Abstract
Introduction:
Three-dimensional measurements of intracranial volume (ICV) can guide clinical management of brain and skull disorders. However, widespread clinical access is limited by the scarcity of software methods for analyzing CT scans, which are more available than MRI images, and the inaccessibility of proprietary and expensive commercial software. The presented method can calculate ICV from CT scans, using an open-source software, 3D Slicer.
Methods:
The open-source workflow was optimized with a data-driven approach to find the optimal parameters for ICV accuracy. The accuracy of the open-source method was determined by comparing it to commercial and proprietary software with CT scans of pediatric hydrocephalic macrocephaly patients with craniosynostosis undergoing total vault reconstruction (N = 5 patients,15 scans).
Results:
An open-source pipeline that combines an initial semi-automatic segmentation of a coronal CT reconstruction with a fully automatic segmentation minimizes the ICV error. The open-source method shows excellent agreement with both the commercial and proprietary software methods (R2 = 0.998 and 95% confidence interval of best-fit line slope: [0.986; 1.047], [0.985;1.066] respectively). The mean percent difference of ICV measurements of the open-source software from the commercial software was −0.56% [95% CI: −1.08%, −0.028%] and from the proprietary software was −0.07% [95% CI: −1.26%, 1.1%]. The mean percent difference of ICV measurements of the commercial software from the proprietary software was 0.36% [95% Confidence Interval: −0.61%, 1.32%].
Conclusions:
This is the first study comparing an open-source method for measuring ICV with commercial and proprietary options. A high degree of fidelity was found, confirming this open-source method as a viable option for clinicians who are looking to incorporate ICV measurements into their practice.
Introduction
Measurements of intracranial volumes (ICV) enable clinicians to quantitatively assess disease pathophysiology, guide treatment, and surgical decision-making. 1-7 Two-dimensional measurements, either made on medical images or by the clinician at the point-of-care, do not accurately estimate 3-dimensional volumes extracted from medical images.8-12 Thus, as a standard of care, clinical teams typically extract 3-dimensional volumes from imaging methods such as computed tomography (CT) or magnetic resonance imaging (MRI). However, 2 barriers exist, limiting widespread adoption. First, most of the available software packages can only measure intracranial volume from MRI scans.13-16 However, CT scans are more widely accessible and frequently used than MRI imaging due to faster image acquisition and lower costs. 17 There exists a paucity of methods using CT scans. Second, many software packages are prohibitively expensive or proprietary to a hospital or company. Some hospitals in low-resource countries or independent practices will not be able to access these tools. Furthermore, when physicians or groups are considering purchasing these products, it is unknown whether open-source packages provide equivalent accuracy. To date, there has not been a comparison of ICV accuracy between open-source, commercially available, and proprietary methods.
In this study, we developed an automated method to calculate intracranial volume from CT scans using 3D Slicer, an open-source software. Furthermore, we compare intracranial volume measurements from CT scans using the open-source software, a commercial software, and a proprietary software in a patient population. The results from this study may inform clinical and research units that are choosing software tools for ICV measurement.
Methods
Selection and Description of Participants
Ethical approval was obtained from the University of Texas at Austin Institutional Review Board. Pre-operative, immediate post-operative, and long-term follow up (2 or 3 years following surgery) CT scans were obtained in 5 hydrocephalic macrocephaly patients with underlying craniosynostosis (5 male patients, average age at surgery = 2.6 years, average weight = 11.7kg) who underwent total vault reconstruction. All CT scans were taken in the axial plane with a constant slice thickness of 1 mm. Coronal and sagittal reconstructions were performed and bone and soft tissue windows were applied, leading to six permutations of each scan.
Data Collection and Measurements
The open-source software, 3D Slicer (Slicer 4.11.20200930), commercial software, Materialise Mimics (MIS 25), and proprietary software, 3D Systems, were used to calculate the intracranial volume. All CT scans (15) were analyzed using open-source and commercial software, whereas only pre-operative and post-operative scans from 4 patients (8 scans) were available for analysis by the proprietary software, for access reasons. Measurements made using the open-source and commercial software were run using a Dell XPS 15 9500 PC with an Intel Core i7-10750H processor by the same operator. The details of the proprietary software method for measuring ICV were not made available.
The open-source pipeline for 3D Slicer proceeded as follows (Supplementary Video 1). CT scans were loaded into 3D Slicer as DICOM images. The chosen sequence was opened in the Segment Editor. A mask containing bone intensities was created using an established Hounsfield Unit (HU) threshold for bone segmentation on CT images. 18 The Islands feature was used to keep only the largest connected component in the mask, which corresponds to the skull, excluding the vertebrae. A 3D dimensional segment was created from the internal cavity of the skull. The entire area surrounding the mask was set to a uniform density, corresponding to air on a CT scan, and the mask was exported as a solid volume. This segmentation is defined as the initial segmentation. This initial segmentation adapts a previously described but untested workflow. 19 However, unlike previous methods, we continued to refine this segmentation further. Next, the internal cranial volume was refined using the Swiss Skull Stripper extension on 3D Slicer. 13 This extension is a fully automated method that uses a reference atlas as an initialization for a level-set that is fitted to the brain-skull border on the patient scan. The initial segmentation was used as the initialization for the refinement. Finally, the total volume of this refined model of the intracranial space was calculated in the 3D Slicer software.
A data-driven method was used to optimize this workflow. The scan sequence and 3D Slicer segmentation method was determined by analyzing a subset of three patient scans (one pre-operative and two post-operative). CT scans were categorized based on window (soft tissue vs bone) or view (axial, coronal, or sagittal) and the initial segmentation was compared to the refined segmentation with the Skull Stripper extension. The absolute error of the ICV measurement in the open-source software using each combination of these variables was computed, using the proprietary software ICV measurement for patient scan as the ground truth. The optimal combination of variables for the open-source method was moved forward for the rest of the analysis on the remaining CT scans.
The commercial software, Materialise Mimics was also used to measure intracranial volume on the same scan sequences as 3D Slicer (Supplementary Video 2). The slices were segmented using a threshold corresponding to bone (226 to maximum value). The mandibular volume was manually split from the mask. The remaining skull volume was exported as an STL part that was imported into the Materialise 3-matic software. The foramen magnum and all other holes in the part were closed by applying a surface normal to the edges of the hole. Then, the entire part was inverted to create a volume in the intracranial space that was measured in the 3-Matic software.
Statistics
All statistical analyses were performed with MATLAB R2020a. Multi-factor ANOVA was performed to identify the effect of CT scan window, scan view, and open-source software segmentation method on the ICV measurement error from the ground truth 3D systems ICV measurement. All effects were treated as fixed effects. Tukey’s test post-hoc analyses were performed for pairwise comparison of each combination of variables.
Random-intercept linear mixed modeling was used to determine the correlation between ICV measurements using open-source, commercial, and proprietary software methods (fixed effect), treating scan timing (pre-operative, post-operative, and long-term follow-up) and individual patient as random effects.
Results
Optimal Scan Parameters and Segmentation Method in Open-Source Software
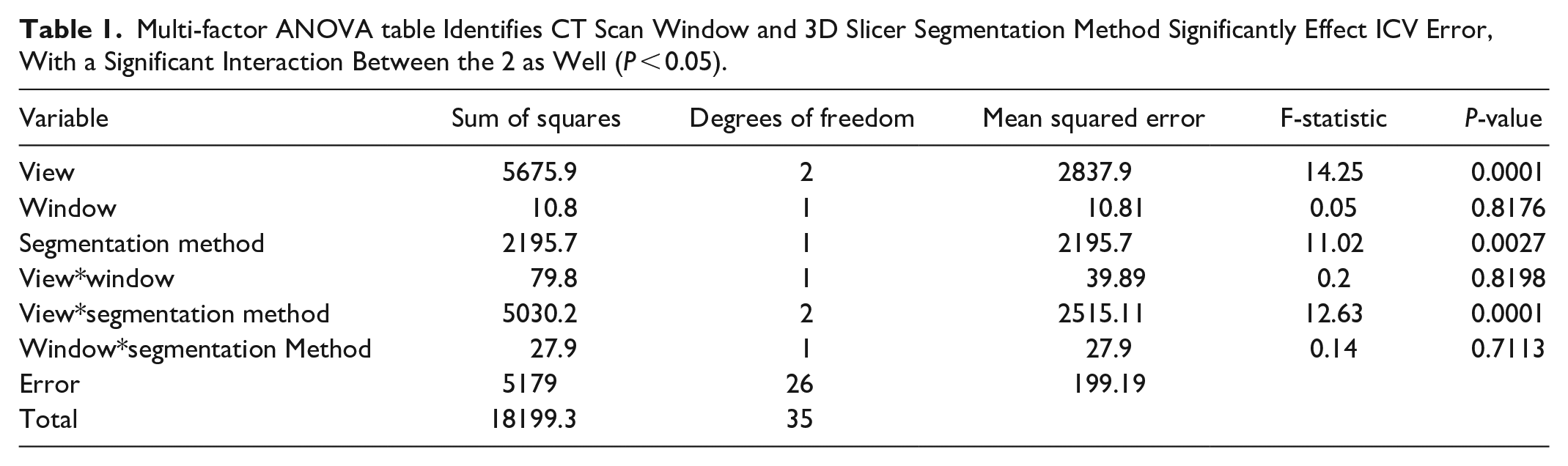
CT scan view and 3D Slicer segmentation method had a significant effect (P < .05) on the ICV measurement error from the proprietary software, whereas CT scan window did not have an effect. There was a significant interaction between CT scan view and segmentation method (Table 1). Post-hoc analysis showed that ICV measurements using the coronal view and the refined segmentation improved the ICV measurement error. Furthermore, the combination of coronal view and refined segmentation significantly improved the ICV measurement error (Figure 1). Based on this, CT scans with a soft tissue window and coronal reconstruction were used for ICV measurements for all patients in both the open-source and commercial software and the refined segmentation method was used for ICV analysis in the open-source software.
Multi-factor ANOVA table Identifies CT Scan Window and 3D Slicer Segmentation Method Significantly Effect ICV Error, With a Significant Interaction Between the 2 as Well (P < 0.05).
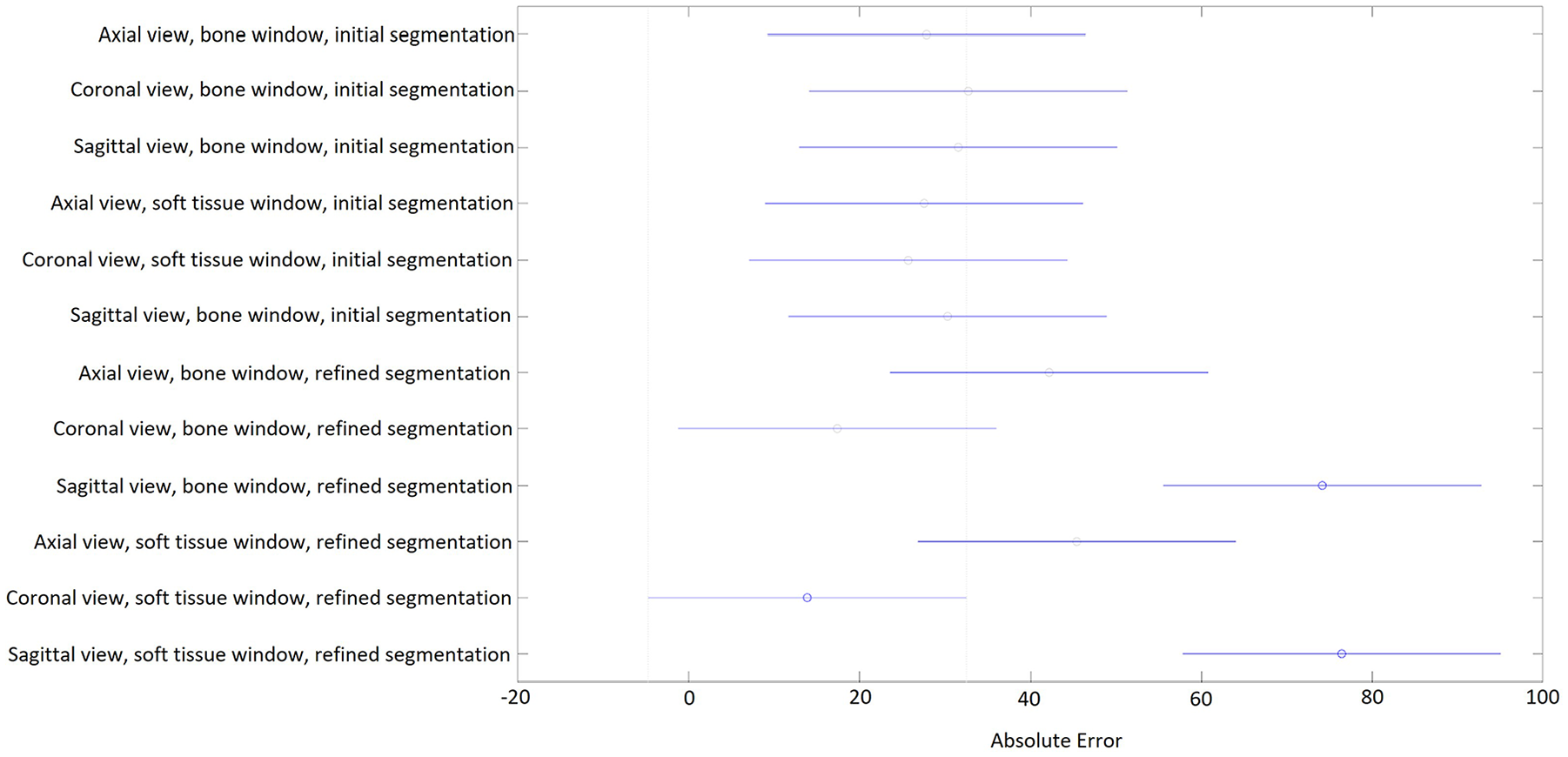
Post-hoc analysis using Tukey’s test shows that refined segmentation technique on the open-source software with coronal view reconstruction, using the soft tissue window had the lowest ICV error compared to the proprietary software. Mean error in cm3 (circles) and 95% confidence interval (bars) are shown.
Comparison of Intracranial Volumes
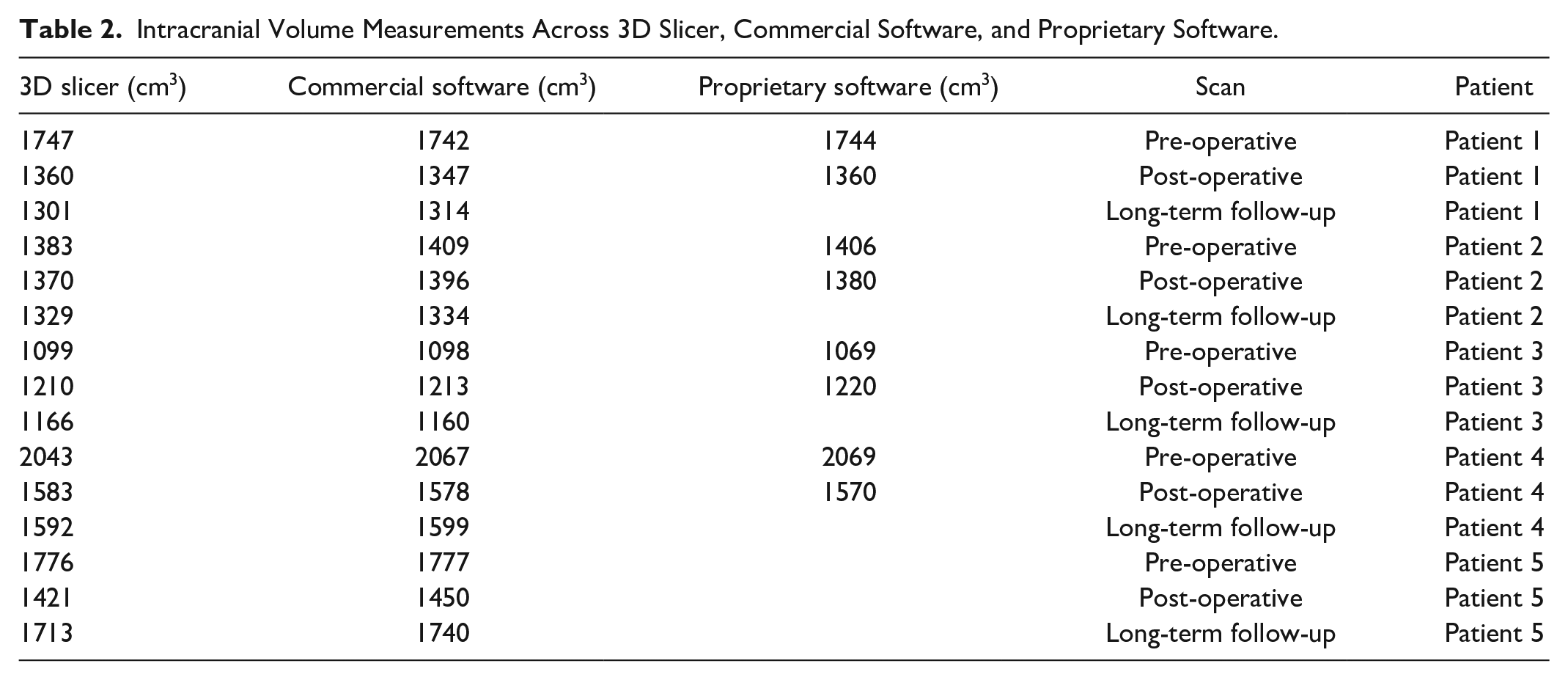
ICV measurements were performed using the open-source software and the commercial software for the 15 CT scans whereas only 8 scans were available for analysis with the proprietary software. Volume measurements are presented in Table 2. The ICV measurements across the 3 types of software were similar. The mean percent difference of ICV measurements of the open-source software from the commercial software was −0.56% (95% CI: −1.08%, −0.028%) and from the proprietary software was −0.07% (95% CI: −1.26%, 1.1%). The mean percent difference of ICV measurements of the commercial software from the proprietary software was 0.36% (95% Confidence Interval: −0.61%, 1.32%).
Intracranial Volume Measurements Across 3D Slicer, Commercial Software, and Proprietary Software.
ICV measurements by different software were also highly correlated (R2 = 0.998 for commercial against open-source software [Figure 2], R2 = 0.998 for proprietary against open-source software [Figure 3], R2 = 0.999 for proprietary software against commercial software [Figure 4]). Furthermore, there was nearly a one-to-one correspondence between ICV values from different software. For the prediction of commercial software ICV values from open-source ICV measurement, the slope of the best-fit line, βopen-source = 1.012 (95% CI: 0.986; 1.047). For the prediction of proprietary ICV values from open-source and commercial measurements, βopen-source = 1.025 (95% CI: 0.985;1.066), βcommercial = 1.016(95% CI: 0.986; 1.047). Power analysis shows that a minimum sample size of 4 scans is sufficient to detect the obtained effect sizes of R2 values with a significance criterion of α = 0.05 and power = 0.8. 20
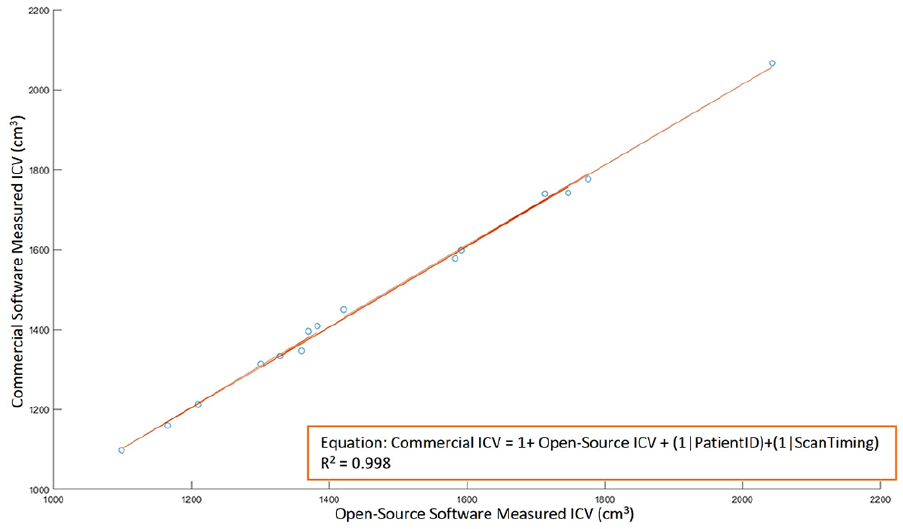
ICV measurements with open-source software and commercial software show good agreement (R2 = 0.998) with linear mixed-effects regression.
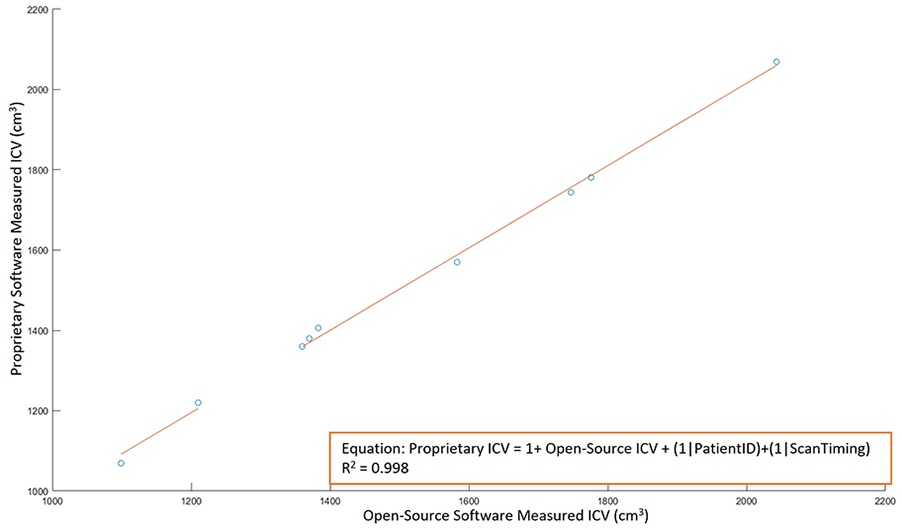
ICV measurements with open-source software and proprietary software show good agreement (R2 = 0.998) with linear mixed-effects regression.
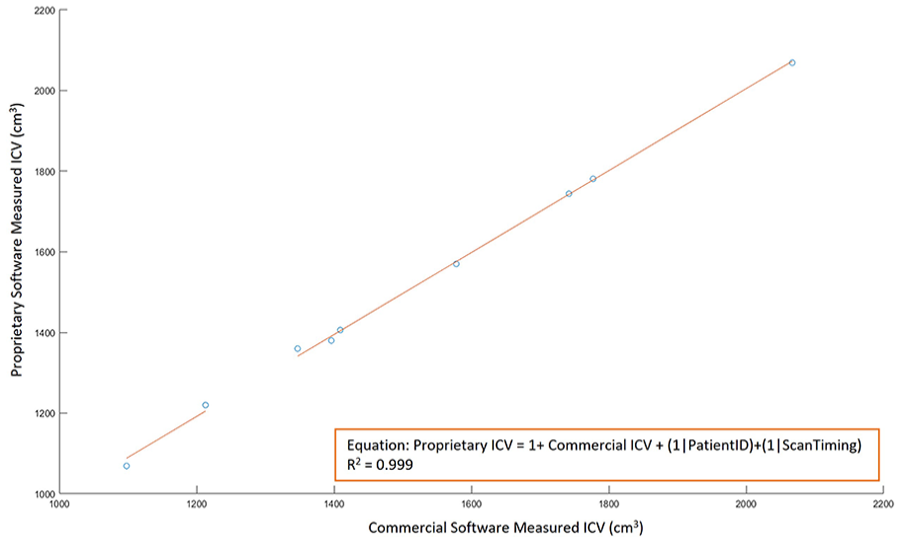
ICV measurements with commercial software and proprietary software show good agreement (R2 = 0.998) with linear mixed-effects regression.
Discussion
Clinical practice can benefit from accurate ICV measurements. For one, ICV measurements can be used as an aid for diagnosis. A clinical series of 84 children with craniosynostosis found that these children had markedly different ICV compared to an age-matched control group. 1 A series of 51 patients with clinically suspected normal pressure hydrocephalus were found to have elevated ICV compared to an age and sex-matched control group. 3 In addition, the deviation from normative values can give clues to the underlying pathology. Smaller ICV in prodromal Huntington’s disease patients may be caused by underlying neurodevelopmental abnormalities, for example. 21 As abnormalities in brain development can cause ICV changes, tracking ICV in children could also assess normal brain development. 22 These normative values and their changes with normal growth and development have been described in the literature. 23
Additionally, ICV measurements can be used for guiding treatment and monitoring outcomes, as an adjunct to other clinical data points, to influence pre-operative decision making. For example, among children who underwent corrective surgery for craniosynostosis, those with recurrent synostosis had a smaller pre-operative ICV compared to those with a favorable outcome, indicating that the synostosis was ongoing at the time of surgery. 1 One prospective series found that patients with syndromic synostosis had a larger increment in ICV following corrective surgery, compared to non-syndromic patients. 5 Another study identified that ICV increased after corrective surgery for patients with metopic or sagittal synostosis but not for unicoronal synostosis. 7 Additionally in this study, age at which the repair occurred was found to be a significant predictor of ICV change. ICV measurements can also be used to track surgical outcomes. A series of 18 patients with craniosynostosis found that corrective craniofacial surgery increased the patients’ ICV and subsequently, their ICV growth rate followed age and sex-matched controls. 4
In this study, we aim to make the measurement of ICV more accessible for clinical use. First, we address an unmet need in the literature by designing a method that utilizes CT scans, which are more common and affordable than MRI images. Second, we utilized an open-source software for ICV analysis. 3D Slicer is an open source software for medical image visualization and analysis through the DICOM format, which has been used for clinical research. Slicer contains core functions for image filtering, registration, segmentation, 3-dimensional model visualization, and support for image-guided therapy. Medical applications of 3D Slicer include quantitative analysis of MRI and PET images for monitoring treatment response of cancers, creating 3D-printed models for surgical planning, brain tumor segmentation, and morphological measurements of anatomical structures.24-27
The quoted cost of commercial software is in the range of tens of thousands of US dollars (Supplementary Table 1) and proprietary software is not available for public use. We validated our method against a gold standard commercial software and a proprietary software, explicitly demonstrating for the first time that an open-source method is equivalent to commercial and proprietary methods for intracranial volume analysis.
The main limitation for this study is that we do not have a ground truth measurement for ICV, even though we show good agreement across multiple methods using the same scans. This suggests that the error in ICV measurement would lie in the fidelity and resolution of the CT scans. Indeed, contrast-enhanced CT scans could be beneficial in intracranial volume segmentation, as demonstrated by one study. 28 However, this method cannot be applied to non-enhanced scans taken at a single time point. Commonly, manual segmentation of the same image, on a per-slice basis, was used as the ground truth in previous studies.29,30 However, this relies on the user’s ability to mark slices with high precision and accuracy and furthermore still relies on the quality of the image itself. It is also possible to create a simulated image with a known intracranial volume. However, it would be challenging to create a simulated model that captures all the possible pathophysiological variations in head shape, limiting the generalizability of the model to patient populations. A previous study compared a technique for manual segmentation from CT scans of dry skulls to the real volume of the skull. 31 However, this option was not available to us for this study.
Additionally, another limitation is that the open-source method is not fully automated, requiring still user interaction. This makes it challenging to load entire cohorts for high-throughput analysis. However, the open-source method can be performed in under 5 minutes per scan (Supplementary Video 1), with only intermittent user interaction in that period, making it acceptable for use within a single patient appointment. Most software tools that can measure ICV automatically are built for MRI scans. Free Surfer (http://surfer.nmr.mgh.harvard.edu/) and SPM (www.fil.ion.ucl.ac.uk/spm) are well-known examples. One study developed an automated method, CTSeg, for non-enhanced CT scans, by extending an SPM toolbox. 32 However, this was not yet validated on patients with cranial deformities.
While CT image acquisition offers benefits compared to MRI in the realm of speed and accessibility, it can also pose a hazard by exposing the patient to ionizing radiation. 33 It is possible to measure ICV without the use of tomography. One study developed a simple conversion between 3-dimensional (3D) photographs and intracranial volume, which could circumvent CT scans altogether. 23 They created a regression model that related the participant’s head volume, which can be measured with 3D photography, to their intracranial volume in a large population of subjects without cranial abnormalities. They achieved a good average error of approximately 4%. However further work will be needed to generalize the relationship between head volume and intracranial volume in different patient populations.
Conclusion
While ICV measurements are a useful adjunct to clinical decision-making, it can be difficult to perform these measurements due to access issues. For one, existing software packages are limited to MRI images, which are less commonly obtained than CT scans. Additionally, many packages are prohibitively expensive. There is a dearth of literature explicitly comparing open-source, commercial, and proprietary software packages for calculating ICV, making it difficult for teams to make an informed purchase. We present a method for ICV measurement from CT scans using an open-source software and validate it against existing commercial and proprietary software. This may be beneficial for improving access to ICV measurements for clinicians worldwide.
Supplemental Material
sj-docx-1-fac-10.1177_27325016241268300 – Supplemental material for Improving Access to Intracranial Volume Measurements for Clinical Use: Open-Source Method using Computed Tomography
Supplemental material, sj-docx-1-fac-10.1177_27325016241268300 for Improving Access to Intracranial Volume Measurements for Clinical Use: Open-Source Method using Computed Tomography by Akhil Surapaneni, Sarah A. Frommer and Raymond Harshbarger in FACE
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics Approval
This study was approved by our institutional review board (Study No. 00001996)
Informed Consent
General informed consent requirement was waived by the institutional review board.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
