Abstract
This study aimed to compare the 3D skull models reconstructed from computed tomography (CT) images using three different open-source software with a commercial software as a reference. The commercial Mimics v17.0 software was used to reconstruct the 3D skull models from 58 subjects. Next, two open-source software, MITK Workbench 2016.11, 3D Slicer 4.8.1 and InVesalius 3.1 were used to reconstruct the 3D skull models from the same subjects. All four software went through similar steps in 3D reconstruction process. The 3D skull models from the commercial and open-source software were exported in standard tessellation language (STL) format into CloudCompare v2.8 software and superimposed for geometric analyses. Hausdorff distance (HD) analysis demonstrated the average points distance of Mimics versus MITK was 0.25 mm. Meanwhile, for Mimics versus 3D Slicer and Mimics versus InVesalius, there was almost no differences between the two superimposed 3D skull models with average points distance of 0.01 mm. Based on Dice similarity coefficient (DSC) analysis, the similarity between Mimics versus MITK, Mimics versus 3D Slicer and Mimics versus InVesalius were 94.1, 98.8 and 98.3%, respectively. In conclusion, this study confirmed that the alternative open-source software, MITK, 3D Slicer and InVesalius gave comparable results in 3D reconstruction of skull models compared to the commercial gold standard Mimics software. This open-source software could possibly be used for pre-operative planning in cranio-maxillofacial cases and for patient management in the hospitals or institutions with limited budget.
Keywords
Introduction
Advances in craniofacial imaging have placed the importance on the 3D reconstruction of the skull model for medical applications. This technology has provided new possibilities to visualize complex medical data through generation of 3D skull models which were used for basic cranial education for medical students, 1 surgical training for surgeons, 2 pre-operative planning,3,4 facial contouring surgery, 5 forensic medicine and dentistry, 6 computer-assisted surgery, 7 maxillofacial prosthesis 8 and craniofacial reconstruction.9–12
Craniofacial reconstruction is commonly performed following head or facial trauma and on cancer patients who have lost part of the bony structures following tumour surgery. In current practice, the reconstruction of craniofacial defects is normally based on bone graft which is shaped to fit the defect. However, clinically, bone graft is limited to a small defect as the graft is taken from the patient’s own bone. With 3D skull model derived from CT data, pre-surgical planning can be done to fabricate an implant from compatible biomaterials such as titanium mesh or methylmethacrylate. 13 Using this technique, bigger and complex defect of the skull can be repaired.
The use of 3D reconstruction and 3D model reduces the possibility of errors during surgery, improves fit, and provides better implant stability. 14 In addition, the main advantage of 3D reconstruction is the possibility to simulate the surgical approach in the operating theatre, usually the supine position with lateral rotation of the head. 15 3D reconstruction also allows simulation of the various surgical approaches in order to select a minimal invasive approach during pre-operative planning.
The emerging of open-source software has shown an increasing trend16,17 because of the main advantage of being free and easily available, which significantly lowers the entry barrier to using it. Abdullah et al. 18 compared 3D reconstruction of segmented mandible between commercial software and open-source MITK software on five CT images. Results of the geometric differences showed average errors of two superimposed 3D models were less than 1% using Hausdorff distance (HD). However, these comparisons were performed on part of craniofacial region but do not cover the whole skull.
Most of the skull segmentation studies have utilised commercial software to create the meshed model or 3D model of the skull from patients’ CT data; for example, Mimics software,19–21 CATIA software 22 and Maxilim® software. 23 The 3D models created using skull segmentation software can be used for pre-operative planning or to design cranial implants. 24 However, most of the software mentioned in the literature is either commercial software or built in-house which were out of reach to researchers without big budgets or facilities.
In the medical imaging field, an accurate, reliable, and yet low-cost 3D imaging software for 3D reconstruction of the skull is crucial as it would have an impact on patient management that an appropriate software for skull segmentation is desirable. In this study, three open-source software, MITK, 3D Slicer and InVesalius were selected for the analysis based on their robustness, visualizations, reliability, and ease of use. Furthermore, this software was able to segment the 2D images and reconstructed them into 3D exportable models that are in STL files. An STL file describes the surface geometry of an object which can be sent to the 3D printers for printing of the skull.
The MITK software was available since 2003. 25 It has now evolved into a software system that can cover all steps of a clinical workflow including data retrieval, image analysis, diagnosis, treatment planning, intervention support and treatment control 26. The interface is easy to use, and the steps are user-friendly, especially to researchers or clinicians without an image processing background. The 3D Slicer software is a cross-platform and open-source software and features a flexible and mature plugin system. The community is very large and active. Compared to MITK, its approach is generally more application-centric though this has been addressed by the possibility of running modules on their own without the application on main window. On the other hand, the InVesalius software was made fully available as a free and open source via the Public Software Portal in November 2007.
The aim of this study is to assess the accuracy of 3D skull models reconstructed with commercial software and three open-source software. Geometric analyses were utilised to assess the differences and similarities between two skull models.
Methods
A sample population from January 2010 until December 2016 with CT head scans was obtained from the Neurosurgery Department, Hospital Universiti Sains Malaysia (USM), Kubang Kerian, Kelantan. Convenience sampling was used for this research. Data were collected from archived images of patients from the Picture Archiving and Communication System (PACS) server at the Radiology Department, Hospital USM. Ethical application was approved by the Ethics and Research Committee USM, reference number USMKK/PPP/JEPeM (259.3[2]).
Fifty-eight patients who had undergone CT scans with the CT images of 1-mm slice thickness and a matrix of 512 × 512 pixels were selected for this study. They were scanned using the Siemens Somatom Definition AS+ 128-slice (Siemens, Erlangen, Germany) at the Radiology Department, Hospital USM. The scans were acquired with 120 kVp. The slice thickness was 1.0 mm. For each patient, we reconstructed the scans with the same set of reconstruction parameters where the field of view (FOV) was 256 × 256 mm2. The pixel size = FOV/matrix (number of pixels) = 256 × 256/512 × 512 = 0.25 mm.
The CT data in DICOM format were imported into the medical imaging software (Materialise Mimics ver 17.0, MITK Workbench 2016.11, 3D Slicer 4.8.1 and InVesalius 3.1) for 3D reconstruction process. All four software were used to construct the 3D images from 2D cross-sectional images that were retrieved from the PACS server. Automatic segmentation of the skull was applied to create the 3D model using thresholding method. All four software went through similar steps in 3D reconstruction process, which is loading the DICOM format, segmentation using thresholding method (226 Hounsfield Unit) for 3D skull model creation as well as post-processing to eliminate the noises. In order to eliminate the noises, region growing method was used in Mimics and MITK, while largest island method and largest surface selection were applied in 3D Slicer and InVesalius, respectively. Finally the obtained 3D model was exported to standard tessellation language (STL) format as shown in Figure 1.
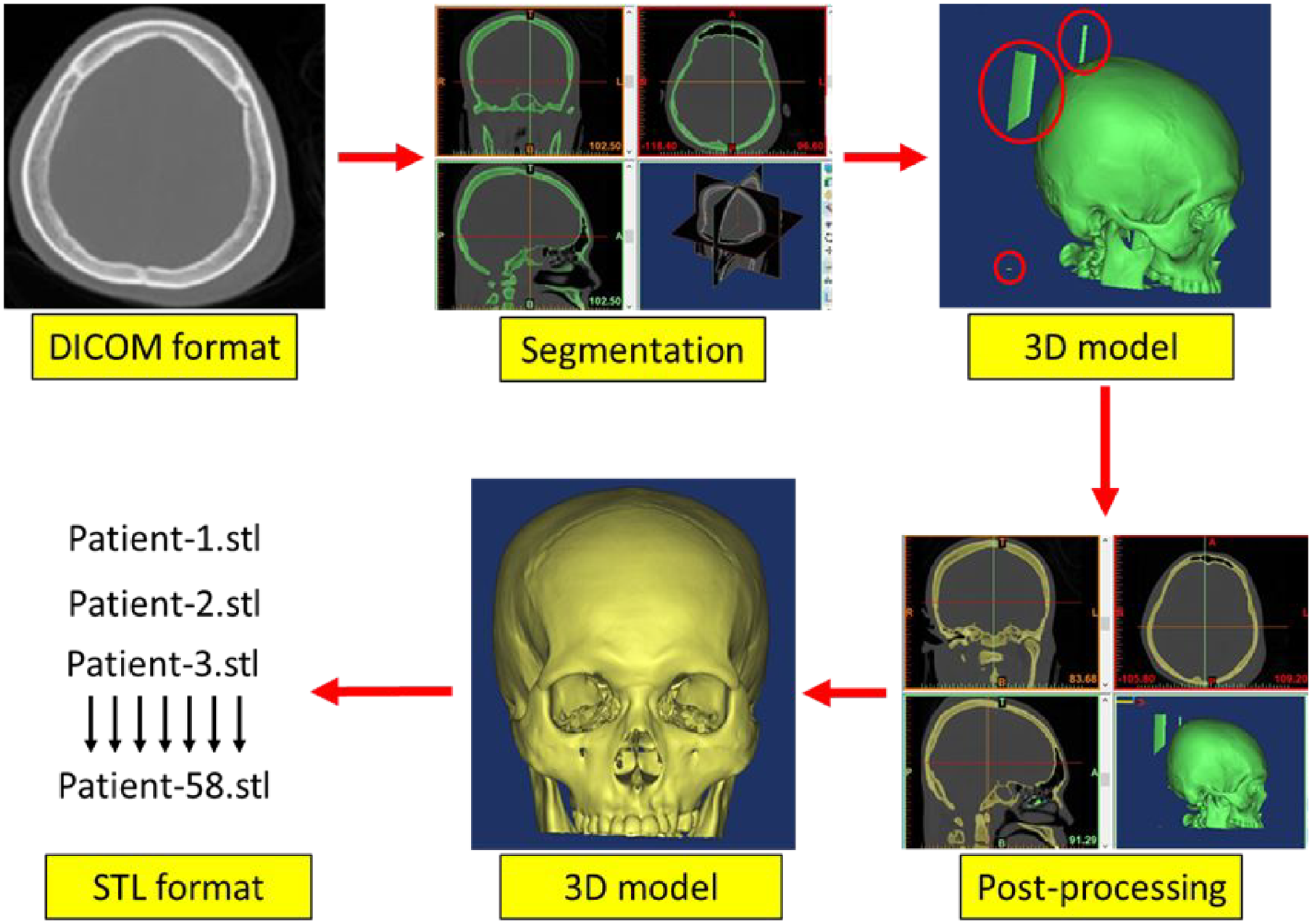
Summary of steps involved in 3D reconstruction of skull model for all four software.
All the data obtained in STL format were compared for accuracy analysis using analysis of variance (ANOVA) followed by Dunnet tests and geometric analyses (HD and DSC). The Statistical Package for the Social Science (SPSS) version 24.0 was used for statistical analyses in this study. Test of normality and homogeneity were carried out and confirmed using Kolmogorov-Smirnov test. One-way ANOVA was used to test for evidence of statistical differences in craniometric measurements of reconstructed 3D skull models. Dunnett tests were performed to find differences between each experimental group and the control group. A statistical significance value was set at p < 0.05.
In summary, four software; a commercial Mimics software and three open-source software (MITK, 3D Slicer and InVesalius) were utilised to reconstruct the 3D skull models of 58 patients as shown in Figure 2.
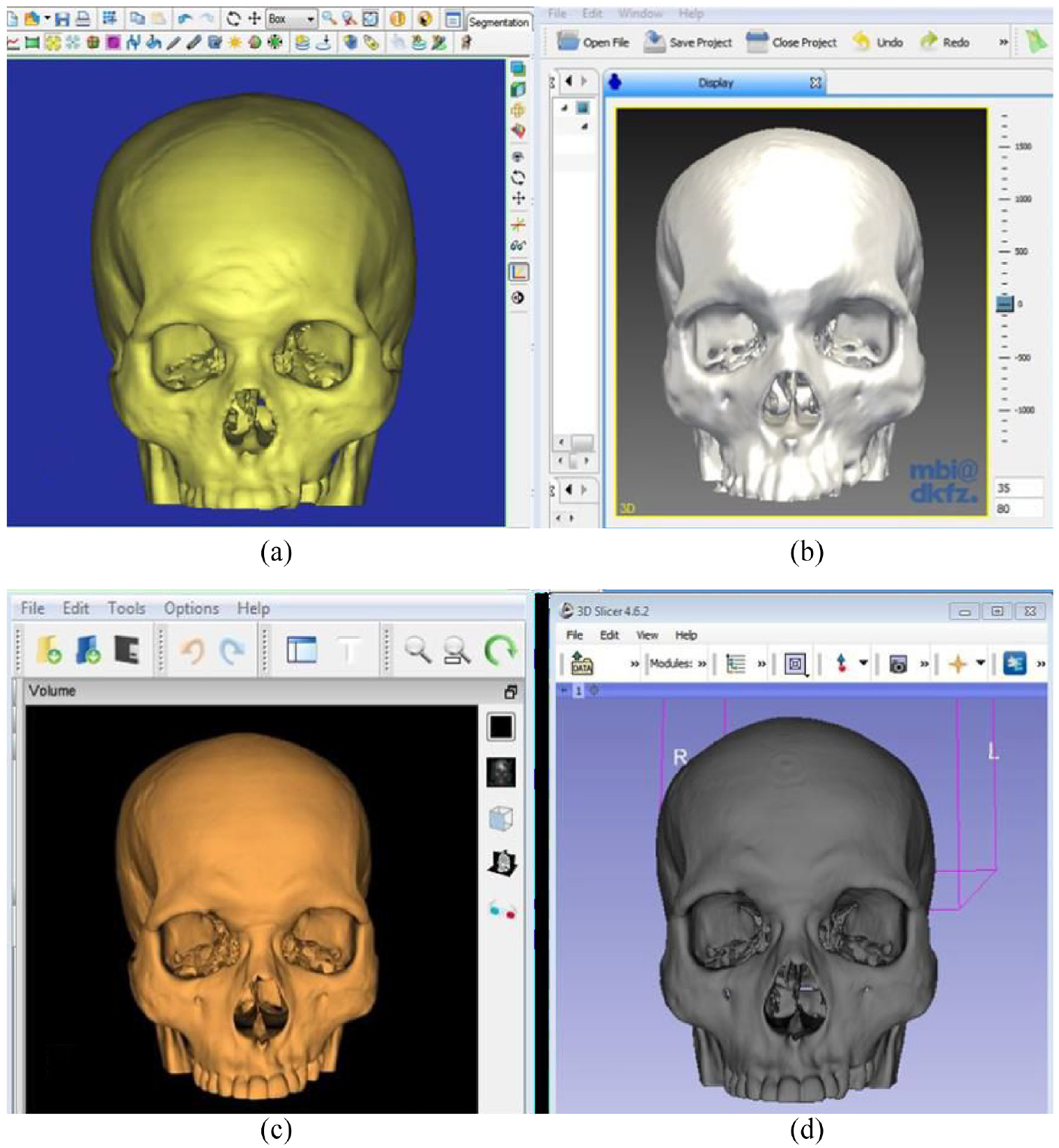
The 3D skull models produced using different software: (a) mimics, (b) MITK, (c) inVesalius and (d) 3D slicer are shown.
The accuracy of the 3D skull model reconstructed using the three open-source software (MITK, 3D Slicer and InVesalius) were compared with 3D skull model reconstructed using the commercial Mimics software. The comparison was made using metrics based on 3D geometry, which were HD and Dice similarity coefficient (DSC) using the open-source CloudCompare v2.8 software following the steps in Abdullah et al. 26 study as shown in Figure 3.
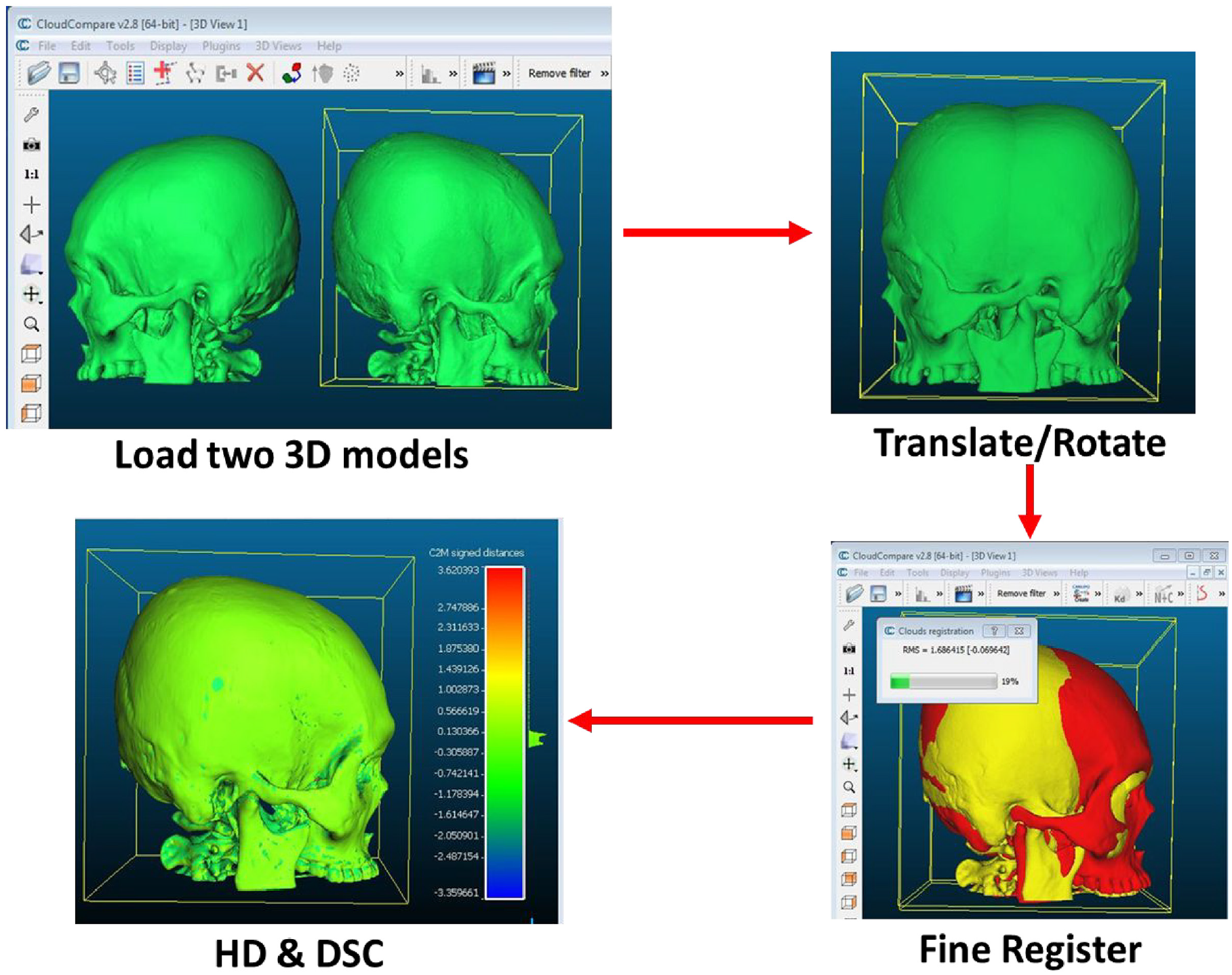
Geometric analyses of HD and DSC using CloudCompare v2.8 software.
Results
The 3D skull models segmented using Mimics software were used as the gold standard in this study. Similar segmentation processes were performed using MITK, 3D Slicer and InVesalius software. Results of the volume of 3D skull models segmented using four different software analysed using one-way ANOVA are shown in Table 1.
The 3D skull volume measurements using four different software.
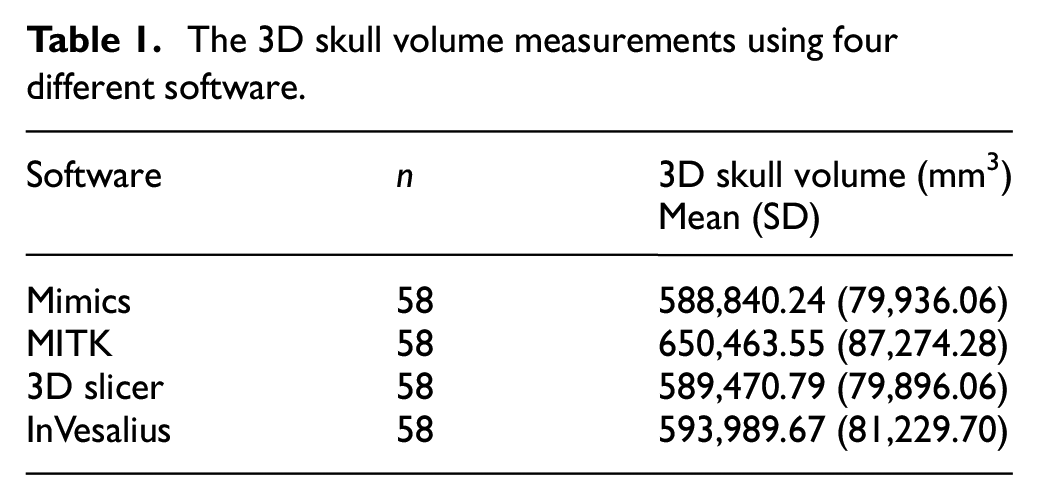
There were no statistically significant differences between volumes of the 3D skull models for all four software determined by one-way ANOVA [F (3, 228) = 1.606, p = 0.189].
Dunnett test comparing the volume of 3D skull models generated from three open-source software (MITK, 3D Slicer and InVesalius) against the control 3D skull models reconstructed using the commercial Mimics software is shown in Table 2.
Dunnett test of 3D skull volume.
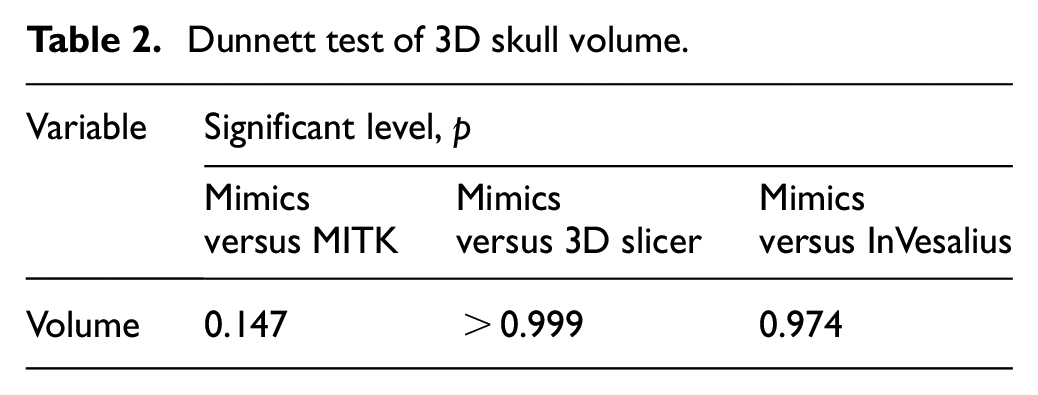

There were no significant differences in comparisons between Mimics and MITK, Mimics and 3D Slicer and Mimics and InVesalius with p > 0.05.
Geometric analyses using HD and DSC are shown in Tables 3 and 4.
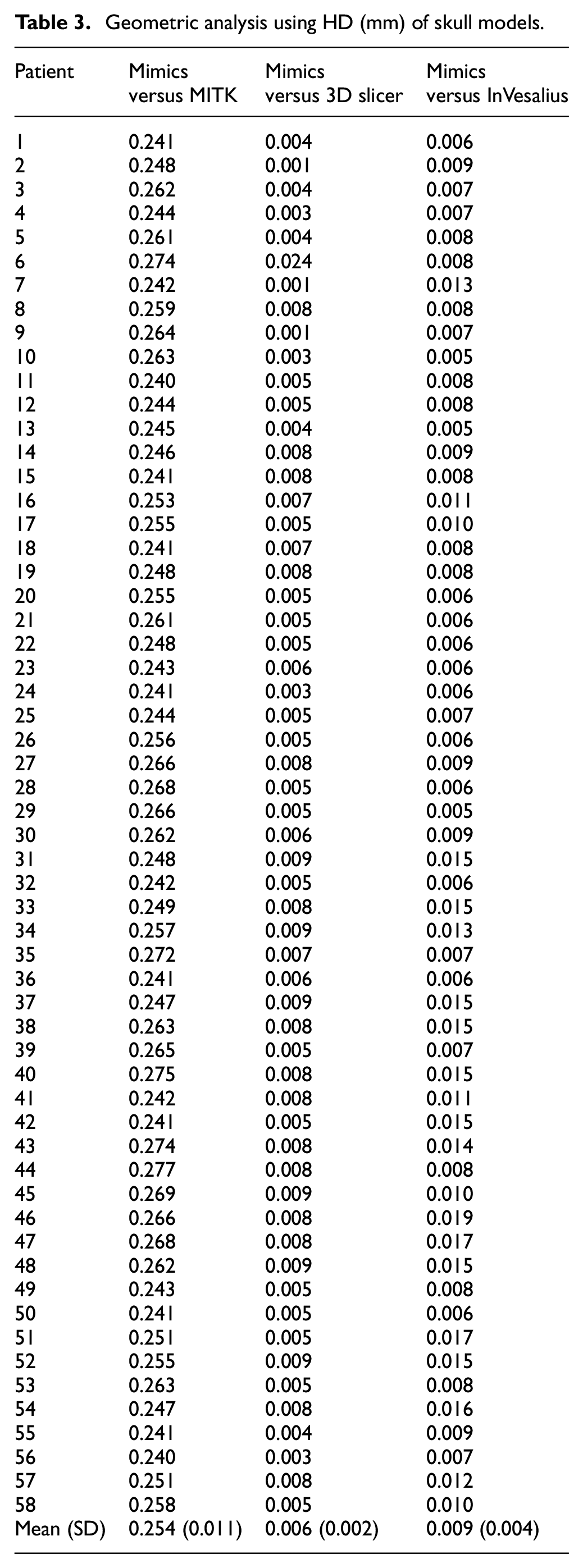
Geometric analysis using HD (mm) of skull models.
Geometric analysis using DSC (mm) of skull models.
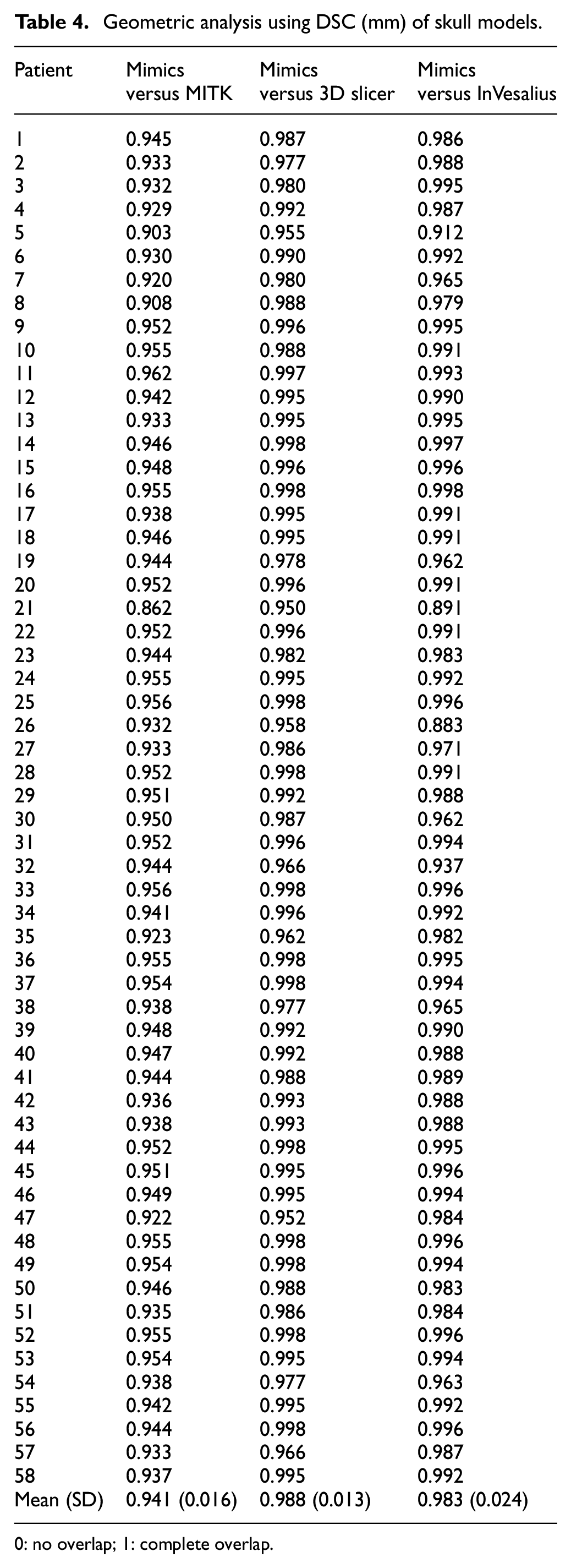
0: no overlap; 1: complete overlap.
Based on HD analysis, the average points distance from Mimics versus MITK were 0.25 mm while for Mimics versus 3D Slicer and Mimics versus InVesalius, there were almost no differences between the two 3D skull models with the average points distance at 0.01 mm.
Based on DSC analysis, the mean (SD) DSC was 0.941 (0.2) or 94.1% similar for MITK versus Mimics, and the mean (SD) DSC was 0.988 (0.01) or 98.8% similar for 3D Slicer versus Mimics, and mean DSC 0.983 (0.2) or 98.3% similar for InVesalius versus Mimics.
Discussion
In this study, the commercial Mimics software as well as open-source software of MITK, 3D Slicer and InVesalius were used to reconstruct 3D skull models from CT scan data. The volume of the 3D skull model reconstructed using the software were initially analysed prior to the geometric analyses using HD and DSC. The volume of the skulls generated from the open-source software were statistically insignificant as compared to the commercial software, which indicating that open-source software could generate accurate 3D models. It should be noted that anatomical structure includes internal cavities and intricate details that volumetric measurement is essential to represent the accurate size and shape of the respected skulls.
The commercial Mimics software used in this study was set as a reference or “gold standard.” It has been reported that HD value less than 3 mm and DSC value above 0.80 were considered adequate and accurate.27,28 The HD value from this study for Mimics versus MITK and Mimics versus 3D slicer were 0.25 and 0.01 mm, respectively. The results suggest that the 3D skull models reconstructed using the open-source MITK software is accurate, or similar to the gold standard. However, the 3D skull models reconstructed using the open-source 3D Slicer software were more accurate as compared to MITK. Another study 26 compared 50 skull models between Mimics versus InVesalius software, the HD value was 0.01 mm which is compatible to the results of 3D Slicer software.
The 3D skull models reconstructed from MITK, 3D Slicer and InVesalius have DSC values of 0.941, 0.988 and 0.983 when analysed against the gold standard. This showed that the skull models were 94.1, 98.8 and 98.3% similar to the gold standard, respectively. These findings showed that 3D skull models reconstructed from the open-source 3D Slicer software were the most accurate compared to other two software. Similarly, Abdullah et al., 26 compared the 3D reconstructed skulls using Mimics and InVesalius on 50 CT images with 97.6% similarity. reAlnaser et al. 29 evaluated seven open-source software for lung segmentation; two of the software were MITK and 3D Slicer. They evaluated the software based on the criteria such as functionality, usability, quality of segmentation and 3D export where the software must be able to export in STL format. Based on their study, MITK scored better than 3D Slicer. They reported MITK as a relatively easy to use and also regarded MITK as more user friendly than 3D Slicer. This study also revealed that MITK is more user-friendly than 3D Slicer. However, there are many learning resources available for 3D slicer which make it beneficial for beginners.
It should be noted that the cost of the commercial software is expensive, and often out of reach for university researchers, especially from public universities with limited grant. Moreover, the purchasing process was quite tedious and took many months for the public university before the software can be utilised, as there were lots of red tape involved. Once purchased, it can only be used by one user at one time (for one license). This would be a major setback for a big group of researchers in a university where they have to queue and wait for their turn to use the software, as the additional license would incur more cost to the university.
One of the advantages of the three open-source software utilised in this study is that the segmentation and post-processing of the images can be applied by an automatic process without manual intervention; therefore, human errors were minimised. Thus, these methods are reliable, repeatable and reproducible by other researchers from other research institutions. However, the involvement of clinicians is necessary for complex cases to identify the region of interest for better clinical outcomes.
Although 58 skulls were successfully reconstructed, limitation such as the presence of CT artefacts of noises required additional processing (Figure 4). In Mimics and MITK software, region growing method was performed to isolate the skull from other unrelated objects. Meanwhile, the largest island method was used in 3D Slicer to isolate the skull. These different methods were used as each software has its algorithm to isolate the different types of hard tissues or objects with a similar range of HU. Region growing method was carried out to select the region of interest by selecting only the connected region with similar pixels value. It removed the detached pixels from the segmentation. This is to ensure that the exact part of interest was selected. On the other hand, largest island algorithm determined the biggest group of connected pixels. Both region growing and largest island methods were useful tools after thresholding method to isolate the skull from other unrelated objects such as gantry and noises.
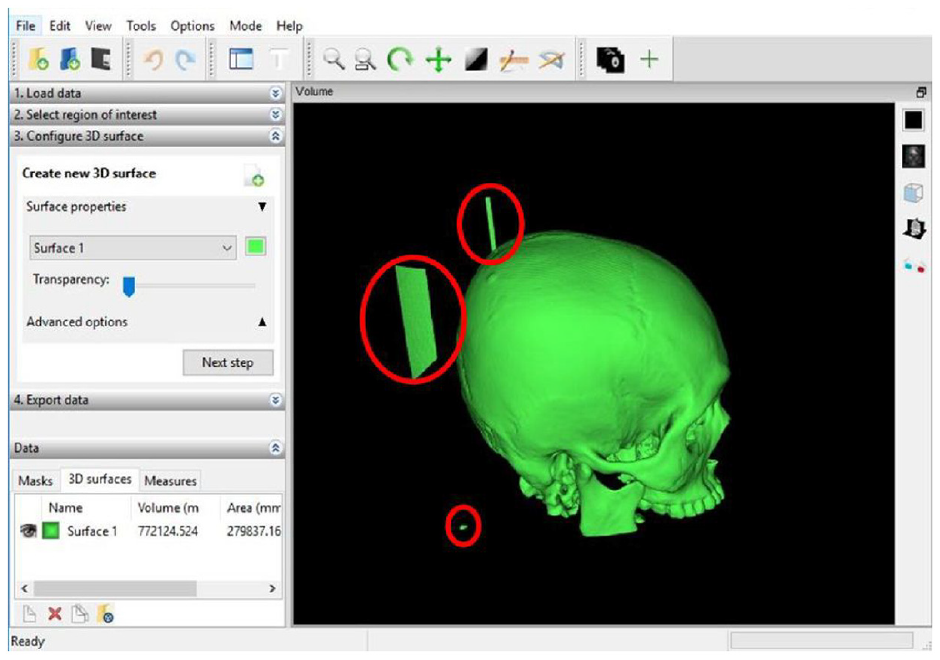
Unrelated objects (in red circles) that need to be removed.
Region growing method was carried out to select the region of interest by selecting only the connected region with similar pixels value. It removed the detached pixels from the segmentation. This is to ensure that the exact part of interest was selected. On the other hand, largest island algorithm determined the biggest group of connected pixels. Both region growing and largest island methods were useful tools after thresholding method to isolate the skull from other unrelated objects such as gantry and noises. Therefore, the algorithm provided in this two open-source software is sufficient to segment the 3D skull models and produced equivalent results when compared with the commercial software. Meanwhile, an almost similar study was conducted where the accuracy of craniomaxillofacial region of mandibular jaw was assessed using Mimics as well as four software of Invesalius, ITK-Snap, Dolphin 3D and Slicer 3D. 30 Their results also revealed that the software were able to define the contour of the mandibular bone which indicated the reliability of the software.
Conclusions
In conclusion, this study confirmed that the alternative open-source software, MITK, InVesalius and 3D Slicer gave comparable results in 3D reconstruction of skull models compared to the commercial gold standard Mimics software. This open-source software could possibly be used for pre-operative planning in cranio-maxillofacial cases and for patient management in the hospitals or institutions with limited budget.
Footnotes
Acknowledgements
We would like to thank Mr. Firdaus Daud from the Radiology Department, Hospital USM for his helpful assistance in this project.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was made possible with funding from TDC Holdings Sdn Bhd through Universiti Sains Malaysia (304.PPSG.6150194/T152).
Data availability statement
The data used to support the findings of this study are available from the corresponding author upon request.
