Abstract
NanoPharos is a FAIR Enabling Resource (type: registry) that aspires to become a FAIR Data Point. It offers FAIR-compliant, ready-for-modelling nanomaterials safety datasets, enriched with molecular/atomistic descriptors, and programmatic or manual export into modelling software.
Name of the FAIR Supporting Resource
NanoPharos: a database for modelling-ready and computational nanomaterials interactions and impacts datasets (http://purl.org/np/RA-7YdatF38SxYdikEs11XiD9rEpYOsXZaTu8o9d1vRp0).
FAIR Supporting Resource type(s)
Registry (data and metadata).
Contributing community
WorldFAIR Nanomaterials Community (http://purl.org/np/RAWg5p8IqzdX0PIYNBpze1iUpyhHXD7JgDQKWxsFNhSZk#NanoCommons).
How NanoPharos supports FAIR
NanoPharos, 1 powered by Pharos Database Solution, 2 is a scalable FAIR Enabling Resource (type: registry) 3 (Figure 1) for nanomaterials environmental health and safety (nanoEHS) datasets. These may be derived from the literature, experiments, or computational models, that is, from first principles (so-called physics-based models) or data-driven modelling including machine learning. NanoPharos aspires to become fully FAIR-compliant, as per the GO FAIR Foundation interpretation 4 of the FAIR Principles, 5 and to act as a FAIR Data Point (FDP). A NanoPharos FAIR Implementation Profile (FAIR Principle R1.3) 6 was developed in the WorldFAIR project, 7 which is updated as new functionalities are implemented.

Diagram of the core components of the NanoPharos FAIR-compliant database and how they map to the FAIR Principles. At the centre is the NanoPharos database, which is connected to essential elements such as GUPRIs for dataset traceability (FAIR Principle F1), comprehensive metadata covering experimental and computational datasets (FAIR Principles F2, F3, F4, I3, R1, R1.2, and R1.3), various access protocols such as application programming interface (API) and KNIME nodes for programmatic and manual interaction (FAIR Principles A1 and A1.1), and licensing mechanisms that ensure proper data reuse and compliance (FAIR Principles R1.1). These elements work together to ensure that NanoPharos is a FAIR–compliant FAIR Enabling Resource of type registry.
NanoPharos datasets are assigned unique identifiers (FAIR Principle F1) in the form of https://db.nanopharos.eu/Queries/Datasets.zul?datasetID=npX, where X is a number in ascending order, in compliance with the Uniform Resource Identifier (URI): Generic Syntax standard (RFC 3986 IETF).
8
The Pharos Database Solution uses the ChemBL database schema,
9
and a slightly modified version, to accommodate the specialised nature of nanomaterials (as both chemicals and particles) and nanoEHS data, has been inherited by NanoPharos. The data schema describes the interlinkages between nanomaterial structure, atomistic characteristics, physicochemical properties, interactions (e.g. exposure routes, pharmacokinetics, and biomolecule binding), and effects (impacts, adverse outcome pathways, toxicity endpoints, alterations in gene expression etc.), thus allowing its translation into a detailed Knowledge Graph. The metadata describing datasets in NanoPharos are defined using a controlled vocabulary based on a semantic model (FAIR Principles F2, I2, I3, and R1.2) and are divided into bibliographic, provenance, and scientific metadata:
Bibliographic metadata: dataset title, description, owner(s), data producer(s), curator(s), contact detail(s), relevant publications, unique IDs, and descriptors, for example, DOI, ORCID, file type, and size. Provenance metadata: description of the methods used to produce the data, date of data production, date of modification (where applicable), and versioning. Scientific metadata: protocols, methods, instruments used, analytical and computational algorithms, and software used and versions. For computational datasets, this includes model documentation using (for example) the Modelling Data templates
10
and the EasyMODA tool.11,12
Metadata and data reporting are performed separately (FAIR Principle F3). When a publication about the dataset exists a direct reference to the respective digital object identifier (DOI) is included in the dataset page (FAIR Principle F1). The datasets’ metadata are recorded as nanopublications using customised templates in nanodash 13 (FAIR Principle F2), with the respective globally unique, resolvable and persistent identifier acting as the metadata's unique identifier (FAIR Principle F1). In this way, metadata can be accessed in formats such as TriG, JSON-LD, N-Quads, and XML (FAIR Principle I1). The nanopublications explicitly include the URI of the data they describe (FAIR Principle F3), as well as the DOI of any publication containing any connected metadata (FAIR Principle I3). The data and metadata available through NanoPharos are publicly available based on clearly defined CC-BY licences (FAIR Principle R1.1), which are flagged both within the database and in nanodash. The licenses related to external publications are those assigned by the respective journals and are not managed by NanoPharos. ID-np23 14 provides an implementation example. Data and metadata, including identifiers, are also indexed in Zenodo, which acts as the metadata's longevity plan (FAIR Principles F4 and A2). This, together with adding the funding information (funder/grant number) to the metadata will allow indexing in OpenAire for data produced under EU framework funding.
NanoPharos is underpinned by an internal semantic model, which describes the implicit meaning of the data in a machine-actionable way (FAIR Principle I3). Finally, NanoPharos uses KeyCloak, an open-source single sign-on (SSO) identity and access management software, as the authentication and authorisation mechanism (FAIR Principle A1.2) for its human interface to access embargoed and sensitive non-publicly available data. Keycloak's extensive functionality (setting up SSO, creating and managing user sessions, and configuring secure connections) ensures that NanoPharos adheres to best practices in cybersecurity.
While NanoPharos is readily accessible online via a human interface, a Representational State Transfer Application Programming Interface (API) is also available 15 (Figure 2), which allows programmatic interaction with other databases and modelling tools (FAIR Principle A1). The API is publicly available and open for interested users to use for data and metadata retrieval (FAIR Principle A1.1). Currently, the API can retrieve specific datasets based on their IDs.

The open and publicly available NanoPharos Restful application programming interface (API) for programmatic access and data retrieval from the NanoPharos database.
NanoPharos provides users with modelling-ready tab-delimited datasets that can be directly imported into computational workflows and software, such as the Isalos Analytics Platform16,17 or KNIME 18 using, for example, the Enalos + KNIME nodes.19,20 In this way, the development of robust and reliable nanoinformatics models, which provide meaningful insights into the characteristics and behaviours of nanomaterials, is streamlined. This process facilitates current analytical needs, while paving the way for future advances in the field, fostering a collaborative and productive research environment.
As described in Section 3, NanoPharos implements several key aspects of the FAIR Principles, promoting optimised data accessibility and reusability. NanoPharos goes beyond the technical interpretation of the FAIR Principles by embracing the scientific FAIRification (SFAIR) principles, tailored for nanomaterials safety assessment. 21 SFAIR do not aim to substitute the original FAIR Principles, but provide non-technical data producers with practical guidelines for implementing these in their everyday practice, an action the FAIR Principes explicitly leave to each community.
NanoPharos considers the complexities and dynamic nature of nanomaterials, including limited batch-to-batch reproducibility (even for commercially provided nanomaterials). 22 Nanomaterials undergo a range of transformations during storage, characterisation, and exposure in toxicology assessments.23–25 The database enables the creation of nanomaterials batches or environmentally transformed variants, which are linked computationally, for example, Rietveld refinement, to a parent nanomaterial. Each batch can be linked to its characterisation and biological assay data, ensuring complete provenance tracking, an essential aspect of data reusability (FAIR Principle R1). The use of the European Materials Registry (ERM) identifier, 26 along with the in-development NanoInChI identifier, provides further granularity in describing the nanomaterial's properties, for example, size, morphology, and crystal structure.27,28 This scientific context ensures data reusability with the highest confidence, fostering the development of nanoinformatics models with excellent data provenance. NanoPharos also enables inclusion of omics data, for example, Saarimäki et al. 29
NanoPharos is designed to capture data related to computational analyses and to support the enrichment of nanomaterials experimental characterisation data with computational descriptors, including atomistic, periodic table, and molecular descriptors (Figure 3). Thus, users can define specific nanomaterials, and enrich the main structure with molecular and atomistic descriptors, which can be linked with a specific ERM identifier. For example, an existing literature-curated dataset having the physicochemical and toxicological data from 14 metal oxide nanomaterials was enriched with 62 atomistic computational descriptors and exploited to produce a robust in silico model for prediction of nanomaterials cytotoxicity based on quantification of membrane damage. 30 The cytotoxicity model 31 was made publicly available and the underpinning dataset, including the calculated descriptors, is available from NanoPharos (via dataset ID = np1). 32

The NanoPharos database provides users with free access to ready-for-modelling datasets. Users can filter the available datasets to meet specific requirements. A dashboard to support the visualisation of the datasets and application programming interface (API) downloads is being implemented.
The NanoPharos database developers are constantly striving to increase its FAIRness and maturity based on the FAIR Principles 5 and the GO FAIR Foundation FAIR Principles interpretations. 4 As stated in section 3, NanoPharos currently supports limited machine (actionable) accessibility and data and metadata retrieval via the APIs at the individual dataset level. A key limitation, currently, of NanoPharos is the lack of semantic annotation of the datasets using FAIR ontologies or structured vocabularies (FAIR Principles I3 and I2). There is also, currently, a relatively modest number of datasets present in the database, but this is expected to increase significantly in 2025, as more functionalities are introduced (see section 6). Another limitation is the fact that the data and metadata contained in NanoPharos have not yet been indexed fully in search engines, limiting its visibility and dissemination potential.
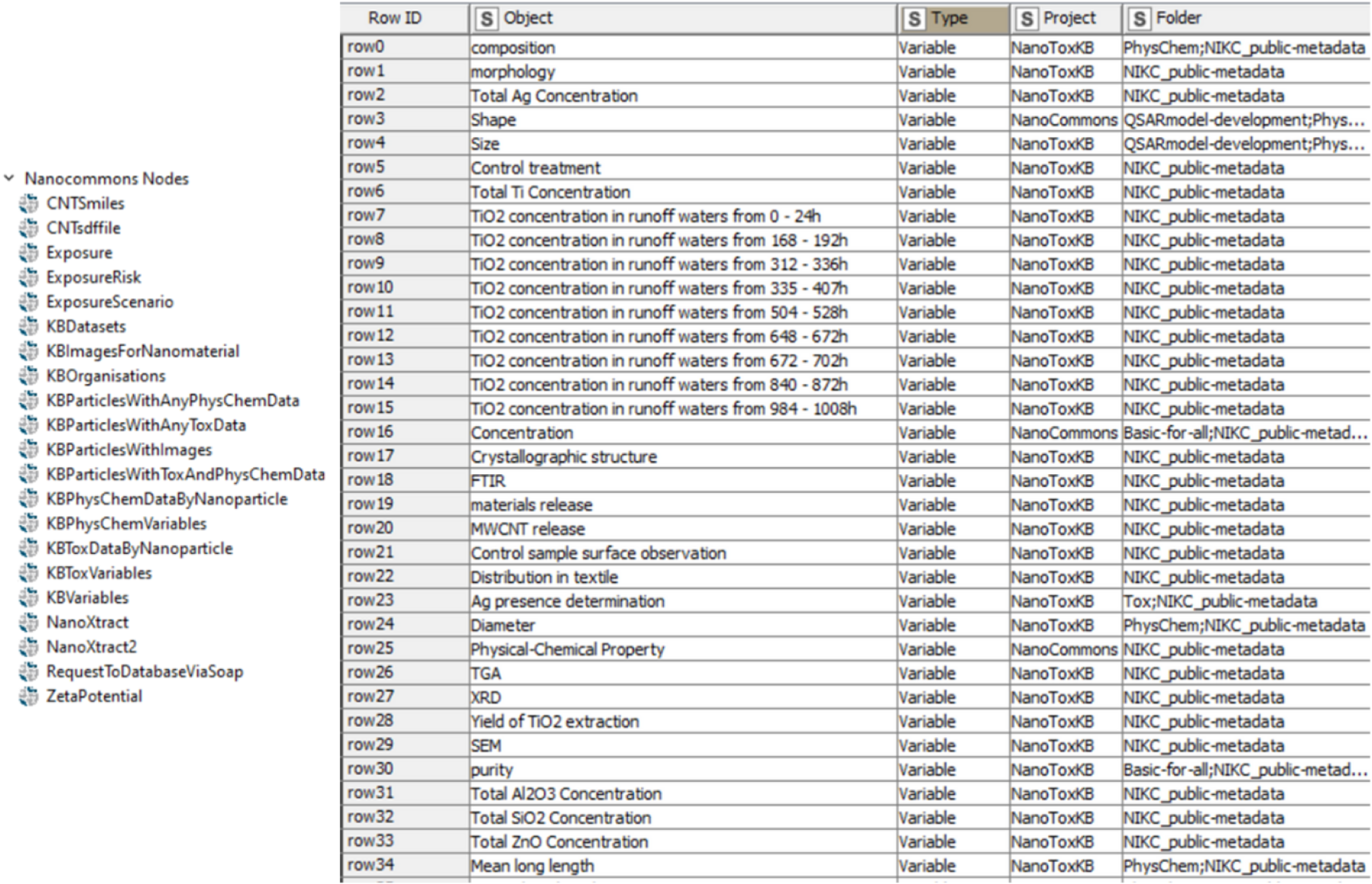
Nanocommons KNIME nodes and application of the Excel Writer node.

Workflow for FAIRification of data via NanoPharos and integration of the nanoinformatics-ready datasets into Computational Modelling Pipelines. Data sources include physics-based models, experimental assessment, image analysis, and literature searches. Datasets are assigned unique identifiers for traceability (FAIR Principle F1) and linked with metadata, including publications (FAIR Principles F2 and R1.2) and semantic models (FAIR Principle I3). Data can be accessed programmatically via application programming interfaces (APIs) or KNIME nodes (FAIR Principle A1.1) or manually in tabular formats, and filling of data gaps to enhance nanomaterials safety research. Image created via canva.com.
Plans exist to enhance NanoPharos’ FAIRness and services by adding features covering the remaining FAIR principles and expanding existing ones. This includes implementing FAIR ontologies or structured vocabularies, for example, eNanoMapper ontology, 33 and Chemical Entities of Biological Interest (ChEBI) ontology, 34 to enable dataset retrieval based on keywords search (FAIR Principles I2 and I3). The API will be complemented with KeyCloak, for authentication and authorisation, to enable access to embargoed or sensitive data based on licensing and specialised permissions (FAIR Principle A1.2). Access to NanoPharos will be further enhanced with KNIME nodes to support programmatic access, structuring 35 and ontological annotation of datasets 36 (as Excel files), thereby facilitating curation and upload of datasets (Figure 4). Available nodes for conversion of datasets into formats such as JSON-LD and XML (FAIR Principle I1) will be implemented for NanoPharos via KNIME and the database's web interface and API. This will enable linking the datasets’ data and metadata with the respective nanoinformatics models that use them, enhancing interoperability and allowing seamless integration with other platforms and tools.
Other improvements underway include the expansion of data and metadata indexing into Zenodo, and where applicable OpenAire, under a single publication (FAIR Principle F4). NanoPharos is currently being mapped to the Common European Research Information Format (FAIR Principles F2 and I3). This implementation will allow the transformation of the NanoPharos datasets into formats other than tabular such as JSON, XML and RDF (FAIR Principle I1), like the metadata availability as nanopublications using RDF. Thus, it will enable automated data and metadata retrieval and integration with external systems that follow similar standards or map to the NanoPharos database (FAIR Principles F2 and I3). This process involves the augmentation of the database with new research findings, predictive accuracies, and ancillary metadata generated during the modelling process.
NanoPharos will also provide a manual and programmatic searchable registry service for data and metadata (FAIR Principle F4), making it easier to retrieve specific datasets and associated metadata. NanoPharos aims to achieve a formal Trusted Repository status under standards such as CoreTrustSeal 37 or ISO16363 38 providing data stewards with confidence in the long-term sustainability, curation, and protection of the NanoPharos datasets and positioning NanoPharos as an FDP for nanosafety and advanced and novel materials data.
Example applications of NanoPharos in data FAIRification and computational workflows
NanoPharos supports data FAIRification and workflow enrichment for experimental and computational data. NanoPharos uses KNIME nodes, as described in WorldFAIR deliverable report D4.1,
39
to integrate, standardise, and FAIRify nanoEHS data, as follows:
Data import and consolidation: Import multiple Excel files with KNIME Excel Reader nodes and combine them using Concatenate, Joiner, or Merge nodes. Data preprocessing and cleaning: Use missing value, duplicate row filter, row splitter, and string manipulation nodes to handle missing values, duplicates, and data inconsistencies. Metadata enrichment: Employ column properties, column renames, and column filter nodes to manage structured metadata, such as descriptions, units, or data types. Data standardisation: Ontology-based curation of datasets is currently being implemented. Data provenance tracking: Use KNIME nodes to record data transformation history and provide workflow audit trails. Data export and sharing: Export resulting FAIR datasets using appropriate Writer nodes (CSV, JSON, and RDF). Workflow documentation and sharing: Document and share full workflows (Figure 5) to facilitate adoption of FAIR practices.
The NanoPharos functionalities are significantly enhanced through the integration with computational tools for generating atomistic and molecular descriptors, for example, ASCOT, 40 NanoConstruct, 41 and Nanotube Construct 42 (Figure 6). These tools provide substantial database enrichment and support the development of machine-learning models. ASCOT is used to generate descriptors that encapsulate the distinct physicochemical attributes of nanomaterials, enriching the database with a more comprehensive analytical depth. 40 NanoConstruct is a toolbox for the digital reconstruction of energy-minimised nanomaterials, based on crystallographic information, and the respective calculation of atomistic descriptors. 41 The Nanotube Construct tool is specialised for generating descriptors for tubular nanostructures, 42 introducing an additional dimension of specificity within NanoPharos. Utilising these data, machine-learning algorithms can more accurately predict essential nanomaterial properties and correlate them with subsequent adverse effects in cells, organisms, humans, and the environment. This integration demonstrates NanoPharos’ adaptability to diverse data types, highlighting its role as a comprehensive, FAIR-aligned nanomaterials research resource. Seamless data exchange with specialised tools (Figure 6) supports various computational workflows, including predictive modelling and nanomaterials behaviour analyses.

ASCOT, NanoConstruct, and Nanotube Construct tools available via Enalos Cloud Platform.
Footnotes
ORCID iDs
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The initial development of NanoPharos was funded via the European Union Horizon 2020 projects NanoCommons (Grant Agreement No. 731032) and NanoSolveIT (Grant Agreement No. 814572). Onward development and further FAIRification efforts were funded via the Horizon Europe WorldFAIR (Grant Agreement No. 101058393) project in which the UK participation was funded by UKRI / Innovate UK via the Horizon Europe guarantee fund (Grant No. 10038665), the Horizon 2020 projects DIAGONAL (Grant Agreement No. 953152), CompSafeNano (Grant Agreement No. 101008099), and the European Union Recovery and Resilience Facility of the NextGenerationEU instrument, through the Research and Innovation Foundation (Project: CODEVELOP-GT/0322/009 3).
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: NanoPharos was initially developed by NovaMechanics Ltd and the University of Birmingham via European Union funding, but ownership, maintenance and onward development have been transferred entirely to the non-profit Entelos Institute to avoid any conflict of interest. The authors have no conflict of interest to report.
