Abstract
Ewing sarcoma (ES) is an undifferentiated round cell sarcoma arising in skeletal and extraskeletal locations usually diagnosed in the second decade of life. It is typically characterized by a monomorphic round blue cell morphology, positive membranous CD99 immunohistochemical staining, and EWSR1 gene fusion with a partner gene from the ETS family. Less frequently, other genes from the FET family of genes substitute for EWSR1. We report a case of oligometastatic Ewing sarcoma with FUS::ERG fusion with an unusual primary location in an older patient. This case highlights the application of ancillary tests, especially next generation sequencing, as an adjunct to the morphological assessment of undifferentiated round cell tumours. We also discuss potential diagnostic pitfalls and literature review of Ewing sarcomas with FUS::ERG fusion.
Introduction
Ewing sarcoma is the prototypical undifferentiated round cell sarcoma reported in skeletal and extraskeletal locations. The current fifth World Health Organization (WHO) categorizes undifferentiated small round cell tumours into Ewing sarcoma, round cell sarcoma with EWSR1-non-ETS fusions, CIC-rearranged sarcoma, and sarcoma with BCOR genetic alterations. 1 These categories emphasize defining these tumours based on their molecular gene rearrangements, although there are pitfalls and limitations associated with molecular tests.2,3 The majority of reported Ewing sarcoma cases exhibit EWSR1 gene rearrangements,1 -4 whereas our case revealed a FUS gene rearrangement.
Case Presentation
Our patient is a 66-year-old male who initially presented with haematuria and was found to have a large 60 mm right renal mass. He subsequently underwent a nephrectomy. Microscopically, the tumour was a round cell tumour arising from the right kidney with positive CD99 immunostaining. However, EWSR1::FLI1 and EWSR1::ERG fusions are typically associated with Ewing sarcoma, and SS18 rearrangements, which are usually observed in synovial sarcoma, were not identified. The initial diagnosis was a round blue cell tumour. At 3 months follow up, the patient chose surveillance as he had planned a 4 month trip, there were uncertain benefits of adjuvant chemotherapy and the diagnosis was unclear at the time. At the time, the patient also underwent molecular testing through the Molecular Screening and Therapeutics (MoST) clinical trial programme at the Garvan Institute of Medical Research, which involves a DNA-based 500 gene hotspot NGS panel, with no actionable mutations identified.
Five years later, he presented with coeliac axis thrombosis and splenic infarction, and was found to have a left lower lung lobe nodule during the investigation. We received a wedge resection of the nodule. Macroscopically, there was a well-circumscribed 10 mm nodule. Microscopically, the nodule showed a round blue cell morphology with cells arranged in sheets, microcystic and organoid patterns (Figure 1A). The cells were relatively uniform with scant cytoplasm, round nuclei and occasional small nucleoli. The stroma was vascular. No necrosis was observed. The morphology and immunohistochemical staining results of the patient were similar to those of the previous pathology, consistent with a metastasis.

High-power views at 400x magnification: (A) Haematoxylin-eosin staining of the lung nodule showing a round blue cell tumour with scattered mitoses and vascular stroma, (B) CD99 immunohistochemistry showing diffuse positive membranous staining of tumour cells, (C) ERG immunohistochemistry showing diffuse positive nuclear staining of tumour cells, (D) NKX2.2 immunohistochemistry showing diffuse positive nuclear staining of tumour cells, (E) ALK D5F3 immunohistochemistry, and (F) PanTRK immunohistochemistry showing variable cytoplasmic and membrane staining with eccentric/paranuclear dots.
An extensive panel of immunohistochemistry from our laboratory and consulting laboratories showed positive staining for CD99 (membranous staining; Figure 1B), ERG (Figure 1C), NKX2.2 (Figure 1D), ALK (D5F3) and pan-TRK - the latter 2 showed variable cytoplasmic and membrane staining with eccentric/paranuclear dot-like staining (Figure 1E and F). S100 was reported to be patchy positive in the lung metastasis and not in the initial tumour (unfortunately the slides could not be located). The tumour cells were negative for cytokeratin markers (pancytokeratin, CAM5.2, EMA, CK7), neuroendocrine markers (CD56, INSM1), other mesenchymal markers (desmin, myogenin, SOX10), vascular markers (CD31, CD34), inhibin, ETV4, STAT6, PAX8, and TTF-1. BCOR immunostaining was equivocal. There were slight differences in staining characteristics among the reports, including CD99 showing nonspecific granular deposits rather than membranous staining and focal perinuclear dot-like staining of CAM5.2 in the initial renal case (however, not all of the initial slides could be retrieved).
A solid tumour next-generation sequencing (NGS) DNA-based panel of 50 genes performed at our laboratory did not detect any of the genes using the Oncomine Precision Assay (Thermo Fischer Scientific, A46291) and was sequenced on the Ion Torrent Genexus Integrated Sequencer using the Ion Torrent GX5 Chip.
The case was reviewed at 3 different laboratories (Douglass Hanly Moir Pathology, Prince of Wales Hospital and Fletcher Consult Service at Brigham, and Women’s Hospital/Harvard Medical School) with the suggestions of FUS, CIC::DUX4 or BCOR::CCNB3 rearranged sarcoma, and GLI1-altered mesenchymal tumour. Further fluorescence in situ hybridization (FISH) analyses performed at the Fletcher Consulting Service revealed no CIC rearrangement; however, FUS rearrangement was found in 96% of the nuclei (Figure 2) using the Vysis LSI FUS (Cen) and Vysis LSI FUS (Tel) probes at 16p11 and with the D16Z3 DNA probe (chromosome satellite II DNA) at 16p11.1-q11.1 (Abbott Molecular). Subsequently, a comprehensive NGS RNA-based fusion panel of 507 genes performed at Monash Pathology confirmed FUS::ERG fusion (Figure 3). The analysis used Illumina MiniSeq with data processing via the Illumina Basespace Sequence Hub. The final diagnosis was metastatic Ewing sarcoma.

FISH analysis showing 2 nuclei with split red–green signals for FUS (white arrows).
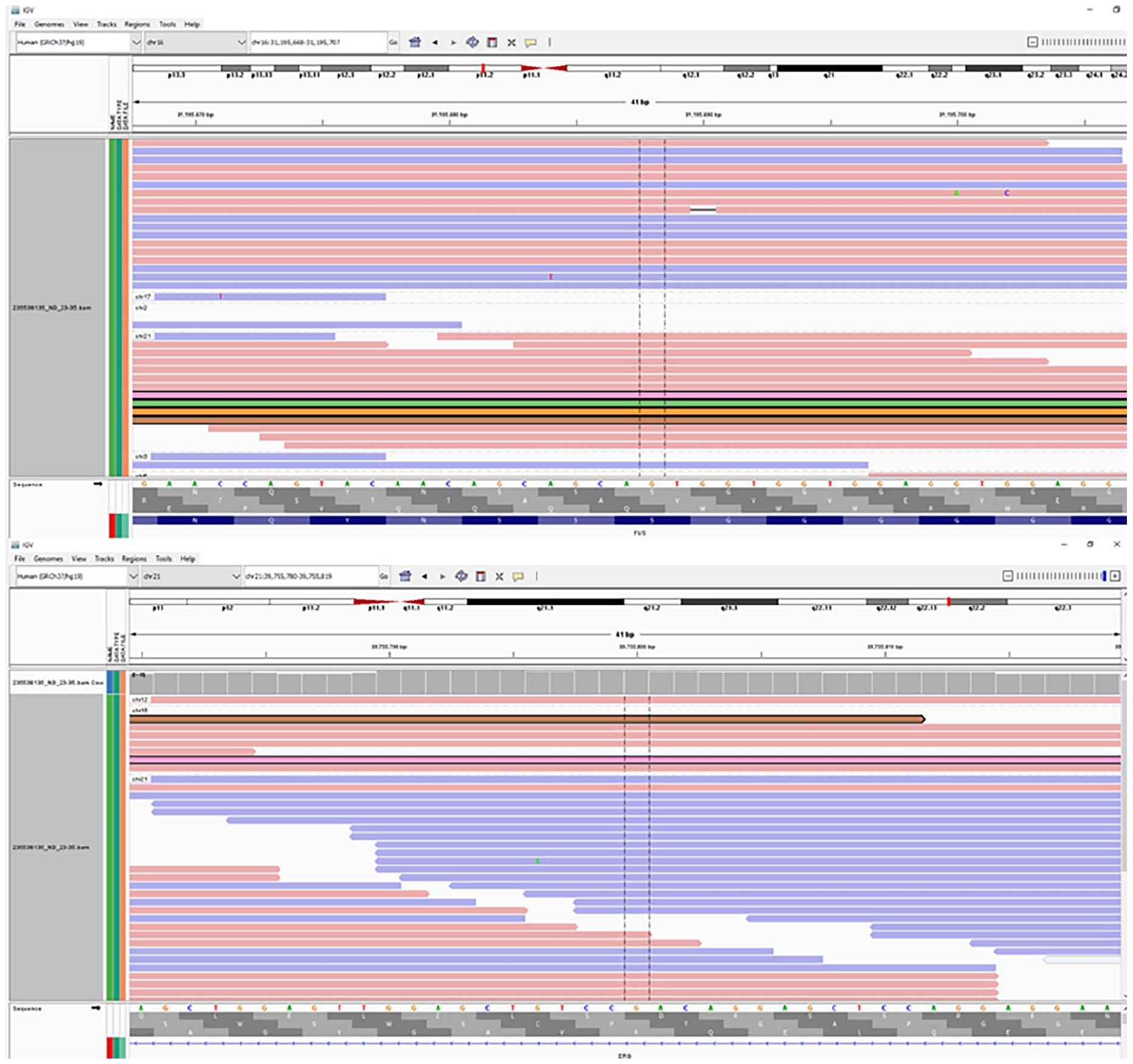
Integrated Genomics Viewer (IGV) from the NGS analysis (performed by Monash Pathology) showing the FUS::ERG fusion transcript. The test performed was an ISO:15189 accredited test and the sequencing QC was optimal with high MAPQ score for both the genes and high fusion call confidence score.
After a multidisciplinary discussion, the patient was managed with surveillance. He remained asymptomatic with no evidence of recurrent or metastatic disease on Positron emission tomography scan at 1-year follow-up.
Discussion
When a pathologist encounters a round blue cell tumour, a long list of differentials is generated, regardless of the site. Diagnosis can be challenging given overlaps in morphology and immunostaining patterns for sarcomas and non-sarcomas. The differentials for the initial tumour was wide and included other lineages such as carcinoma (including neuroendocrine carcinoma), melanoma, germ cell tumour and lymphoid tumours. The sarcoma differentials included synovial sarcoma, solitary fibrous tumour, rhabdomyosarcoma, CIC-rearranged tumour, BCOR-rearranged tumour, GLI1-altered mesenchymal tumour, NTRK-rearranged tumour. Desmoplastic round cell tumour and Wilm’s were considered given the location however the demographics are usually younger.
ES is a relatively rare and aggressive sarcoma with a prototypical round blue cell morphology, 1 although there is a histologic spectrum. 3 The WHO 2020 diagnostic criteria requires a small round cell morphology and CD99 membranous staining. The demonstration of FET-ETS gene fusions is desired but not required, 1 notably because FISH can produce false negative results.2,3 The fusions can be detected by a variety of molecular tests, including fluorescence in situ hybridization (FISH), reverse transcription polymerase chain reaction, and next-generation sequencing (NGS).2 -4
ES is defined by gene fusions involving 1 member of the FET protein family of genes and a member of the ETS family of transcription factors. The FET protein family of genes is functionally interchangeable and includes FUS, EWSR1 and TAF15.1,2,4,5,8 ETS genes include FLI1, ERG, ETV1, ETV4, and FEV.4,5 The majority of reported Ewing sarcoma cases exhibit EWSR1::FLI1 (~85% of cases) or EWSR1:: ERG (5%-10% of cases) fusions.1 -4
ES with FUS gene rearrangements are rare in the literature. The most common reported fusion partner is ERG. The other fusion partners were ETV4, FEV and NFATC2 (the latter of which is a non-ETS gene and thus would not be Ewing sarcoma with FET-ETS fusion according to the WHO 2020 diagnostic criteria).2,6 There is a study reporting 35 cases with EWSR1/FUS::ERG; however, the specific number of FUS::ERG cases was not provided. 5 We are aware of a recent FUS::FLI1 fusion in a local case of ES. 7 In our literature review, we were mindful that terminology may have changed since the publication of older articles, such as Askin’s tumour, which was previously used for ES of the chest wall in the previous WHO classification. 1
There are 11 reported cases of ES with FUS::ERG fusions in our literature review (Table 1). There was 1 other renal primary case and 1 other metastatic case. The t(16;21)(p11;q22) translocation results in FUS::ERG fusion.2,4,8 There are too few known cases of ES with the same translocation to determine if it is of clinical significance and compare the clinical and pathologic features of these cases with those of classic ES cases with the EWSR1::FL1 fusion.2,8 Other interesting characteristics of this case are the patient’s age and primary site location. ES is usually diagnosed in the second decade of life and is rare in individuals older than 30 years. Cases in people above 30 years old are noted to arise in soft tissues rather than the bone.1,3 Interestingly, nested growth is noted to be in the extraskeletal and middle-aged to older adults, 3 as in our case. The only other reported renal ES with FUS rearrangement was in a 3-year-old girl. 4
Summary of characteristics from a literature review of Ewing sarcoma patients with confirmed FUS::ERG fusion.

Staining characteristics were not provided.
Abbreviations: F, female; IHC, immunohistochemistry; M, male; N/A, information not available. Age is shown in years.
There is limited clinicopathological data of renal Ewing sarcoma, however case series and case reports have reported poorer outcomes compared to their bone and soft tissue counterparts, and often presented at extrarenal spread and metastasis at initial presentation.5,9 -12 In 1 systematic review, 12 38.6% of the patients had metastatic disease and 10.7% had locally advanced disease at initial presentation. The overall survival reported in this review was 33.5 months. In 1 study with 35 EWSR1/FUS::ERG cases, 5 there were also associated higher rates of metastasis at presentation and worse overall survival, likely due to a greater incidence of metastatic spread at presentation in less common subsets. However, both study and review did not specify the specific number of cases in which FUS, not EWSR1, was rearranged. There is no consensus guideline on management of renal ES, however adjuvant chemotherapy given at the time of localized disease seemed to have favourable outcomes in previous studies and review.9 -12 Interestingly, our case had an indolent course despite no adjuvant chemotherapy given, with the recurrence presenting 5 years post the initial presentation.
Although our molecular knowledge of ES has progressed, morphology is still crucial for diagnosis. FUS rearrangements can also be observed in other sarcomas, notably myxoid liposarcoma, sclerosing epithelioid fibrosarcoma, and low-grade fibromyxoid sarcoma. There are also reported cases of undifferentiated round cell sarcoma with FUS rearrangement in the literature; however, according to the latest WHO classification, these cases could be ES or Round cell sarcoma with EWSR1-non-ETS fusion, depending on the fusion partner gene.1,6
There is no current immunohistochemistry that is entirely specific to ES. However, the combined positive immunostaining of CD99, NKX2.2 and PAX7 is highly specific, irrespective of the morphology and fusion genes. 3 Unfortunately, we did not have access to PAX7 immunostaining to confirm this. Strong ERG immunostaining has been reported to be a useful marker for previous EWSR1::ERG cases, 13 although ERG can be detected in other entities and is not entirely specific. 2 ERG immunostaining has not been reported in previous FUS::ERG cases. In our case the tumour cells were positive for ERG immunohistochemistry, however the result was interpreted with the overall morphology, other immunostaining characteristics, and molecular analysis. Unfortunately FUS immunohistochemistry was not available at the time to determine if there is potential diagnostic application.
Interestingly our case showed some ALK D5F3 and panTRK immunostaining. However, no NTRK or ALK gene rearrangement was detected by in-house Oncomine Precision Assay panel and the 500 gene fusion panel from Monash. One explanation could be due to cross-reactivity of ALK D5F3 and panTRK antibodies with the products of FUS/ERG gene fusion. Another explanation could be due to modification at the post-transcription level of ALK and NTRK. There is no data to currently support these theories, however panTRK immunohistochemistry has had reported false positives in tumours without the NTRK rearrangements. 14 A lot of the data on ALK D5F3 have been targeted at lung cancers and this has been shown to also have false positives.15,16 We considered the staining of these immunohistochemical stains to be of uncertain significance, given lack of data in solid tumours.
At the time of the initial presentation in the kidney, no FUS FISH probe was available. NGS was also not covered by Medicare at the time and a RNA fusion panel was not available in Australia. However, if a similar case occurred now and if there was sufficient tissue, we would request a RNA fusion panel to cover a wider range of genes, unless the morphology and immunohistochemical features strongly favoured a diagnosis that could be readily confirmed with FISH (ie, if SS18-SSX immunohistochemistry was positive then FISH for SS18 gene rearrangement for synovial sarcoma would be preferred). For sarcomas RNA fusions are more important as the diagnostic criteria under WHO show different categories for round cell tumour depending on the fusion partner for EWSR1 rearranged tumours: Ewing sarcoma or Round cell sarcoma with EWSR1-non-ETS fusions.
Conclusion
The correct diagnosis of undifferentiated round cell tumours is crucial given their different prognoses and management strategies. Awareness of noncanonical genes and advancements in molecular pathology will hopefully prevent misdiagnosis of ES. Our case shows that ES should be a differential for round cell tumours at unusual sites and/or in older patients. If an EWSR1 gene rearrangement is not detected, further testing for other FET genes is warranted when positive CD99 immunostaining and morphology are suggestive of ES. ERG immunohistochemistry is another potentially useful marker for detecting this fusion partner. Our approach to undifferentiated round cell tumours may adapt to new technological advances, improvement to test limitations and cost-benefit ratios, and emerging molecularly defined entities.
Footnotes
Acknowledgements
We would like to thank Dr. Will Anderson from the Fletcher Consulting Service at Brigham and Women’s Hospital/Harvard Medical School for consulting on this case. We would also like to thank Prof. Paola Dal Cin from the Fletcher Consulting Service at Brigham and Women’s Hospital/Harvard Medical School for their FISH images.
Frequently Used Abbreviations
ES = Ewing sarcoma; ETS = E26 transformation specific; EWSR1 = Ewing sarcoma breakpoint region 1/EWS RNA binding protein 1; FISH = fluorescence in situ hybridization; FUS = fused in sarcoma; NGS = next generation sequencing; WHO = World Health Organization
Author’s Note
The author Dan T. Nguyen has since moved to Department of Anatomical Pathology, Prince of Wales Hospital, Randwick, New South Wales, Australia.
Ethical Considerations
Not applicable
Consent for Publication
Consent was obtained from the patient
Author Contributions
DTN wrote the main manuscript text, prepared figure 1 and table 1, and obtained patient consent. MRQ is the main supervisor and provided guidance on the content, revised the manuscript, and provided images part of figure 1. TD also supervised and provided guidance on the content. FM contributed to the diagnosis and provided guidance on the content. PD provided guidance on the molecular aspects and prepared figure 3. YCL contributed to the diagnosis and management of the patient.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Not applicable
