Abstract
The intracellular microsporidian parasite Loma salmonae affects salmonids of the genus Oncorhynchus and is a significant cause of economic losses in pen-reared Chinook salmon (O. tshawytscha) in British Columbia. Loma salmonae infection is easily recognized by the xenomas that form in the gills, but early stages of infection are difficult to detect in histologic sections. In situ hybridization (ISH), using an L. salmonae-specific digoxigenin-labeled single-stranded DNA probe, was used to detect the parasite during the early stages of infection. Loma salmonae was detected in the gut mucosal epithelium as early as 24 hours postexposure (PE), and it localized in the lamina propria of the intestine within 24 hours of infection. After the parasite was detected in the lamina propria, dividing merogonic stages in infected cells in the heart were detected by ISH as early as 2 days PE, providing the first evidence of parasitaemia and hematogenous distribution of this parasite in infected blood cells. The parasites inside the infected cells appeared to be undergoing merogony as they passed through the heart, indicating that proliferation may start at the site of infection, before the parasite arrives to the gills for their final developmental phase. This is the first time that L. salmonae passage through the intestinal wall and migration to the heart has been visualized; however, the identity of the cells harboring the parasite has yet to be determined.
Loma salmonae, an obligate intracellular parasite of the gills of salmonid fish of the genus Oncorhynchus, 25 is the causal agent of microsporidial gill disease in chinook (O. tshawytscha) and coho (O. kisutch) salmon and in rainbow trout (O. mykiss). 5,15,17,19,22,38 Salmon infected with L. salmonae have abundant xenomas full of spores in their gills 10,25 and experience high mortality rates and increased susceptibility to other infectious diseases. 37 The economic losses attributed to infections with L. salmonae are considerable 6 and are caused by the rupture of mature xenomas and the resultant formation of granulomatous inflammation in the gill. 12 The lack of approved drugs to control L. salmonae infections in the aquaculture industry in Canada 4,26,36 makes it necessary to elucidate aspects of the parasite's life cycle to find ways to stop the progression of disease.
Infection with L. salmonae occurs following germination of spores in the gut and the infection of a gut epithelial cell. 35 Until recently, very little was known of the parasite's life cycle prior to the formation of xenomas. Recent studies 32 using the polymerase chain reaction (PCR) have shown that L. salmonae is able to move rapidly (3 days postexposure [PE]) from the intestine to the heart, appearing in the gills of infected fish 2 weeks PE at 15 C. However, it has been difficult to visualize these early stages of infection. An in situ hybridization (ISH) protocol developed for the diagnosis of L. salmonae presporogonic stages in the gills of infected fish 31 was used in this study with the objective of visualizing these early stages. ISH involves the anatomic localization of labeled RNA or DNA molecules that hybridize with complementary target RNA or DNA sequences in the cell, 39 allowing the visualization of infectious agents. 28
The L. salmonae-specific probe for ISH 31 was developed from L. salmonae-specific primers 10 that recognize the internal transcribed spacer regions (ITS) and portions of the gene encoding the small subunit (SSU) and the large subunit (LSU) of the ribosomal DNA sequence of L. salmonae. DNA-based diagnostic methods such as ISH 31 are especially suited for the study of pathogen infections because of their specificity and sensitivity. The uniqueness of the DNA sequences of the host and the pathogen allow recognition of the pathogen DNA in the host regardless of the stage present. 40
This report documents the first use of the L. salmonae-specific DNA probe in an ISH protocol for the detection of the very early stages of the parasite in the intestine of experimentally infected fish.
Materials and Methods
Sample population and experimental design
Naive, size-graded, diploid juvenile rainbow trout (∼20 g) were acquired from a commercial hatchery on Prince Edward Island, Canada, with no previous history of L. salmonae infections. All procedures were performed according to the guidelines of the Canadian Council on Animal Care. 1 The 100 juvenile rainbow trout used for the study were kept in a 100-liter circular fiberglass tank at 15 C. The trial was conducted over a 1-week period.
Fish infection and sampling
Fish were infected by gastric intubation with finely cut xenoma-laden gill filaments from rainbow trout previously infected with L. salmonae. All infected gills were harvested from fish euthanatized with an overdose of benzocaine. The cartilaginous gill arch was removed and the remaining gill filaments were finely cut until only a pulp remained. The pulp was mixed with an equal volume of distilled water. The inoculum was taken up into a 1-ml syringe fitted with a 5-cm piece of tubing of approximately 5 mm in diameter. Each fish was given 0.1 ml of the pulp.
Because of the small size of the fish, whole heart, stomach, and small and large intestine were sampled from five fish at 1, 2, 4, 8, 12, 18, 24, 30, 36, 42, 48, 60, and 72 hours PE and 4, 5, 6, and 7 days PE. Every sample was fixed in 4% paraformaldehyde (Sigma) in phosphate-buffered saline (PBS) for 40 minutes at room temperature and then stored at 5 C in 70% ethanol until processed further. The tissues were dehydrated in ethanols (70%, 90%, 95%, and 100%) and embedded in paraffin in an automatic tissue processor. The stomach and small and large intestine were placed longitudinally to allow examination throughout their length. Three sections 6 µm thick were cut and placed on slides that had been pretreated with 3-aminopropyltriethoxysilane (Sigma) and dried for 2 hours at 40 C.
ISH
A single-stranded DNA (ssDNA) probe was used. The conditions for probe generation and a detailed protocol for the ISH have been described elsewhere. 31 Dried tissue sections were deparaffinized in xylene, rehydrated in a descending ethanol series (100%, 95%, and 70%), and washed in distilled water. The tissues were equilibrated in TES buffer (50 mM Tris-HCl, 10 mM ethylenediaminetetraacetic acid, 10 mM NaCl, pH 7.4) for 10 minutes before treatment with proteinase K (15 µg/ml in TES) for 15 minutes at 37 C. Proteolysis was halted with two washes in PBS with 2 mg/ml glycine (Fisher Scientific) for 5 minutes each. Tissue sections were immersed in aqueous 20% acetic acid at 4 C for 15 seconds and then washed in two changes of distilled water for 10 minutes each. Tissues were briefly postfixed in 0.4% paraformaldehyde in PBS for 5 minutes, rinsed in distilled water, and air dried.
A hybridization mix was made by mixing 5 µl 100× Denhardt's solution (2% Ficoll 400 [Pharmacia], 2% polyvinylpyrrolidone, 2% bovine serum albumin [BSA; Fisher Scientific]), 50 µl 50% dextran sulfate (Sigma), 10 µl 100 mg/ml sperm DNA (Boehringer Mannheim), 100 µl 20× standard saline citrate (SSC: 3 M NaCl [Sigma], 0.3 M sodium citrate [Sigma], pH 7.0), sterile deionized water to 250 µl, and 250 µl formamide (Sigma) with 10 ng of digoxigenin (DIG)-labeled probe. One hundred microliters of this mix was applied to the permeabilized tissue sections. The slides were covered with hybri-slips (Sigma), placed onto a preheated baking tray, and incubated at 90–95 C for 3 minutes to denature the target DNA. The slides were then incubated at 40 C overnight in a humidified chamber.
After hybridization, the sections were washed twice with 4× SSC for 5 minutes at room temperature. The slides were equilibrated in Tris-buffered saline (TBS: 50 mM Tris-HCl [Fisher Scientific], 100 mM NaCl [Sigma], pH 7.2) for 5 minutes and blocked with TBT (TBS, 3% BSA [Fisher Scientific], 0.5% Triton X-100 [Fisher Scientific]) at room temperature for 10 minutes. The sections were incubated with alkaline phosphatase (AP)-conjugated anti-DIG Fab fragments (Boehringer Mannheim) diluted 1:600 in TBT for 1 hour. After incubation, the slides were washed twice in TBS for 5 minutes. The signal (purple precipitate) was developed by applying 200 µl of a buffer containing 45 µl nitroblue tetrazolium (Boehringer Mannheim) and 35 µl bromo-chloro-indolyl-phosphate (Boehringer Mannheim) in 10 ml of 100 mM Tris-HCl, 100 mM NaCl, 50 mM MgCl2 (Sigma), pH 9.5, and incubating the slides for 5 hours in the dark. Color development was terminated by washing in distilled water for 5 minutes. The slides were counterstained with 10% methyl green, air dried, and mounted with Permount (Fisher). Sections were viewed and photographed with an Olympus BX50 photomicroscope.
Negative controls included naive fish tissue hybridized with the labeled probe, infected fish tissue hybridized with unlabeled probe or with hybridization buffer without probe, and infected fish tissue hybridized with the labeled probe but without incubation with the AP-conjugated anti-DIG Fab fragments.
Results
Hybridization of the probe to paraffin-embedded sections of fish tissue infected with L. salmonae yielded strong color development and no hybridization to normal fish tissue. Hybridization signals were observed as purple precipitates in target tissues, indicating the binding of the labeled probe to L. salmonae DNA. Negative controls showed no hybridization signals in the sampled organs (Fig. 1), validating the probe specificity in infected fish; no background or nonspecific binding of reagents occurred.
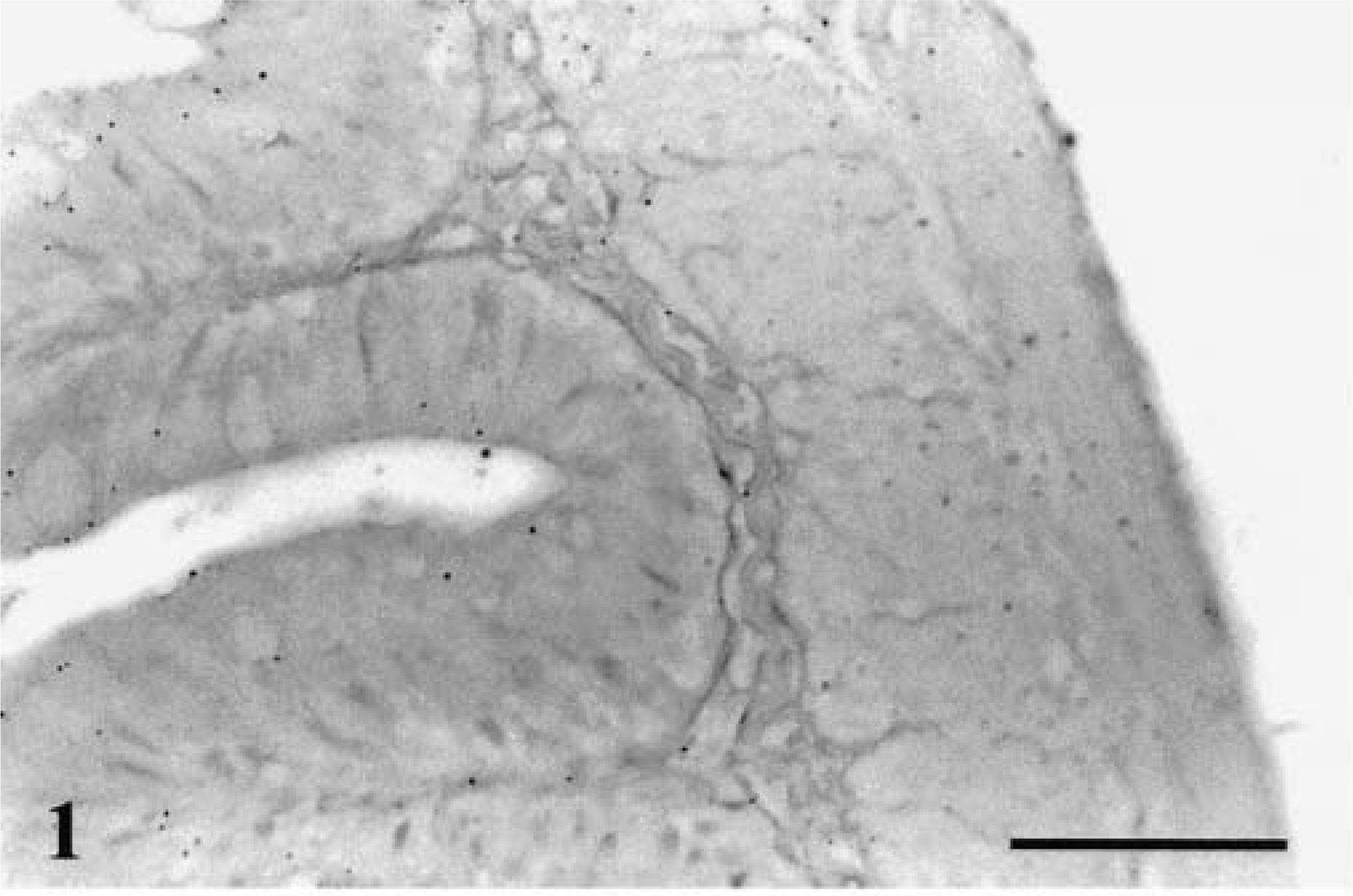
Small intestine; rainbow trout. No hybridization is observed. ISH with L. salmonae ssDNA DIG-labeled probe and AP-conjugated anti-DIG Fab fragments, methyl green counterstain. Bar = 100 µm.
Parasite DNA was identified in stomach content material as early as 2 hours PE and in close association with the mucosal epithelium of the stomach at 4 hours PE (Fig. 2). Other than these observations, no other stages were ever detected in the stomach region.
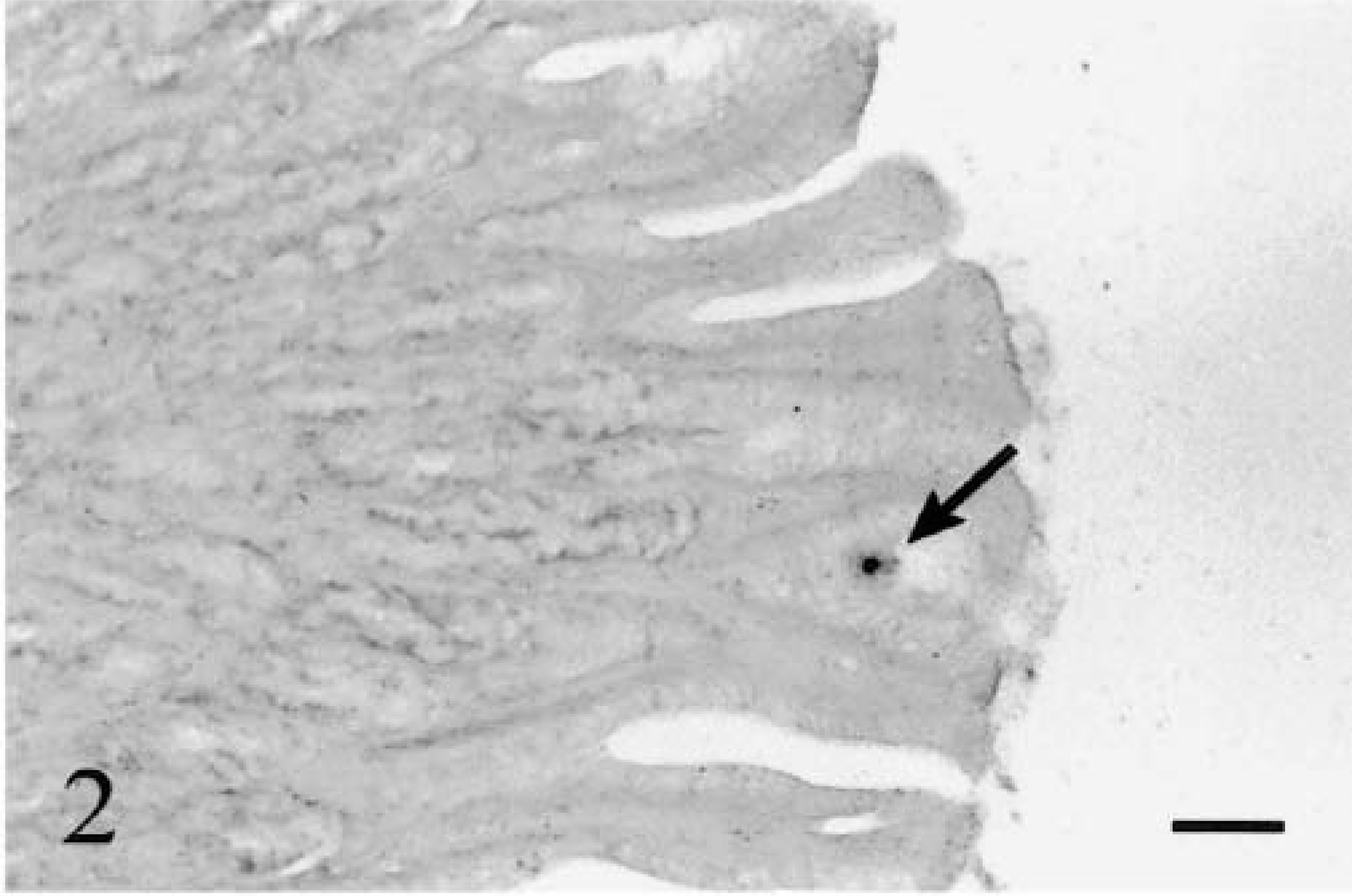
Stomach; rainbow trout infected with L. salmonae, 4 hours PE. A sporoplasm (arrow) is in association with the mucosal epithelium of the stomach. ISH with L. salmonae ssDNA DIG-labeled probe and AP-conjugated anti-DIG Fab fragments, methyl green counterstain. Bar = 100 µm.
In the intestine, parasite DNA could be detected in epithelial cells and lamina propria of the small intestine as early as 12 hours PE (Fig. 3) and could still be detected at 24 hours PE (Fig. 4). In the large intestine, positive signals could be observed as early as 12 hours PE in the lamina propria and were still visible at 24 hours PE (Fig. 5). These signals were not detected in sites other than the gastrointestinal mucosa.
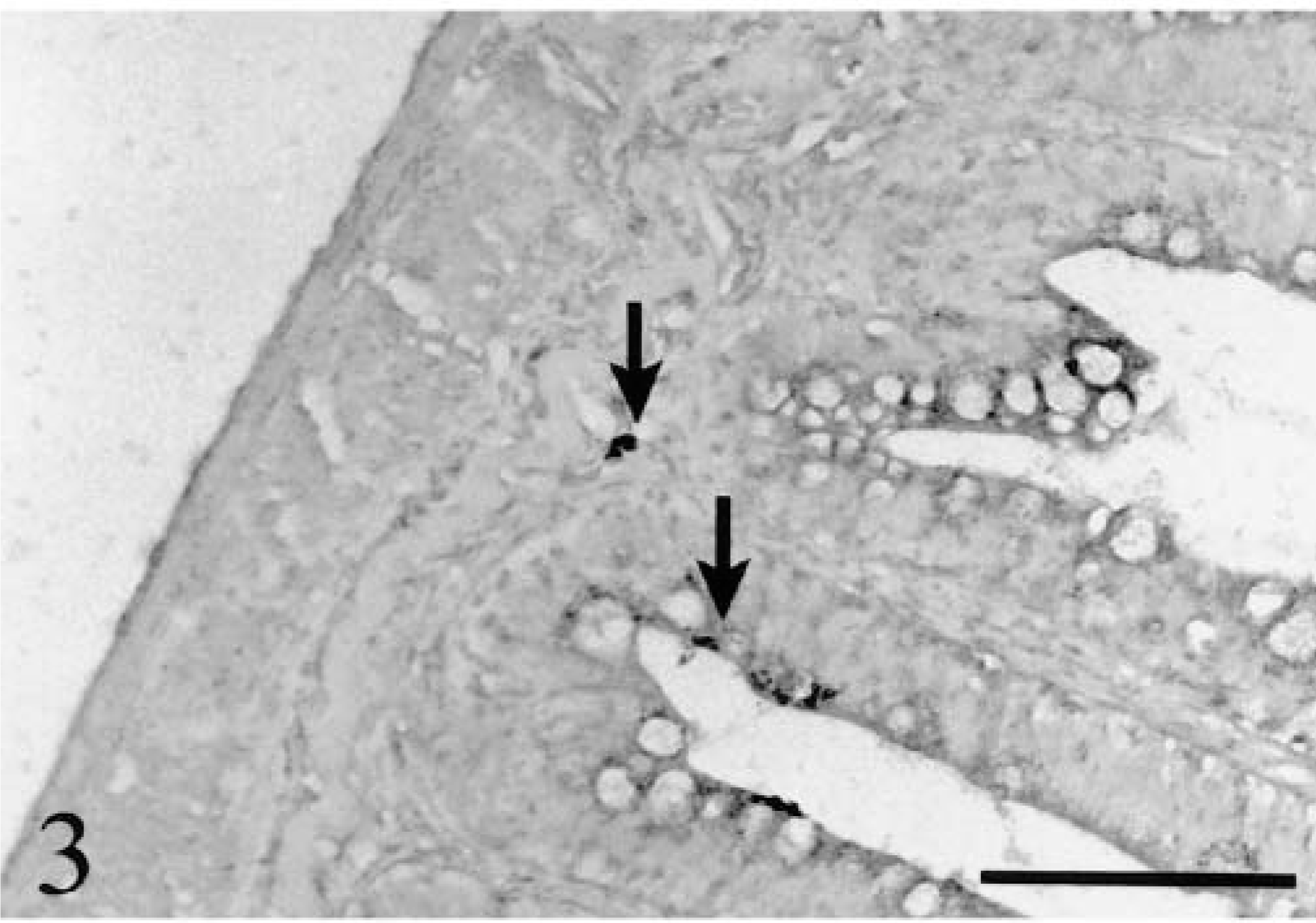
Small intestine; rainbow trout infected with L. salmonae, 12 hours PE. Early stages of infection (meronts) are at the lamina propria level or close to the intestinal lumen in enterocytes (arrows). ISH with L. salmonae ssDNA DIG-labeled probe and AP-conjugated anti-DIG Fab fragments, methyl green counterstain. Bar = 100 µm.
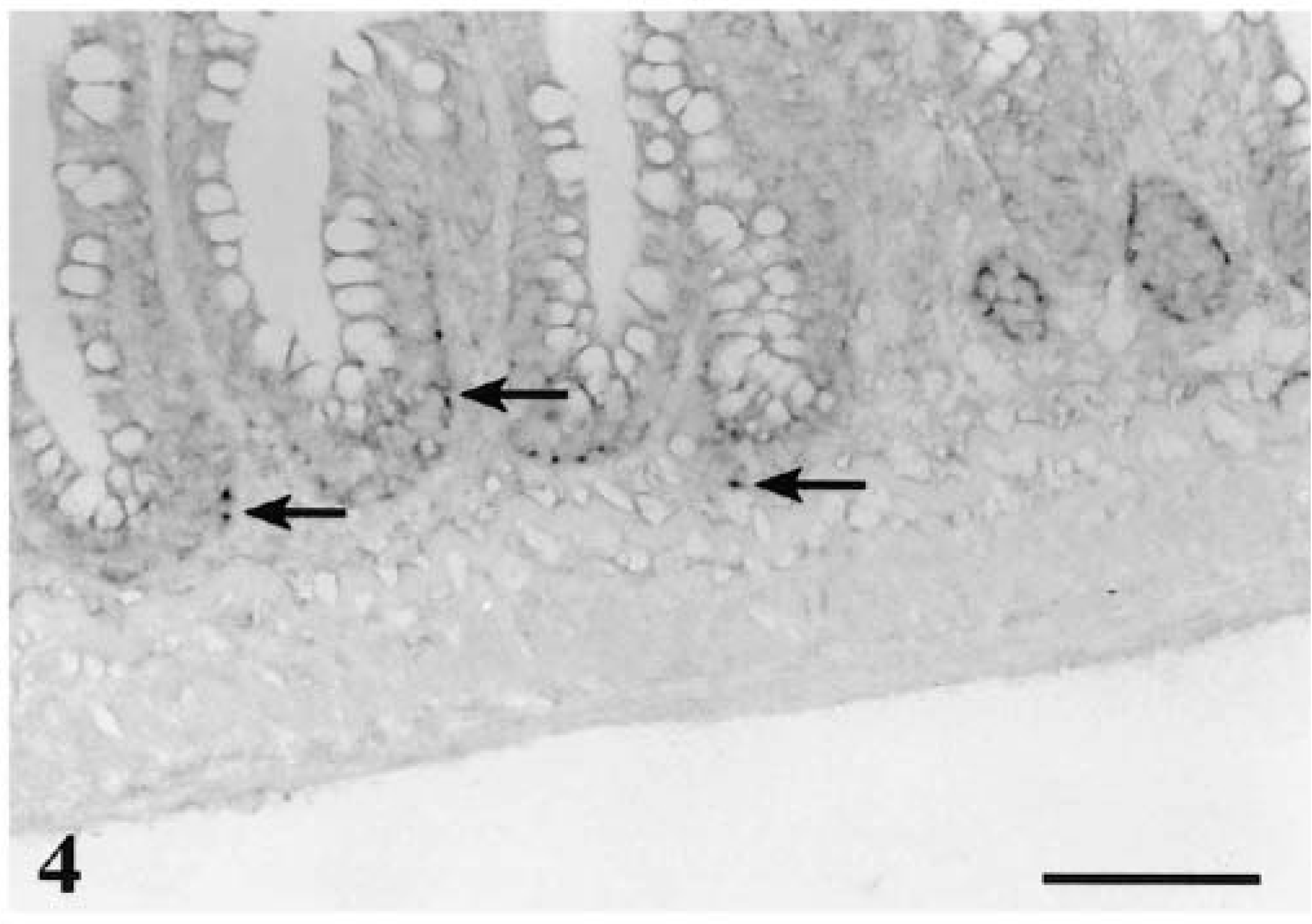
Small intestine; rainbow trout infected with L. salmonae, 24 hours PE. A large number of early stages (meronts) are associated with the lamina propria or the base of enterocytes (arrows). ISH with L. salmonae ssDNA DIG-labeled probe and AP-conjugated anti-DIG Fab fragments, methyl green counterstain. Bar = 100 µm.
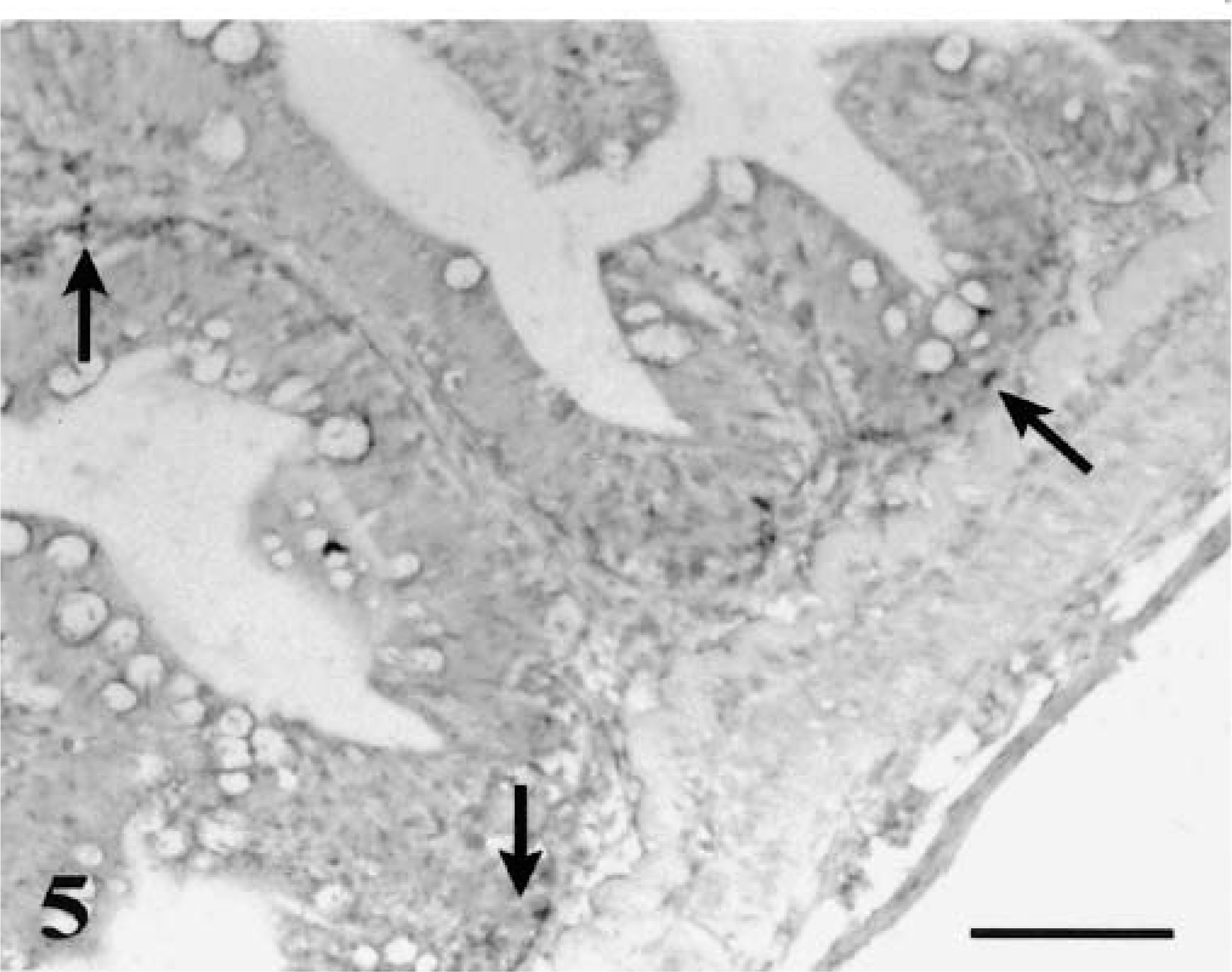
Large intestine; rainbow trout infected with L. salmonae, 12 hours PE. Numerous early stages (meronts) are at the lamina propria level (arrows). ISH with L. salmonae ssDNA DIG-labeled probe and AP-conjugated anti-DIG Fab fragments, methyl green counterstain. Bar = 100 µm.
As early as 2 days PE, infected cells with what appeared to be dividing stages were detected in the heart ventricle. These stages were better visualized at 5–6 days PE and were associated with cells mixed with the blood cells (Fig 6a) or in the ventricular endocardial lining (Fig 6b).
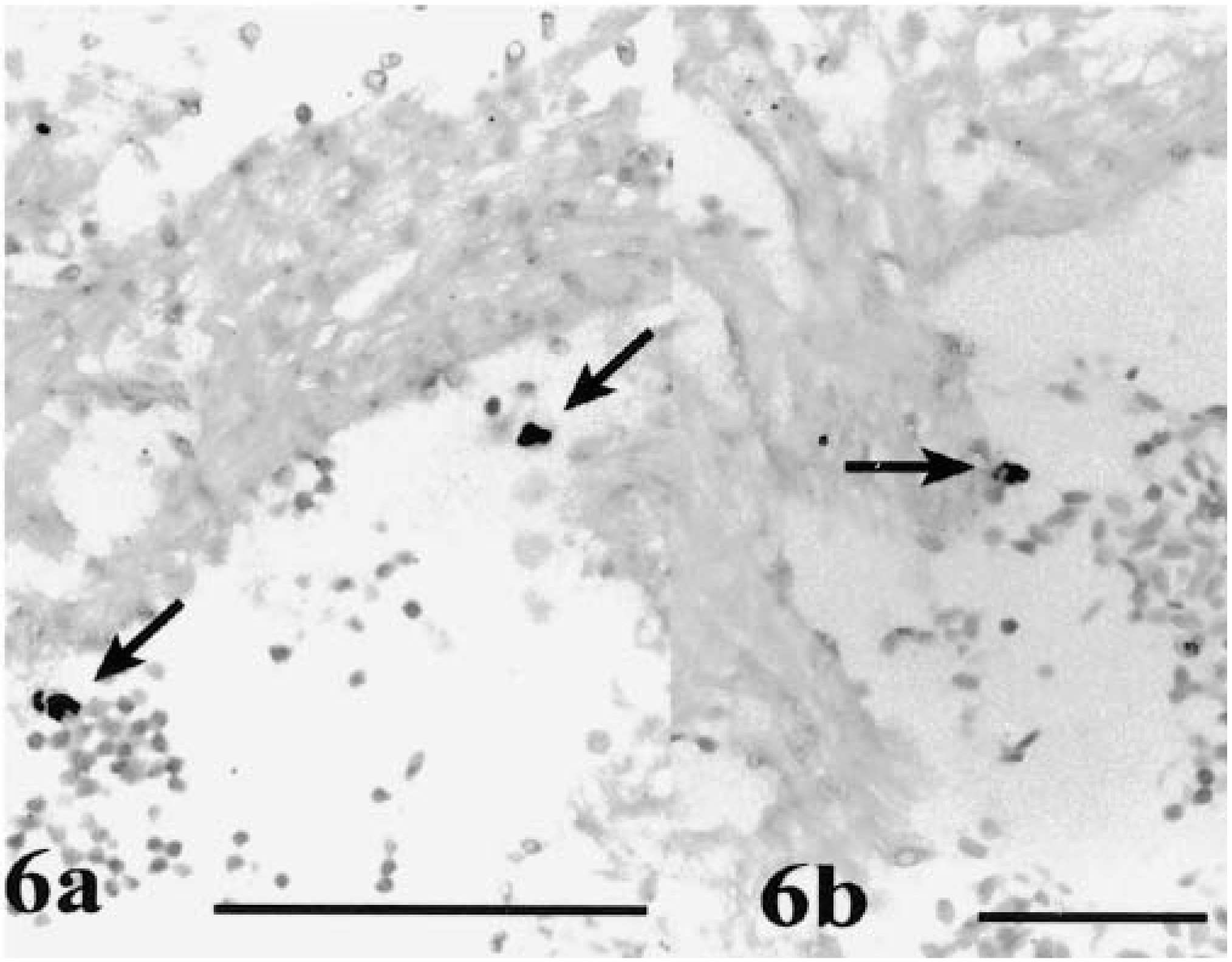
Heart ventricle; rainbow trout infected with L. salmonae. Fig. 6a At 5 days PE, infected cells (arrows) with early merogonic stages of L. salmonae are among noninfected cells in the blood. Fig. 6b At 6 days PE, an infected cell (arrow) is in association with the ventricular endocardial epithelium. ISH with L. salmonae ssDNA DIG-labeled probe and AP-conjugated anti-DIG Fab fragments, methyl green counterstain. Bars = 100 µm.
Discussion
Early stages of L. salmonae were localized in rainbow trout as the parasite passed through the intestinal wall and as infected cells passed through the heart.
Most of the molecular biology techniques in use for the detection of microsporidian pathogens in humans or commercially important animals rely extensively on PCR, 13 which can accurately detect the presence of a pathogen although it does not provide information on the localization of the target pathogen within the organ. This problem can be overcome with the use of in situ PCR 27 or ISH, 43 both of which are very sensitive approaches that detect the presence of a pathogen within a single cell among tens of thousands of uninfected cells, allowing precise localization of a pathogen at the tissue, cellular, and subcellular level. 14,27,29 The hybridization signals confirm the presence of complementary DNA, which may be (but is not always) synonymous with the presence of a whole organism. ISH has been used for the differentiation of viral strains in crustaceans, 11 the detection of early stages of infection of myxosporidia in fish and their tubificid hosts, 3 the diagnosis of microsporidian-like Enterocytozoon bieneusi in immunosuppressed humans, 42 and the tracking of phage DNA through the intestinal wall of rodents. 34
The usefulness of the ISH assay for the detection of L. salmonae in the very early stages of infection in the intestine and heart was tested. By combining the advantages of morphologic observation with molecular information at the level of nucleic acid sequences, 42 this method allowed the visualization of infected host cells in the heart that had not been observed before. Although this technique cannot differentiate between vegetative and live stages, the detection of dividing stages within the infected cells in the heart suggests that the parasite was viable. To our knowledge, results of this investigation are the first to track a fish microsporidian, in this case L. salmonae, as it is transported through the host after penetration of the body. Conditions used in the ISH protocol for this study resulted in optimal visualization and therefore localization of the parasite. Although ssDNA probes do not need denaturation prior to hybridization, addition of the probe before tissue denaturation may have resulted in the strong signals observed, as has been reported previously. 2 However, the technique has its limits; for example, the use of paraffin-embedded sections and light microscopy does not allow resolution of the specific cell types that are host to the parasite, especially when the hybridization signal is so intense that it precludes the visualization of the morphology of the infected cell.
Putative early developmental forms of L. salmonae previously were revealed in the intestine of infected chinook salmon by routine histology with hematoxylin and eosin (HE) staining, 35 and the portal of entry of L. salmonae was assumed to be the intestinal wall. However, in studies of human intestinal microsporidia, in which comparisons between ISH and other more conventional histologic methods have been made, a variety of parasite stages were detected with ISH that were missed when the sections were stained with HE or Giemsa. 18,20,21,41 The structure seen in the intestine of chinook salmon and believed to be a sporoplasm 35 may be another structure; observation of sporoplasms or meronts by conventional techniques in the host tissue is very difficult unless the parasite has entered the sporogonic phase. The development of ISH in this study has allowed for the identification of the initial stages of L. salmonae in the stomach lumen as early as 2 hours PE and in close association with the epithelial surface of the stomach 4 hours following exposure to mature spores, although no sporoplasms were seen inside the cells after this time.
Loma salmonae sporoplasms that penetrate the gut epithelium of the fish host appear to migrate to the lamina propria, and the results suggested that there was not a single preferential site for infection because positive signals (possibly sporoplasms) were detected in both the small and large intestine. From this site, they presumably access the blood flow and are able to move to other parts of the body of the fish. Although infection begins as the sporoplasm penetrates the mucosal epithelium of the intestine, 35 it is not known with certainty whether the sporoplasms or the meronts are capable of moving between cells. The results of this study unequivocally demonstrate for the first time that L. salmonae spores infect the fish through the gut. L. salmonae then migrates to the lamina propria, from where it seems to be transported to other parts of the body inside infected cells. The putative transport cells have not yet been identified, but, as in the case of Eimeria, they could be intraepithelial lymphocytes 8 or T cells 16,30 or migratory cells such as monocytes. 23 How these cells become infected is not known, but they might become infected during phagocytosis of the parasite in the lamina propria of the intestine. 9 Microsporidia (E. bieneusi) observed in the lamina propria of infected monkeys had the appearance of being within macrophages, suggesting the possibility of hematogenous spread of the parasite. 41 This hypothesis had not been experimentally demonstrated, but our results provide the first evidence of L. salmonae transport through the body of the host fish within infected cells found in the blood.
The results of this study suggest that in L. salmonae, as with some myxosporidians 24 and other microsporidians, 7 merogony (the proliferative phase that leads to the formation of a high number of meronts 9 ) begins in sites other than the site of xenoma formation. The infective sporoplasm seems to have initiated the early dividing (merogonic) stages in the infected host cells by the time they were detected in the heart (5 days PE), although the proliferative phase is completed at the site of sporulation in the gills, as has been observed in a previous study. 31 Furthermore, the ability of the parasite to divide within these cells, which do not appear to destroy it, suggests that the signals are not produced by degraded pieces of DNA but probably by the whole organism. This finding suggests that L. salmonae is transported from the site of germination to the site of sporulation inside infected cells. Whether these cells are the original infected enterocytes or are monocytes infected by the movement of the meronts from cell to cell is not known. Spore passage from the intestine to the blood circulation by the action of some phagocytic cell has been described for Encephalitozoon cuniculi, 7 and the investigators proposed that the first cycle of growth occurred in these phagocytic cells. This mechanism allows the movement of the parasite to organs with a high blood flow. Our results confirm this type of movement for L. salmonae; cells harboring the dividing parasite were detected as they passed through the heart of the infected fish. Evidence of these stages in transport cells had not been observed previously. Studies to determine the specific identity of the infected cells should be undertaken. The ISH technique adapted for transmission electron microscopy with the aid of a monoclonal antibody specific for different fish leukocyte subpopulations could be used to better localize these early stages of development so that their relationship with other cells and, more importantly, the identity of the cells that are used as host and for transportation could be determined.
Loma salmonae was previously considered relatively nonpathogenic to salmonids in fresh water but is now recognized as a cause of high morbidity and mortality in Pacific and chinook salmon in Canada. 33 The development and use of an L. salmonae-specific probe allowed definition of various aspects of the life cycle that have been difficult to visualize because of the lack of sensitive methods of detection and localization of the initial stages of infection. ISH is a sensitive and specific indicator of microsporidial infection in fish. In this study, it was used to reveal stages of the parasite that would have been difficult to appreciate with conventional histochemical techniques. This higher sensitivity and specificity of ISH is not unexpected because whereas conventional techniques are used to detect specific anatomic structures, such as the spore wall, ISH is capable of detecting the DNA of the parasite, which is present in all life stages. Although ISH has been utilized to investigate the pathogenesis of human microsporidia, 21 its use in studies of microsporidial infections in fish had not been previously reported.
Footnotes
Acknowledgements
Funding for this work was received through the Natural Sciences and Engineering Research Council of Canada Strategic Grant Program.
