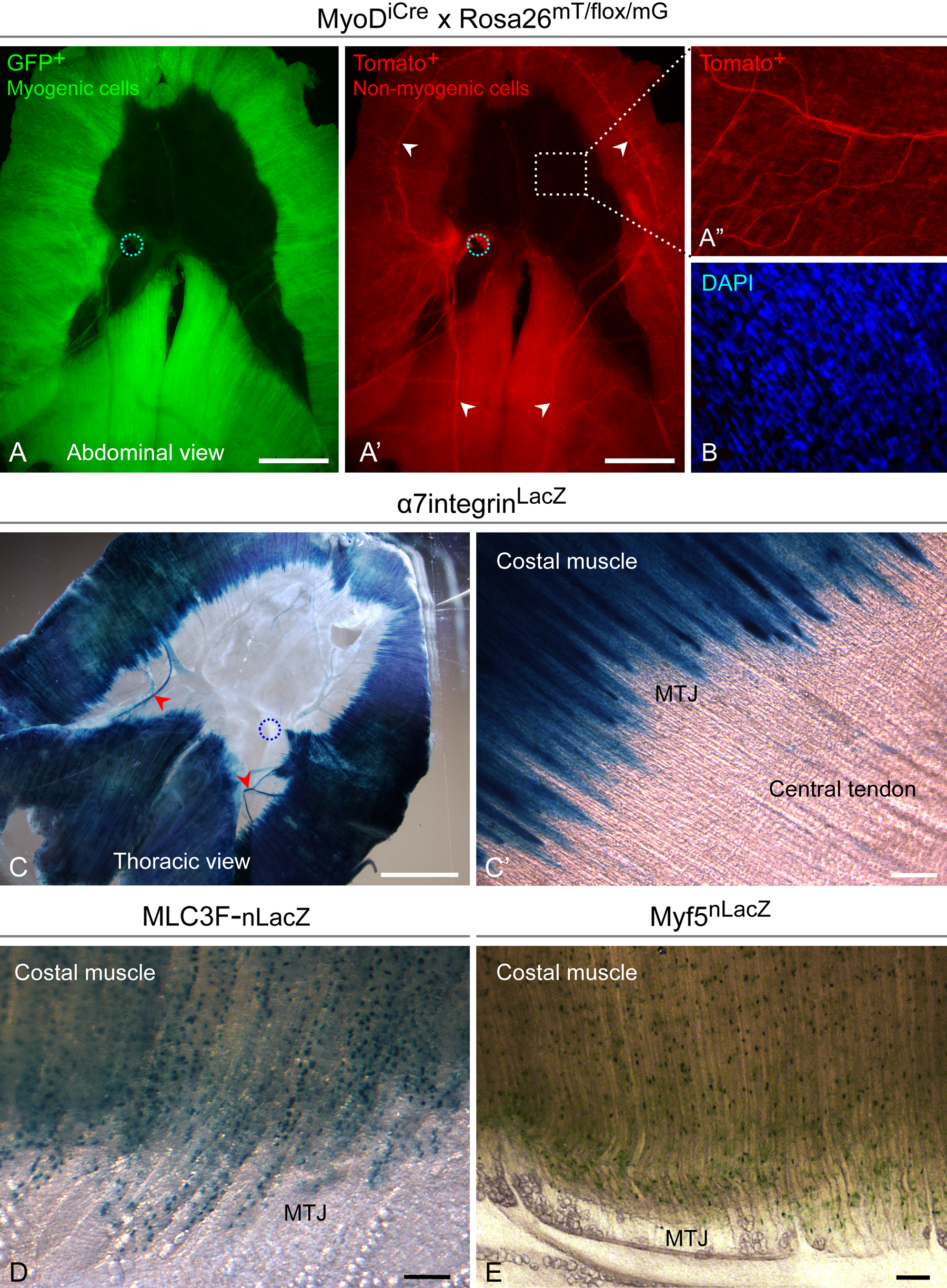
Skeletal muscle–related reporters identify features of the diaphragm domains, viewed by direct fluorescence (A–B) or following X-gal staining (C–E). (A, A′) MyoDiCre-driven GFP expression identifies myofibers (and their satellite cells), while the interstitial non-myogenic cells are distinguished based on red fluorescence. (A′) White arrowheads indicate the phrenic nerves. (A′′) Higher magnification is needed to observe the faint red fluorescence of the central tendon. (B) The cellular nature of the central tendon is confirmed by DAPI counterstaining. (C) a7integrin-driven LacZ expression identifies the myofibers (and satellite cells), as well as the phrenic arteries (red arrowheads). The venous drainage system is seen in the central tendon (discussed later). (C′) Higher-magnification view showing interdigitations of myofibers at the myotendinous junction (MTJ). (D, E) MLC3F-nLacZ and Myf5nLacZ reporters, expressed by myofiber nuclei and satellite cells, respectively. Additional details about Myf5nLacZ reporter expression in satellite cells are presented in Supplemental Fig. S1. (D, E) Round cells seen at the periphery of the central tendon are perivascular fat cells that line the venous arcade, a portion of which is shown in panel E. Scale bars, 5 mm (A, A′, C), 100 µm (C′, D, E).
Figure 2C demonstrates uniform expression of α7integrin-driven LacZ in both muscle domains. As previously published, α7integrin is expressed by myofibers and their associated satellite cells and also by vascular smooth muscle (Burkin and Kaufman 1999; Flintoff-Dye et al. 2005; Rooney et al. 2009). Indeed, herein LacZ expression is seen in the phrenic arteries as they emerge from the crural domain, crossing the central tendon and into the costal domain (Fig. 2C, red arrowheads). The strong staining of the muscle domains, however, does not permit distinction of additional vascular elements within the muscle tissue. Most notably is the clear definition of the myotendinous junction observed in diaphragms from α7integrinLacZ mice (Fig. 2C′). The individual myofibers do not all terminate at a single distinct boundary between muscle and tendon; rather, there is a transitional zone between these domains. This observed interdigitation, which is considered to provide a stronger cohesion between muscle and tendon, is in agreement with what was reported for an example in the dog, based on gross anatomy (Hwang et al. 2005).
We also looked at two commonly used myogenic reporter mice, MLC3F-nLacZ and Myf5nLacZ, to define the diaphragm muscle domains (Fig. 2D, E). In the MLC3F-nLacZ transgenic mouse (also referred to as 3F-nlacZ-E mice), regulatory elements of muscle-specific myosin light chain 3F (MLC3F) drive β-gal expression in myofiber nuclei but not in satellite cells (Kelly et al. 1995; Beauchamp et al. 2000; Day et al. 2010). In the Myf5nLacZ mouse, one of the Myf5 alleles was modified to direct LacZ expression (Tajbakhsh et al. 1996), resulting in β-gal protein in the nuclei of satellite cells (Beauchamp et al. 2000; Day et al. 2010). Figure 2 (D, E) depicts specific X-gal staining of myofiber nuclei (MLC3F-nLacZ) and satellite cells (Myf5nLacZ) in the costal domain. Similar staining patterns were observed in the crural domain (not shown). These patterns are in agreement with previous results obtained with limb muscles (Beauchamp et al. 2000; Day et al. 2010). Notably, with both reporter mice, one can notice the interdigitation between the muscle and tendon domains (Fig. 2D, E) as described above using the α7integrinLacZ mouse. Also observed at the muscle-tendon transitional zone are channels of round cells. As discussed later, these are fat-containing cells based on Oil-red-O staining.
Collectively, the series of studies summarized in Fig. 2 demonstrate common reporter expressions in the two muscle domains and a clear demarcation of the transitional zone where the myofibers and tendon connective tissue meet.
Detection of the Arterial and Capillary Networks Within the Diaphragm
To analyze the diaphragm vasculature, we used the XLacZ4 transgenic mouse in which there is a specific β-gal reporter expression in the nuclei of vascular smooth muscle (Klinghoffer et al. 2001; Tidhar et al. 2001; Kirillova et al. 2007; Armulik et al. 2011). Classically, the vascular contractile cells (also known as “mural cells”) are subdivided into smooth muscle cells and pericytes (Diaz-Flores et al. 2009; Armulik et al. 2011). Ultrastructural and immunohistological studies have elucidated the gradual transition from the smooth muscle phenotype in terminal arterioles and venules to the pericyte phenotype associated with capillaries and postcapillary venules (Holley and Fahim 1983; Joyce et al. 1985; Diaz-Flores et al. 1991). With few exceptions, most studies utilizing the XLacZ4 mouse have used this transgene to trace retinal microvasculature. Thus, herein, we first established that the XLacZ4 reporter mouse is useful for monitoring mural cells in skeletal muscles (Supplemental Fig. S2, panel A). We then used the reporter to delineate the diaphragm vasculature as detailed below.
Defining the arterial supply
Two whole diaphragms from XLacZ4 mice stained for β-gal are shown in Fig. 3A, B. The arterial network clearly stands out in these images due to the higher density of smooth muscle (X-gal+) cells in arteries compared with veins. This has allowed us to identify the three main sources of arterial supply (phrenic, internal thoracic, and intercostal arteries) and their arborization. In the first diaphragm preparation (Fig. 3A), the aorta with its associated adventitia and the surrounding rib cage with intercostal muscles have not been removed. This permitted detection of the phrenic arteries emerging from the aorta, as well as the intercostal arteries entering the diaphragm. However, details of the costal and crural domains are better represented in the second preparation (Fig. 3B), where the diaphragm has been further dissected. Our main observations are recapitulated in a schematic view of the diaphragm arterial network shown in Fig. 3C. The phrenic arteries, emerging directly from the aorta, run anterior and lateral through the crural muscle toward the costal muscle while giving off branches that irrigate the crural muscle (Fig. 3A). Subsequently, each of these phrenic arteries divides into medial and lateral branches (Fig. 3A, red asterisk). The lateral branches enter the costal muscle, while the medial branches run anteriorly along the margin of the central tendon. Under close examination, an anastomosis between the two medial branches of the phrenic arteries and the respective internal thoracic arteries can be seen (shown schematically as two red dotted lines in Fig. 3C). These internal thoracic arteries represent a second source of arterial supply arising from the anterior part of the costal muscle (Fig. 3A–C, light orange arrowheads). A third arterial supply originates from the intercostal arteries arising from the lateral part of the costal muscle (Fig. 3A–C, orange double arrowheads). Overall, this heavily branched and dense arterial system of the diaphragm muscle is distinctive from any of the limb muscles we have examined. Additionally, as seen in Fig. 3A (zoom-in view), there is only limited arterial supply to the diaphragm tendon.
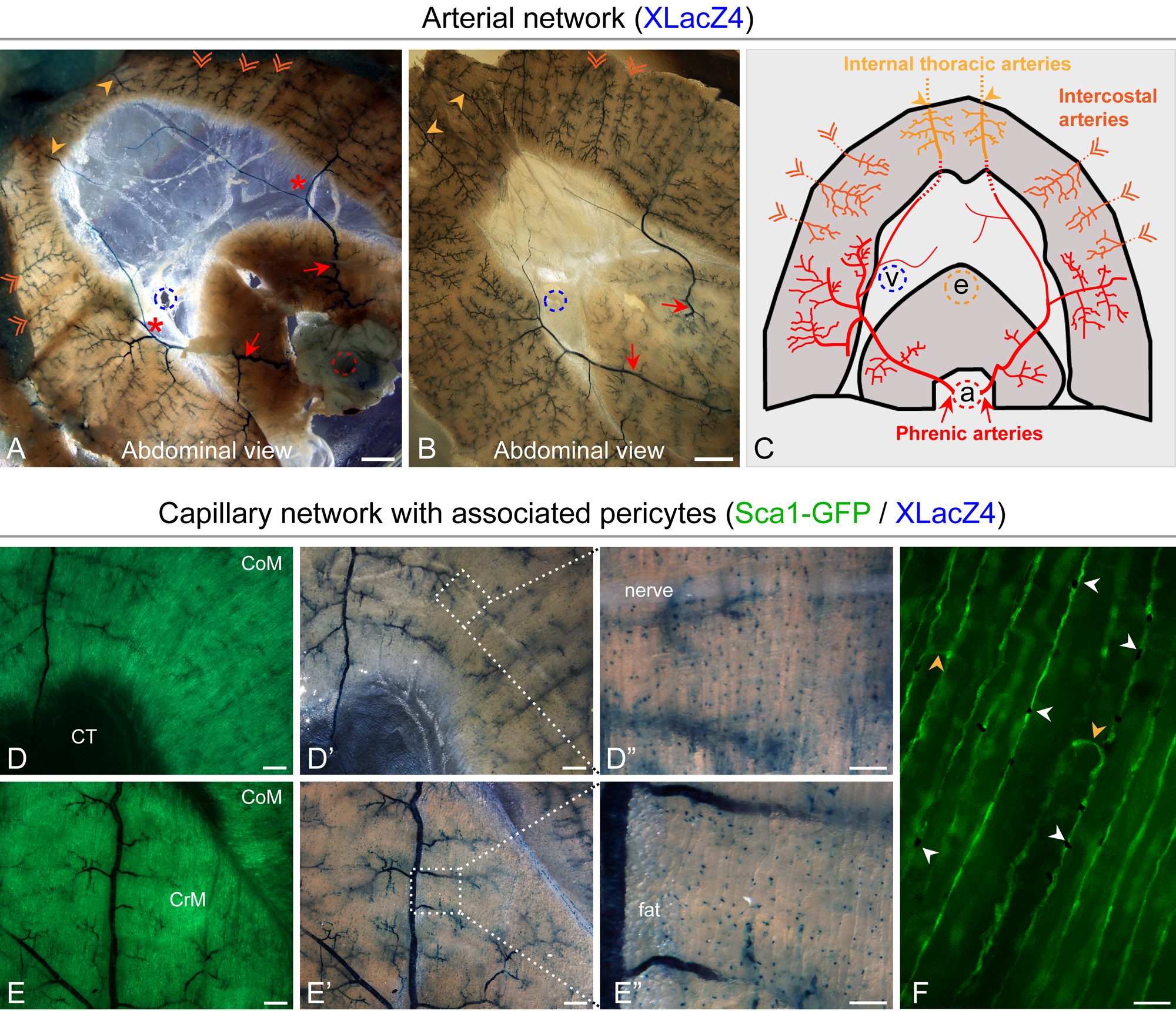
Identification of the major arterial networks and their schematic representation (A–C) as well as the description of the capillary system with associated pericytes (D–F) in diaphragms from XLacZ4 or XLacZ4/Sca1-GFP reporter mice. The XLacZ4 transgene expresses β-gal reporter in nuclei of mural cells (vascular smooth muscle cells and pericytes) and the Sca1-GFP transgene identifies the capillary endothelium. All images were obtained following X-gal staining. (D, D′ and E, E′) Paired fluorescent and phase images of costal and crural muscle domains, depicting the dense capillary network (GFP+, panels D, E) and arterial trees (X-gal+, D, D′ and E, E′), as well as the presence of numerous solitary XLacZ4+ pericytes (X-gal+, panels D′, E′) that are more obvious in the higher-magnification views (D′′, E′′). (F) A higher-magnification fluorescent image showing capillaries (GFP+) with associated solitary pericytes (black dots) in costal muscle. White arrowheads point to pericytes in close proximity with capillary endothelium, while orange arrowheads identify anastomoses between capillaries running parallel to the myofibers. Notably, perivascular fat can be seen associated with the venous arcade in panel D′ and with the artery in panel E′′ (further discussed in Fig. 4). Scale bars, 1 mm (A, B, D, D′, E, E′), 100 µm (D′′, E′′), 50 µm (F).
The aforementioned three main arterial networks have been described in general anatomy books only in terms of external blood supply to the diaphragm without description of the arterial arborization within the diaphragm. Furthermore, the few published studies on this subject have contributed only partial details about the vasculature within the diaphragm. The most complete study has been carried out on the dog diaphragm using acrylic color injection techniques (Comtois et al. 1987). An earlier study also defined some of the arterial supply components in rat and bat diaphragms (Beck and Baxter 1960). Here, using the XLacZ4 transgene, we were able to provide what is, to the best of our knowledge, the most comprehensive view available to date of the arterial network organization within the mouse diaphragm.
Defining the capillary network
Along with the XLacZ4 reporter for mural cell detection, we used the Sca1-GFP transgenic reporter for direct tracing of the capillary endothelium, following our initial findings that this transgene is expressed strongly in skeletal muscle and retina endothelium (Day et al. 2007; Kirillova et al. 2007; Xia et al. 2010). Using the double transgenic Sca1-GFP/XLacZ4 mouse, we concluded that both the crural and the costal muscle domains harbor a rich capillary network running along the myofibers (Fig. 3D, E), and numerous XLacZ4+ solitary cells (Fig. 3D′–E′′). These solitary cells were confirmed as pericytes by their proximity to the capillary endothelium (Fig. 3F, white arrowheads). The image in Fig. 3F also depicts some transverse anastomoses (orange arrowheads) that interconnect parallel capillaries. Notably, the high density of the solitary pericytes along the capillary endothelium in diaphragm musculature is in stark contrast with the low pericyte density in the tibialis anterior muscle (Supplemental Fig. S2, panel A) and other limb muscles we have examined (gastrocnemius, EDL). Additional examples of diaphragm muscle depicting XLacZ4 expression in mural cells through the gradual transition from arteries to capillaries are shown in Supplemental Fig. S2, panel B. The higher-magnification images of costal and crural muscle domains (Fig. 3D′′, E′′) further outline the numerous pericytes situated along the myofibers.
The appearance of the XLacZ4 pericytes in these images resembles that of the satellite cells in diaphragm muscle from Myf5nLacZ mice (Fig. 2E). However, while pericytes and satellite cells can be found in very close proximity to each other, they occupy distinctly different locations. Pericytes and their associated endothelium are surrounded by a common basal lamina and reside in the connective tissue between individual myofibers, while satellite cells are located underneath the myofiber basal lamina (Yablonka-Reuveni 2011). We have never detected XLacZ4 pericytes in the satellite cell niche when analyzing adult mice. We also verified in ex-vivo studies (freshly isolated myofibers, myofiber and primary cultures, and clones) that resident satellite cells and their progeny do not express the XLacZ4 transgene at any time (data not shown).
Collectively, the present analysis with Sca1-GFP/XLacZ4 reporters provides a more contemporary means (GFP fluorescence) to detect the dense capillary bed in the diaphragm, in accordance with previously published findings for the mouse diaphragm (Simionescu et al. 1978b). Furthermore, to the best of our knowledge, the present study represents the first detailed analysis at the light microscopy level of diaphragm pericyte location and density.
Defining the Venous Network Within the Diaphragm
When the diaphragm was isolated from the mouse, prevalent vascular sinuses were observed in the central tendon (Figs. 1A, 2C, 2E, and 3A, zoom-in view). These were still filled with blood when freshly isolated unfixed diaphragms were examined (Fig. 4A). We have concluded that this vascular system in fact corresponds to the main route of venous drainage formed by the left and right phrenic veins and joining the inferior vena cava (Beck and Baxter 1960; Simionescu et al. 1978a, 1978b). Notably, the presence of a prevalent venous system in the central tendon was mentioned in the early study of the primate diaphragm (Blair 1923), but no images were provided for this aspect of the report. Additionally, two other accessory routes of venous drainage were reported within the lateral half of the costal muscle domain, positioned anatomically parallel to the arterial distribution and converging into the internal thoracic and intercostal veins (Martin et al. 1983; Boczkowski et al. 1990). Given the limited descriptions that we have been able to gather across the few studies mentioned above, we have concluded that no comprehensive reports exist about the diaphragm venous drainage, whether in human anatomy textbooks or in studies done with experimental animals, including mice. Here, we provide a more detailed summary of the main drainage route formed by the phrenic veins running within the central tendon, as well as additional information about intramuscular venous organization.
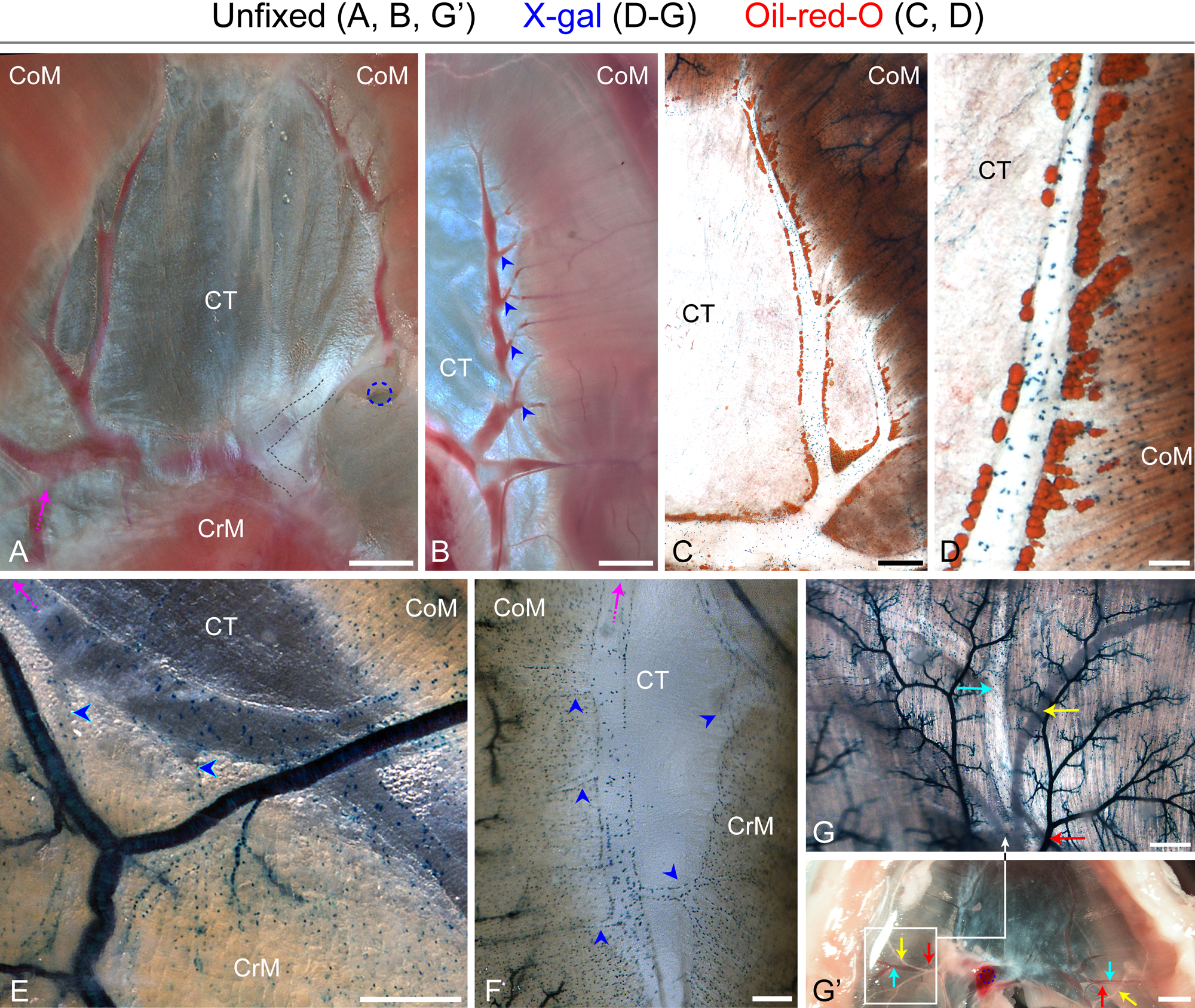
Identification of the major venous drainage system in diaphragms from XLacZ4 mice as viewed directly after isolation (A, B, G′) or following X-gal staining combined with Oil-red-O staining (C, D) or X-gal staining by itself (E–G). (A) In this thoracic view, the left and right phrenic veins form a clearly distinguishable arcade on each side of the central tendon; these arcades join (black dotted lines) and drain into the inferior vena cava. (B–D) Details of the venous arcade (abdominal views). (B) Blue arrowheads point to multiple tributaries coming from the costal muscle domain and joining the main phrenic vein. (C, D) Lower- and higher-magnification images of Oil-red-O staining, which reveal the presence of perivascular fat cells along the venous arcade. (E, F) Abdominal and thoracic views, respectively, of the left dorsal region of the diaphragm depicting the crural and costal muscle venous drainage; blue arrowheads point to tributaries coming from the muscle domains and draining into the venous arcade. In panels A, E, and F, the magenta arrows point to the same vein. (G) Details of the venous network inside the costal muscle (abdominal view); the venous system displays a distinctive pattern of scattered XLacZ4+ cells, while the arterial supply is detected in this image based on strong X-gal staining. Arrows: cyan arrow indicates a large tributary of the right phrenic vein; red arrow indicates a ramification of the left phrenic artery; and the yellow arrow indicates the phrenic nerve. (G′) A lower-magnification image indicating the localization of the field shown in panel G. Arrows inside the framed field and in the contralateral side in panel G′ point to vascular and nerve structures according to the color scheme in panel G. Scale bars, 2 mm (G′), 1 mm (A, B), 500 µm (C, E, F, G), 100 µm (D).
As illustrated in Fig. 4A, the left and right phrenic veins form a well-delineated venous arcade on each side of the diaphragm along the myotendinous junction within the central tendon. These two arcades are interconnected through a large anastomosis that drains into the inferior vena cava. This transverse vein is partly covered by a thick supporting aponeurosis reducing the clarity of this connecting channel in the image (Fig. 4A black dotted lines; see also Fig. 3A, zoom-in view, for a clearer image when tissue is fixed). Figure 4B depicts clearly the many tributaries (still filled with blood) arising from the costal muscle domain, crossing through the myotendinous junction, and joining the main phrenic vein in the central tendon domain (blue arrowheads). Staining of the whole diaphragm with Oil-red-O reveals clearly the presence of fat-containing cells along this venous arcade as illustrated in the left diaphragm leaflet view (Fig. 4C). Figure 4D shows a higher magnification of this perivascular fat and also depicts scattered XLacZ4+ cells throughout the vein wall. These XLacZ4+ cells appear in low density, typical of the venous wall and in stark contrast to the denser smooth muscle layer in the arterial wall discussed earlier (Fig. 3 and Supplemental Fig. S2, panel A).
The typical pattern of scattered XLacZ4+ cells in veins facilitated the detection of venules coming from the inside of the crural muscle (Fig. 4E, F). Hence, Fig. 4E shows a main tributary vein (magenta arrow), part of the venous arcade, running along the crural muscle and collecting blood from this domain as indicated by the presence of collecting venules (pointed out by blue arrowheads). Moreover, Fig. 4F depicts collecting venules (blue arrowheads) coming from the crural and the costal muscle domains, converging into the same larger vein (magenta arrow).
The ability to clearly detect the venous system with the aid of the XLacZ4 reporter has also permitted us to gain further insights into finer structural details. In particular, we conclude that the collecting venules, which arise from the muscle domains and drain into the venous arcade, can be divided into two main groups. The first category consists of venules that arise from the portion of muscle located just near the myotendinous junction (Fig. 4E, F), the shorter ones potentially representing postcapillary venules (Fig. 4F). A second category is composed of collecting venules that appear to originate from the peripheral regions of the muscle domains (Fig. 4B, blue arrowheads). The most obvious example is shown in Fig. 4G′ (diaphragm abdominal view), where each main branch of the left and right phrenic veins (cyan arrows) courses from the costal muscle domain near its peripheral margin, following the respective phrenic artery branch (red arrows). A detailed higher-magnification view of the right branch is provided in Fig. 4G, where this branch and its tributaries spread within the lateral half of the costal muscle domain, indicating that the venous arcade also participates in the drainage of the lateral half of the diaphragm. Additionally, we noticed that some of these venules anastomose with the venous networks, draining to the intercostal and the internal thoracic veins, as previously mentioned for the rat and dog diaphragm (Comtois et al. 1987; Boczkowski et al. 1990).
Collectively, we have provided novel insights about the venous arcade, drawing a clear picture of the organization of this prominent venous system formed by the phrenic veins located within the central tendon, the manner by which it receives numerous affluents arising from the costal and crural muscle domains and its interconnection with other venous networks.
Gene Expression Profiles in the Different Diaphragm Domains
Gene expression profiles of the different muscle and tendon domains were compared by semi-quantitative RT-PCR analysis (Fig. 5A). At first, we aimed to determine if the satellite cell genes (Pax3 and Pax7) and the muscle differentiation genes (MRF4, myogenin, myostatin) were expressed at a common level in the two muscle domains, indicative of a common myogenic signature (as observed with muscle reporters mice, Fig. 2). Indeed, the results shown in Fig. 5A (wildtype) depict a common expression signature of the myogenic-associated genes in the costal and crural muscles.
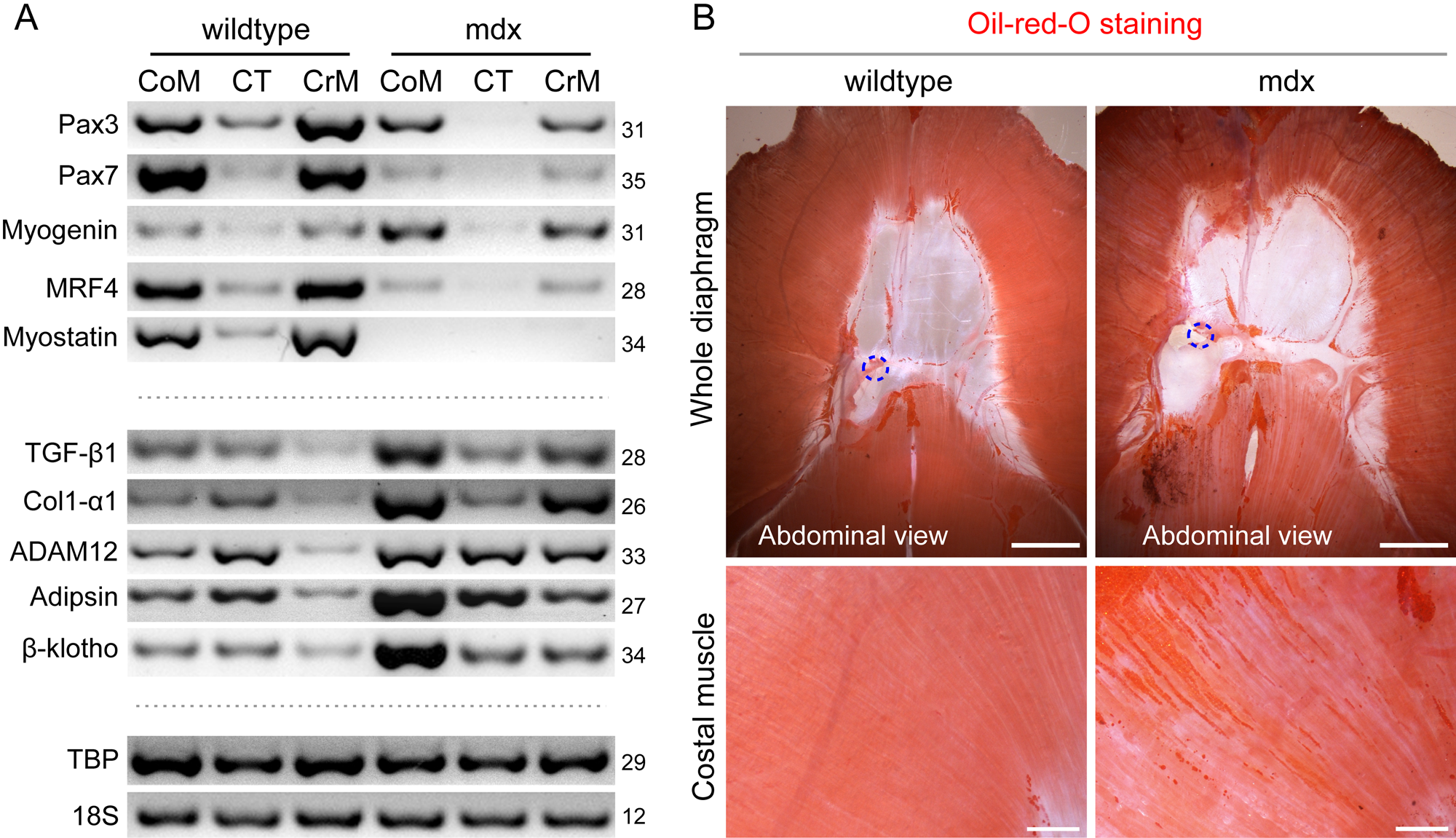
Comparison of gene expression signatures across the muscle and tendon domains of diaphragms from wildtype versus mdx mice. To ensure that the mdx diaphragm exhibited signs of advanced dystrophy, all mice used in this study were 12–14 months old. (A) A representative semi-quantitative RT-PCR analysis of the costal muscle (CoM), crural muscle (CrM), and central tendon (CT) domains for expression levels of muscle-related genes (Pax3, Pax7, myogenin, MRF4, myostatin) and additional genes relevant to fibrosis and adiposity (TGF-β1, Col1-α1, ADAM12, adipsin, β-klotho). Expression of TBP and 18S served as quality and loading controls. The number of PCR cycles is indicated per each gene at the right side. For each experiment (n=4), two littermate mice were used in parallel for each preparation the costal and crural muscles were processed and analyzed individually per each mouse, but the central tendon was pooled from two mice due to the reduced amount of tissue recovered from this domain. (B) Oil-red-O-stained diaphragms from wildtype and mdx mice are shown with detected fat deposits, in parallel with the gene expression analysis. The higher-magnification images of costal muscle depict channels of intramuscular fat that are prevalent in the mdx muscle. Scale bars, 5 mm (B, whole diaphragm panels), 1 mm (B, costal muscle panels).
MRF4 gene expression was examined in the present study as a measure of overall myogenicity in both whole tissues (Fig. 5A) and cell culture (Fig. 6). We specifically selected MRF4 as a common myogenic regulatory differentiation marker because it is the only myogenic regulatory factor that is expressed at a high level in adult myofibers, while MyoD, Myf5, and myogenin expression levels are relatively lower (Hinterberger et al. 1991; Garry et al. 2000; see also Supplemental Fig. S3, panel A). In accordance with these published studies, Fig. 5A depicts a relative reduced expression of myogenin compared with MRF4 in the adult diaphragm.
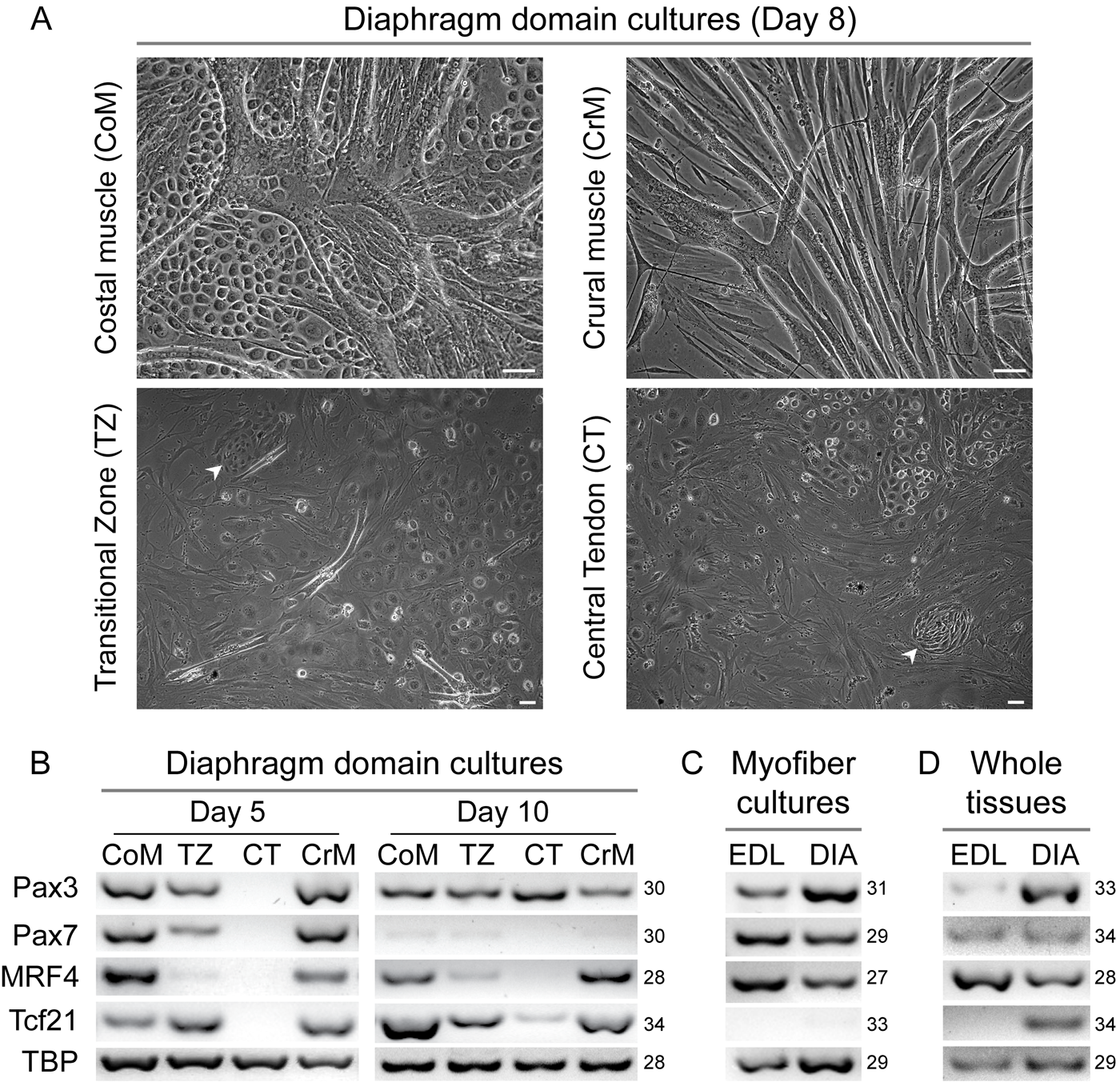
Morphology and gene expression signatures of cultured cells isolated from diaphragm domains. In this analysis, in addition to the three domains detailed in Fig. 5, we analyzed the transitional zone (TZ), a narrow strip carefully isolated at the muscle–tendon borders of both muscle domains. (A) Representative images of day 8 cultures from CoM, CrM, TZ, and CT domains. In addition to myogenic cells that fuse into myotubes, the domain cultures exhibited a range of other cell types, as discussed in the Results section. Scale bars, 50 µm. (B–D) Semi-quantitative RT-PCR analyses of: (B) day 5 and 10 cultures from the diaphragm domains; (C) control day 7 cultures of pure myogenic cells obtained from isolated extensor digitorum longus (EDL) and diaphragm (DIA) myofibers (each RNA preparation was established from 13 myofiber cultures harvested together); (D) whole EDL and diaphragm (DIA) tissues. TBP expression served as a quality and loading control. Numbers on the right side of panels B–D indicate the number of PCR cycles.
The comparative analysis of myostatin, a muscle-specific growth regulatory member of the TGFβ super family, was of particular interest, as it was not necessarily expressed at a common level in all muscles; for example, soleus muscle was shown to display very little expression (Supplemental Fig. S3, panel A), in agreement with an earlier comparison between quadriceps, tibialis anterior, EDL, and soleus (Carlson et al. 1999). A trace expression of the myogenic genes detected in the tendon domain (Fig. 5A) was likely contributed by the muscular tissue at the myotendinous junction; but, notably, small atypical muscle patches sometimes detected within the tendon domain could also potentially contribute to such a low-level myogenic gene expression (Blair 1923; our observation with mouse diaphragm).
Here, we also evaluated myogenic gene expression in diaphragm domains isolated from mdx mice in parallel to the wildtype analysis to determine if myogenicity was equally affected in both muscle domains of the mdx diaphragm (Fig. 5A). In the current study, to ensure that the mdx diaphragm exhibited signs of advanced dystrophy, mice used were 12-14 months old (mdx and control wildtype). Obvious signs of the pathology were noted upon diaphragm isolation, such as muscle thinning and calcification foci.
As shown in Fig. 5A, the expression level of the satellite cell gene Pax7 was severely reduced in the mdx muscle domains. This decline is likely due to depletion of myogenic progenitors (and not merely transcript dilution of mRNA from inflammatory and fibrotic cells). The poor myogenicity of cultures derived from diaphragms of mdx mice was also indicative of a decline in myogenic progenitors (data not shown). Surprisingly, Pax3 expression did not decline as drastically as Pax7 expression when the muscle domains between mdx and wildtype diaphragm were compared (Fig. 5A). This observation suggests that apart from satellite cells, there is a second Pax3-expressing cell population that is not depleted in muscle domains of mdx diaphragms.
Additional evidence that muscle wasting is occurring in both costal and crural domains of mdx mice was illustrated by the drastic decline in MRF4 and myostatin expression levels (Fig. 5A), which are expressed in healthy adult muscles at relatively high levels (Supplemental Fig. S3, panel A). In contrast and unexpectedly, myogenin expression was elevated in the mdx muscle domains (Fig. 5A). Typically, in a healthy state, myogenin expression is upregulated upon differentiation of satellite cell progeny, coinciding with Pax7 and Myf5 down regulation and cell cycle exit (Shefer et al. 2006; Day et al. 2009; Yablonka-Reuveni and Day 2011) but is detected only at relatively low levels in adult muscle (Fig. 5A) (Hinterberger et al. 1991; Garry et al. 2000). Nevertheless, this increased myogenin expression in mdx diaphragm while MRF4 expression declines may reflect persistence of myogenin-expressing activated satellite cells during the ongoing degeneration/regeneration rounds occurring in the mdx environment (Garrett and Anderson 1995; Yablonka-Reuveni and Anderson 2006). While formation of myogenin-expressing nascent myotubes might still occur (Garrett and Anderson 1995), myofiber maturation (associated with MRF4 expression) and satellite cell renewal (associated with Pax7 expression) appear to be severely affected by the diseased muscle environment. Furthermore, the increase in myogenin expression along with the decline in myostatin observed in mdx muscle domains in the present study is in agreement with the reported mdx gastrocnemius microarray study (Tseng et al. 2002).
Differently from myostatin, the expression level of its related family member, TGF-β1, is upregulated in the muscle domains from mdx mice (Fig. 5A), in agreement with TGF-β1 involvement in the muscle fibrosis associated with the mdx condition (Turgeman et al. 2008). A parallel increase was also observed in the expression level of collagen type I α1 chain (Col1-α1), also known to be involved in mdx- associated muscle fibrosis (Turgeman et al. 2008). Here, we also demonstrate an increase in ADAM12 expression level in the muscle domains of mdx diaphragm (Fig. 5A). ADAM12, a member of the ADAM (a disintegrin and metalloprotease) family, has been implicated in both skeletal muscle myogenesis and adipogenesis (Kawaguchi et al. 2002; Kurisaki et al. 2003). Initially, overexpression of ADAM12 was proposed to alleviate the mdx pathology (Kronqvist et al. 2002). However, in later studies, it was shown that, in fact, the regeneration process is significantly impaired in ADAM12 overexpressing transgenic mice (Jorgensen et al. 2007).
A comparison of whole diaphragms from wildtype and mdx mice stained for Oil-red-O (Fig. 5B) has further drawn our attention to diaphragm adiposity. We observed an increase in fat deposits within the muscle domains associated with the mdx phenotype (Fig. 5B). This intramuscular fat is found lined in parallel with myofibers as well as appearing in larger patches of fat deposits. Also noticeable in both wildtype and mdx diaphragms is the perivascular fat lining the venous system in the central tendon (as previously discussed, Fig. 4C, D). In parallel to this morphological analysis, we noted an increase in the expression level of adipsin, an adipocytokine and a well-established fat marker that shows increased levels in liver fibrosis (Tsukamoto 2005; Li et al. 2008; Matsubara et al. 2009).
We also investigated, in this mdx versus wildtype comparison, the status of α- and β-klotho gene expressions. The klotho genes code for transmembrane proteins that have been implicated in a range of metabolic functions and signaling pathways (Kuro-o et al. 1997; Fon Tacer et al. 2011). When undertaking the current klotho expression analysis, we were influenced by the reports that α-klotho (also named “klotho”) suppresses TGF-β1 signaling and reduces renal fibrosis (Doi et al. 2011; Sun et al. 2011). Additionally, both α- and β-klotho have been implicated in the regulation of adiposity based on their expression pattern and elucidated molecular functions (Ito et al. 2000; Mizuno et al. 2001; Chihara et al. 2006). We have also noted in our studies that β-klotho (but not α-klotho) was highly expressed in white and brown fat tissues (Supplemental Fig. S3, panel A). Published information on klotho gene expression in skeletal muscle is, however, scarce. A multi-tissue quantitative PCR survey reported low expression levels of α- and β-klotho genes in skeletal muscle (quadriceps and tongue) (Fon Tacer et al. 2011). In agreement with the latter study, in our survey that included control tissues with high α- and β-klotho expressions (Kuro-o et al. 1997; Ito et al. 2000), we detected only low-level expression of the klotho genes in some muscles, with only β-klotho being expressed in the diaphragm (Supplemental Fig. S3). Similar to our finding with the whole diaphragm, only β-klotho was detected in the mouse diaphragm domains (Fig. 5A). Albeit detected at low levels, persistent results were obtained from all mice tested, ranging from young to old age. In the context of mdx mice, we observed a global increase in the expression levels of β-klotho in diaphragm muscle domains (Fig. 5A).
In all, the status of klotho gene expression in mdx pathology is distinct from that of α-klotho expression in renal fibrosis. Our results show that α-klotho is not detected in diaphragm muscle (whether in wildtype or mdx mice), while β-klotho, detected at relatively low levels in wildtype diaphragm, shows an increase in mdx diaphragm, concomitant with the muscle pathology of enhanced fibrosis and adipogenicity. We have initiated cell culture studies aiming to identify the cell type(s) contributing to klotho gene expression in mouse muscles (unpublished). Expression of the two klotho genes was found to be extremely low in primary myogenic cultures and without any specific pattern that could link the klothos to specific phases of myogenesis. Our pilot studies of FACS sorted myogenic and non-myogenic cell populations have indicated that the expressed klotho genes are contributed by the non-myogenic cells; α-klotho expression remains low regardless of time in culture while, interestingly, β-klotho expression was upregulated in correlation with the emergence of adipogenic cells (data not shown).
Myogenic Progeny from Costal and Crural Domains Exhibit a Common Gene Expression Signature in Cell Culture
Our initial goal was to further compare the different diaphragm domains by establishing primary cultures and analyzing the progression toward myogenic differentiation by morphology (e.g., myotube formation) and gene expression. Cells were dissociated from each of the diaphragm muscle and tendon domains and cultured using our standard conditions for primary myogenic cultures. Here, we also subsectioned and investigated in culture the diaphragm “transitional zone,” a narrow strip at the border of the muscle domains and the central tendon. Figure 6A demonstrates the cultures of the four subsections: costal muscle (CoM), crural muscle (CrM), transitional zone (TZ), and central tendon (CT). By day 8, many of the myogenic progeny had already fused into myotube networks that are prevalent in the costal and crural domain cultures (Fig. 6A). The TZ culture contained only residual myotubes with the dominant cells displaying a non-myogenic phenotype, while the CT culture comprised essentially non-myogenic cells. The nodule-like cell foci seen in the TZ and CT cultures (Fig. 6A, white arrowheads) are discussed in greater detail later.
Cultures from the muscle domains also contained non-myogenic cells defined by being negative for the myogenic markers Pax7, MyoD, myogenin, and sarcomeric myosin when analyzed by immunostaining (Shefer et al. 2006; Danoviz and Yablonka-Reuveni 2012). Such cells are typically co-isolated with satellite cells when primary myogenic cultures are prepared from any muscle group. However, we observed in the costal muscle domain cultures numerous patches of unique non-myogenic cells (Fig. 6A) exhibiting a phenotype that we have never observed in myogenic cultures from limb or head muscles. These unique cells were also noticed in cultures from the other domains but were especially prominent in the costal culture, with large islands of tightly packed cells surrounded with well-developed myotubes (Fig. 6A). The cells first appeared round and refractive and quickly developed large patches with a cobblestone pattern. In contrast with fibroblastic-like cells commonly encountered in primary cultures, these patches seemed to have an epithelial-like appearance. We propose that these diaphragm-specific cells may represent mesothelial cells, derived from the pleural and peritoneal mesothelium, a monolayer of epithelium that covers the abdominal and thoracic surfaces of the diaphragm (Mutsaers 2004; Wang et al. 2010). Indeed, morphologically, these cells observed here in the domain cultures resemble what has been previously published for mesothelial cells isolated from the parietal pleura (Jaurand et al. 1981).
We further investigated diaphragm domain cultures by semi-quantitative RT-PCR with the main goal of comparing the molecular signature of the myogenic progeny in the different domains (Fig. 6B). Similar to the whole tissue study (Fig. 5A), attention was given especially to the Pax3/7 genes, while MRF4 gene expression was used as a marker for myogenic differentiation. While both Pax3 and Pax7 genes are known to be expressed in diaphragm satellite cells (Day et al. 2007), we have focused here specifically on comparing the myogenic progeny from the costal versus crural muscle domains.
As summarized in Fig. 6B, Pax3 and Pax7 were highly expressed in both muscle domain cultures by day 5 (Fig. 6B). At that time, Pax3 and Pax7 were also detected at relatively lower expression levels in the TZ cultures, contributed by residual myogenic cells. Pax7 expression declined considerably by day 10, likely due to a depletion of progenitors upon differentiation and fusion into myotubes, while the non-myogenic cell population continued to expand. Surprisingly, Pax3 expression did not exhibit such a strong decline in the muscle and TZ domain cultures. However pure myogenic cultures derived from myofibers (Supplemental Fig. S1, panels C, E) demonstrated that both Pax3 and Pax7 were expressed at relatively high levels after 7 days in culture (Fig. 6C).
Based on these myofiber culture results, the detection of Pax3 expression without a similar Pax7 expression in the costal and crural muscle domain cultures indicated the existence of a second Pax3-expressing population. Furthermore, Pax3 expression was even upregulated by day 10 in the central tendon domain culture, which harbors neither myogenic traits (Fig. 6A) nor MRF4 expression (Fig. 6B). This late-onset Pax3 expression in the central tendon effectively demonstrates the existence of a non-myogenic Pax3-expressing population. Such a second population was already hypothesized earlier in this study based on detecting Pax3 signals in mdx diaphragm domains when the Pax7 signal diminished (Fig. 5A). Ideally, one would wish to further explore the existence of myogenic and non-myogenic Pax3-expressing cells by immunostaining with antibodies to Pax3 and Pax7. However, despite concentrated attempts by ourselves and others (reviewed in Yablonka-Reuveni 2011), immunodetection of Pax3 in myogenic culture from adult muscle still remains a challenge at the present time; differently, Pax7 immunostaining has been a successful endeavor.
We also examined Tcf21 gene expression in diaphragm domain cultures (Fig. 6B). Like the myogenic regulatory factors (MyoD, Myf5, MRF4, myogenin), Tcf21 is a class II bHLH transcription factor, but its expression is not muscle-restricted; rather, it is distributed in several cell lineages (Acharya et al. 2011). Our interest in Tcf21 was prompted by earlier studies showing its expression by myogenic cells from jaw muscle but not in the limb (EDL) muscle (Ono et al. 2010). Hence, we wished to explore the status of Tcf21 expression in the diaphragm domains. By day 5, we detected Tcf21 expression in all cultures, except for that of the central tendon. Subsequently, by day 10, the CT culture also exhibited Tcf21 expression, albeit at a lower level compared with the other domain cultures (Fig. 6B). However, Tcf21 was not detected in diaphragm or EDL myofiber cultures when these were composed entirely of myogenic cells (Fig. 6C). Differently, diaphragm myofiber cultures (but not EDL ones) that contained non-myogenic cells (Supplemental Fig. S1, panels D, F) did exhibit Tcf21 expression (data not shown). Additionally, in agreement with the results for mixed myofibers cultures (i.e., containing both myogenic and non-myogenic cells), Tcf21 was detected only in whole diaphragm and not in EDL muscle (Fig. 6D). Given these collective analyses, we concluded that Tcf21 expressed in muscle domain cultures was contributed by non-myogenic cells that are present in both diaphragm cultures and whole muscle.
Diaphragm Domain Cultures From Pax3Cre × Rosa26mT/flox/mG Mice
The aforementioned expression analyses showing a “retained” expression of Pax3 within the mdx muscle domains (Fig. 5A) as well as an increase in the expression of Pax3 with time in culture from all diaphragm domains (Fig. 6B) led us to the hypothesis of the existence of non-myogenic Pax3-expressing cells. The goal of this study was to potentially distinguish these non-myogenic Pax3-expressing populations, by using a Pax3Cre reporter mouse. As anticipated due to the embryonic origin of the diaphragm muscle (in accordance with early reports about Pax3-mediated reporter expression in limb myogenic cells; Schienda et al. 2006), myoblasts and myotubes exhibited the Pax3Cre-driven GFP reporter expression in the current diaphragm study (Fig. 7A, A′). The majority of the non-myogenic populations noted in Fig. 6A (i.e., fibroblast-like cells and mesothelial cells) were found negative for Pax3Cre as demonstrated by the sole expression of the red reporter in such cells (Fig. 7B–C′). Surprisingly, the nodules detected initially in Fig. 6A (TZ and CT cultures) were characterized by the expression of Pax3Cre-driven GFP (Fig. 7D, D′). Consequently, we examined closely the cultures from all four domains and detected GFP+ nodules across all cultures (as shown next in Fig. 8). These nodules, which were enlarging over time, appeared more distinctive in the TZ and CT cultures; possibly with the initial low density of these cultures and the absence of “dominating” myotubes, such nodules have more opportunity to develop. The nodules that developed in the diaphragm domain cultures were confirmed to be void of skeletal myogenic traits. They were negative for Pax7 and MyoD proteins (determined by immunostaining) and non-reactive with X-gal when cultures were prepared from the reporter mice, Myf5nLacZ and MLC3F-nLacZ (Fig. 7E). These nodules were also void of ancestral expression of MyoD when cultures were prepared from MyoDiCre × Rosa26mT/flox/mG mice (Fig. 7F–F′′).
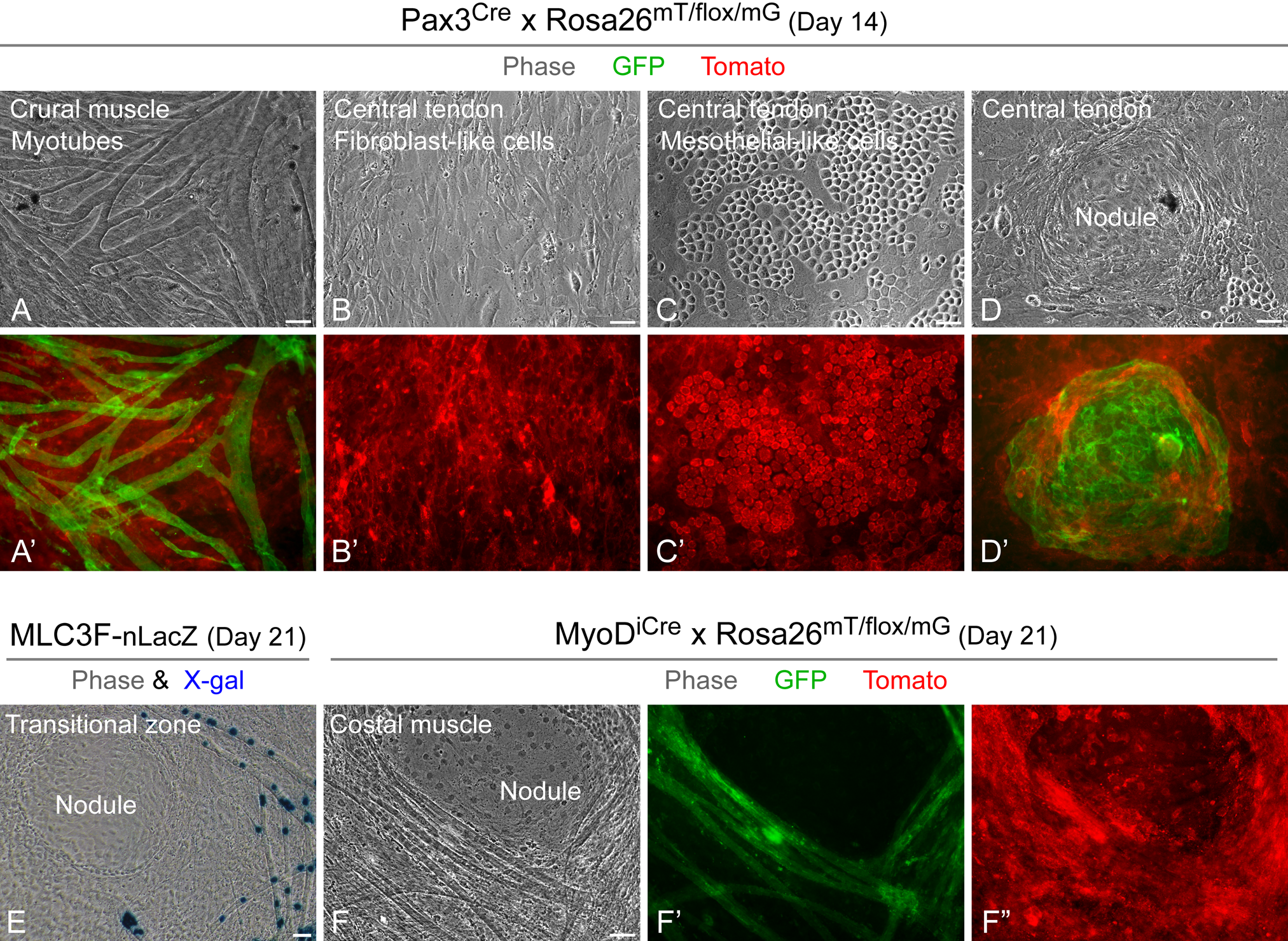
Identification of cell populations expressing Pax3Cre-driven GFP in live diaphragm domain cultures. (A–D′) Representative images of day 14 cultures prepared from the crural muscle and central tendon domains of Pax3Cre x Rosa26mT/flox/mG mouse. In addition to myotubes that were found to be GFP+ as anticipated, non-myogenic cells growing as nodules were also GFP+, whereas all other kinds of non-myogenic cells were Tomato+. (E–F′′) Representative day 21 control cultures prepared from (E) the transitional zone domain of MLC3F-nLacZ mouse (fixed culture for X-gal staining) and (F–F′′) the costal muscle domain of MyoDiCre x Rosa26mT/flox/mG mouse (live culture); LacZ and MyoDiCre-driven GFP expressions are restricted to myogenic cells. Scale bars, 50 µm (A–F′′).
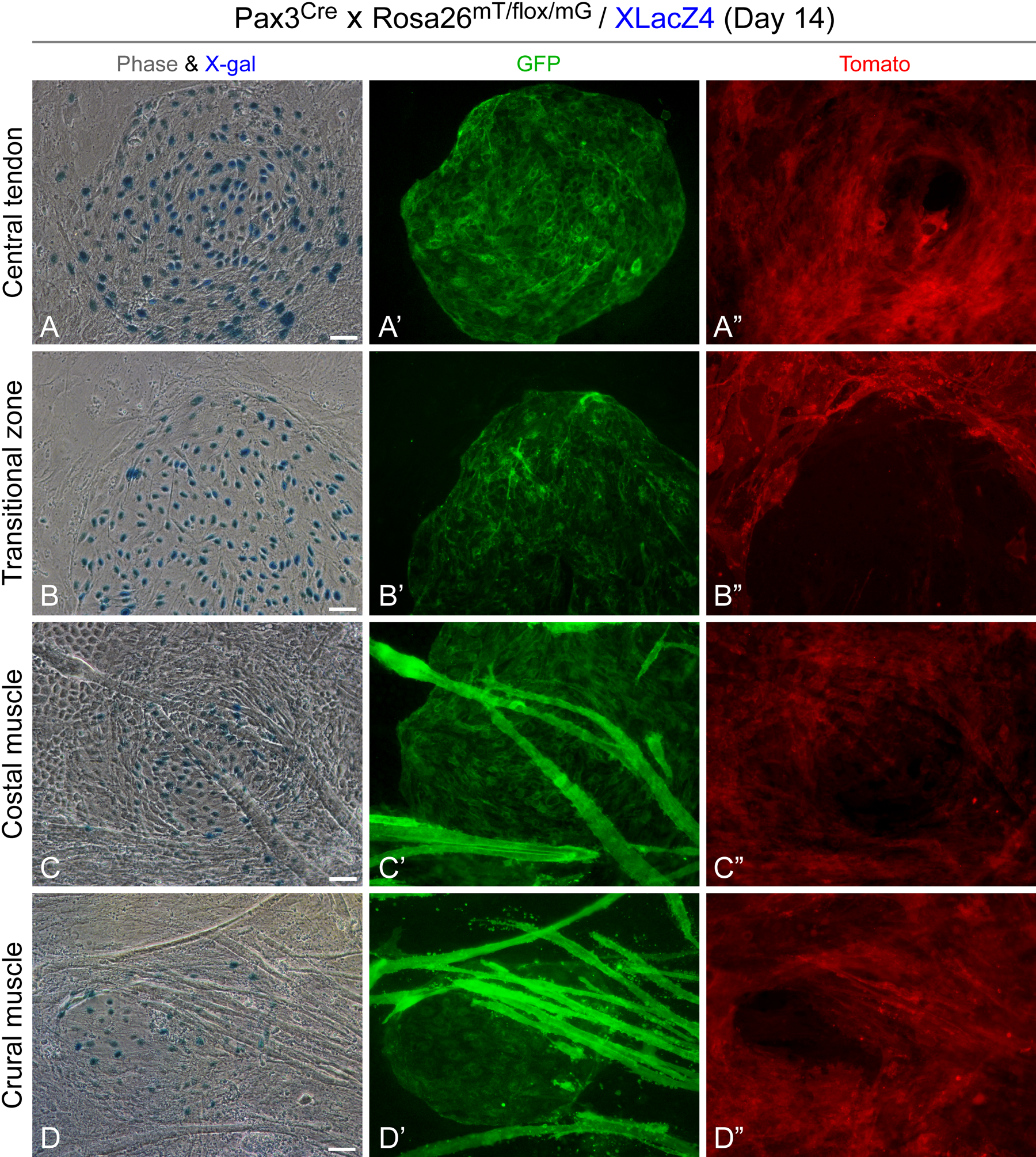
Characterization of the nodules in diaphragm domain cultures by dual expression of Pax3Cre-driven GFP and the XLacZ4 transgene. Cultures of the four diaphragm domains were prepared from a Pax3Cre x Rosa26mT/flox/mG x XLacZ4 mouse. (A–D′′) Representative paired fluorescent and phase images of nodules from the four domain cultures after fixation in 2% paraformaldehyde and staining with X-gal at day 14. The apparent double fluorescent reporter expression at the periphery of the nodules is not within the same cells and is due to outgrowth of the enlarging nodules over Tomato+ cells and/or growth of Tomato+ cells under the nodules. Scale bars, 50 µm (A–D′′).
Utilizing triple heterozygous Pax3Cre × Rosa26mT/flox/mG × XLacZ4, we further demonstrated that many of the Pax3Cre-driven GFP+ nodules harbored XLacZ4+ cells (Fig. 8). Occasionally, XLacZ4 cells appeared scattered individually, lacking GFP expression and resembling what we have observed also in primary cultures from limb muscles (data not shown). However, the majority of XLacZ4+ cells were localized in the observed nodules. Nodules in CT and TZ cultures were more uniform with regard to the expression of XLacZ4 by culture day 14 (Fig. 8A–B′′), while GFP+ nodules in cultures from muscle domains were not fully positive for XLacZ4 by that time point (Fig. 8C–D′′). This distinction might again be reflective of the different niches in which the nodules are developed when cultures have minimal or no myotubes compared with cultures from the myogenic domains.
Based on the dual expression of Pax3Cre-driven GFP and XLacZ4 transgene, we propose that the nodules reflect clonal growth of vascular mural cell-related progenitors. Indeed, a recent study demonstrated that some vascular smooth muscle cells in the adult mouse express Pax3 (Goupille et al. 2011). However, Pax3Cre-driven GFP expression, as seen in the present study, may represent ancestral expression of the reporter and not necessarily de novo activation of the Pax3 gene. In fact, studies on the ontogeny of vascular smooth muscle cells suggest multiple origins, one of which is the neural crest (Le Lievre and Le Douarin 1975; Yablonka-Reuveni et al. 1995; Armulik et al. 2011), a well-recognized Pax3-expressing lineage (Engleka et al. 2005). Furthermore, the progenitors contributing to the establishment of these nodules can potentially be multipotential adult stem cells of neural crest origin and not necessarily represent differentiated progeny (Hauser et al. 2012). We propose, however, that the observed Pax3Cre-driven GFP+ cells may represent true Pax3 upregulation upon vascular mural cell lineage determination in the cell cultures. The source for this process is potentially the mesothelial cells, especially as there are already examples of such an origin for vascular mural cells in the gut, lung, and liver (reviewed in Armulik et al. 2011). Coronary vessel mural cells in the heart also appear to develop from the mesothelium (Armulik et al. 2011), and interestingly, Tcf21 was reported to be involved in this differentiation process (Ishii et al. 2007).
We are presently designing experiments to establish the origin and function of the Pax3Cre+/XLacZ4+ cells detected unexpectedly in the present study of diaphragm domain cell cultures. Additionally, in view of the aforementioned implication of Tcf21 in the epithelial-mesenchymal transition of mesothelial cells, a potential link among mesothelial cells, Tcf21-expressing cells, and Pax3Cre+/XLacZ4+ cells will be closely examined in the diaphragm.
Concluding Remarks
The diaphragm (Greek: dia = across, phragma = wall) is an important organ vital for life-supporting functions and is certainly in many respects unique among the muscles of the body. The oldest known record concerning the diaphragm goes back to the Iliad of Homer in his descriptions of wounds suffered by Greeks and Trojans (9th century BCE). The diaphragm continued to fascinate Greek philosophers and physicians for many centuries with theories ranging from the metaphysical nature of the diaphragm to its physiological role as a somatic organ (Fritts 1976; Derenne et al. 1994). Interestingly, the ancient recognition of the diaphragm as the seat of emotions, linked with the soul and the process of thoughts, prevails at the present time in alternative medicine and especially in yoga where the conscious diaphragmatic breathing supports emotional equilibrium (Jaloba 2011). With the accomplishment of Galen (130–200 CE), the anatomical and physiological knowledge about the respiratory system reached a complete comprehension that is still considered correct in modern times (Fritts 1976; Derenne et al. 1995). Almost all of this knowledge was then lost over the next few hundred years, to be only rediscovered during the Renaissance. In particular, sketches from the notebooks of Leonardo da Vinci depict the diaphragm as a hollow spoon held upside down with inspiratory and expiratory points (Fritts 1976). By the middle of the 18th century, the muscularity of the diaphragm and its role in respiration and other diverse functions had been well recognized as was evident in the writings of Albrecht von Haller (reviewed in Fritts 1976). The 20th-century pioneering work of Duncan Blair about the diaphragm central tendon (Blair 1923) and that of Felix Beck and James Baxter about the diaphragm blood supply (Beck and Baxter 1960) cannot be overlooked either when considering the history of diaphragm research.
Here, we have provided a comprehensive structural and gene expression analysis of the mouse diaphragm muscle and tendon domains, based on whole tissue and cell culture studies. Our study may add only little new knowledge compared to what has been gleaned during many centuries of diaphragm research. Nevertheless, our “rediscovery” of old knowledge about the diaphragm anatomy, combined with new microanatomy insights reported herein, is significant for future investigations on the diaphragm basic biology and pathological conditions.
Footnotes
Acknowledgements
We thank our team members for their important contributions to this study. Maria Elena Danoviz and Andrew Shearer contributed some of the mouse strains used in the present study. Yunfei Li provided excellent technical support with diaphragm cultures.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants to Zipora Yablonka-Reuveni from the National Institutes of Health (AG021566; AG035377; AR057794) and the Muscular Dystrophy Association (135908).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
