Abstract
Abstract
Background
Lamivudine (LAM)-resistant hepatitis B virus (HBV) with mutations in the polymerase region frequently appears after long-term use of LAM. Several methods allowing detection of mutant strains (YIDD, YVDD) have been reported, but they have no quantitative characteristics. In this study, we explored a unique approach for quantification of each mutant strain.
Methods
A method for detection and quantification of wild and mutant strains was developed using realtime polymerase chain reaction and type-specific minor groove binder (MGB) probes, and tested in patients with chronic hepatitis B before and after additive treatment with adefovir dipivoxil (ADV).
Results
A good correlation was confirmed in HBV DNA quantity obtained between the YMDD-specific MBG probe assay and Amplicor HBV Monitor assay results (P < 0.001), linear between 3 and 9 log copies/mL serum. Of 109 samples from patients with chronic hepatitis B tested by both these assays and conventional direct sequencing, 90 (88.2%) showed identical results. The assays successfully detected and quantified a single type of mutant in three of four patients with additive ADV treatment, and also two coexisting mutant types (YIDD and YVDD) in the remaining patient.
Conclusions
Our specific and sensitive method for detection and quantification of HBV DNA with the wild-type YMDD motif and its two mutant forms (YIDD and YVDD) appears to be clinically useful, especially in patients with multiple mutant HBV infections.
Introduction
Lamivudine (LAM) was initially synthesized as an anti-human immunodeficiency virus (HIV) drug and was later found to inhibit hepatitis B virus (HBV) proliferation as well as to decrease copy number of HBV DNA and to improve liver function in hepatitis patients. However, in many cases HBV gains tolerance to LAM after long-term use, 1–3 requiring the use of interferon in addition to conventional hepato-protective drugs. 4 Recently, another novel nucleoside analogue, adefovir dipivoxil (ADV) was reported to have strong antiviral effects on LAM-resistant mutants. 5–9 The active centre of reverse transcriptase of LAM-resistant HBV has a lower affinity for LAM than that of wild-type HBV, 10 the most common mutations in the YMDD motifs in association with tolerance to LAM being YIDD (rtM204I) and YVDD (rtM204V) affecting the tyrosine-methionine-aspartate-aspartate (YMDD) motif (amino acid 203–amino acid 206) in domain C of the polymerase region. 10–12 HBV infection can occur with not only one, but also a mixture of viruses; but the detection and quantification of an individual HBV type is difficult, especially when the viral load differs greatly among the infected virus types. 13,14 In this study, we have developed a unique approach to measuring mutant HBV by realtime polymerase chain reaction (PCR) using type-specific minor groove binder (MGB) probes, and applied the method to patients with chronic hepatitis B receiving additive treatment with ADV.
Methods
Patients
Four patients with chronic hepatitis B who had been treated with LAM and were later found to have breakthrough hepatitis associated with the appearance of mutated HBV were enrolled in this study. The patients initially received 100 mg of LAM every day, and then were additionally administered 10 mg of ADV once daily. At the beginning of the combination therapy of LAM and ADV, the YMDD motif mutation was found in all four cases; two also featured YIDD and the other two YVDD. The average duration of the combination therapy was 149 days (112–189 days), and the average HBV load at the beginning of the combination therapy was 7.44 log copies/mL (6.74–8.73 log copies/mL). Informed consent for participation in this study was obtained from all patients.
Extraction of hepatitis B virus DNA
Viral DNA was extracted from 200 μL of serum using a QIAamp DNA Blood Mini kit (QIAGEN GmbH, Hilden, Germany) according to the manufacturer's instructions.
Quantification of hepatitis B virus DNA
To measure the quantity of HBV DNA, AMPLICORTM HBV MONITOR (Roche Diagnostics, Tokyo, Japan) was used. HBV DNA in the samples that exhibited high viral load beyond the upper limit of detection (7.6 log copies/mL) were re-examined by the 100-fold dilution method, which was validated previously. 15 The overall range of our measurements spanned between 2.6 and 9.6 log copies/mL.
Type-specific minor groove binder probe assay
To measure the quantity of the wild-type strain (YMDD) and the two mutant types (YIDD and YVDD), we developed a type-specific TaqMan MGB probe assay. For this purpose, three reaction mixtures, each containing one type-specific probe, were prepared and the quantity of HBV DNA was measured by a realtime PCR method using an external standard. One mixture contained 1 × Platinum Quantitative PCR SuperMix-UDG (Invitrogen, Carlsbad, CA, USA) (0.75 U of Platinum Taq polymerase, 20 mmol/L of Tris–HCl [pH 8.4], 50 mmol/L of KCl, 200 μmol/L of each dNTP), 0.4 μmol/L of a sense primer (HBV-MGB2-S: 5′-TCCTATGGGAGTGGGCCTC-3′), 0.4 μmol/L of an anti-sense primer (HBV-MGB-AS: 5′-GTACAGACTTGGCCCCCAAWAC-3′), 0.4 μmol/L of a type-specific TaqMan MGB probe (YMDD: 5′-FAM CTTTCAGTTATATGGATGAT MGB-3′, YIDD: 5′-FAM CTTTCAGTTATATTGATGAT MGB-3′, YVDD: 5′-FAM TTTCAGTTATGTGGATGAT MGB-3′) and 4.0 mmol/L MgCl2. The total volume was adjusted to 25 μL by adding 5 μL of a DNA sample (or external standard) and DNase- and RNase-free water. Realtime PCR was performed using a 7500 RealTime PCR System (Applied Biosystems, Foster City, CA, USA). The PCR conditions were as follows: 50°C for 2 min and 95°C for 2 min for the initial period and then 95°C for 25 s and 62°C for 60 s for 45 cycles. Regarding analytical conditions for the ‘baseline’, three cycles for ‘start’ and 15 cycles for ‘stop’ were used. The threshold concentration was designated as 0.005 for both YMDD and YIDD and 0.02 for YVDD. HBV DNA concentrations were presented as log copy numbers in 1 mL of serum (log copies/mL).
External standard
In order to construct three external standards for the three type-specific reactions, PCR products from each of the three strains were obtained using the primer sets of HBV-MGB-2 and HBV-MGB-AS, and then inserted into the pGEM T Easy Vector (Promega, Madison, WI, USA). Each plasmid was linearized by SacI digestion and copy numbers were assessed by absorption spectrophotometry and adjusted to 2 × 105, 2 × 103, 2 × 101 and 2 × 100 copies/mL. Five microlitres of each concentration was added to the reaction mixture as described above.
Amplification of the hepatitis B virus DNA polymerase region by polymerase chain reaction
The YMDD motif was analysed by direct sequencing after amplification of the HBV DNA polymerase region, performed in 25 μL of a reaction mixture containing 1 × PCR Master Mix (Promega) (50 U/mL of Taq DNA polymerase in a supplied reaction buffer [pH 8.5], 400 μmol/L of each dNTP, 3 mmol/L MgCl2), 0.4 μmol/L of a sense primer (HBV-1-S: 5′-TGGCTATCGCTGGATGTGTCT-3′), 0.4 μmol/L of an anti-sense primer (HBV-3-AS: 5′-CGGGCAACGGGGTAAAGG-3′) and 5 μL of sample DNA. The reaction was performed with a GeneAmp™ 9700 (Applied Biosystems) at 94°C for 2 min for the initial incubation, 94°C for 30 s for denaturing, 57°C for 30 s for annealing, 72°C for 30 s for extension for 40 cycles and a final extension step of 72°C for 7 min. The PCR product of 794 bp was confirmed on a 2% agarose gel.
Amplification of the hepatitis B virus DNA polymerase region by nested polymerase chain reaction
When any PCR product was not observed by the above method, nested PCR was performed using 1 μL of the first PCR product with a sense primer (HBV-5-S: 5′-CCCTCWGTTGCTGTACAAAACCT-3′) and an anti-sense primer (HBV-1-AS: 5′-AAAGGCATTAAGGCAGGATA-3′) under the same conditions as for the first round of PCR. The nested PCR product of 505 bp was confirmed by agarose gel electrophoresis.
Sequence analysis of the YMDD motif
PCR products were purified using a QIAquick PCR purification kit (QIAGEN GmbH) before sequence analysis using a BigDye Terminator cycle sequence kit version 3.1 (Applied Biosystems) with HBV-5-S as the sequence primer. After the reaction, the sample was treated with a DyeEX 2.0 Spin kit (QIAGEN GmbH), and was electrophoresed in an ABI PRISM genetic analyzer 310 (Applied Biosystems). The amino acid sequence was determined with DNA sequencing analysis software version 3.0 (Applied Biosystems). When mixed bases were read on the same position, the nucleotide sequences were determined according to all visible peaks and translated into amino acids.
Results
Linearity
The linearity of HBV DNA quantification by the present method was confirmed by measuring serial dilution of plasmids carrying PCR products of YMDD, YIDD and YVDD strains. The dilution curve was linear between 1.0 and 7.0 log copies for each strain (Figure 1).
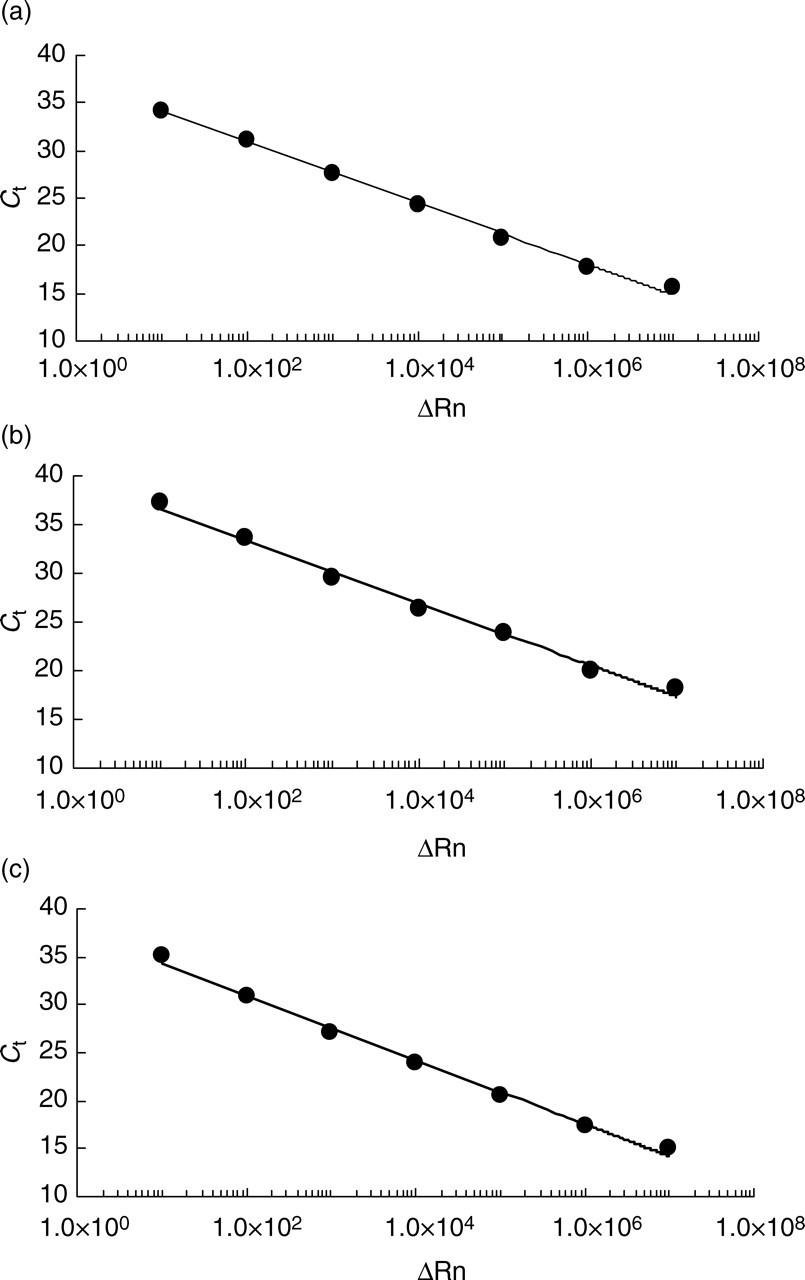
Dilution curves by titration of linearized plasmids. Type-specific TaqMan minor groove binder probe assays using a plasmid corresponding to YMDD motif (a), YIDD (b) and YVDD (c)
Specificity
In order to study the specificity of the YMDD-specific HBV DNA quantification, the three reaction mixtures, each containing one type-specific probe, were added with DNA (6.0 log copies) from the YMDD strain, and measurement was conducted as described above. When the combination of probe and HBV DNA type was matched, an increase in delta-normalized reporter (ΔRn) beyond 0.05 was observed. In the case of a mismatched probe, the intensity was below 0.02 (Figure 2). For YIDD and YVDD strains, the same procedures were performed, and comparable results were obtained.
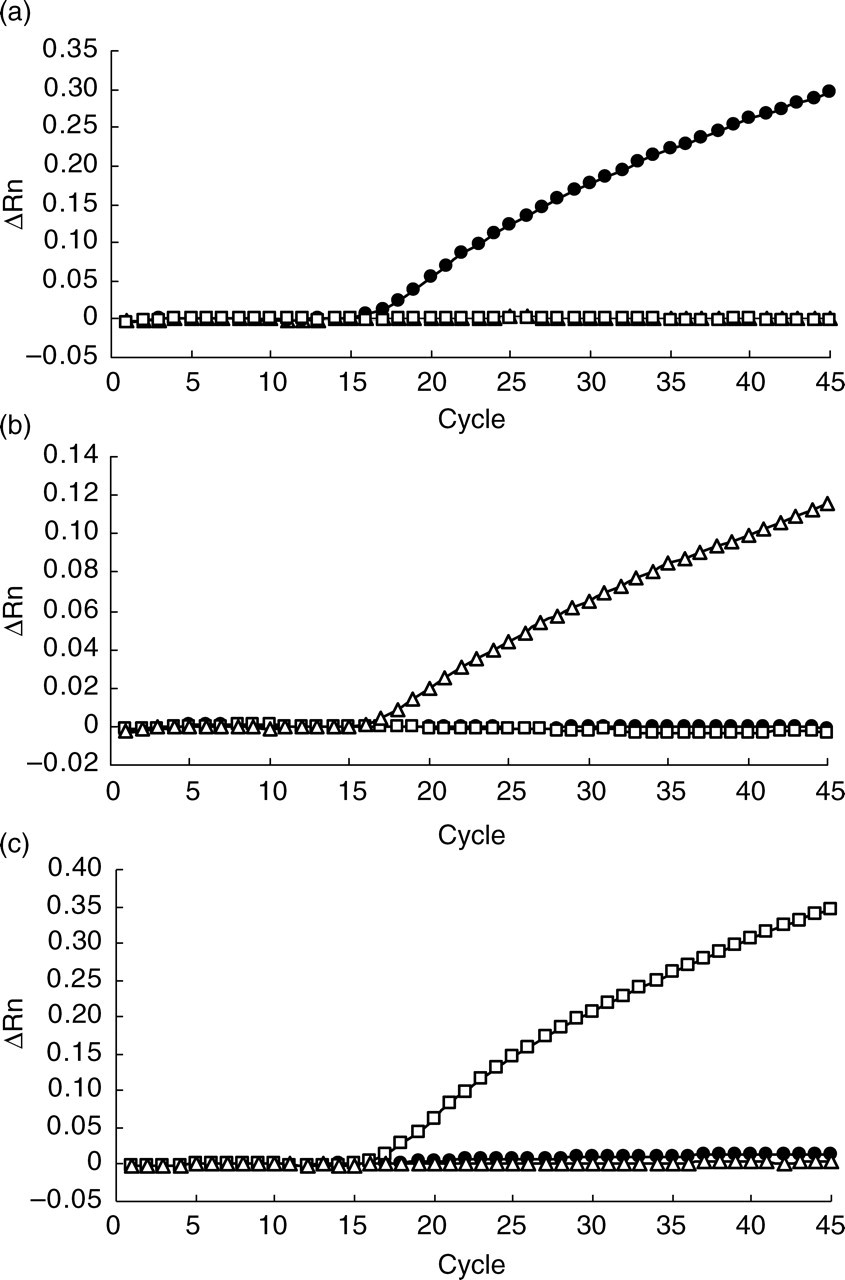
Specificity of YMDD (a), YIDD (b) and YVDD (c) specific TaqMan minor groove binder probe assays. Symbols represent linearized plasmids corresponding to YMDD (•), YIDD (▵) and YVDD (□)
Reproducibility
When HBV DNA quantification was carried out 10 times for the same sample, the coefficients of variation were 1.0% for YMDD, 2.2% for YIDD and 1.9% for YVDD.
Comparison with a commercial kit
The present method was compared with a commercial PCR kit (AMPLICORTM HBV MONITOR) using serum samples from patients (n = 14) with chronic hepatitis B who were not treated with LAM. Because all patients had YMDD strains but no YIDD or YVDD strains, only values for the YMDD strain were compared. A strong correlation was found between the two assays (y = 0.972x + 0.214, r = 0.98, P < 0.001).
Sensitivity
Two of the three linearized plasmid solutions, containing sequences corresponding to YMDD, YIDD or YVDD, were mixed at mixing ratios of 100:100 (7.6 log copies each), 100:10, 100:1, 10:100 and 1:100. The resulting three sets of plasmid mixtures (YMDD vs. YIDD, YMDD vs. YVDD and YIDD vs. YVDD) were then measured. The plasmid corresponding to YMDD (or YIDD) was detectable and measurable down to 1% of YIDD and YVDD (or YMDD and YVDD). The plasmid corresponding to YVDD was detectable to 1% of YMDD and YIDD, but measurable only at 10%.
Comparison with conventional direct sequencing
Both the MGB probe assay and direct sequencing were performed for 109 chronic hepatitis B patients on LAM therapy (Table 1). HBV DNA was detected in 102 (93.6%) of the samples, and identical results were obtained for 90 (88.2%) patients with both methods. In the other 12 samples, direct sequencing resulted in overestimation in 10 cases (9.8%) and underestimation in two (2.0%), in comparison with the type-specific MGB probe assay.
A comparison of hepatitis B virus mutant types obtained by type-specific TaqMan minor groove binder (MGB) probe assay and direct sequencing
ND, not detected
Clinical course
The HBV load was measured after combined therapy with LAM and ADV in the four patients who had LAM-resistant mutant HBV(s) (Figure 3). In three (cases 1–3), a single mutant strain was detected by direct sequencing, and the same type was detected and quantified by type-specific MGB probe assay. In the other case (case 4), suggested by direct sequencing to have multiple mutants, both YIDD and YVDD were individually detected and successfully quantified by the type-specific MGB probe assays. At the beginning of the combination therapy, YVDD mutants were found by this method to be predominant, but YIDD mutants had become more common by day 91. In all cases, the wild-type HBV remained below the detection limit at all time points during the study.
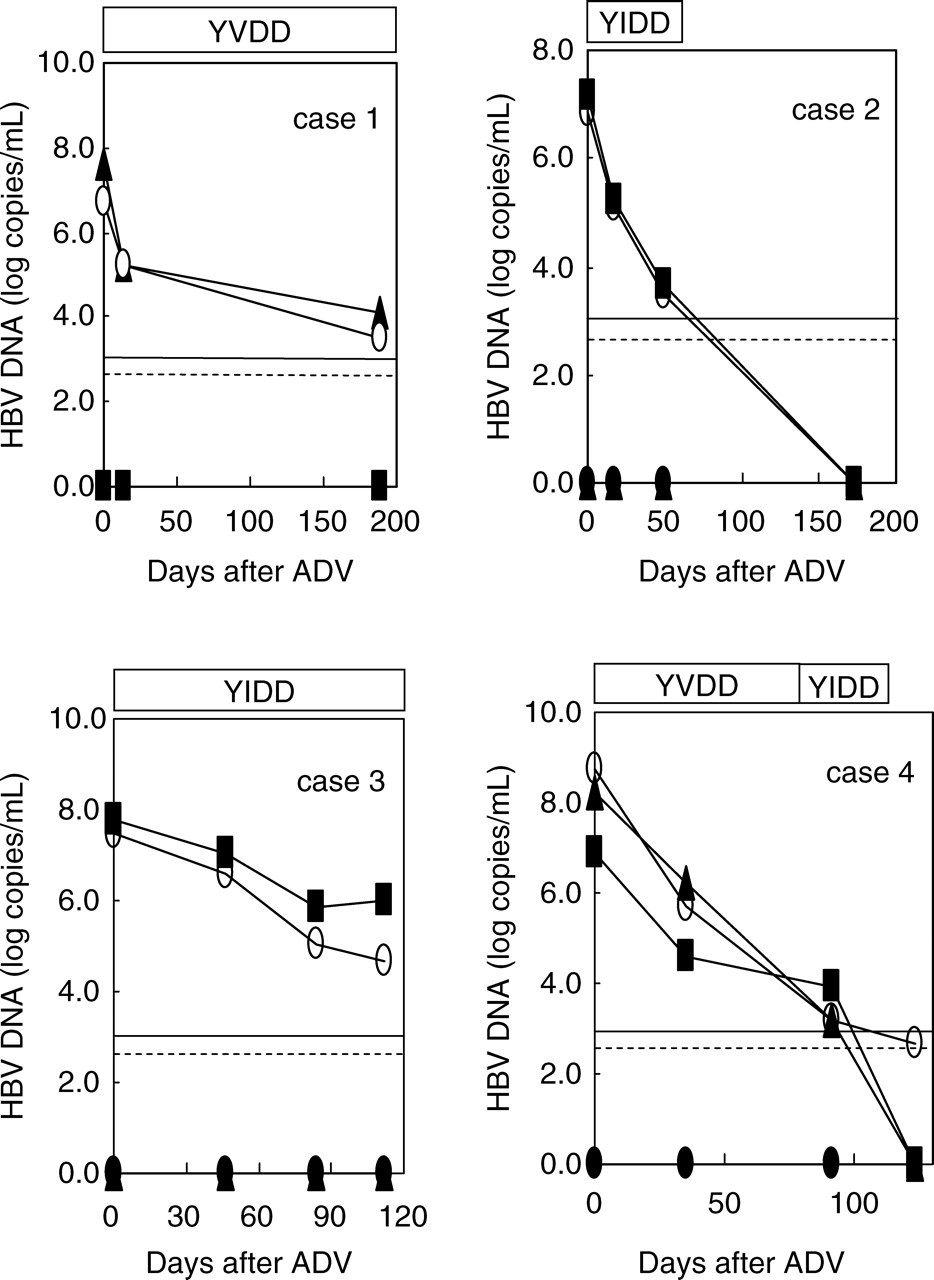
Change in the quantity of serum hepatitis B virus (HBV) DNA monitored by type-specific TaqMan minor groove binder (MGB) probe assay in patients treated with adefovir dipivoxil. Horizontal lines represent the detection limits of type-specific TaqMan MGB probe assay (– – –) and Amplicor HBV Monitor (—). Symbols represent YMDD DNA (•), YIDD DNA (▪), YVDD (▴) and Amplicor HBV Monitor (○)
Discussion
Natsuizaka et al. 16 earlier reported that monitoring the pattern of YMDD motif mutations is useful because changes, such as an accumulation of a specific strain or a change of dominant strain type, can help predict the prognosis. Several methods for detecting mutant HBV strains that are resistant to LAM have been reported. Of the different possibilities, the PCR–restriction fragment length polymorphism method, 17 line probe assay, 18 peptide nucleic acid-mediated PCR 19 and direct sequencing assay are used only for qualitative purposes. The dominant strain may be predicted by reading the peak height or peak area of an electropherogram of direct sequencing or by observing the density of PCR products on gel electrophoresis. Direct sequencing, however, cannot quantify mutant strains because electropherograms are essentially qualitative, and also because no standard method is available for the evaluation of mixed bases. Amino acid sequences cannot be identified when mixed bases are observed in the same codon.
In our present study, quantification of each strain of HBV DNA proved possible with realtime PCR using a unique TaqMan MGB probe for each mutant strain. TaqMan MGB probe was constructed with a fluorescent reporter dye at the 5′ end and with a non-fluorescent quencher and MGB at the 3′ end. A non-fluorescent quencher is thought to contribute to lower noise in comparison with a fluorescent quencher. The MGB in the probe, on the other hand, elevates the melting temperature (T m) through binding to the double-stranded DNA composed of the probe oligonucleotide and the single-stranded target DNA. For this reason, the MGB probe gives a significant change in T m even when only a single mismatch exists in the target, and thus can be used for single nucleotide polymorphism analysis. 20 As the YIDD and YVDD motifs were both generated from the YMDD motif by single base mutations (M204I: ATG → ATT, M204V: ATG → GTG), this property of the MGB probe was advantageous for our assay. Actually, when the TaqMan MGB probe was changed from a regular one in our experiment, we confirmed that the T m value was elevated from 40.4–46.2°C to 63.1–65.3°C using the MGB probe. The latter temperature zone meets the requirement of realtime PCR.
Punia et al. 21 reported a method for quantification of LAM-resistant HBV DNA by realtime amplification refractory mutation system PCR. In comparison with their method, our approach provides a better absolute detection limit: 3 vs. 4 log copies/mL of serum. However, regarding relative detection limits, they reported 0.01% of total HBV for YIDD, while ours were 1.0% for YMDD and YIDD, and 10% for YVDD. A possible explanation for the poor relative detection limit of our method is that the specific binding of the probe to the target strain suffers from competition by other strains that are inferior in affinity for the probe but present at a larger copy number than the target. Another realtime PCR assay based on melting curve analysis using fluorescent biprobe hybridization has been reported. 22 This assay gives the ratio of copy numbers of HBV mutants to the total HBV, but not the concentration of HBV mutants. As T m is influenced by any type of mismatch under the probe, a mismatch adjacent to the YMDD sequence can affect the measurement.
Using conventional sequencing, we overestimated the number of strains in 9.8% of the studied samples and underestimated in 2.0%, when compared with our proposed method. The overestimated samples possessed mixed bases within a codon, presumably due to the presence of multiple mutants or to a simple drift of the baseline in electropherograms. The underestimation with our sequencing method might possibly be due to its low sensitivity relative to our proposed method.
When we applied our method to patients with chronic hepatitis B who received additional ADV after the emergence of LAM-resistant mutants, we could successfully quantify the mutant types in cases 1–3 in good accordance with the results of AMPLICOR™ HBV MONITOR and direct sequencing. The merit of our method, however, was confirmed best in case 4, whose dominant HBV was determined as a YVDD strain by direct sequencing at the start of ADV administration; but was found by our method to shift to YIDD and YVDD strains in variable proportions during the course of treatment. Briefly, the YVDD was about 1 log larger than the YIDD at the beginning of ADV administration; while on day 90, the YIDD was about 1 log larger than the YVDD. This change in the strain type during the treatment is of interest because it is reported that tolerance to LAM differs depending on the mutant type. 23 If YVDD were more susceptible to ADV than YIDD, the observed change in the strain type in case 4 would be well explained. A previous report using molecular modelling which provided evidence that the mutant type is more susceptible to ADV than the wild type supports this possibility. 24
In conclusion, the present method can provide type-specific and quantitative information on multiple HBV strains quickly and reliably. Because drug resistance is a major hazard in anti-HBV therapy, this method might be useful for clinical purposes to monitor change in HBV mutant type in patients treated with anti-HBV drugs.
