Abstract
Background
Circulating nucleic acids were discovered more than 60 y ago. With the recent developments in the study of circulating nucleic acids, its application in the diagnostic field has increased. The objective of this study was to assess the usefulness of the quantification of cell-free plasma DNA (CF-DNA) concentration in the diagnosis of infections in febrile patients and as a prognostic marker in septic patients.
Methods
Concentrations of CF-DNA, procalcitonin (PCT) and C-reactive protein (CRP) were measured in 110 febrile patients who were clinically diagnosed with fever of unknown origin, localized infection, sepsis or septic shock.
Results
Concentrations of CF-DNA increase according to the severity of the infection. The best cut-off point for predicting infection was 2800 GE (genome equivalents)/mL (sensitivity: 95.0%; specificity: 96.7%) and 14,000 GE/mL for sepsis prediction (sensitivity: 77.8%; specificity: 94.6%). Higher concentrations of CF-DNA were found in exitus septic patients than in survivors. The diagnostic efficiency of CF-DNA was similar to PCT and higher than CRP in infectious processes.
Conclusions
Normal concentrations of CF-DNA can exclude the presence of an infection in febrile patients, and very high concentrations (>10-fold over the normal reference range) stratify the severity of infections, showing a high prognostic value to predict mortality in the absence of other causes for elevated CF-DNA.
Introduction
Fever is one of the most non-specific signs of illness and is considered to be a protective response against a large number of infectious and non-infectious diseases. It is one of the most frequent consultation causes not only at emergency departments (ED), 1,2 but also in hospitalized patients, mainly at intensive care units (ICU). 3,4
The onset of fever in critically ill patients is a source of concern and priority, because a non-on-time diagnosed infection can rapidly progress into sepsis, a common cause of mortality and morbidity. Although an early diagnosis of sepsis is very important for effective therapies to be used, 5 such diagnosis can be difficult because clinical signs and the behaviour of laboratory markers are often similar to those associated with different severities of systemic inflammatory response syndrome, both from infectious and non-infectious ethiology. Traditional markers, such as body temperature, heart rate, respiratory rate or white blood cell count, are non-specific. Microbiological culture is the reference method, but it is not considered a fast marker for diagnosis and, moreover, false-negative results can be obtained in patients receiving antibiotherapy. Biochemical markers, such as interleukins, C-reactive protein (CRP) and procalcitonin (PCT), have improved considerably the correct classification of infectious processes, 6,7 but an intensive research is still being carried out on new markers. 8–11
A worse prognosis of sepsis is expected if multiorganic dysfunction syndrome (MODS) occurs. Several mechanisms have been proposed to explain the cell and tissue damage associated with MODS, such as a high liberation of proinflammatory mediators, as well as a noticeable increase in cell apoptosis. 12–15 Cell-free plasma DNA (CF-DNA) is comprised of acellular DNA fragments circulating in peripheral blood, with cell apoptosis being proposed as a potential source of this type of DNA. 16,17 Recently, high concentrations of CF-DNA have been reported in a large number of clinical conditions such as cancer, trauma, burns, myocardial infarction, stroke, etc. 18–23 Elevated levels of nucleosomes 24 and CF-DNA have also been detected in the blood of septic 25,26 and critically ill patients. 27,28
The aim of this study was to ascertain the performance of a new marker of cellular apoptosis and tissue damage, CF-DNA, in the diagnosis of infectious diseases compared with other classical markers of infection such as PCT and CRP, as well as to assess the potential prognostic value of CF-DNA for predicting sepsis and mortality.
Methods
Patients
A prospective collection was made of blood samples from 110 febrile syndrome patients, aged 44 ± 20 y (mean ± standard deviation), with a mean evolution time of 19 h (range: 2–120 h). Thirty-six per cent of the patients (n = 40) were attended at the ED of the Hospital Universitario Central de Asturias, and the other 70 were in-patients, either at ward or ICU. All the patients were sequentially recruited whenever they were diagnosed for acute febrile syndrome. Sixty per cent of the patients were men. Clinical data such as symptoms at admission, associated complications, SOFA score (Sepsis-related Organ Failure Assessment) 29 and mortality were obtained from the medical records of the patients.
Eighty out of the 110 febrile patients were diagnosed with infection, sepsis or septic shock, within 72 h after drawing the blood sample, according to the criteria of the international Consensus Conference of the American College of Chest Physicians and The Society of Critical Care Medicine.
30
Four groups of patients were established:
Group 1: Fever without infection or fever of unknown origin (FUO) (n = 30); Group 2: Local infection confirmed by microbiological culture (n = 26); Group 3: Sepsis or severe sepsis (n = 28); Group 4: Septic shock (n = 26).
None of the ED patients developed sepsis, 17 were diagnosed with FUO and 23 with a localized infection. Therefore, all the 54 sepsis patients were in-patients.
Collection and preanalytical processing of the samples
Blood samples were drawn in EDTA-K3 tubes and processed within two hours after venipuncture. A first centrifugation was carried out at 1800
Quantification of cell-free plasma DNA by realtime PCR
CF-DNA was quantified by realtime quantitative polymerase chain reaction (qPCR) amplification of the β-globin gene in an ABI Prism 7000 Sequence Detection System thermocycler (Applied Biosystem, PE, Foster City, CA, USA). The following sequences were used as primers: 354-F (5′ GTGCACCTGACTCCTGAGGAGA 3′) and 455-R (5′ CCTTGATACCAACCTGCCCAG 3′). Target sequences were amplified in a 10-μL reaction volume containing 3 μL of DNA solution, 5 μL of AbsoluteTM QPCR SYBR® Green ROX Mix 2X (AB Gene, ABGene House, Epsom, UK) and 300 nmol/L of each oligonucleotide primer. The thermal profile was carried out as follows: a first denaturation step at 95°C for 15 min, followed by 45 cycles at 95°C for 15 s and 62°C for one minute. Human genomic DNA was extracted and quantified by absorbance measurement at 260 nm, and then serial diluted samples of this solution were used for preparing a six-point calibration curve (10,000, 5000, 1000, 100, 10 and 5 pg/μL). Both calibrators and samples were analysed in triplicate, and a blank reaction was included in every run.
Several negative controls were included in every analysis. A conversion factor of 6.6 pg of DNA per cell was used to express the results as genome equivalents (GE), defined as the quantity of a particular DNA sequence in one diploid cell. 31 DNA concentration was expressed as GE/mL of plasma.
The observed CF-DNA results were compared with those recently described by our group, 32 with the same procedure, in a normal reference population (group 0) and which ranged from 120 to 1800 GE/mL (median: 620 GE/mL;interquartile range: 390–1100 GE/mL).
PCT measurement
Plasma PCT was measured by Time-Resolved Amplified Cryptate Emission technology, on a Kryptor® analyser (BRAHMS, Hennigsdorf, Germany), with a functional sensitivity of 0.06 ng/mL. Expected PCT values in a non-infectious process are lower than 0.5 ng/mL.
CRP measurement
CRP was measured by an immunoturbidimetric assay on a Modular® P (Roche Diagnostics, GmbH, Mannheim, Germany) chemistry analyser. The reference limit for this method is less than 0.5 mg/dL.
Statistical analysis
The analysis of the results was performed with the statistical application MedCalc®9.2 for Microsoft Windows. Since non-Gaussian distributions were observed, non-parametric tests were used as necessary (Mann-Whitney U test, Kruskal–Wallis test, Spearman rank correlation and point-bi-serial correlation). A P value <0.05 was considered statistically significant (before applying Bonferroni correction, if needed).
Results
As shown in Figure 1, concentrations of CF-DNA seem to increase according to the severity of the infection. Statistically significant differences were observed among CF-DNA levels when all five groups were compared (Kruskal–Wallis test; P < 0.05). Non-parametric post hoc comparisons, Mann-Whitney U test corrected by Bonferroni, revealed that CF-DNA concentrations were not significantly different between healthy donors (group 0) and patients with non-infectious causes of fever (group 1). Neither were there any significant differences between groups 3 and 4 (sepsis and shock septic patients, respectively). However, significantly higher values of CF-DNA were observed in groups 3 and 4 compared with the rest of the groups, as well as group 2 compared with groups 0 and 1, respectively (Mann-Whitney U test, corrected by Bonferroni; P < 0.01). Concentrations of CF-DNA moderately correlated with PCT and CRP levels (r = 0.720 and 0.585, respectively; P < 0.0001).
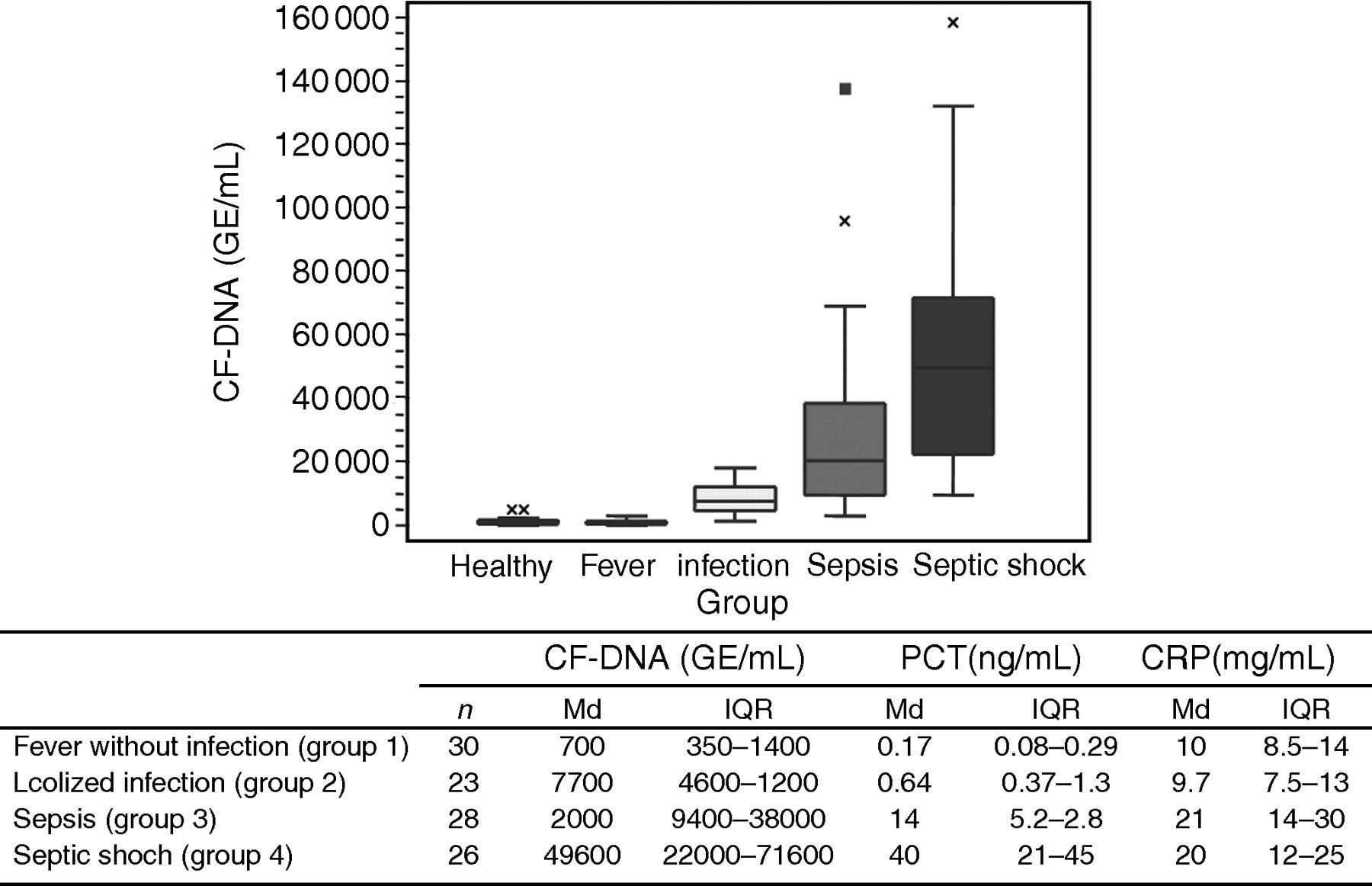
Concentrations of CF-DNA (GE/mL), PCT (ng/mL) and CRP (mg/dL) in healthy individuals, and in patients with fever, infection, sepsis and septic shock. Md: median; IQR: inter-quartile range
With regard to PCT and CRP, differences were also found among all the groups of febrile patients (Kruskal–Wallis test; P < 0.05) (Figure 1). PCT concentrations were significantly higher in group 3 than in group 2, as well as in group 2 compared with group 1, median values of PCT being 14, 0.64 and 0.17 ng/mL, respectively (Mann-Whitney U test, corrected by Bonferroni; P < 0.005). No significant differences were found in PCT concentrations between groups 3 and 4. CRP did not show significant differences between groups 1 and 2, median values being 10 and 9.7 mg/dL, respectively, as well as in groups 3 and 4 (median: 21 and 20 mg/dL, respectively). Nevertheless, significantly higher CRP values did appear when comparing septic patients (groups 3 and 4) with non-septic febrile patients (groups 1 and 2) (Mann-Whitney U test, corrected by Bonferroni; P < 0.005).
Receiver operation characteristics curves were constructed to compare the diagnostic efficiency of the three biochemical markers to predict infection in febrile patients. The best cut-off points for predicting infection were 2800 GE/mL for CF-DNA, 0.47 ng/mL for PCT and 11 mg/dL for CRP, based on the areas under the curve (AUC) of 0.99, 0.95 and 0.69, respectively. The sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV) and likelihood ratio for positive (+LR) and negative (−LR) results are shown in Figure 2.
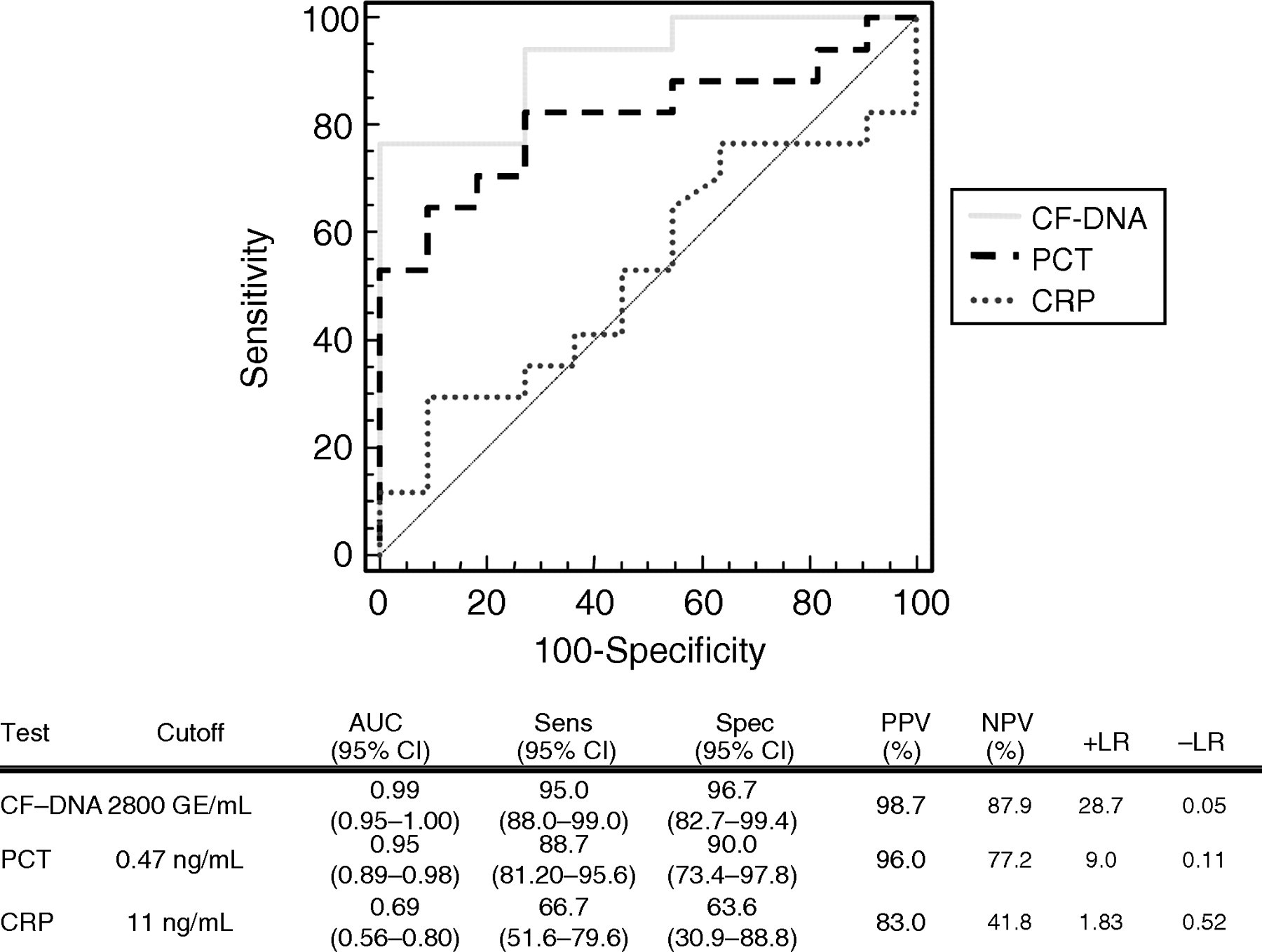
CF-DNA, PCT and CRP diagnostic efficiency to predict infection in febrile patients. Sens: sensitivity, Spec: specificity, PPV: positive predictive value, NPV: negative predictive value, +LR: Positive likelihood ratio, and −LR: Negative likelihood ratio
CF-DNA, PCT and CRP prediction of sepsis in febrile patients
Fifty-four out of the 110 patients developed sepsis. A PCT concentration of 1.8 ng/mL showed a sensitivity of 100%, specificity of 87.5% and the highest diagnostic efficiency of sepsis, showing an AUC of 0.99. The best cut-off point of CF-DNA to predict sepsis was 14,000 GE/mL. Using this CF-DNA cut-off point, 86% of febrile patients were correctly classified as either septic or non-septic patients, whereas PCT and CRP correctly diagnosed 94% and 82% of the patients, respectively (Figure 3). The highest +LR corresponded to CF-DNA, with a value of 14. However, because of the high number of patients who stayed in ICU, the prevalence of sepsis in the studied febrile population was high. In this cohort of patients, it was observed that showing a plasma CF-DNA concentration higher than 14,000 GE/mL in a febrile patient meant a 93% prognostic value of developing sepsis.
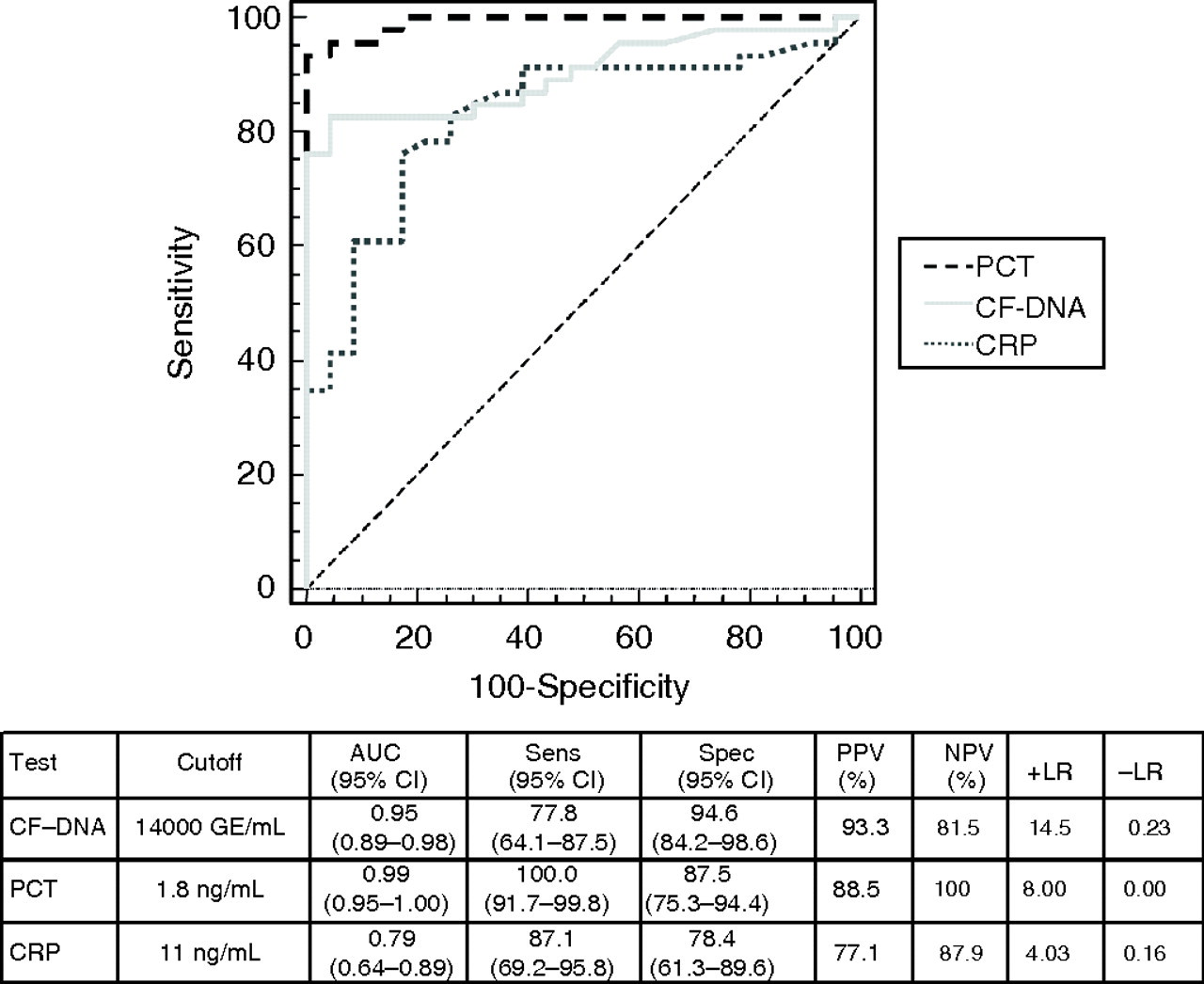
ROC curves comparing the discriminant power of CF-DNA, PCT and CRP to diagnose sepsis in febrile patients (P < 0.0001)
Prognostic value of CF-DNA in septic patients
Sixteen patients died during the period of study, which means a global mortality rate of 14%. Considering only the 54 patients with sepsis or septic shock, the observed mortality rate rises to 30%. CF-DNA concentration slightly correlates to the following variables: mortality rate (r = 0.427; P < 0.005), SOFA score calculated on admission (r = 0.301; P < 0.005), length of stay at ICU (r = 0.401; P < 0.005), need of mechanical ventilation (r = 0.294; P < 0.05) and requirement for further surgery (r = 0.335; P < 0.05). No significant correlation was observed between CF-DNA concentrations and the length of hospitalization and the ionotropic support.
No significant differences were observed between survivors and septic patients who died in regard to PCT, CRP, age, sex and treatment. However, significantly higher concentrations of CF-DNA were found in exitus patients than in survivors, with median values of 58,000 GE/mL (n = 16) and 35,000 GE/mL (n = 38), respectively (Mann-Whitney U test, P < 0.001). CF-DNA concentrations higher than 36,000 GE/mL predicted mortality in septic patients with an efficiency of 77%. The sensitivity, specificity, PPV, NPV and likelihood ratios are shown in Figure 4. According to these results, a CF-DNA concentration higher than 36,000 GE/mL in a septic patient increases the probability of death from 30% to 52%, while concentrations below this cut-off point decrease this probability by 14%.
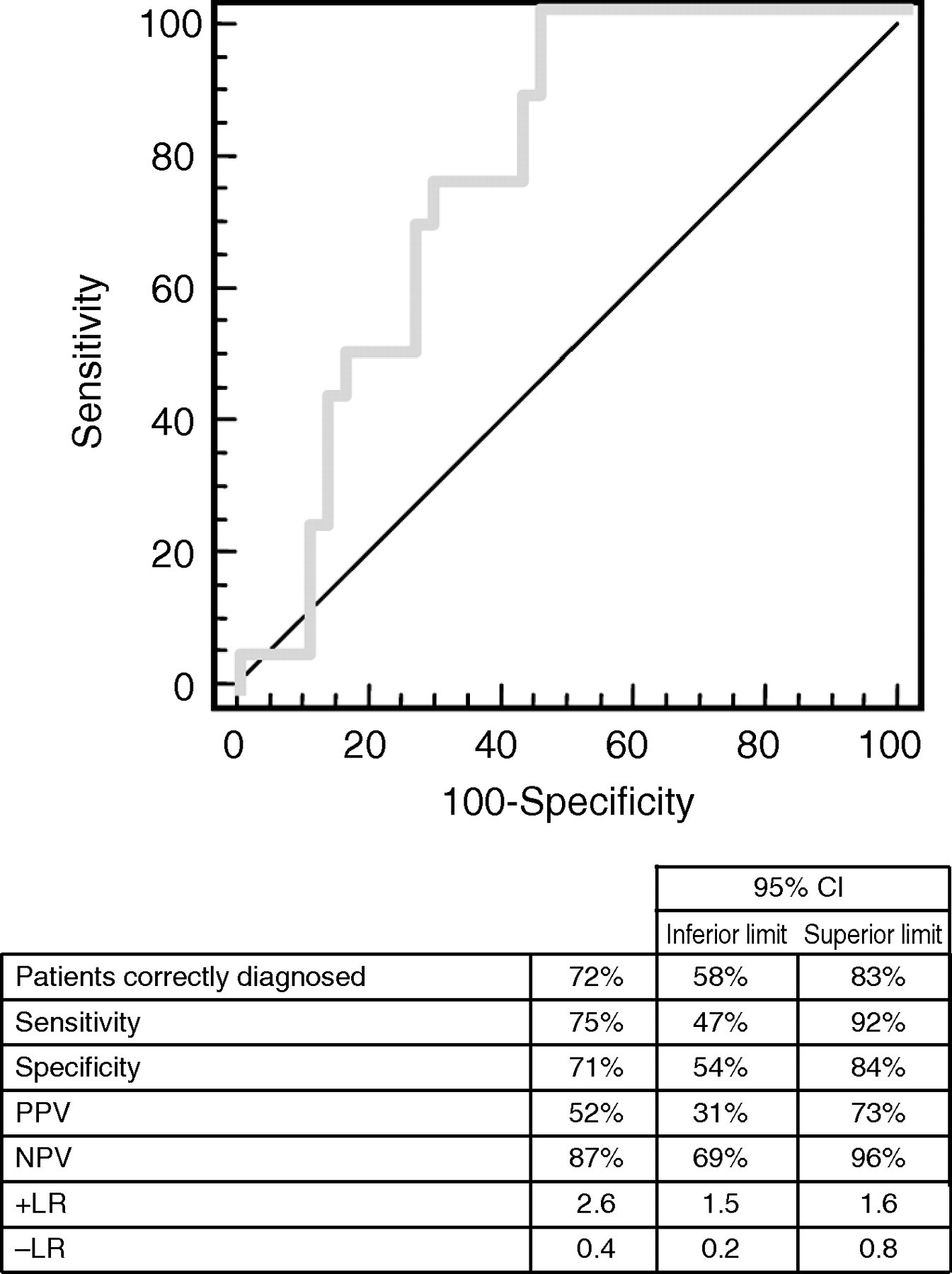
ROC curve of CF-DNA to predict death in sepsis patients
Discussion
The results obtained in this study show that there is an increase in CF-DNA concentration proportional to the severity of the infection (higher increases in sepsis than in localized infections, and in septic shock than in sepsis). Moreover, the highest concentrations of plasma DNA were observed in septic patients who died, suggesting a prognostic value for this marker. Cell apoptosis is one of the potential sources of CF-DNA. Initial observations indicated that free DNA recovered from plasma consisted of DNA fragments ranging from <0.5 to 21 kb. Subsequently, it was described that most of DNA circulates as mononucleosomes, protecting DNA from degradation; hence, internucleosomic degradation of chromatin, characteristic of apoptosis, was suggested as a potential source of CF-DNA. 17 In fact, a correlation between levels of CF-DNA and plasma levels of typical cellular apoptosis markers in lung cancer patients has been described. 33 Therefore, CF-DNA can be considered as a direct marker of cell apoptosis. In addition, in 2004 Brinkmann et al. 34 found that when neutrophils are activated by bacterial endotoxins, proinflammatory cytokines or pharmacological agents, they released molecular complexes formed in part by DNA. 35 Several studies have shown the usefulness of cell apoptosis quantification to establish the prognosis of septic patients. Le Tulzo et al. 36 described a marked increase in the apoptosis grade of circulating lymphocytes in patients with septic shock compared with critically ill patients without sepsis and to healthy volunteers. They concluded that apoptosis of lymphocytes increases rapidly in patients with septic shock, leading to a deep and persistent lymphopaenia, associated to poor outcome. Similar results were found by Hotchkiss et al., 37 comparing septic patients with critically ill patients without sepsis, as they observed a correlation between lymphocyte apoptosis grade and sepsis severity.
The diagnosis of infection is not always easy as a localized infection involves a moderate response of the host and non-specific clinical manifestations are very often present, especially in elderly people, children or immunodeficient patients. Nowadays, the most useful biochemical markers for bacterial and fungi infections are PCT and CRP. According to our findings, CF-DNA shows an efficiency similar to PCT and better than CRP for diagnosis of infection. The slightly increased sensitivity found in our study may be due to the fact that patients with localized bacterial and viral infections, and therefore without raised PCT concentrations, were included. However, those same patients did show an increase of CF-DNA concentrations. The high specificity of CF-DNA observed in infectious processes is somewhat odd because of the wide variety of diseases or conditions that may elevate plasma concentrations of this marker. The most acceptable reason is that only 10 out of the 110 patients studied had some concomitant disease that could increase the CF-DNA concentrations (4 postsurgical patients, 3 polytraumatic people, 3 oncological patients and 1 patient with an autoimmune disease), and all of them developed an infection. Therefore, it might be interesting to expand this part of the study including more patients with this type of pathology to verify such a high specificity. Our results agree with those previously reported by Castelli et al., 7 describing PCT as the best biochemical marker for predicting sepsis.
Besides its prognostic efficiency for hospitalized patients at risk of sepsis, this is the first description of the usefulness of CF-DNA in the diagnosis of infection and sepsis in febrile patients. Our data also agree with previous studies carried out in critically ill patients in ICU, finding higher CF-DNA concentrations in non-surviving than in surviving ICU patients. 26,28 We have also found higher circulating plasma DNA concentrations in exitus than in surviving septic patients, both groups showing significantly higher CF-DNA concentrations than those found in critically ill patients. Therefore, the degree of cell damage seems to be higher in septic patients, with several affected organs or tissues, than in critically ill patients with diseases affecting a single organ. Saukonnen et al. 25 studied 255 patients with severe sepsis and septic shock, finding a median CF-DNA value of 8000 GE/mL. These authors also found higher concentrations of CF-DNA in exitus than in surviving septic patients, describing a sensitivity and specificity for ICU death prediction of 67%, with a cut-off value of 12,000 GE/mL. However, in our study, the median CF-DNA concentration observed in septic patients (20,000 GE/mL) was 10-fold the upper limit of the reference range described by our method in healthy population, 32 but quite different from their data. Moreover, we found considerably higher concentrations of CF-DNA in septic exitus patients than the same authors (58,000 versus 16,000 GE/mL). In fact, we observed similar concentrations in localized infections to those they described for septic patients. Unfortunately, since Saukonnen et al. 25 did not include a control group of healthy individuals in their study, it is difficult to know whether the observed differences could be explained by differences in plasma DNA extraction and quantification protocols or by other factors present in the selected patients. For instance, the different blood drawing times could partially explain these differences. In our study, samples were obtained at diagnosis time, when the fever was rising, whereas Saukonnen's study used blood samples taken at admission. Perhaps a later stage of infection in our study could at least partially explain our data. However, since the diagnostic efficacy, correct classification rate and positive likelihood ratio are similar, methodological differences could be not the only but probably the main cause of such discrepancy in the values.
Finally, we have seen a moderate association between CF-DNA concentrations, PCT and CRP, and SOFA score. Previous studies have also shown a correlation between CF-DNA and CRP, 28 SOFA score, 28 interleukin 6, 24 interleukin 8, 24 elastase, α 1-antitrypsin 24 as well as plasminogen activator inhibitor type 1. 24 However, other groups described no correlation between all these markers. 26
In conclusion, infectious processes, particularly sepsis, markedly increase CF-DNA concentrations, reflecting the cell apoptosis status and DNA secretion by polymorphonuclear cells activated during this process. This is probably a protective response, in order to reduce any excessive immune response to the infection. Apoptotic induction increases with infectious severity. Therefore, normal concentrations of CF-DNA could exclude the presence of an infection in febrile patients. A moderate elevation of CF-DNA concentrations might support a diagnosis of infection in febrile patients, if other causes for increased levels of DNA are not present. In these cases, high concentrations of CF-DNA can stratify the severity of infections, from localized infections to septic shock, with prognostic value of exitus. At present, the CF-DNA clearance mechanism is not precisely known, but a very short half-life of less than 30 min has been described. 38,39 This fast metabolization can be interesting in terms of diagnosis, correlating the findings to the current pathology. With the protocol described, CF-DNA can be quantified within 5–6 h after blood extraction. In fact, superfast qPCR can reduce the analysis time by half, and this could potentially make it useful as a new routine biochemical marker.
DECLARATIONS
