Abstract
Objective
To compare maternal plasma with serum for measuring markers currently used in first and second trimester screening for Down's syndrome.
Setting
A laboratory-based investigation of two sample types in assays used in prenatal screening for Down's syndrome.
Methods
A paired data-set included both plasma and serum from 101 pregnant women. A nested case/control data-set included only plasma samples from 34 first and 23 second trimester Down's syndrome pregnancies, each matched with six euploid controls. Analyte levels were measured and converted to multiples of the median (MoM).
Results
In the paired data-set, each of the five analytes (alphafetoprotein, unconjugated estriol, human chorionic gonadotropin, inhibin-A and pregnancy-associated plasma protein A) in serum and plasma was highly correlated (r > 0.970) and after conversion to MoM, the resulting distributions were equivalent (P > 0.7). In the matched data-set, plasma-based median MoM levels in cases were consistent with the published serum counterparts for all markers.
Conclusions
This study provides strong evidence that current serum-based prenatal screening can be performed equally well using plasma samples. This may prove useful, especially if secondary screening using a DNA-based test requires maternal plasma.
Introduction
Evidence has accumulated to show that molecular-based testing can significantly improve the process of prenatally diagnosing Down's syndrome. In 1997, free circulating DNA of fetal origin was identified in maternal plasma. 10 Since then, several groups have quantified the amount of circulating chromosome 21 fragments in maternal plasma by next-generation sequencing methods, allowing the identification of pregnancies with Down's syndrome and other common aneuploidies.11-16 In a recent collaborative study, we reported the results of a blinded clinical validation of a laboratory-developed test (LDT).16–18 Overall, 98.6% of the 212 singleton Down's syndrome pregnancies were identified and 0.2% of the 1471 matched controls were falsely positive; testing failed in 13 additional control samples (0.8%). High performance was also found for trisomy 18 and trisomy 13. 17
Several issues currently limit the use of next-generation sequencing as a first-line prenatal screening test. Clinical validation studies of these LDTs have enrolled patients already identified as being at high risk (e.g. prenatal test screen positive, abnormal ultrasound findings and advanced maternal age) and having invasive diagnosis, so that proof of fetal karyotype is available. Such testing is labour-intensive, costly and turn-around time is up to 10 days. Next-generation sequencing, however, can be used now as an intermediate step before invasive diagnostic testing among those pregnancies at high risk of Down's syndrome or other common aneuploidies. In this way, the offer of invasive procedures to those who are falsely positive by screening can be reduced by up to 95%, increasing the overall safety of prenatal diagnosis while maintaining high detection.
Most next generation sequencing methods tested so far have required the use of ethylenediaminetetraacetic acid (EDTA) plasma to allow for DNA extraction, while current first-line screening tests rely on a serum sample. We speculated that EDTA plasma might be an acceptable sample type for measurement of the routine prenatal screening markers (i.e. PAPP-A, AFP, uE3, hCG and InhA). If so, the collection of a single tube type could be used for both the initial screening test as well as the secondary screening by next generation sequencing, simplifying the process. This would also avoid the delay and anxiety created in informing a woman about an initial high-risk result. This latter advantage could also be realized by initially collecting a plasma and a serum sample, but this would complicate processing and storage.
Two sample sets were utilized. The first was a set of paired maternal serum and plasma samples, collected at the same time, from low-risk women undergoing routine first and second trimester prenatal screening. These plasma samples were further divided into low- and high-speed supernatants to determine whether centrifugation speed affected the measurement of the ‘serum’ markers (because next-generation sequencing of free DNA in maternal plasma requires a high-speed centrifugation to ensure removal of all cell debris). A second nested case/control set of residual plasma samples was available from women with known euploid and Down's syndrome pregnancies from our cohort study originally collected to validate a next-generation sequencing methodology.
Methods
Paired Plasma And Serum Samples
Women presenting to phlebotomy for routine prenatal serum screening in a clinic setting at Women & Infants Hospital (WIH) were asked to donate an additional tube of EDTA plasma at the same time for research purposes. The study was approved by the Institutional Review Board for Human Studies at WIH and all participating women signed the consent. Paired serum and plasma samples were collected from 101 women presenting for routine prenatal screening, 50 in the late first trimester and 51 in the early second trimester. Since the opportunistic sampling occurred before routine screening, the samples were representative of the general pregnant population.
Both the serum and plasma samples were transported at room temperature by courier from the phlebotomy station to the laboratory at WIH, within 24 hours. Upon receipt, a random subset of samples were centrifuged at low speed (2500 g), with an aliquot of all plasma samples subject to a high-speed spin (15,500 g). A 1-mL aliquot of the plasma spun at high speed was immediately stored at -80°Cto allow for quantification of fetal DNA at the Sequenom Center for Molecular Medicine (SCMM, San Diego, CA, USA). All remaining samples were stored in the refrigerator until testing. Assays were completed within three days of blood draw, at which time any residual sample was stored at – 20°C.
A subset of plasma samples having both low- and high-speed spins were compared with serum and to each other using correlation analysis. For each analyte, paired serum and plasma test results were compared using Bland-Altman analysis.19,20 Gestational age-specific reference data were calculated and results converted to multiples of the median (MoM). The MoM results were analysed via a probability plot to determine the range over which the logarithmically transformed data fit a Gaussian distribution. A standard deviation of the log MoM values was computed using data from the probability plot; 95% confidence intervals were computed on a logarithmic scale assuming a Gaussian distribution. Statistical significance was two sided at the P = 0.05 level.
Nested Case/Control Sample Set
A second group of maternal EDTA plasma samples, stored frozen at — 80°C, was available from women who had participated in an international clinical validation study to determine the ability of a maternal plasma-based DNA test to identify Down's syndrome. 16 This study was conducted under institutional review board approval (or equivalent) at each site and at the Coordinating Centre at WIH in Rhode Island. Samples had been drawn into EDTA plasma tubes immediately prior to invasive testing, kept refrigerated and processed by high-speed centrifugation within six hours (average 2 hours). Each patient's sample set was assigned a unique study code and aliquots were stored at — 80°C for approximately two years prior to testing for screening marker levels.
To evaluate the measurement of screening markers in plasma, a nested case – control set was developed. To obtain unbiased estimates of the plasma marker parameters, women referred for diagnostic testing due to positive serum screening results were excluded. Samples with less than 0.5 mL of plasma and samples from women whose consent did not allow for additional studies outside of the original study aims were also excluded. Only women who were part of the clinical validation study were included, as the DNA fetal fraction was already available. A total of 57 separate pregnancies with Down's syndrome (34 with first trimester and 23 with second trimester samples) satisfied inclusion criteria and each corresponding sample was matched to six euploid samples (controls) based on gestational age (nearest week; same trimester), maternal race (self-declared) and time in freezer storage (within 1 month). Upon review, five euploid samples were found to have maternal plasma AFP levels above 3.0 MoM. Subsequent investigation revealed pregnancies diagnosed with gastroschisis (11.9 MoM), anencephaly (6.7 MoM), urinary system defect (5.5 MoM), and multiple anomalies including cystic hygroma, echogenic bowel, and cardiac defects (3.1 MoM). No further information was available for the fifth case (5.0 MoM). The indication for all five was an abnormal ultrasound and all plasma test results for these five samples were omitted from further analyses.
The control samples were used to generate gestational age-specific reference ranges and maternal weight adjustment equations 21 with results expressed for each marker as weight-adjusted MoMs. Separate probability plots were generated for cases and controls for each analyte after a logarithmic transformation. Linear regression was applied to derive the standard deviations using the linear portion of the probability plot. The median and log mean MoM values in pregnancies with Down's syndrome were derived separately for each marker. The resulting plasma median MoM levels in cases were compared with published parameters based on serum measurements.22,23
Assays
Levels of AFP, uE3, total hCG and InhA in second trimester serum and plasma samples, and PAPP-A in the first trimester, were measured using Beckman Coulter (Chaska, MN, USA) DxI immunoassays following the manufacturer's instructions. Coefficients of variation were <10% across the range of patient values. For a subset of first trimester plasma samples, the low-speed/high-speed comparison for PAPP-A utilized measurements from an earlier, laboratory-developed manual enzyme-linked immunosorbent assay using analyte-specific reagents from Beckman Coulter, Inc.
The proportion of cell-free DNA derived from the fetus/placenta in the maternal plasma samples (fetal fraction) was determined from testing done at the Sequenom Center for Molecular Medicine (San Diego, CA, USA) as part of a clinical validation study.16,17 The methodology relies on differentially methylated markers between the mother and the placenta. 24 Results are expressed as a percentage (fetal/total x 100). To obtain reliable DNA test results, the fetal fraction should be between 4% and 50%. 14
Results
Paired Sample Set: Second Trimester
A matched comparison of Down's syndrome marker measurements in plasma and serum samples
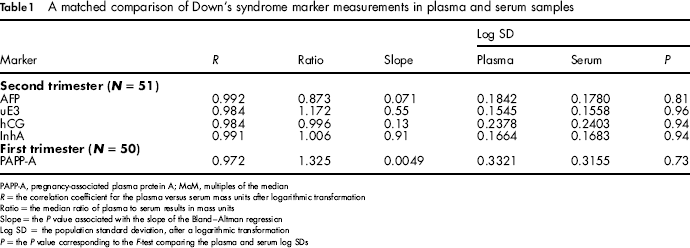
PAPP-A, pregnancy-associated plasma protein A; MoM, multiples of the median
R = the correlation coefficient for the plasma versus serum mass units after logarithmic transformation
Ratio = the median ratio of plasma to serum results in mass units
Slope = the P value associated with the slope of the Bland-Altman regression
Log SD = the population standard deviation, after a logarithmic transformation
P = the P value corresponding to the F-test comparing the plasma and serum log SDs
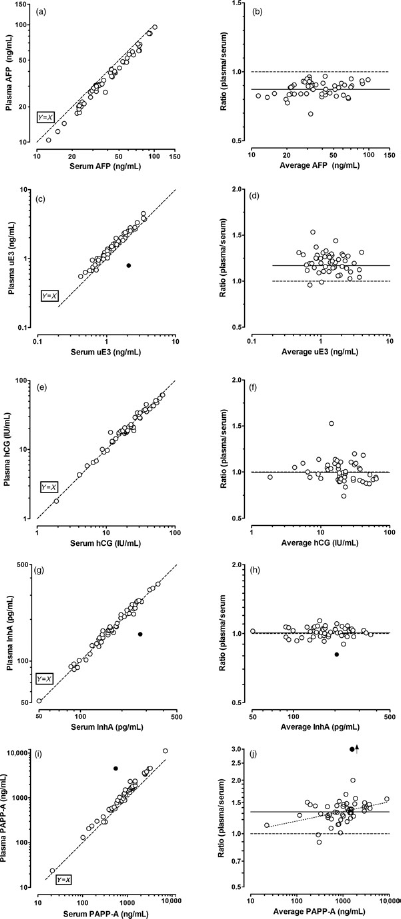
Analysis of measurements in paired plasma and serum samples. A serum and a plasma sample were collected from 51 women in the early second trimester enrolled during routine prenatal screening. Filled circles indicate outlying values not included in the analyses. (a) A comparison of plasma AFP results has high correlation (r = 0.992). (b) The corresponding Bland Altman plot for the same AFP data. The average ratio is 0.73, with no statistically significant change in that ratio over the range studied (P = 0.079). (c and d) The same comparisons, but for uE3 measurements. The correlation is high (r = 0.985) and the average ratio is 1.173, with no significant change over the range studied (P = 0.55). (e and f) The data for hCG measurements. The correlation is high (r = 0.984) and the average ratio is 0.996, with no significant change over the range studied (P = 0.13). (g and h) The data for InhA measurements. The correlation is high (r = 0.991) and the average ratio is 1.006, with no significant change over the range studied (P = 0.91). In the first trimester, 50 women were sampled. (i and j) The data for PAPP-A measurements. The correlation is high (r = 0.992) and the average ratio is 1.325. However, there is significant trend towards higher ratios with higher PAPP-A values (P = 0.005). AFP, alphafetoprotein; uE3, unconjugated estriol; hCG, chorionic gonadotropin; InhA, inhibin-A; PAPP-A, pregnancy-associated plasma protein A
The same series of analyses for AFP measurements were also performed for each of the other second trimester markers (uE3, hCG and InhA) in the same paired samples. The correlation coefficient of results in split samples after low- and high-speed centrifugation was 0.97 or greater for these markers (data not shown). High correlations and at most, minor proportional differences, were found for uE3, hCG and inhibin A. Figure 1c-h and rows 2 through 4 of Table 1 summarize the results for the uE3, hCG and InhA plasma versus serum comparisons. Medians and probability plots for these markers are also available online (Supplementary Figure S1b-d).
Paired Sample Set: First Trimester
Paired plasma and serum samples were collected from 50 women at the time of routine first trimester or integrated screening (median gestational age 11.7 weeks, range 10.0-14.0). The correlation coefficient of PAPP-A results in split samples after low- and high-speed centrifugation was 0.990 (data not shown). Results of the analyses for first trimester PAPP-A measurements in the matched serum and plasma samples are shown in Figures 1i and j, and summarized in the last row of Table 1. PAPP-A medians and probability plots are available online; see Supplementary Figure S2). Plasma and serum results for PAPP-A were highly correlated (r = 0.972). Although PAPP-A values were significantly higher in plasma (+32.5%) than in serum, the population distributions did not differ. First trimester hCG measurements were not subject to analysis, as second trimester hCG measurements had already been examined.
Aliquots of the 101 plasma samples collected prospectively as part of the paired study were tested for fetal fraction. These samples had been transported to the laboratory according to routine protocols for prenatal serum screening; without regard for DNA integrity. The median time from sample collection to freezer storage was 22 hours (range 1–95 hours), during which samples remained at room temperature. The median fetal fraction was 6%, with 30% of the samples having a fetal fraction less than 4%.
Nested Case/Control Sample Set: Second Trimester
Median gestational age was 17.2 weeks, (range 14.2-21.5 weeks). Gestational age-specific median levels were established using the 133 control (euploid) samples for AFP, uE3, hCG and InhA and used to convert both case and control results to MoM. Weight adjustment was performed using an established set of in-house formulas, modified for the observed average weight in the present study group (156 pounds). The resulting weight-adjusted MoM levels were then examined via probability plots (Supplementary Figure S3). The corresponding medians and logarithmic means (and 95% confidence intervals) in Down's syndrome cases for all four second trimester analytes are shown in Table 2 in the first four rows. These are compared with published serum estimates.22,23 All data points were used to calculate the hCG and InhA parameters, but for AFP parameters in cases, data were limited to 5th-95th centile data. For uE3 measurements, control data were limited to between the 5th and 95th centiles and cases were limited to between the 10th and 90th centiles. Truncation limits for AFP, uE3, hCG and InhA were 0.4-3.0 MoM, 0.4-2.0 MoM, 0.4-5.0 MoM, and 0.3-5.0 MoM, respectively, equivalent to those reported in the published literature.22,23
A comparison of five marker levels in Down's syndrome pregnancies in first and second trimester plasma compared with published levels in serum
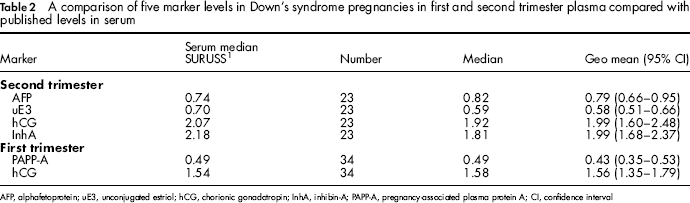

AFP, alphafetoprotein; uE3, unconjugated estriol; hCG, chorionic gonadotropin; InhA, inhibin-A; PAPP-A, pregnancy-associated plasma protein A; CI, confidence interval
Nested Case/Control Sample Set: First Trimester
Median gestational age was 12.4 weeks, (range 8.4-14.4). Median data and weight adjustment formulas were established using the 204 control (euploid) samples for PAPP-A and total hCG (average weight 145 pounds). These measurements were then used to convert results to weight-adjusted MoM levels. These data were also examined via probability plots (Supplementary Figure S3). The median and logarithmic mean levels (and 95% confidence intervals) in Down's syndrome cases for both first trimester analytes are contained in the last two rows of Table 2. These were comparable with published serum estimates.22,23,25
Discussion
Prenatal screening for Down's syndrome, recommended by the American College of Obstetrics and Gynecology for all pregnant women, is commonly achieved using first (PAPP-A and hCG) and/or second trimester (AFP, uE3, hCG and inhibin A) serum markers in combination with first trimester ultrasound measurement of NT. Introduced in the past year, DNA-based tests with markedly superior screening performance require collection of blood as EDTA plasma. Results from the study presented here show that an EDTA plasma sample could be used for measurement of the conventional ‘serum’ markers. Analysis of matched serum and plasma showed a high correlation for each marker. These data suggest that plasma is an acceptable sample type for conventional multiple marker screening, as well as for the newer DNA-based test. However, to use plasma in place of serum for screening, appropriate reference data must be established. Median equations are commonly calculated and updated in prenatal screening laboratories when, for example, a new assay method or reagent lot change occurs. Our comparisons showed that relatively modest adjustments in medians would be needed to accommodate plasma. The largest difference was observed for PAPP-A, where plasma levels were, on average, about 30% higher than in serum. The median MoM levels for first trimester PAPP-A and second trimester AFP and uE3 measurements in plasma were appropriately low in Down's syndrome cases, while second trimester hCG and inhibin A MoM levels were high. Plasma measurements of uE3 were somewhat lower than expected.
Women categorized as high risk for Down's syndrome can now be offered secondary screening by next-generation sequencing. An advantage of sequencing in high-risk women is that the false-positive rate can be reduced from 3% or more using conventional screening tests, to 0.5% or lower with sequencing, while maintaining virtually all detected cases. Thus, many fewer women will be offered diagnostic testing and be faced with the risk of a procedure-related pregnancy loss. The use of plasma for conventional screening may provide simpler testing for patients and laboratories. Plasma may be drawn for conventional screening and an aliquot stored for cell-free fetal DNA extraction and analysis. The samples for women with initial screen positive results could then be sent reflexively for DNA sequencing, and only reported as high risk after that test was complete. The number of couples caused anxiety by a screen positive results would also be reduced. Furthermore, results may be provided earlier in pregnancy, especially if sequencing was performed subsequent to first trimester screening protocols. In the future, DNA testing may be offered as a first-line screening test. In this setting, women experiencing test failures would not be offered immediate diagnostic testing, but would be offered conventional testing instead. Such testing may also utilize existing plasma samples rather than delaying the process to collect a new sample.
One limitation of the proposed use of plasma for screening is that, when drawn into standard EDTA plasma collection tubes, the samples must be handled in an expedited manner to allow for extraction of high-quality free DNA. Serum is currently transported to testing laboratories by use of courier or express mail, with delivery times in the range of 24 hours, as observed at present. It is acceptable for serum samples to remain at ambient temperatures during this time 26 for the measurement of Down's syndrome markers. However, this handling is not acceptable for DNA; 30% of our routinely processed samples yielded insufficient DNA for sequencing. DNA has been shown to deteriorate progressively with time in plasma16,27 leading to clinical implementation of a six-hour limit for processing plasma samples for free DNA isolation. This limitation can be obviated by use of DNA stabilizing plasma tubes (Cell-Free DNA BCT, Streck, Omaha, NE, USA). These tubes also contain EDTA and at least one of the commonly used testing platforms (DELFIA chemistry, PerkinElmer, Waltham, MA, USA) includes a Europium reagent for its time-resolved fluorescence immunoassays, which would be chelated and thus inactivated when EDTA was present. In addition, since PAPP-A is a zinc-metalloprotease, EDTA may sufficiently disrupt the PAPP-A structure to reduce or eliminate proper antibody binding in certain other immunoassays. This appears not to be a problem with the PAPP-A assay used in the present study. Thus, similar PAPP-A validation studies would need to be performed for other assay platforms.
Prenatal screening also typically includes an assessment of risk for open fetal defects in the second trimester. This is commonly performed using either fetal ultrasound or maternal serum AFP measurement. We excluded five samples in our study with high maternal plasma AFP levels. Two of these had a documented open fetal defect (anencephaly or gastroschisis). Therefore, plasma AFP measurement will most likely be acceptable for screening of open fetal defects.
In summary, EDTA maternal plasma is reliable in measuring the markers commonly used in current first and/or second trimester Down's syndrome screening. The use of plasma in conventional multiple marker screening may be convenient option to allow for reflexive DNA testing for fetal aneuploidy.
Footnotes
We are grateful to Jennifer Hubbard (Research Assistant, Division of Research, Women & Infants Hospital) for her assistance with patient recruitment, and to the Molecular Diagnostics Development Department at Sequenom Center for Molecular Medicine, San Diego, CA, for determining the fetal fraction of circulating cell free DNA in the plasma samples from the paired study. We thank Sequenom, Inc. for allowing use of residual plasma samples collected as part of their external validation study, staff at the Enrollment Sites, and the women who agreed to participate.
