Abstract
Background
Cancer patients with a Lewis (a−b−) phenotype have no carbohydrate antigen 19-9 (CA19-9) in their serum. However, we found a small but distinct elevation in the serum CA19-9 level in three cancer patients with the Lewis-negative phenotype. Here, we investigated the reason of such phenomena.
Methods
Six cancer patients with a Lewis-negative phenotype were selected by very low CA19-9 concentrations: three showed a small elevation (Group A) and the other three showed no elevation (Group B) in the serum CA19-9. We investigated the difference by analyzing the Lewis/Secretor genotypes.
Results
All of the six patients with a Le (a−b−) phenotype were genuine Le-negative genotypes: four individuals were homozygous for le1 (le59,508 ), one patient was compound heterozygous for le1 (le59,508 ) and le2 (le59,1067 ) and one patient was compound heterozygous for le1 and le202,314 . As for the Secretor gene, the three patients in Group B were homozygous for Se2 (one patient) or compound heterozygous for Se2 and sej (two patients), while the patients in Group A were all homozygous for sej genotypes.
Conclusions
Even genuinely Le-negative patients, who genetically lack the Le enzyme and theoretically never produce CA19-9, occasionally show a slight increase in serum CA19-9 level when they are homozygous for Se-negative genotypes and suffer from advanced cancer with overproduction of glycans as precursors of CA19-9. Although such cases are not frequent, we should be acquainted with the correlation between serum CA19-9 values and genotypes of Lewis and Secretor genes.
Introduction
Carbohydrate antigen 19-9 (CA19-9), which is equivalent to the sialyl Lewis antigen, is a well-known tumour marker for pancreatic cancer, colorectal cancer, gastric cancer and several types of adenocarcinomas. Healthy individuals with an Lea-positive secretor status may show physiologically elevated CA19-9 values in all types of secretions (e.g. saliva, bronchial/gastric secretions, bile juice, etc.), in contrast to patients with a Lewis (a−b−) status, who have little or no CA19-9 in their serum. 1 Usually, CA19-9 is detected at low concentrations (<37 kU/L) in the serum of healthy individuals.
The Lewis histo-blood group system consists of two major antigens – Lea and Leb – and three common phenotypes – Le (a−b−), Le (a+b−) and Le (a−b+). The Lea antigen is synthesized from a type 1 precursor substrate by the Lewis-encoded α-(1,3/1,4)-fucosyltransferase (Fuc-T III; FUT3), while the Leb antigen is synthesized from a type 1 H substrate. There are two alleles at the Lewis locus: Le, which encodes a functional fucosyltransferase and le, which encodes a non-functional enzyme. The le59,508 and le59,1067 alleles confer very low enzymatic activity as compared with the Le and Le59 alleles. 2 The Lewis phenotype is determined by the combination of the Lewis and Secretor genotypes. The Secretor enzyme is α-(1,2)-fucosyltransferase (Fuc-T II) and encoded by the Secretor (FUT2) gene. An individual homozygous for the le allele has neither the Lea nor the Leb antigen. Secretors have at least two functional Se alleles, while non-secretors are homozygous for non-functional se alleles. In particular, a partially inactive se allele named sej was found in about 40% of the Japanese population. 1,3 Several polymorphisms in the Lewis and Secretor genes have been reported. 1–9 In the Japanese, Lewis-negative and Secretor-negative allelic frequencies are about 30% and 45%, respectively. 1
Recently, we found a small but distinct elevation in serum CA19-9 level in a patient with end-stage colorectal adenocarcinoma. Some textbooks have described that Le-negative patients did not show the elevation of serum CA19-9 values even if they suffered from pancreatic or gastrointestinal cancer. Therefore, we tried to reveal why Le-negative cancer patients showed a tiny increase of serum CA19-9 values. We additionally selected cancer patients with lower serum CA19-9 levels than expected from other tumour markers, such as carcinoembryonic antigen (CEA). These patients expressed erythrocyte Lewis-negative phenotypes. To define the molecular basis for this discrepancy, we investigated these cancer patients by both immunohistochemical and molecular analyses, and identified the relationship between serum CA19-9 levels and the Lewis/Secretor genotypes.
Materials and methods
Samples
Patients were selected by very low CA19-9 concentrations (less than 1 kU/L) in their serum samples compared with other laboratory and clinical properties during the time course of their medical treatment in Hamamatsu University Hospital in January 2009 to May 2010. Blood and cancer tissue were obtained from the enrolled patients. Written informed consent was obtained from each patient. This study was approved by the Institutional Review Board of the Hamamatsu University School of Medicine (18–28).
Measurement of tumour marker concentrations in serum samples
The serum CA19-9 concentrations were measured using an electrochemiluminescense immunoassay (ECLIA) on a Modular Analytics E170 module (Roche Diagnostics Co, Tokyo, Japan), a chemiluminescence enzyme immunoassay on Lumipulse f (Fujirebio Co, Tokyo, Japan) and a radioimmunoassay (TFB, Inc, Tokyo, Japan). All of the three CA19-9 assays used NS19-9 monoclonal antibody as a detection antibody. The serum CEA and DU-PAN-2 concentrations were analysed by an ECLIA (Roche) and an enzyme immunoassay (Kyowa Medex Co, Tokyo, Japan), respectively. The cut-off values in serum for CA19-9, CEA and DU-PAN-2 were defined as 37 kU/L, 5 μg/L and 150 kU/L, respectively.
Lewis blood typing
Lewis hemagglutination assays were performed with blood grouping reagent anti-Lea and anti-Leb antibodies (Ortho-Clinical Diagnostics, Inc, Raritan, NJ, USA) and with Seraclone anti-Lea and anti-Leb antibodies (Biotest Medical Diagnostics GmbH, Dreieich, Germany).
DNA sequencing analysis and genotyping of the Lewis and the Secretor genes
Genomic DNAs were extracted from the blood samples with a QIAamp DNA Blood Maxi kit (QIAGEN, Tokyo, Japan). The coding exons of the Lewis genes were amplified by polymerase chain reaction (PCR) and directly sequenced with ABI BigDye Terminator Ready Reaction Mix (Applied Biosystems, Foster City, CA, USA) and an ABI 3130 Genetic Analyzer (Applied Biosystems). Information regarding the PCR primers is available upon request. The Lewis genotyping was also confirmed for the 59T>G, 508G>A and 1067T>A polymorphic sites, by PCR-restriction fragment length polymorphism as described in the previous report. 1,9 The Secretor gene was also analysed by PCR and direct sequencing.
Immunohistochemical staining
Cancerous colon tissue and accompanying normal mucosa were fixed in buffered 10% formaldehyde, embedded in paraffin and then cut into 4-μm sections. Immunohistochemical staining was performed with a Dako ChemMate EnVision™ kit (Dako, Glostrup, Denmark) and Dako Autostainer (Dako). The sections were dewaxed, incubated with methanol containing 3% H2O2 in methanol at room temperature (RT) for five minutes to block endogenous peroxidase activity, immersed in 0.01 mol/L citrate buffer, pH 6.0, heat-treated at 95°C in a water bath for 40 min and washed three times with distilled water. The sections were then incubated at RT for 30 min with monoclonal antibodies (mAbs) of 116-NS-19-9 against CA19-9 (M3517; Dako; diluted 1:50), SPM522 to Lewis a (ab64099; Abcam, Tokyo, Japan; diluted 1:10) or 2-25LE to Lewis b (ab3968; Abcam; diluted 1:200). A subsequent reaction was carried out using second antibodies conjugated to peroxidase-labelled dextran polymer (EnVision™/HRP, Dako) at 37°C for 30 min. After development with 3,3′-diaminobenzidine tetrahydrochloride at RT for about five minutes, the sections were counterstained with haematoxylin.
Results
Clinical and laboratory data (excerpted) of the patients
RK, rectal cancer; LC, lung cancer; GC, gastric cancer; ECLIA, electrochemiluminescence immunoassay; CLEIA, chemiluminescent enzyme immunoassay
Characteristic laboratory data are excerpted and shown from past to latest
Figure 1 and Table 2 show the representative DNA sequencing results and the summaries of DNA sequencing and genotyping for Lewis and Secretor genes. All of the six patients were homozygous for Le-negative genotypes. Four individuals (patient nos 3, 4, 5 and 6) were homozygous for le1 (le59,508
), and one patient (no. 2) was compound heterozygous for le1 (le59,508
) and le2 (le59,1067
). One patient (no. 1) was compound heterozygous for le1 (le59,508
) and for 202T>C (Trp68Arg) and 314C>T (Thr105Met) (highly possibly le202,314
). These two missense mutations have previously been reported as genetic polymorphisms, rs812936 and rs778986, respectively. One patient (no. 6) was heterozygous for 732C>T (Tyr244Tyr) which is identical to the silent mutation reported as rs1960111 in the NCBI SNP database.
Representative DNA sequencing results of a patient (patient no. 1) for Lewis and Secretor genes. Patient no. 1 was heterozygous for the four missense mutations (L20R, W68R, T105M and G170S) in the Lewis gene and possibly compound heterozygous for le1 (le59,508
) and le202,314
. The patient was also heterozygous for 235G>A (Gly90Arg) in the Secretor gene Nucleotide substitution and genotype in Lewis and Secretor genes *Bold letters indicate substituted nucleotides or genotypes with activity reduction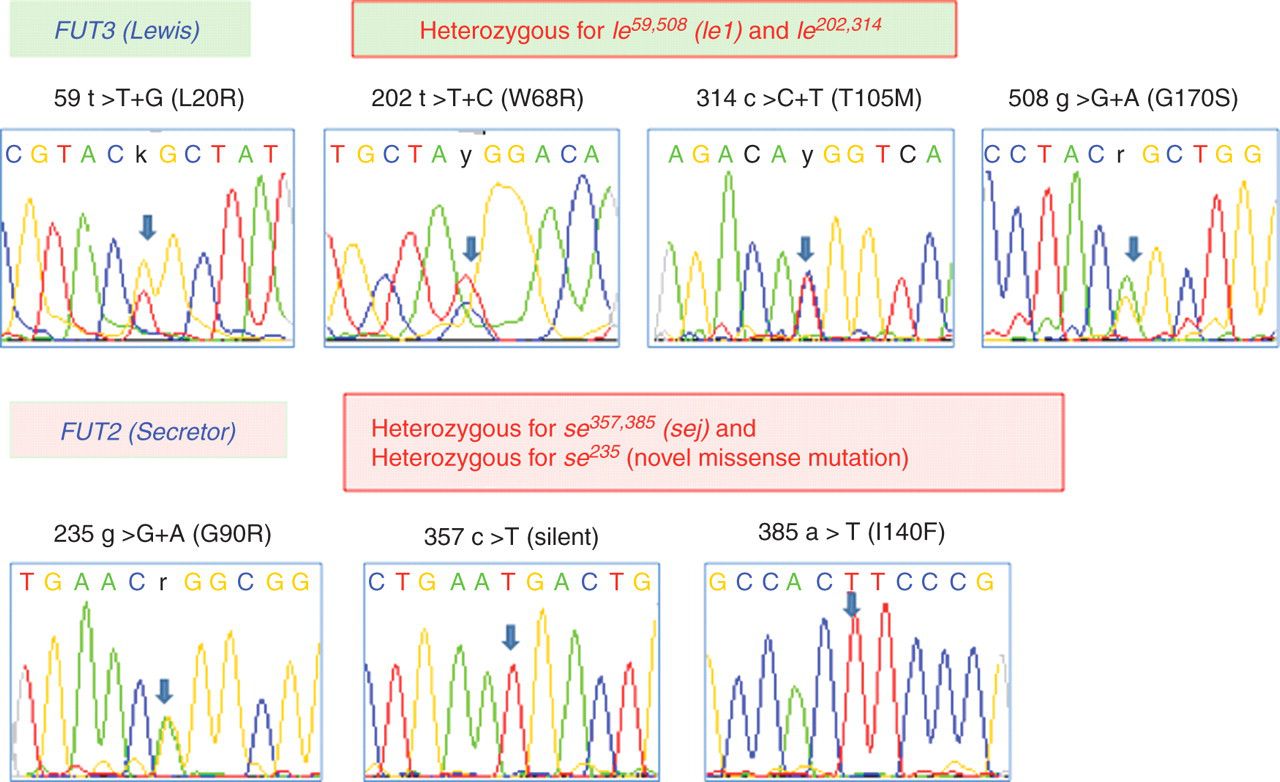
One patient (no. 6) was homozygous for Se2, two patients (nos 4 and 5) were compound heterozygous for Se2 and sej, and three patients (nos 1, 2 and 3) were homozygous for the sej genotype. Patient no. 1 was also heterozygous for 235G>A (Gly90Arg) in the Secretor gene. This missense mutation was not found in the NCBI database.
Immunohistochemical staining with anti-CA19-9, anti-Lea and anti-Leb mAbs was performed on sequential sections of normal and cancerous tissue specimens from the five patients. The representative immunohistochemistry results and the summary are shown in Figure 2 and Table 3. No staining with any of the mAbs was observed in patient nos 1, 4 and 6. Weak positive staining with anti-CA19-9 and anti-Lea was observed in patient nos 2 and 3. The patients in Group B seemed to show relatively weak staining.
Representative haematoxylin–eosin (HE) staining and immunohistochemistry with anti-CA19-9, anti-Lea and anti-Leb monoclonal antibodies (mAbs) (colon, ×20). Sequential sections of normal and cancerous tissue specimens from the patient (patient no. 1) were shown. No staining was observed with any of the mAbs; however, weakly positive staining with anti-CA19-9 was observed in only a small part of a glandular cavity and faintly positive staining was also observed in a quite small portion of a normal tubular gland (shown by arrows) Summary of the immunohistochemical staining NA, not analysed (no specimens taken)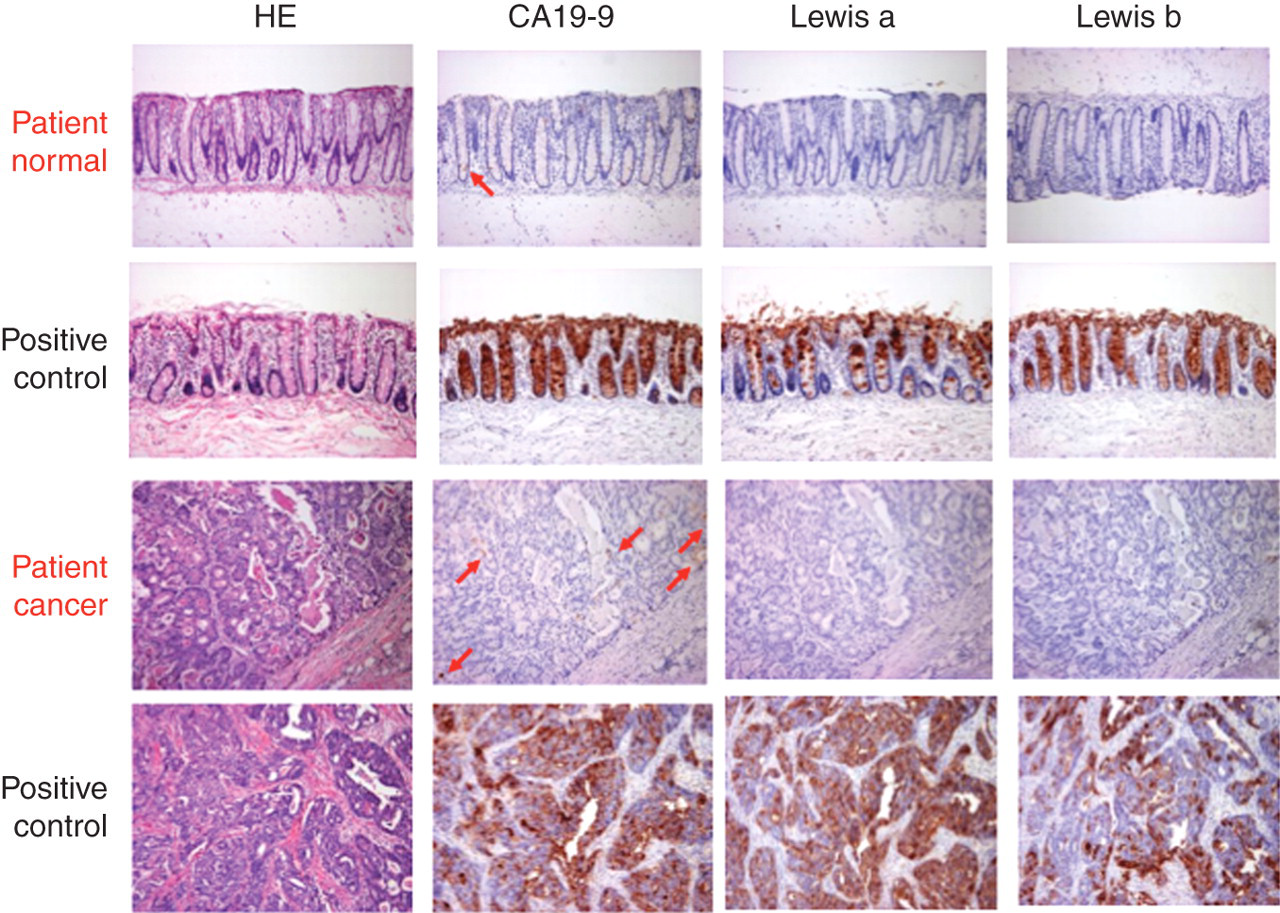
Discussion
In the present study, we noticed that some patients showed an unexpected discrepancy in serum CA19-9 and CEA values and investigated how serum CA19-9 values could be slightly or moderately elevated in patients with Lewis-negative phenotype by using genotyping analysis. According to the Lewis (FUT3) genotypes of the six patients, no increase of CA19-9 in serum should be observed in all of the six patients.
Narimatsu et al. 1 reported the effect of dose levels of the Le and Se genes on the serum levels of CA19-9 and DU-PAN-2 in normal individuals and cancer patients. The participants were separated into nine groups according to their Le and Se genotypes. The Le-negative individuals (homozygous for Le function loss mutation; le/le genotype) showed undetectable CA19-9 values in their serum samples. There have been some reports showing that pancreatic and colorectal cancer patients with the Le blood group-negative phenotype tend to show negative CA19-9 values in serum. 10,11 Despite this tendency, there have been some exceptional patients who showed the Le-negative blood group phenotype but apparently had positive CA19-9 values. 12–14 Such discrepant phenomena were only detected by phenotyping but were not detected by genotyping because the adsorption of Le-active glycolipids to red blood cells is inhibited in such patients. 14 The genuinely Le-negative patients, who genetically lack the Le enzyme, were found to be homozygotes with null Le alleles (le/le). They must never show the expression of the sLea antigen in colorectal cancer tissue 1 or a positive CA19-9 value in serum. 5 Therefore, Narimatsu and colleagues 2,3 have demonstrated that Le genotyping is the most accurate method for distinguishing genuinely Le-negative patients from false-negative patients. Narimatsu et al. 1 have also presented that the Se gene dosage clearly affects the CA19-9 values and that the se/se individuals (non-secretors) apparently showed higher CA19-9 values than did the secretors in Le-positive individuals.
Compared with the previous reports, the findings in our present study seem very interesting. The six patients can be divided into two groups; Group A: patient nos 1, 2 and 3 showed CA19-9 values close to zero in a healthy state, but the values increased slightly with cancer progression; Group B: patient nos 4, 5 and 6 showed nearly zero values of CA19-9 in both a healthy state and during cancer progression. All of the six patients were homozygous for null Le alleles. This means that they are genuinely Le-negative patients, who genetically lack the Le enzyme and, theoretically, never produce CA19-9.
1,5
However, Se genotypes showed a striking difference. The three patients in Group A were homozygous for sej, which is an activity deficiency mutant, while the three patients in Group B had a wild Se allele heterozygously or homozygously. Figure 3 shows a schematic presentation of one possibility to explain how the CA19-9 values occurred in these patients. We interpreted that the large amount of carbohydrate substrate produced by carcinogenesis was transferred to CA19-9 via trace FUT3 activity due to a FUT3 functional mutation/polymorphism. The pathway producing CA19-9 could also be accelerated by the Secretor genotype of patient nos 1, 2 and 3, who were homozygous for sej, which is a mostly inactive se allele. A similar large amount of carbohydrate substrate could cause the eminent increase of the DU-PAN-2 value. On the contrary, the patients in Group B had both the wild-type Se allele and active Se enzyme, and then the pathway might preferentially flow to H-type 1 synthesis, resulting in much less CA19-9 production. The hypothesis seems likely and suitable; however, it is not supported by the direct experimental data. Considering the extremely high CEA concentrations in the serum samples of the patients, tumour products may be present in the circulation, which interfere with the immunoassays. In short, we would like to suggest that the phenotype difference between Groups A and B originated mainly from the residual activity of secretor enzyme and additionally from the degree of CEA elevation possibly corresponding to the tumour burden.
A hypothetical scheme for biosynthetic pathways for Le antigens and type 1 chain in genetically Le-negative individuals. In genetically and genuinely Le-negative individuals, the CA19-9 values in serum samples are nearly zero in a healthy state (H). On the other hand, when the individuals suffer from cancer, large amounts of carbohydrate substrate are produced by carcinogenesis and excessive sLec (DU-PAN-2) is produced (C). The accumulated sLec is transferred to CA19-9 via trace FUT3 activity due to a FUT3 functional mutation/polymorphism, and therefore the CA19-9 level becomes slightly elevated (C). The increase in the level of CA19-9 might be much stronger if the individual is homozygous for se mutant genotype and has low Secretor (FUT2) activity (Group A; patient nos 1, 2 and 3) in contrast with Group B with high FUT2 activity (patient nos 4, 5 and 6) shown as (B)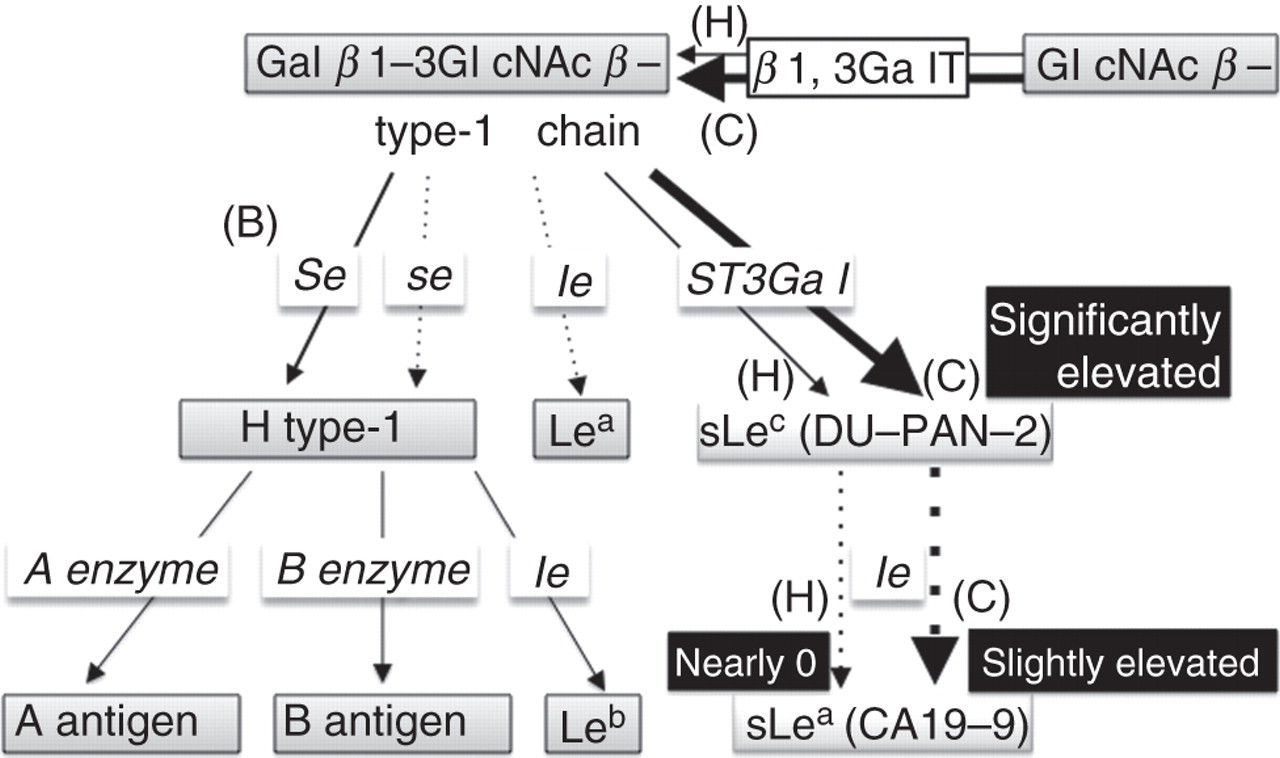
Mollicone et al. 4 reported Indonesian Lewis-negative individuals and found that the le59 (T59G; Leu20Arg) is located in the enzyme's transmembrane domain and that the substrate affinity of the expressed enzyme expressed in COS-7 cells was essentially identical to that of the wild type. However, the le1067 (le2) (T1067A; Ile356Lys) is situated in the enzyme's catalytic domain, and the mutation inactivates the enzyme activity. Nishihara et al. 2 also reported that the T1067A mutation reduced Lewis enzyme activity less than 10%, whereas the G508A mutation made the enzyme completely inactive. Moreover, Nishihara et al. 8 reported that (1) the catalytic domain of the le59 enzyme is in the transmembrane domain and does not affect the folding of the catalytic domain, and (2) Le508 (le1; Gly170Ser) or le1067 (Ile356Lys) is in the catalytic domain and affects the folding of the catalytic domain.
The two missense mutations, le202 (T202C; W68R amino acid substitution) and le314 (C314T; T105M amino acid substitution) that were originally described in a Swedish population, 6 have not been found as single isolated point mutations so far. The functional contribution of each of these two mutations to the Lewis negative phenotype was investigated by Elmgren et al. 7 by transfection of the chimeric FUT3 constructs. There was no decrease in enzyme activity or in immunoreactivity of transfected cells with the C314T mutation. On the other hand, large decreases in enzyme activity and immunoreactivity were observed on transfected cells with the T202C mutation. In addition, neither enzyme activity nor immunoreactivity for Lewis antigens were detected on transfected cells with both of the T202C and C314T mutations. Sialyl Lea epitopes were also decreased in the cells with the T202C mutation, and were much evidently decreased in the cells with both of the 202C and C314T mutations. It is noteworthy that a very weak activity (less than 0.5% of the wild-allele activity) was observed for the double mutations. Orntoft et al. 5 also reported that homozygous individuals with the combination of 202/314 mutations were Lewis-negative on erythrocytes and had no enzyme activity in saliva. Transfection experiments of the mutation combination into COS7 cells showed absence of both Lea structure and FUT3 enzymatic activity. In the report, plasma sialyl Lea (CA19-9) levels in cancer patients with several types of FUT3 mutations were also shown. The plasma CA19-9 levels in the cancer patients homozygous or compound heterozygous for le202,314 were very low. We can therefore say that our patient no. 1, who was combined heterozygous for le59,508 (le1) and le202,314 (homozygous for Le-negative genotypes) and notably homozygous for sej, suffered from advanced cancer and had exceptionally high CEA levels, and we conclude that CA19-9 might be generated from excessive glycan produced by cancer cells and distributed into serum.
Point mutations inactivating the Le gene are ethnic-group-specific. In Japan, the frequencies of the Le alleles, Le, le1, le2 and le3, are 68.9%, 24.8%, 5.8% and 0.5%, respectively. 1 The mutations frequently found in the Scandinavian population, i.e. T202C and C314T, have not been found in the Le (a−b−) individuals in the Japanese population. 1 The Lewis allelic frequencies are summarized in the report by Park et al. 9 The le202,314 mutation was observed in 0.140 of Caucasians, 0.080 of Africans, 0.031 of Chinese and 0 of Japanese. Patient no. 1 in this study might be the first case with the le202,314 found in Japan.
CA19-9 has been used as a biomarker for pancreas cancer, colorectal cancer, gastric cancer and any other adenocarcinoma. Many investigations have shown that CA19-9 is a useful marker for cancer screening and is a prognostic factor during treatment. Recently, preoperative and postresection levels of serum CA19-9 were investigated and the results proved that the prognosis was predictable. 15,16 In such reports, Le-negative phenotype or genotype is considered. Kawai et al. 17 reported on an association of serum CA19-9 levels and smoking. Taken together, serum CA19-9 levels can be influenced by both genetic and acquired factors.
In conclusion, even genuinely Le-negative patients, who genetically lack the Le enzyme and theoretically never produce CA19-9, occasionally show a slight increase in serum CA19-9 level when they are homozygous for Se-negative genotypes and suffer from advanced cancer with a production of excessive glycan as precursors of CA19-9. Although such cases are not frequent, serum CA19-9 values should be interpreted on a one-by-one basis, and we should be acquainted with the correlation between serum CA19-9 values and genotypes of Lewis and Secretor genes.
DECLARATIONS
