Abstract
To disentangle genetic and environmental contributions to individual differences, classical twin models (CTMs) are predominantly used. Prior research revealed several conditions under which CTMs spawn biased heritability and environmentality estimates, including the simultaneous relevance of nonadditive genetic and shared environmental influences, the presence of assortative mating, and passive gene-environment covariance. Nuclear twin family models (NTFMs), adding information from parents of twins, can handle these conditions. Little is known, however, on the actual impact of such conditions considering the empirical properties displayed by various personality-related phenotypes. The present study therefore investigated differences between NTFMs and CTMs in terms of error-adjusted heritability and environmentality across 93 personality-relevant constructs based on data from 636 twin pairs and 320 parents. Overall, NTFMs resulted in significantly lower heritability and higher environmentality estimates, suggesting an overall overestimation of heritability in CTMs. These trends pertained to about two-thirds of the investigated constructs, but reversely directed changes were detectable under specific conditions. We discuss the extent to which the assumptions underlying CTMs might bias conclusions regarding genetic and environmental contributions to personality differences and demonstrate benefits of NTFMs. Furthermore, we provide user tutorials for implementing CTMs and NTFMs based on the open-source and user-friendly statistical program JASP.
Keywords
To disentangle the genetic from environmental differences in personality-relevant constructs, behavior genetic models make use of data from twins and family members. In this manner, they allow for a decomposition of the phenotypic differences (i.e., observed or measured variance) into genetic and environmental components and thus enable estimations of heritability and environmentality (Neale & Maes, 2004). The majority of behavioral genetic research relies on classical twin models (CTMs) based on information from monozygotic (MZ) and dizygotic (DZ) twin pairs (Polderman et al., 2015). Crucially, previous research demonstrated several conditions under which the constrained assumptions underlying CTMs result in biased estimates of genetic or environmental variance (Keller & Coventry, 2005; Keller et al., 2010).
The inclusion of information from further relatives has been proposed to mitigate such biased estimates. Within the nuclear twin family model (NTFM; Heath et al., 1985), information from twins’ parents in addition to data from twins themselves are considered. As such, the NTFM was elaborated as an appropriate compromise between accuracy and simplicity (Keller et al., 2010). Empirical studies comparing the estimates of genetic and environmental variance resulting from NTFMs as opposed to CTMs predominantly focused on specific phenotypes only (e.g., Bleidorn et al., 2018; Wolfram & Morris, 2023). Besides phenotype-specific comparisons between NTFM- and CTM-based heritability estimates and a review about two decades ago (Coventry & Keller, 2005), an overall empirical picture on the extent to which NTFMs indeed provide more accurate estimates of genetic and environmental contributions to individual differences in various personality-related phenotypes is lacking. Against this background, the current study examined differences between NTFMs and CTMs regarding heritability and environmentality estimates across 93 personality-relevant constructs. Firstly, we expound some basic concepts of behavioral genetics and corresponding empirical findings. Then, we explain assumptions underlying CTMs and NTFMs before introducing the current study with more details.
Some Basic Concepts and Findings of Behavioral Genetic Research
In quantitative behavioral genetic research (see Knopik et al., 2017, for an introduction), the concept of heritability refers to the proportion of phenotypic variance in a given sample that can be attributed to genetic differences. For one thing, such genetic differences can arise from additive genetic factors, meaning that multiple gene variants add up in their effects on phenotypic variance. The proportion of phenotypic variance due to additive genetic variance only is usually referred to as narrow-sense heritability. The broad-sense heritability further incorporates nonadditive genetic variance, which can result from interaction effects of gene variants within same gene loci (allelic dominance deviation) or between different gene loci (gene-by-gene interaction or emergenesis). From previous empirical findings it can be derived that gene-by-gene interaction rather than dominance deviation contributes to the variance in complex polygenic characteristics (Polderman et al., 2015; Vukasović & Bratko, 2015). In the current work on personality-relevant constructs that are proposed to be polygenic, we thus operationalized nonadditive genetic factors in terms of gene-by-gene interaction (see also Kandler, Zapko-Willmes et al., 2021). As MZ twins are genetically identical, they share all additive and nonadditive genetic factors that contribute to differences between unrelated individuals. All first-degree relatives, including DZ twins, show, on average, 50% overlap of additive genetic factors and no overlap of nonadditive genetic factors due to gene-by-gene interaction effects (Lykken, 2006).
Likewise, the concept of environmentality can be defined as the proportion of phenotypic variance attributable to differences in environmental experiences or contextual circumstances. These can be shared by different family members and thus contribute to their similarity. For example, they can be shared exclusively by family members within generations (e.g., siblings) or by all family members within and across generations (i.e., between parents and their offspring). Consequently, the so-called nonshared environmental experiences that are specific to each individual would only contribute to the dissimilarity between any family members.
Empirical behavioral genetic findings connote substantial genetic influences on variance in complex personality-relevant constructs (see Polderman et al., 2015, or Kandler, Zapko-Willmes et al., 2021, for overviews). 1 Mentioning some examples, broad-sense heritability estimates of core personality traits usually range between H2 = .40 to .60, while the environmental variance appears to be mainly due to individual-specific factors (Kandler & Papendick, 2017; Vukasović & Bratko, 2015). Individual differences in social values or attitudes seem to be somewhat less genetically driven but suggest some environmental influences shared by family members instead (Kandler et al., 2014; Nguyen et al., 2021).
Most of such behavioral genetic findings rely on CTMs. As these models do not include parental information, they cannot consider that parents might provide familial rearing environments according to their own and thus in part to their children’s genetic predispositions. Such a nonrandom link between genetic and family environment factors, referred to as passive gene-environment covariance (Plomin et al., 1977), could contribute to individual differences in personality-relevant characteristics as indicated by prior investigations (e.g., Briley et al., 2018; Wolfram & Morris, 2023).
Furthermore, parents of twins might choose one another not completely at random but based on their phenotypic (dis)similarity, referred to as assortative mating (Heath & Eaves, 1985). Assortative mating is related to genetic and environmental contributions to personality differences: It might either induce that twins’ parents are, to some extent, genetically similar to one another, increasing the probability of their DZ twin children (but not of their genetically identical MZ twins) being genetically similar beyond 50%, or it might enhance the non-genetic within-family resemblance (see also Briley et al., 2019). Both of these circumstances would affect differences between individuals in an entire population. Meta-analytic findings indicate considerable assortative mating for various constructs, ranging up to about r = .50 for some values and attitudes (Horwitz et al., 2023).
Previous studies demonstrated that including parental information in twin models resulted in somewhat different conclusions on the genetic and environmental contributions to the variance in specific constructs, such as self-esteem (Bleidorn et al., 2018), educational attainment (Wolfram & Morris, 2023), or core personality trait domains (Boomsma et al., 2018; Kandler, Richter, & Zapko-Willmes, 2019). These findings hint at the relevance of extending behavior genetic models beyond data from twins only, considering additional potentially contributing factors, such as shared environmental influences in the presence of nonadditive genetic influences, passive gene-environment covariance, and assortative mating (see Coventry & Keller, 2005, for a review).
Classical Twin Models
Using information from MZ and DZ twins reared together, CTMs allow to estimate sources of individual differences based on comparisons of the observed covariances within these two different types of twin pairs (Neale & Maes, 2004). Thus, CTMs rest upon the different genetic relatedness of MZ as opposed to DZ twins, as well as the assumption that environmental experiences shared by twins contribute equally to the similarity within both MZ and DZ pairs (Kendler et al., 1993). Against this background, two main derivations can be drawn from the covariances of MZ and DZ twins reared together. First, only individual-specific (nonshared) environmental experiences can account for differences within MZ twin pairs. Second, differences between the covariances of MZ and DZ twin pairs can be solely attributable to genetic factors, so that larger differences imply larger genetic influences on the variance in a given phenotype.
(Co)Variances Underlying Classical Twin and Nuclear Twin Family Models.
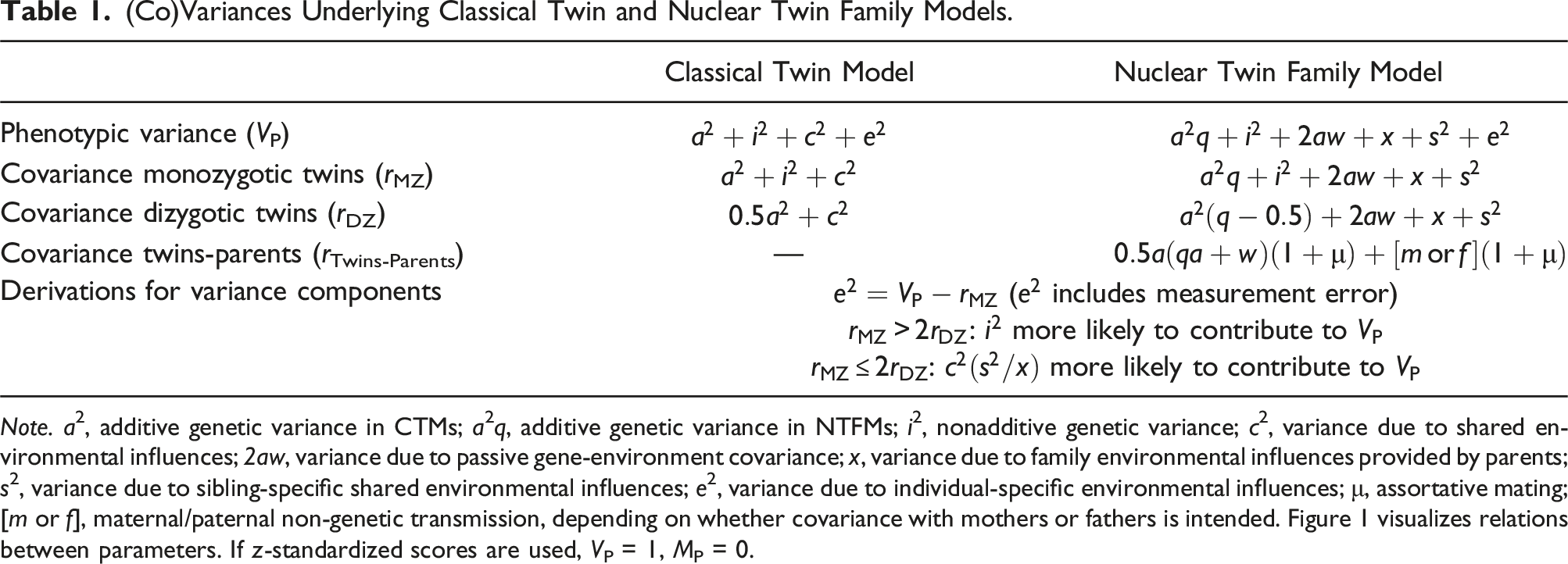

Note. a2, additive genetic variance in CTMs; a2q, additive genetic variance in NTFMs; i2, nonadditive genetic variance; c2, variance due to shared environmental influences; 2aw, variance due to passive gene-environment covariance; x, variance due to family environmental influences provided by parents; s2, variance due to sibling-specific shared environmental influences; e2, variance due to individual-specific environmental influences; μ, assortative mating; [m or f], maternal/paternal non-genetic transmission, depending on whether covariance with mothers or fathers is intended. Figure 1 visualizes relations between parameters. If z-standardized scores are used, VP = 1, MP = 0.

Classical Twin and Nuclear Twin Family Structural Equation Models. Note. Violet rectangles represent observed variance for each group of family members. Pink circles refer to genetic factors and green circles to environmental factors. A (a), additive genetic factors (effects); I (i), nonadditive genetic factors (effects) due to gene-by-gene interaction; C (c), shared environmental factors (effects); S (s), sibling-specific environmental factors (effects); FE, environmental factors due to family environment provided by parents; E (e), individual-specific environmental factors (effects); w, covariance between A and FE; μ, assortative mating; m/f, maternal/paternal non-genetic transmission; q, variance of A; x, variance of FE. Variances of all other latent variables are fixed to 1. Bold parameters indicate parameters set differently for monozygotic (before the slash) and dizygotic twins (after).
Nuclear Twin Family Models
Including information from parents in addition to MZ and DZ twins, NTFMs are based on the covariance between twins and their parents in addition to the covariances within twins (see Table 1) that are informative on genetic and environmental sources of variance (Heath et al., 1985). The structural equation model in Figure 1(b) includes the different components that can contribute to the phenotypic variance. In addition to A, I, and E components that are also specified in CTMs, NTFMs further disentangle C into environmental influences due to sibling-specific experiences that contribute to twin-similarity only (S), or due to family environmental circumstances provided by the parents that increase the similarity among all family members (FE). Also, NTFMs consider passive gene-environment covariance as a nonrandom link between A and FE factors (w) as well as effects of assortative mating (μ).
NTFMs go beyond the restrictive assumptions underlying CTMs in different respects (see also Bleidorn et al., 2018; Kandler et al., 2024, for overviews; and Keller et al., 2009, for details). First, nonadditive genetic factors can be estimated in the presence of shared environmental influences, assuming the latter to be attributable to either sibling-specific or family-environmental effects. Fundamentally, in CTMs, if both nonadditive genetic and shared environmental influences constitute relevant sources of variance, but only one of them is considered in the model, their respective contributions would be underestimated, whereas additive genetic variance would be overestimated (Keller & Coventry, 2005). Since NTFMs are based on three different covariances, two within twins and one between twins and their parents, 2 to estimate four factors of family resemblance (A, I, S, and FE), they also require one of the three I, S, or FE factors to be set at zero. This restriction, however, only pertains to the specific origins of shared environmental variance so that such constraints result in comparably less biased heritability and environmentality estimates (see Keller et al., 2009, for details).
Second, as parental information is not involved in CTMs, they cannot cover potential effects of assortative mating. Positive assortative mating induces genetic similarity of parents, increasing the overlap of additive genetic factors in DZ twins beyond 50%. In CTMs, however, the correlation of A factors is fixed to be r = .50 within DZ twins. The only way through which CTMs can address the extent to which DZ twins are genetically more similar than 50% thus runs via the perfectly correlated C factors. Consequently, CTMs can underestimate the contribution of genetic factors to the variance in case of positive assortative mating, while overestimating shared environmental influences (Keller et al., 2010). In NTFMs, assortative mating is considered as the co-path μ between both parental phenotypes. To account for the possibility that assortative mating impacts the additive genetic variance (q), the parameter q is defined by a nonlinear constraint. That is, this parameter is not estimated but determined by other estimated parameters and their interrelations (Keller et al., 2009). Given that in general the variance of the latent factors equals 1, the additive genetic variance would be higher than 1 if
Likewise, assortative mating can impact the variance attributable to family environmental influences (x), and the covariance (w) between additive genetic and family-environmental factors. The latter condition involves a third limitation of CTMs. As passive gene-environment covariance increases the similarity within MZ and DZ twins to a similar degree, its presence would result in overestimations of shared environmental influences in CTMs (Briley et al., 2018, 2019; Purcell, 2002). In NTFMs, the covariance w between the A and FE variables can be interpreted in terms of passive gene-environment covariance, since w forms a function of the parental phenotype that is genetically linked to the twins’ phenotype and associated with the environment provided by the parents (Kandler, Richter, & Zapko-Willmes, 2019).
The variance attributable to the family environment (x) results from maternal (m) and paternal (f) transmission. Both contributions can be different from one another, as there are two different covariances between either mothers or fathers and twins underlying NTFM-based estimations (see Table 1). Thus, x results from tracing the pathways between both parental and the twins’ FE variables, forming a further nonlinear constraint:
The Current Study
Prior research demonstrated mainly three crucial conditions under which the less constrained assumptions underlying NTFMs enable less biased estimates of heritability and environmentality than do CTMs: the simultaneous relevance of nonadditive genetic and shared environmental influences, the presence of assortative mating, and passive gene-environment covariance (Keller et al., 2010). Within the current study, we aimed at gaining a broad picture across a representative set of 93 personality-relevant constructs on the extent to which NTFMs and CTMs come to different conclusions regarding heritability and environmentality.
Applying genetically informative structural equation models to empirical data from twins and their parents, we tested whether the genetic and environmental contributions to the variance were estimated significantly different by NTFMs and CTMs, considering unreliability due to random error of measurement. Furthermore, we sought to demonstrate the consequences of the aforementioned crucial conditions regarding their impact on heritability and environmentality estimates for selected personality-relevant constructs. In addition, we initially implemented genetically informative structural equation models with the use of the open-source and user-friendly statistical program JASP (JASP Team, 2024). Therefore, we intended to provide a corresponding introduction and supply user tutorials for CTMs and NTFMs based on JASP as part of the current work.
Methods
Transparency, Openness, and Reproducibility
We provide information on the measures used, report criteria for data inclusions, and describe our statistical analyses. In this respect, we adhere to the Journal Article Reporting Standards (Appelbaum et al., 2018). The current study was not preregistered, and we did not make specific hypotheses. Data analytic scripts, codes, and reduced anonymized input data files as well as supplemental material files are openly available: https://osf.io/db465/. The entire data set is available as Scientific Use File that can be requested after completing a Data Use Agreement via the homepage of the SPeADy project: https://www.speady.de/contact/?lang=en. The data were used by Kandler et al. (2024) and parts of it in six former studies (see Supplement A). All studies pursued different foci not demonstrating differences between CTMs and NTFMs.
Sample
The sample was drawn from the Twin Family Study of the Study of Personality Architecture and Dynamics (SPeADy; Kandler, Penner, et al., 2019). The SPeADy research project has been approved by local ethics committees. Each participant provided written informed consent. We used data from complete twin pairs, encompassing 573 pairs of the first wave of data collection to which 63 pairs that newly participated at the second wave were added. Overall, the sample thus comprised data from 636 complete twin pairs, aged between 14 and 88 years (Mage = 38.45), with 343 being MZ (78% female) and 293 DZ (49% female same-sex; 38% opposite-sex). This sample of twin pairs was used as a basis for both CTM and NTFM analyses. For the NTFM analyses, we additionally considered data from mothers (aged 37–87 years) for 203 pairs and data from fathers (aged 35–85 years) for 117 pairs. Data from both mothers and fathers were available for 99 pairs (16%), so that 122 pairs (19%) offered data from one parent. Consequently, the entire sample comprised data from N = 1592 different individuals. The sample can be seen as heterogeneous with regard to sex, age, or education, but not as representative for the German population.
Measures
All constructs have been assessed as part of a survey questionnaire encompassing several instruments for measuring personality-relevant characteristics (Kandler, Penner, et al., 2019). In the current work, we used all the constructs assessed within SPeADy, which cover 50 measures of personality traits and corresponding facets based on the Big Five trait taxonomy and the HEXACO model, six major life goals, six interests, five moral concerns, 19 value-priorities, six self-schemas, and religiousness. In this manner, we obtained a broad range of constructs that relate to individual differences in personality-related aspects (Kandler et al., 2014). By means of self-reports, participants evaluated their individual level of agreement to the respective items on Likert-scales of different length. Measures of the 93 constructs were assessed for measurement invariance across twins and parents (see Kandler et al., 2024). More detailed information on the applied measures and reliability estimates can be found in Supplement B.
Analyses
General Procedures
Initially, the data were corrected for age and sex differences using linear regression procedures (McGue & Bouchard, 1984) and the z-standardized residuals were further used. Therefore, the entire phenotypic variance to be decomposed in our analyses equals 1. Genetically informative structural equation models were performed using JASP, Version 0.18.3 (JASP Team, 2024). For both CTMs and NTFMs we used the full sample of complete twin pairs (N = 636). By fitting all models on the basis of full information maximum likelihood estimation (Little & Rubin, 2019), we used all available data for model parameter estimates. For each construct, we estimated the intra-class correlations (ICCs) within MZ and DZ twin pairs as well as between twins and parents. We used the ratios of these twin correlations as one criterion to decide whether to consider nonadditive genetic or shared environmental variance components in the respective models (according to the heuristics stated in Table 1). As a second criterion for model selection, we compared the fit indices of models including different variance components. In particular, we compared two CTMs excluding either I or C (i.e., ACE and AIE models) and three NTFMs either excluding I or S or constraining m and f to be zero, thus, excluding FE (i.e., ASFE, AIFE, and AISE models). These models were compared descriptively by the Akaike Information Criterion (AIC; Akaike, 1987) and the Bayesian Information Criterion (BIC; Schwarz, 1978). We used those models offering the descriptively best fit as indicated by lower AIC and BIC values.
Classical Twin and Nuclear Twin Family Modeling
CTMs and NTFMs were specified as two-group models comprising either data from MZ or DZ twins (and parents). Any factor loadings, intercepts, and means were constrained to be equal across zygosity groups and family members. Usually, variances of the latent variables were fixed to 1, so that the genetic and environmental contributions could be ascertained by squaring up the respective path coefficients. Exceptions form the latent variables A (with variance defined by q; contribution by a2q) and FE (with variance and contribution defined by x) in the NTFMs. For plausible estimations, we constrained path coefficients related to variance components to positive values. We provide a detailed explanation on the implementation of NTFMs and CTMs in JASP at: https://osf.io/db465/.
Between-Model Comparisons of Genetic and Environmental Variance
To obtain heritability and environmentality estimates corrected for (possible) attenuation, we computed sum-scores per construct and model considering the internal consistency coefficient McDonald’s ω of their respective scale. A construct’s heritability was thus defined as the amount of reliable phenotypic variance attributable to any genetic sources (i.e., broad-sense heritability, H2) including additive and nonadditive factors as well as the contribution of assortative mating (i.e., q–1):
We treated each construct as a specific case and ran paired sample t-tests as well as Wilcoxon signed-rank tests based on the 93 variables as units to evaluate whether the heritability or environmentality estimates differed significantly in central tendency between CTMs and NTFMs. Furthermore, we compared additive and nonadditive genetic variance between both models. Environmentality tests were not distinguished further, as potential between-model differences of Env2 can be solely attributable to estimates of shared environmental sources (since e2 always equals VP–rMZ; see Table 1). To identify if the results replicate depending on whether or not personality-relevant constructs are conceptualized in a narrow or rather broad sense, we specified two different categories and tested for between-model differences within these categories separately. For one thing, we considered narrow-sense (or potential core) personality traits, including Big Five and HEXACO personality trait domains and facets (n = 50). For another thing, we considered broad-sense (or potential surface) personality characteristics, including life goals, interests, moral concerns, value-priorities, self-schemas, and religiousness (n = 43). Details on this conceptual differentiation between core and surface characteristics of personality can be found in Kandler et al. (2014) or Kandler, Penner, et al. (2019). Based on the results, we scrutinized the impact of crucial conditions for differences between CTM-based and NTFM-based heritability or environmentality estimates.
Results
General Results
The ICCs for twins and parents are reported in Supplement C. Supplemental Table D1 depicts the fit indices of the differently reduced models, highlighting those that were finally used to evaluate the genetic and environmental variance estimates. Mostly, NTFMs excluding shared environmental sources due to family transmissions fitted the data best (n = 68), from which for 58 constructs CTMs excluding C were applied. For 22 constructs, NTFMs excluding sibling-specific shared environmental sources provided the best fit, from which 21 were specified as CTMs excluding C. For three constructs, NTFMs and corresponding CTMs excluding nonadditive genetic factors offered the best fit to the data.
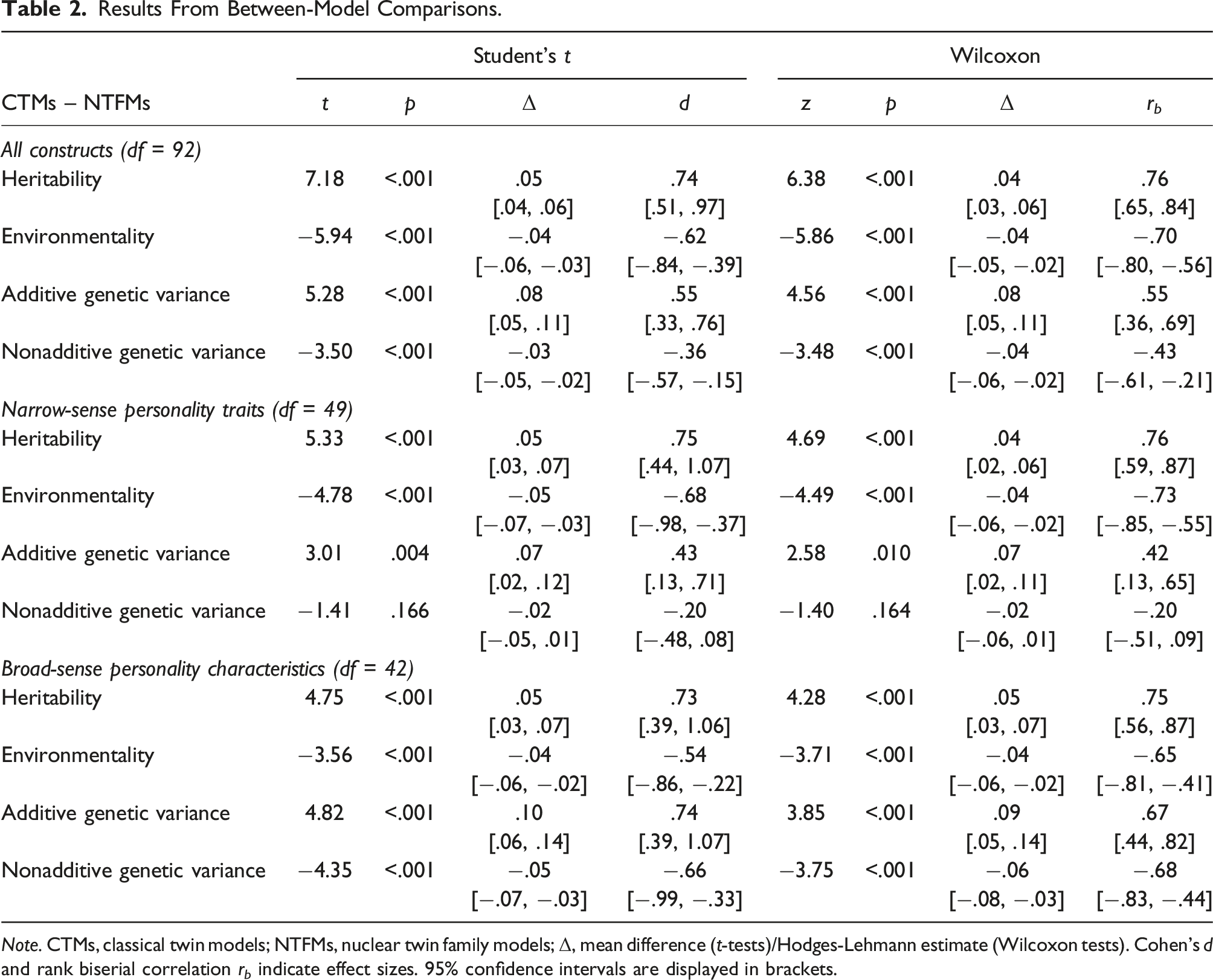
Between-Model Comparisons of Heritability and Environmentality
Figure 2 illustrates the error-adjusted heritability and environmentality estimates for the 93 constructs based on CTMs and NTFMs and their relation to one another. The entire parameter estimates and variance components are provided in Supplement D and E. Across all constructs, t-tests and Wilcoxon tests indicated significant mean and median differences for both heritability and environmentality of medium effect sizes (see Table 2). In particular, the CTM-based heritability was significantly higher in central tendency (5%) compared to the NTFM-based heritability. Likewise, the CTM-based environmentality was significantly lower (−4%). The higher CTM-based heritability primarily pertained to the additive genetic variance, as t-tests and Wilcoxon-tests implied equally directed between-model differences of 8%. By contrast, nonadditive genetic variance was estimated lower by CTMs (−3%). Similar between-model differences of comparable effect sizes were detectable when examining narrow-sense core personality traits or broad-sense personality characteristics separately: In both categories of personality constructs, t-tests and Wilcoxon tests indicated significantly higher CTM-based heritability estimates across the respectively included constructs, primarily due to different estimates of additive genetic variance, and significantly lower CTM-based environmentality estimates (see Table 2). Error-Adjusted Heritability and Environmentality Estimates. Note. CTM, classical twin model; NTFM, nuclear twin family model; H2, heritability; Env2, environmentality. Estimates are corrected for attenuation/inflation due to measurement error. Pink (H2) and green (Env2) lines indicate between-model differences in line with the overall trend. Gray lines indicate differences <1%, black lines indicate reversely directed differences. Results From Between-Model Comparisons. Note. CTMs, classical twin models; NTFMs, nuclear twin family models; Δ, mean difference (t-tests)/Hodges-Lehmann estimate (Wilcoxon tests). Cohen’s d and rank biserial correlation r
b
indicate effect sizes. 95% confidence intervals are displayed in brackets.
In line with the overall trends across all constructs, lower (higher) NTFM-based heritability (environmentality) estimates at least as high as the mean difference of 5% (−4%) were implied for 43 (41) constructs, and of at least 1% for 63 (61) constructs. Twelve (14) constructs showed reversely directed differences. Please note that these numbers should imply a descriptive overview on the observable trends rather than (statistical) inferences. Both heritability and environmentality estimates were highly correlated between CTMs and NTFMs across the 93 personality variables (see Figure 2).
Regarding the three crucial conditions previously outlined as being associated with biases in CTMs (please find an overview in Supplemental Table D4), assortative mating was significant for 30 constructs (range: μ = −.19 to .61), whereas passive gene-environment covariance accounted for at least 1% of the variance in 14 constructs (range: 1%–17%). For 38 constructs the simultaneous relevance of nonadditive genetic and shared environmental sources was indicated, as both explained variance (≥5%) in corresponding NTFMs.
Based on six constructs, we exemplified the differences between CTMs and NTFMs regarding the estimates of genetic and environmental variance components (see Figure 3). Furthermore, Figure 3 depicts the ICCs of the different dyads of family members that are informative on the covariance structure on which the estimations of CTMs or NTFMs are based. The six constructs were selected as they are well suited to demonstrate the effects that the three crucial conditions or their combinations yielded on heritability and environmentality estimates depending on the model used (see Discussion for more details). Equal to Figure 3, the variance components of all 93 constructs are shown in Supplemental Figures E1–E6. Standardized Variance Components and Intra-Class Correlations for Specific Constructs. Note. Pink and green bars depict different variance components. Black and white bars depict ICCs for different dyads of family members (i.e., for MZ, monozygotic twins; DZ, dizygotic twins; M-T, mothers-twins; F-T, fathers-twins). CTM, classical twin model; NTFM, nuclear twin family model; a2, additive genetic variance in CTMs; a2q, additive genetic variance in NTFMs; i2, nonadditive genetic variance; c2, shared environmental variance; 2aw, variance due to passive gene-environment covariance; x, variance due to family environmental influences provided by parents; s2, sibling-specific shared environmental variance; e2, individual-specific environmental variance; μ, assortative mating. Estimates are corrected for attenuation/inflation due to measurement error. Supplement D provides uncorrected estimates and confidence intervals. *p < .05.
Discussion
Based on empirical data across 93 personality-relevant constructs, NTFMs resulted in significantly lower error-adjusted heritability and higher environmentality estimates than CTMs. While these trends pertained to about two-thirds of constructs, reversely directed differences were detectable under specific conditions. In the following, we discuss the differences between CTM-based and NTFM-based estimates and their consequences for conclusions on genetic and environmental contributions to individual differences.
Between-Model Differences in Heritability and Environmentality
As the NTFM-based heritability estimates were significantly lower for most personality-relevant constructs investigated in our study, the relevance of genetic contributions to personality differences appeared to be overestimated by CTMs, overall. This is in line with prior empirical findings (Coventry & Keller, 2005). Initially, the mean difference of 5% suggests that CTMs suffice to gain an idea on whether genetic factors explain phenotypic variance at all, supporting the derivations by Keller et al. (2010). Nonetheless, for several constructs the heritability estimates differed more considerably between NTFMs and CTMs than implied by the mean difference of 5%. This is crucial, as one does not necessarily know in advance if the sources of variance in the examined constructs might induce biased conclusions on genetic and environmental variance if relying on findings from CTMs only. The six examples in Figure 3 illustrate the potential impact that specific conditions might have for heritability and environmentality estimates.
For instance, the twins’ ICCs of the HEXACO trait domain agreeableness (Figure 3) implied the inclusion of nonadditive genetic factors (I) in the CTM. Derivable from the applied NTFM including sibling-specific shared environmental factors (S) in addition, however, both sources explained considerable variance (>10%). As the used CTM excluded shared environmental factors (C) in the presence of I, the entire resemblance of DZ twins could only be accounted for via the a-path (Figure 1(a)). Consequently, the resulting H2 estimate reflected, in part, variance attributable to shared environmental influences.
It has been argued that the application of CTMs excluding I might circumvent that heritability estimates are inflated (Keller & Coventry, 2005). However, the second example of the trait facet flexibility outlines the consequences of not considering I, albeit of relevance, in CTMs. Since nonadditive genetic influences increase the covariance among MZ but not DZ twins, this difference could only be addressed via the a-path – also spawning up-biased heritability estimates. As such, the covariance structure of MZ and DZ twins that is derivable from ICCs and underlies CTMs does not indicate that one source of variance indeed approaches zero (Ozaki et al., 2011).
A reversely directed difference between CTM-based and NTFM-based heritability estimates was found, for instance, for tradition values. Significant positive assortative mating derivable from the NTFMs can explain this trend (Briley et al., 2019; Keller et al., 2010). Since assortative mating increases the genetic similarity within DZ twins beyond 50%, it reduces differences between the ICCs of MZ and DZ twins. In the applied CTM, the extent to which the DZ exceeded half the MZ twin resemblance was thus covered via the c-path – but was in fact not much driven by shared environmental influences. Correspondingly, the NTFM considering assortative mating yielded a substantially higher heritability.
Negative assortative mating appeared to contribute to down-biased CTM-based estimates of additive genetic variance as well, but did not impact broad-sense heritability estimates, which can be seen in the example of the Big Five trait domain neuroticism. Derivable from the nonlinear constraint defining q, DZ twins share less of additive genetic factors as a consequence of negative assortative mating, increasing the difference between the ICCs of MZ and DZ twins. The used CTM accounted for this via the i-path – reducing the relative proportion of additive genetic variance. Including parental information and thus the covariance between twins and parents, the NTFM could consider that parents and any child always share 50% of additive genetic factors, as opposed to DZ twins (see Keller et al., 2009, for details) – enabling to approach the relevance of additive genetic variance more accurately.
Across all constructs, we found the highest assortative mating for religiousness but detected no between-model differences in terms of additive genetic variance. This demonstrates that assortative mating might impact biased CTM-based estimates of additive genetic variance under the condition that the latter explains considerable variance in general. As it can be seen in Figure 3, the additive genetic contributions to variance (indicated by a2/a2q) were larger for tradition values than for religiousness. Besides, the impact of assortative mating could be covered by variance due to non-genetic familial transmission. In NTFMs, positive assortative mating increases x, leading to lower proportions attributable to additive genetic variance. Accordingly, our findings for religiousness demonstrate that different conditions could balance each other out regarding biases between CTM-based and NTFM-based estimates.
Looking at the Big Five trait domain extraversion, the twins’ ICCs and CTM-results suggested that additive genetic factors contribute to 32% of phenotypic variance. The NTFM-results, however, implied that the primary amount of genetic variance was due to nonadditive genetic factors and that the entire DZ twin similarity was attributable to familial environmental transmission. Given that additive genetic factors would influence the covariance between twins and mothers or twins and fathers to a similar degree, the differences detectable between the corresponding ICCs support this implication.
Passive gene-environment covariance was indicated for only about 15% of investigated constructs, but was also related to between-model differences. Since passive gene-environment covariance increases the resemblance within MZ and DZ twins to a similar degree (Plomin et al., 1977), this condition can be compensated in CTMs via the c-path. For religiousness, the NTFM allowing for a more fine-grained examination of the sources of non-genetic twin-similarity showed that the CTM-based environmentality did not reflect the amount of variance solely attributable to environmental influences. Instead, c2 was confounded with variance in fact attributable to the covariance between A and FE factors (i.e., 2aw). This, in turn, can be regarded as reflecting variance due to genetic predispositions becoming salient by environmental circumstances (Kandler, Zapko-Willmes et al., 2021).
Implications of Between-Model Differences in Heritability and Environmentality
Previous research mainly elaborated the overestimation of additive genetic variance in CTMs compared to NTFMs (Coventry & Keller, 2005; Keller & Coventry, 2005). Accordingly, the found between-model differences regarding estimations of the broad-sense heritability were primarily due to additive genetic factors. This becomes particularly relevant considering the discourse on gaps between heritability estimates derived from behavioral genetic twin studies and alternative ways to approach heritability, such as genome-wide association studies (Bourrat & Lu, 2017). Our results imply that CTMs, on which most heritability estimates rely, might convey an up-biased picture of the genetic foundations of personality differences, which might contribute to – albeit not explain – this discrepancy. This underlines that CTM-based estimates should be viewed as rough approximations only.
Similar between-model differences were implied when considering narrow-sense traits (incl. Big Five and HEXACO trait concepts) and broad-sense characteristics of personality (incl. values, motives, self-schemas, and interests), separately. Hence, the conceptual differences elaborated for these two categories of constructs (Kandler et al., 2014, 2024) appear to be not systematically related to the extent of biases in CTMs compared to NTFMs. Instead, these trends support overall overestimations of genetic contributions to personality differences by CTMs. Furthermore, as those constructs showing a simultaneous relevance of nonadditive genetic and shared environmental sources consisted roughly half of narrow-sense traits and half of broad-sense characteristics, this condition seems to be no unique feature for the one or the other. This aligns with those few studies comparing NTFM-based and CTM-based estimates of specific core traits (i.e., neuroticism; Boomsma et al., 2018) and other personality-relevant characteristics (i.e., self-esteem; Bleidorn et al., 2018). In our results, the relevance of passive gene-environment covariance did not differ depending on whether or not conceptualizing the personality-relevant constructs in a broad sense. However, prior research outlined that passive gene-environment covariance might play a greater role for educational or cognitive constructs (Briley et al., 2018; Wolfram & Morris, 2023). Of those constructs for which assortative mating was significant, two thirds could be considered as broad-sense personality characteristics. This aligns with meta-analytic findings which suggest that assortative mating might be most relevant for values or social attitudes (Horwitz et al., 2023). Our results also hinted at assortative mating for various value-priorities or moral concerns.
Altogether, our findings indicate that specific properties displayed by a given construct do not always come along with an overestimation or underestimation of heritability and environmentality by CTMs since, empirically, various properties and their interplay can account for genetic and environmental variance estimates simultaneously. The benefits of NTFMs do not imply that their application exceeds the use of CTMs throughout all occasions. Notably, our findings on high between-model correlations of heritability and environmentality estimates indicate that comparable inferences on the relations between the amount of genetic and environmental variance of different constructs can be drawn from both. Hence, CTMs might be as useful as NTFMs in multivariate applications. According to the consideration by Briley et al. (2019), we emphasize that the merits of specific models and thus the choice which one to use strongly depends on the research question of interest. Therefore, we provided user tutorials for both CTMs and NTFMs with the use of JASP for the research community (see https://osf.io/db465/).
Limitations and Future Research
Although NTFMs have been found to provide a better approximation of heritability and environmentality than CTMs, the incorporation of data from other family members might further increase the accuracy of the estimates (Keller et al., 2010). For example, the Stealth or the Cascade model make use of data from twins’ spouses and children (Keller et al., 2009). As this additional information enhances the covariance structure underlying such models, they enable a more fine-grained disentanglement of the sources of phenotypic differences and within-family similarities. However, this is often accompanied by a lower precision of parameter estimates: Stealth and Cascade models require sufficient information from each group of family members, which is rarely accomplished. As such, NTFMs were elaborated as an appropriate compromise between spawning less biased estimates (than CTMs) and requiring more complex data collections (than Stealth and Cascade models; Keller et al., 2010). Nonetheless, the extent to which more extended behavior genetic models come to similar or different conclusions than NTFMs should prospectively be evaluated using empirical data.
Moreover, NTFMs are based on the assumption that genetic and environmental contributions to individual differences are equal for the parental generation and the offspring generation (Keller et al., 2009). For various personality-relevant constructs, however, genetic and environmental variance was stressed to shift throughout development, implying different heritability estimates for children and their parents (Kandler, Bratko, et al., 2021). Due to reasons of model identifications, these trends can hardly be considered in NTFMs. Related to this, the models discussed in the present study cover the general limitations that they estimate the net contributions of genetic and environmental variance to individual differences. Hence, they do not cover additional types of gene-environment interplay, which might still be confounded with heritability and environmentality estimates (Briley et al., 2018). Future studies including quantifiable measures of the environment and their interaction with genetic factors (Purcell, 2002) are thus required to investigate how these circumstances impact NTFM-based estimates. Likewise, both discussed models assume that shared environmental factors contribute equally to the similarity within MZ and DZ twins (Kendler et al., 1993). However, MZ twins might share more environmental experiences than DZ twins, which could lead to an overestimated heritability in twin models.
Finally, we focused on the differences between CTMs and NTFMs regarding the relative contributions of certain genetic and environmental variance components to personality differences. Hence, we did not primarily focus on the statistical significance of specific parameter estimates. Prior research, however, demonstrated that the sample size and the ratios of differently related family members might impact the statistical power to detect certain genetic and environmental effects – and these impacts might differ between CTMs and NTFMs (Medland & Keller, 2009). It could be interesting for future research to test whether the presently found between-model differences of CTMs and NTFMs vary as a function of the sample size or the sample structure.
Conclusion
The current study extended previous research by demonstrating that CTMs and NTFMs come to significantly different conclusions regarding heritability and environmentality estimates considering empirical information on 93 personality-relevant constructs. Broad-sense heritability and especially additive genetic variance were overestimated while environmentality was underestimated by CTMs. Not only the possibility to estimate shared environmental and nonadditive genetic influences simultaneously but also the possibility to consider the potential presence of assortative mating or passive gene-environment covariance constitute factors making the application of NTFMs beneficial over and above CTMs. Altogether, when aiming at deciphering genetic and environmental contributions to individual differences, the inclusion of information from parents in addition to twins appears to enable not only a more fine-grained differentiation of the single sources of variance but also a more accurate approach to estimate broad-sense heritability and environmentality.
Supplemental Material
Supplemental Material - Benefits of Nuclear Twin Family Models for Estimating Heritability: An Investigation of 93 Personality-Relevant Constructs Using JASP
Supplemental Material for Benefits of Nuclear Twin Family Models for Estimating Heritability: An Investigation of 93 Personality-Relevant Constructs Using JASP by Jana Instinske and Christian Kandler in Personality Science
Supplemental Material
Supplemental Material - Benefits of Nuclear Twin Family Models for Estimating Heritability: An Investigation of 93 Personality-Relevant Constructs Using JASP
Supplemental Material for Benefits of Nuclear Twin Family Models for Estimating Heritability: An Investigation of 93 Personality-Relevant Constructs Using JASP by Jana Instinske and Christian Kandler in Personality Science
Footnotes
Author Note
The handling editor of this research paper was Dr. Cristian Zanon, Federal University of Rio Grande do Sul, Brazil.
Acknowledgments
We thank the whole SPeADy team, in particular the (former) team members Yannik Wiechers, Alexandra Zapko-Willmes, Julia Richter, Jantje Bollmann, Michael Papendick, Angelika Penner, Elif Yalcin, Annika Overlander, Sophie Walter, Jana Willemsen, Corinna Eickes, Fynn Plugge, Hannah Sarnizei, Paula Wundersee, Rebecca Gruzman, Pauline Hirche, Julia Schneider, Kai Tippelt, and Felix Butt for their important contributions to the research conducted within the course of the SPeADy research project. Furthermore, we thank the study participants for taking the time to participate in the SPeADy research project.
Author Contributions
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present research based on the SPeADy research project that was supported by grants from the German Research Foundation [grant number: 272981829] awarded to Christian Kandler.
Data accessibility statement
All material (i.e., data analytic scripts and codes) used for this research paper as well as supplemental material files are openly available and can be accessed and downloaded at: ![]() . Similarly, the anonymized input data files that are necessary for the model analyses to reproduce can be found there: https://osf.io/db465/.
. Similarly, the anonymized input data files that are necessary for the model analyses to reproduce can be found there: https://osf.io/db465/.
Supplemental Material
Supplemental material for this article is available online. Depending on the article type, these usually include a Transparency Checklist, a Transparent Peer Review File, and optional materials from the authors.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
