Abstract
Background:
Preeclampsia (PE) is a complication of pregnancy characterized by hypertension, with limited therapeutic options and variable treatment response. Identifying novel drug targets is urgently needed.
Method:
We performed Mendelian randomization (MR) to explore potential drug targets for PE using data from the UK Biobank (nCase = 184, nControl = 361,010) and the FinnGen database (nCase = 7,377, nControl = 211,957). Genetic instruments for pQTLs were obtained from five proteomic studies. Bayesian colocalization analysis and summary-data-based MR (SMR) analysis were performed to assess the causal relationship between two related signals (protein levels and PE risk). We further conducted single-cell type expression analysis and phenome-wide MR. In addition, a protein–protein interaction network of key genes was constructed via the GeneMANIA website.
Results:
At a significance level of p < 5 × 10−8, MR analysis revealed seven protein-PE pairs. Gene prediction indicated that KDEL (Lys-Asp-Glu-Leu) Containing 2 and Keratin 18 (KRT18) were positively correlated with the risk of PE, whereas the other five proteins (Complement Factor B [CFB], FYN Proto-Oncogene [FYN], RAN Binding Protein 1 [RANBP1], Amphoterin-Induced Gene and ORF 1 [AMIGO1], and Arginase 2 [ARG2]) were negatively correlated. None of the five proteins had reverse causality. Bayesian colocalization analysis verified the positive correlation of CFB (coloc.coloc.susie PPH4 = 1) and KRT18 (coloc.coloc.abf PPH4 = 0.74) with PE. The genes encoding the proteins KRT18, FYN, ARG2, and RANBP1 were distributed in specific cell types within PE tissues from patients with PE. Moreover, the examination of single-cell localization provided insights into the extensive distribution of the KRT18 gene, which is highly expressed in the villous cytotrophoblast (VCT) and extravillous trophoblast (EVT) populations.
Conclusion:
Our MR analysis suggested that the plasma proteins KRT18, FYN, RANBP1, AMIGO1, and ARG2 had causal effects on PE risk. These findings indicated that these five proteins might be promising druggable targets for PE and warrant further clinical therapy, especially KRT18.
Introduction
Preeclampsia (PE) is a pregnancy-specific multisystem disorder with an incidence of ∼2–5% that can be diagnosed by new-onset hypertension and one other PE-associated symptom or sign (e.g., proteinuria or widespread end-organ injury) and typically occurs after 20 weeks of gestation.1,2 PE contributes significantly to maternal and perinatal morbidity and mortality worldwide, highlighting an emerging field of research. PE can be classified into two major subtypes: (a) early-onset PE (delivery at <34 weeks gestation), which is primarily attributed to uteroplacental ischemia, and (b) late-onset PE (delivery at ≥34 weeks gestation), which is predominantly associated with a metabolic crisis leading to an imbalance between fetal requirements and maternal resources. 3 Although the pathophysiological mechanisms of PE remain uncertain, several triggers have been identified in previous studies. The placenta supports the fetus throughout pregnancy; however, abnormal placental development marked by shallow invasion of trophoblast cells into the uterine wall is considered a leading cause of PE. 4 Immune-mediated placental dysfunction and systemic endothelial injury are the primary pathological factors of PE. Soluble fms-like tyrosine kinase-1 (sFlt-1), placental growth factor (PlGF), endoglin, and microRNAs are potential biomarkers for the prediction and diagnosis of PE, providing opportunities for personalized monitoring and intervention strategies. A prognostic study conducted by Zeisler et al. revealed that an sFlt-1/PlGF ratio ≤38 can accurately predict the development of PE within 1 week, with a negative predictive value of 99.3%. 5 Moreover, the INHBA, OPRK1, and TPBG genes were found to be associated with PE, and a predictive model was established. 6 Despite these advancements, early diagnostic tests and effective treatments for PE should be tested.
Plasma proteins, which can originate from any organ, cell, or even from the mother through placental contributions from the mother, participate in a range of biological processes (BPs), including signaling, inflammation, and transportation.7,8 Sarosh et al. demonstrated that circulating plasma antiangiogenic factors such as sFlt-1 were effective biomarkers for risk stratification and PE screening. 9 Single-nucleotide polymorphisms (SNPs) link protein levels to genetic loci, and publicly available genome-wide association studies (GWASs) have identified protein quantitative trait loci (pQTLs) associated with a multitude of plasma proteins. These pQTLs reflect the circulating levels of plasma proteins and can be used to explore the causality between plasma proteins and PE. Moreover, Mendelian randomization (MR) has been widely used for potential biomarker screening and drug target development. Owing to advances in genetic instruments using SNPs identified by GWASs, MR analysis based on the integration of GWAS and pQTL data has promoted the development of novel therapeutic strategies for many diseases.
In this study, our analysis aimed to evaluate the causal effect of plasma proteins on PE and to identify potential biomarkers for PE. First, we used MR to identify plasma proteins potentially associated with PE risk using pQTL data from five large-scale proteomic studies. The primary findings were subsequently further validated using Bayesian colocalization, summary-data-based MR (SMR), and heterogeneity in dependent instruments (HEIDI) analysis. Moreover, single-cell type expression analysis was performed, and we identified specific cell types on the basis of target protein-coding genes that were enriched in PE tissues. We then constructed a protein–protein interaction (PPI) network to further investigate the therapeutic potential of our plasma protein biomarkers.
Materials and Methods
Study design
Supplementary Figure S1 displays the entire study design. To summarize, we utilized pQTL data from five large-scale proteomic studies and employed a two-phase (discovery and replication) proteome-wide MR framework to investigate the associations of pQTLs with PE.10–14 The private patient data from datasets are deidentified. Bayesian colocalization, SMR, and HEIDI tests were utilized to validate the causal links between protein biomarkers and PE. Furthermore, single-cell type expression analysis was conducted to identify the specific cell types in which target protein-coding genes were enriched in PE tissues. Finally, a PPI network was constructed using the identified protein biomarkers to further investigate potential therapeutic targets. 15
Data source
The PE data were derived from the UK Biobank (UKBB, https://www.ukbiobank.ac.uk/) and the FinnGen database (https://r10.finngen.fi/). The UKBB dataset (study code O14) includes data from 184 PE patients and 361,010 controls of European ancestry (http://biobank.ctsu.ox.ac.uk/crystal/field.cgi?id=O14). The FinnGen dataset, coded O15_PREECLAMPS, includes data from 7,377 PE patients and 211,957 controls of European ancestry (https://storage.googleapis.com/finngen-public-data-r10/summary_stats/finngen_R10_O15_PREECLAMPS.gz). In our MR analysis, the FinnGen dataset was used for the discovery phase, and the UKBB dataset was used for the replication phase. Within the FinnGen dataset, genetic variants related to PE with a p value <5 × 10−8 and with linkage disequilibrium (R2 < 0.001) were selected as instrumental variables for the reverse MR analysis of PE.
Protein-related MR studies
The pQTLs from the five proteomic studies mentioned were utilized for the selection of genetic instruments. The platform ID of each protein was mapped to gene symbols and unified using annotations from the original studies and manual examinations. Subsequently, SNPs were mapped to the human genome build 37 (GRCh37) to standardize genome coordinates. Genetic instruments and proteins were selected using the following criteria: (i) SNPs related to any protein were selected based on a stringent significance threshold (p < 5 × 10−8); (ii) Limited to the complex linkage disequilibrium structure, SNPs and proteins within the major histocompatibility complex (MHC) region (chr6: 26.0–34.0 Mb) were excluded 15 ; (iii) The independent pQTLs were identified for each protein using linkage disequilibrium clustering (R2 < 0.001); and (iv) The strength assessment of the genetic instruments was performed with R2 and F-statistics, where R2 is the proportion of protein level variability explained by each genetic instrument, and instruments with an F-statistic less than 10 were filtered out. For proteins that appeared multiple times in the studies, the one with the highest sum of R2 values was chosen. According to the following criteria, genetic instruments were further divided into two types: cis- or trans-pQTLs. A pQTL was considered cis when the leading SNP was within 1 Mb of the transcription start site of the protein-coding gene. However, those outside the aforementioned region were considered trans-pQTLs.10,12,14 Ultimately, all cis-pQTLs identified in the studies were selected as our instrumental variables, incorporating a total of 7,468 cis-pQTLs and 2,968 unique plasma proteins into the analysis.
The two-sample MR package was used for the MR analysis. 16 For proteins with only one genetic instrument, the Wald ratio method was employed to calculate the log odds change in PE risk per standard deviation (SD) increase in circulating protein levels, using the instrument serving as a proxy. For proteins with multiple genetic instruments, the inverse variance weighted (IVW) method was applied to obtain MR effect estimates. If proteins had multiple genetic instruments, we employed the IVW method to estimate MR effect sizes. To determine heterogeneity among genetic instruments, Cochran’s Q test was conducted. In addition, we performed further analyses, including simple median, weighted median, and MR–Egger analyses, to address potential pleiotropy.
MR analysis was conducted on identified proteins using PE GWAS data from the FinnGen database and UKBB, initially using p < 0.05 as the threshold for preliminary significance. A meta-analysis of the MR data from two sources was conducted employing either a random effects model or a fixed effects model. The model choice was based on dataset heterogeneity: A random effects model was used when heterogeneity was present, and a fixed effects model was applied when heterogeneity was absent. For multiple testing correction, we performed Bonferroni correction, setting the significance level at p.adj <0.05. All analyses were conducted with R software version 4.1.2.
Colocalization analysis
To calculate whether two associated signals (protein levels and PE risk) are consistent with a shared causal variant, rather than a noncausal variant due to linkage disequilibrium, Bayesian colocalization analysis was performed using protein data and FinnGen PE GWAS data utilizing the “coloc” package. 17 Five hypotheses of colocalization analysis provide a comprehensive framework for investigating the genetic underpinnings of both protein levels and PE risk at the genomic locus: (i) no causal variant affects protein levels or PE risk (H0); (ii) a causal variant affects only protein levels (H1); (iii) a causal variant affects only PE risk (H2); (iv) two distinct causal variants affect protein levels and PE risk independently (H3); and (v) a shared causal variant affects both protein levels and PE risk (H4).
Bayesian colocalization analysis was employed to calculate posterior probabilities for the five hypotheses mentioned above, evaluating whether the two traits share a single genetic variant. If a protein is linked to multiple pQTLs, colocalization analysis is conducted separately for each pQTL, focusing on the one exhibiting the most compelling evidence of colocalization. The analysis utilized default parameters, with prior probabilities set as follows: p1 = 1 × 10−4 (for SNP association with protein), p2 = 1 × 10−4 (for SNP association with PE), and p12 = 1 × 10−5 (for SNP association with both protein and PE).
In our study, we rigorously confirmed hypothesis 4 (PPH4), which suggests the presence of a shared genetic variant affecting both protein levels and PE within a specific genomic region. We employed two widely recognized algorithms, namely, coloc.abf and coloc.susie, to evaluate the extent of colocalization. Strong colocalization was defined as PPH4 >80%, moderate colocalization as PPH4 >60% and <80%, and weak colocalization as PPH4 <60%. A gene was considered to show evidence of colocalization if it demonstrated a gene-based PPH4 >60%, as determined by at least one of the algorithms.
SMR analysis
SMR analysis was further performed to verify the causal correlation between proteins and PE. Furthermore, we conducted the HEIDI test, employing multiple SNPs within a specific genomic region, to distinguish proteins associated with PE risk due to shared genetic variation from those influenced solely by linkage disequilibrium. Both the SMR and HEIDI tests were performed by SMR software (version 1.3.1). A significance threshold of p < 2.38 × 10−3 (0.05/21) was established for the SMR analysis. A p value >0.05 in the HEIDI test indicated that the observed association between the protein and PE was not driven by linkage disequilibrium. However, due to database limitations, not all proteins were subjected to HEIDI testing.
Downloading and preprocessing of single-cell sequencing data 18
The Gene Expression Omnibus (GEO) database hosts an extensive collection of single-cell sequencing data. In this study, the single-cell sequencing dataset GSE173193 containing data from a total of 8 PE samples was acquired from the GEO database. This dataset included data from 2 PE placental tissue samples and 2 normal placental samples. The single-cell raw data from GSE173193 were imported using the Seurat package (version 4.2.0) in R. 19 Initially, cells and genes of low quality were filtered out using the following criteria: (i) cells expressing fewer than 200 genes were removed, and (ii) genes not detected in any cells were discarded. Cells with a gene expression count ranging from 200 to 9,000 and cells with mitochondrial gene percentages under 20% were maintained. Moreover, cells with less than 90,000 unique molecular identifier counts were maintained. The “Normalize Data” function within the Seurat R package was used for data normalization. Following normalization, highly variable genes in single cells were identified by balancing the relationship between average expression and dispersion. Subsequently, principal component (PC) analysis was performed, and the significant PCs were utilized as inputs for graph-based clustering. The Harmony method was employed to eliminate batch effects across different samples. For clustering, the FindClusters function, which is based on a clustering algorithm optimizing shared nearest neighbor modularity, was utilized to produce 21 clusters across 25 PC components at a resolution of 0.4. t-Distributed stochastic neighbor embedding (t-SNE) was then performed using the “Run t-SNE” function. Cell clustering was visualized using t-SNE-1 and t-SNE-2.
PPI network analysis
The GeneMANIA website (http://genemania.org) facilitates the prediction of relationships between functionally similar genes and central genes, encompassing PPIs, protein–DNA interactions, pathways, physiological and biochemical reactions, coexpression, and colocalization. 20 In this study, a PPI network of key genes was constructed via the GeneMANIA website. Subsequently, Gene Ontology (GO) analysis was performed on the key genes and their interacting genes using the “clusterProfiler” R package.
Phenome-wide MR studies
To investigate the potential side effects of four drug targets, we employed gene expression as the exposure factor and utilized disease summary statistics from the UKBB (n ≤ 420,531) as outcomes for a comprehensive phenome-wide MR analysis. The UKBB Disease GWASs were conducted using the scalable and accurate implementation of generalized mixed model (SAIGE) method, which addresses imbalanced case–control ratios. Due to statistical power considerations, we selected 851 traits (diseases) other than PE for phenome-wide MR analysis, each with more than 100 cases. Summary statistics for disease-associated SNPs were downloaded from the SAIGE GWAS (https://www.leelabsg.org/resources). Subsequently, MR analysis was performed using either the IVW or Wald ratio method with identical parameters, leveraging pQTLs. A causal effect with a false discovery rate (FDR) <0.05 was deemed statistically significant.
Results
Proteome-wide MR analysis identified seven circulating plasma proteins associated with PE
All the genetic instruments had F-statistics greater than 10, indicating strong instrument strength (Supplementary Table S1). Using the Wald ratio or IVW methods, and after the Bonferroni correction (p.adj < 0.05), a total of seven proteins were found to be significantly associated with the risk of PE (Table 1). Gene prediction indicated that KDEL (Lys-Asp-Glu-Leu) Containing 2 (KDELC2) and Keratin 18 (KRT18) were positively correlated with the risk of PE, whereas the other five proteins (Complement Factor B [CFB], FYN Proto-Oncogene [FYN], RAN Binding Protein 1 [RANBP1], Amphoterin-Induced Gene and ORF 1 [AMIGO1], and Arginase 2 [ARG2]) were negatively associated with the risk of PE, suggesting that lower levels of these five proteins are linked to a greater risk of PE (Fig. 1). These associations were generally consistent across additional analyses including the simple median, weighted median, and MR–Egger analyses. In addition, the CFB protein was found to be heterogeneous and pleiotropic, the KDELC2 protein was found to be neither heterogeneous nor pleiotropic (pheterogeneity > 0.05, pleiotropy > 0.05), and the heterogeneity and pleiotropy of the remaining proteins could not be tested due to data structure limitations (Table 2). The results of the proteome-wide MR within the discovered protein set are presented in Supplementary Table S1.
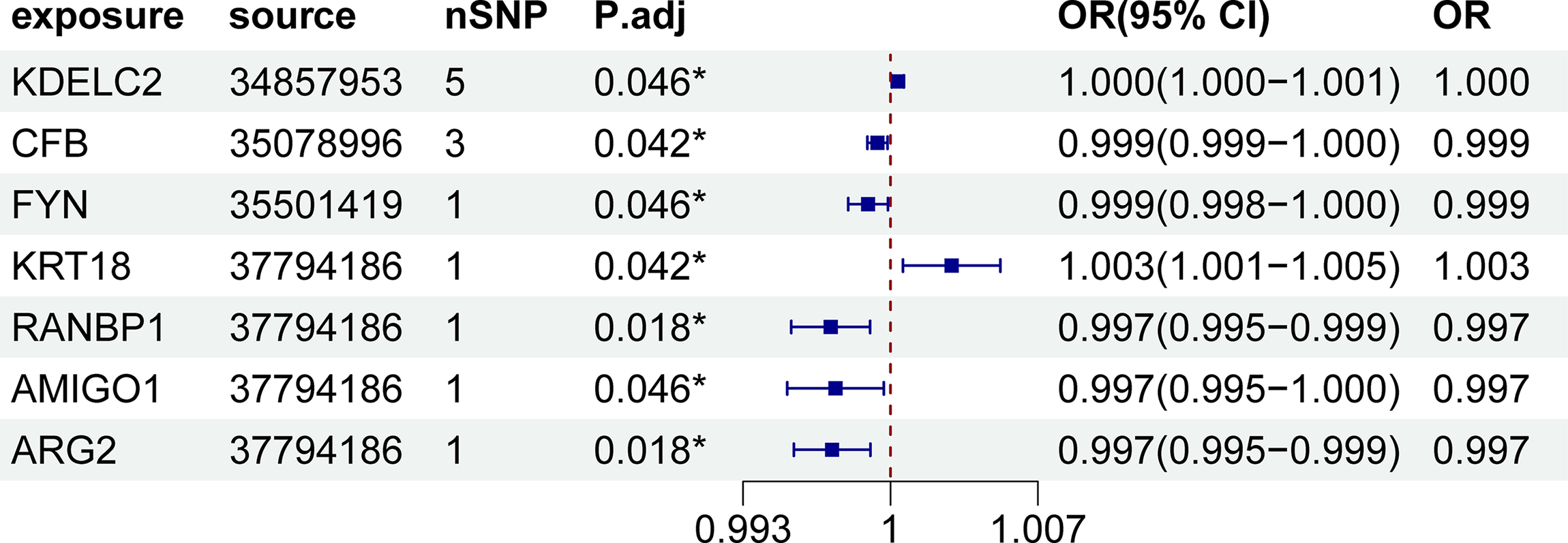
The meta-analysis results of proteome-wide MR analysis.
The Results of the Proteome-Wide MR

AMIGO1, Amphoterin-Induced Gene and ORF 1; ARG2, Arginase 2; CFB, Complement Factor B; FYN, FYN Proto-Oncogene; HEIDI, heterogeneity in dependent instruments; KDELC2, KDEL (Lys-Asp-Glu-Leu) Containing 2; KRT18, Keratin 18; MR, Mendelian randomization; RANBP1, RAN Binding Protein 1; SMR, summary-data-based Mendelian randomization.
The Heterogeneity and Pleiotropy Testing for CFB and KDELC2

PE, preeclampsia.
In a meta-analysis combining data from the two sources, several proteins were found to be significantly associated with PE risk. The odds ratios (ORs) for PE per SD increase in the levels of proteins predicted by gene analysis, with 95% confidence intervals (CIs), were as follows: the OR for KDELC2 was 1.000 (95% CI: 1.000–1.001), that for CFB was 0.999 (95% CI: 0.999–1.000), that for FYN was 0.999 (95% CI: 0.998–1.000), that for KRT18 was 1.003 (95% CI: 1.001–1.005), that for RANBP1 was 0.997 (95% CI: 0.995–0.999), that for AMIGO1 was 0.997 (95% CI: 0.995–1.000), and that for ARG2 was 0.997 (95% CI: 0.995–0.999).
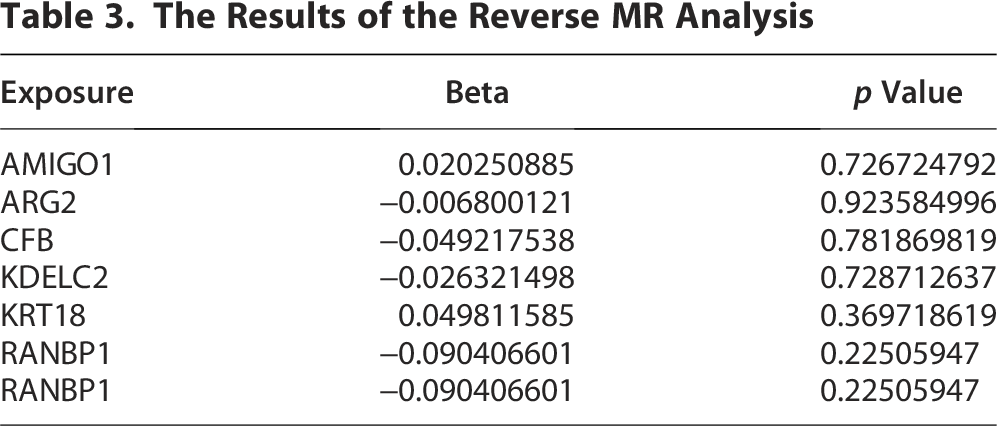
In the reverse MR analysis, no association was detected between genetic susceptibility to PE and the levels of these seven proteins (Table 3).
The Results of the Reverse MR Analysis
Colocalization analysis supports the causal relationship between two proteins and PE
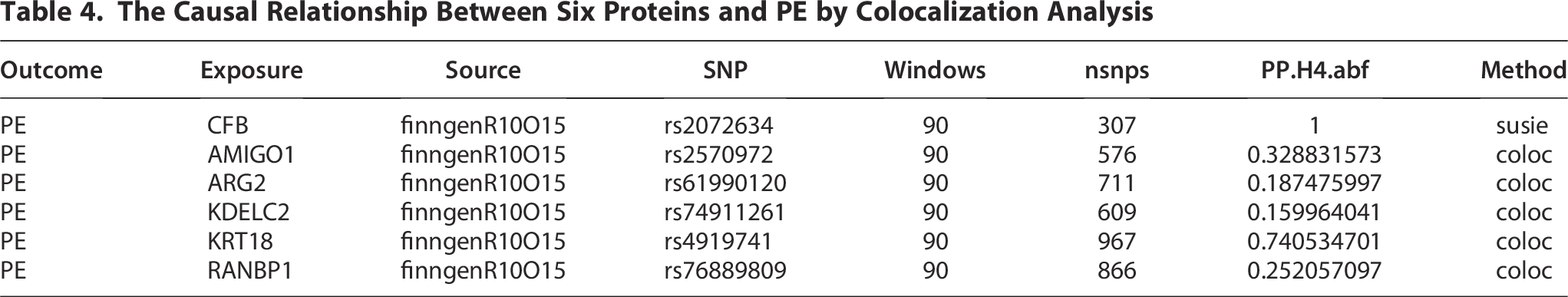
Among the seven potential causal proteins identified by proteome-wide MR, one protein (FYN) lacked complete summary-level data, rendering it untestable through colocalization analysis. Among the remaining six proteins, strong evidence of genetic colocalization supported the causal relationship of two proteins (CFB with a coloc.susie PPH4 of 1 and KRT18 with a coloc.abf PPH4 of 0.74) with PE under various priors and windows. This suggests a high probability of shared single causal variants between CFB protein levels and PE risk and a moderate probability of shared multiple causal variants between KRT18 protein levels and PE risk (Table 4).
The Causal Relationship Between Six Proteins and PE by Colocalization Analysis
The SMR and HEIDI tests validated six pathogenic proteins
To further validate the observed findings, SMR and HEIDI tests were conducted on the seven proteins with complete summary-level data. All proteins except KDELC2 passed the SMR test (p.adj < 0.05). Due to the lack of sufficient SNPs, proteins other than CFB could not undergo HEIDI testing. Based on the evidence, these proteins were categorized into three tiers. One protein (KRT18) that passed all tests was categorized as Tier 1 (Table 1). The five proteins that either failed the colocalization analysis or HEIDI test or could not be tested due to a lack of data (AMIGO1, ARG2, CFB, FYN, and RANBP1) were categorized as Tier 2 proteins (Table 1). The KDELC2 protein, which failed the meta-analysis, colocalization analysis, and HEIDI test, was categorized as Tier 3 (Table 1).
Research on cell type-specific expression in PE tissues
To investigate whether the genes encoding the six circulating proteins were enriched in specific cell types within PE tissues, we further conducted single-cell type expression analysis using single-cell RNA sequencing data from the GEO database. The cells were clustered into 21 clusters and further classified into nine cell types (granulocyte, villous cytotrophoblast [VCT], macrophage, extravillous trophoblast [EVT], myelocyte, thymus/natural killer [T/NK] cell, monocyte, syncytiotrophoblast [SCT], and B lymphocyte [B cell]). Figure 2B shows that the genes encoding protein KRT18 were primarily distributed in VCT and EVT cells (Fig. 2C). The genes encoding for proteins ARG2 and RANBP1 were mainly distributed in VCT cells (Fig. 2D and E). The gene encoding for protein FYN was predominantly found in T/NK cells (Fig. 2F). The genes encoding the proteins AMIGO1 and CFB were not enriched in any specific cell type in the single-cell dataset (Fig. 2G and H).
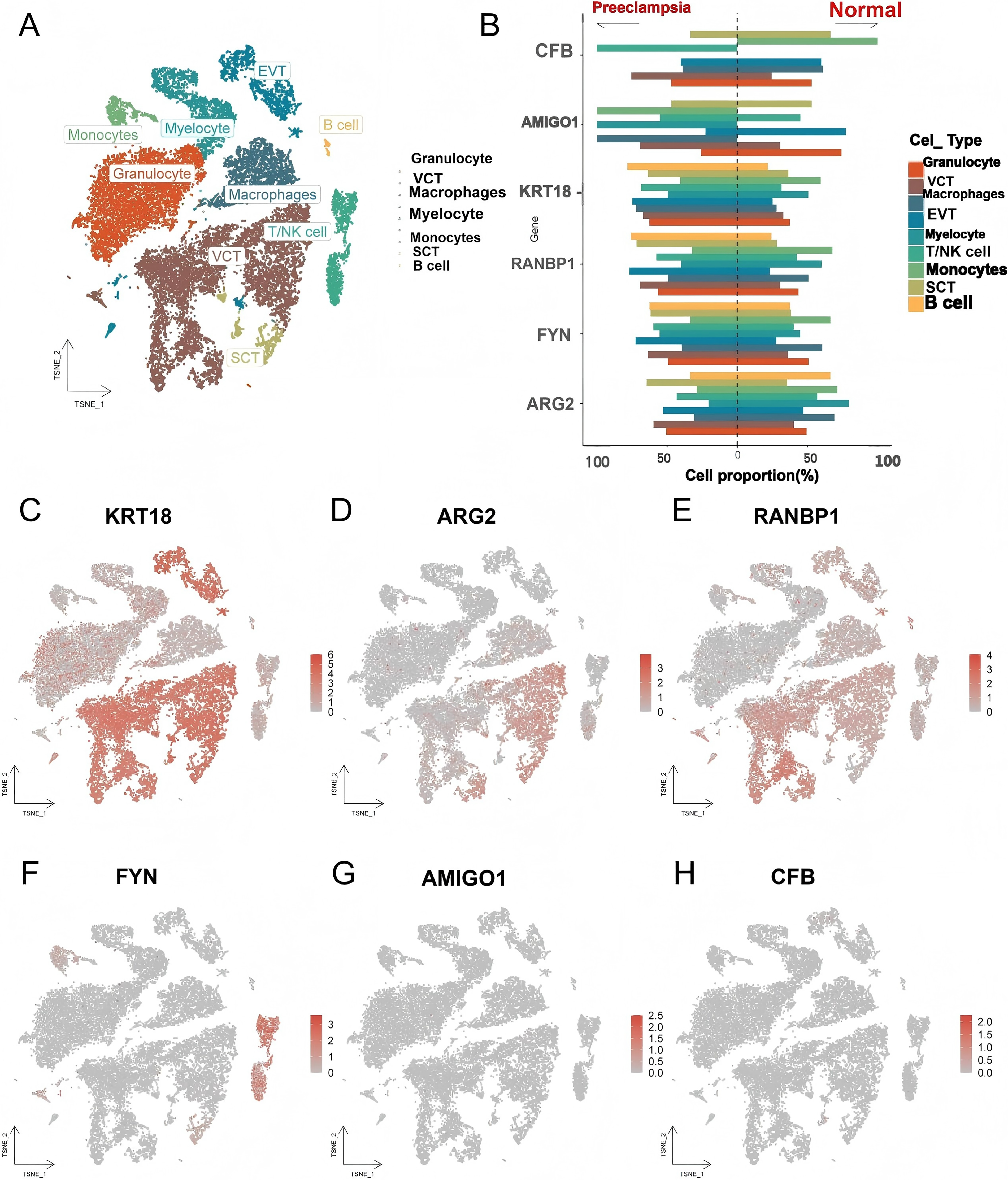
The single-cell type expression of protein-encoding genes identified through proteome-wide MR in PE tissues.
PPI network
We constructed a PPI network for circulating proteins using the GeneMANIA database (Fig. 3A). To further investigate the functions of the characteristic proteins, we performed GO enrichment analysis on a total of 26 proteins, including 6 circulating proteins and 20 proteins associated with the 6 circulating proteins. The GO enrichment results revealed that BPs, such as negative regulation of chemokine production and response to

The protein interaction network.
The Results of PPI Network

PPI, protein–protein interaction.
MR analysis of GWASs on the identified PE drug target proteins and other diseases
We evaluated whether the expression of the six drug target proteins associated with PE plays a role in other diseases. Thus, a broader MR screening was conducted across 851 non-PE diseases or traits using the UKBB (Supplementary Table S2). CFB was significantly associated with digestive system diseases; lower levels of CFB were related to celiac disease (OR = 0.255457) and intestinal malabsorption (OR = 0.302310), while higher levels of CFB were associated with ulcerative colitis (OR = 1.760720) (Fig. 4 and Supplementary Table S3). No other diseases were found to have a significant association with these drug target proteins (FDR 0.05), and the summary results are presented in Supplementary Table S3.

Displays a Manhattan plot of the phenome-wide MR results for AMIGO1, AARG2, CFB, FYN, KRT18, and RANBP1. Note: In the phenome-wide MR results, the vertical axis represents p values. Each point represents a disease trait, and different colors signify the MR results for different expressions.
Discussion
To our knowledge, this is the first MR analysis to identify drug targets for PE based on pQTL data from five large-scale proteomic studies. Here, we identified seven proteins as candidate targets for PE, including KDELC2, KRT18, CFB, FYN, RANBP1, AMIGO1, and ARG2. Except for CFB and KDELC2, the five other proteins were shown to be associated with PE via MR analysis, further suggesting the reliability of the methods used in this study.
The observed modest ORs for proteins associated with the risk of PE imply that individual genetic influences are limited. This observation corresponds with the polygenic characteristics of complex diseases such as PE, in which the disease development is attributed to the cumulative impact of numerous variants with small effects across various biological pathways.21,22 The presence of small effect sizes does not diminish their biological importance; instead, it may indicate a widespread distribution of risk within regulatory networks. Notably, proteins exhibiting modest genetic influences can still represent valuable therapeutic targets, particularly if they are situated at crucial regulatory points (for instance, upstream signaling nodes or network hubs) where slight adjustments could trigger significant downstream effects. A case in point is KRT18, which, despite its limited effect size, is abundantly expressed in trophoblasts and plays a vital role in regulating essential processes such as cell adhesion and apoptosis, highlighting its potential relevance in pharmacotherapy. To achieve robust causal inference, we adopted a comprehensive multi-tiered analytical strategy. This strategy included MR, Bayesian colocalization, SMR/HEIDI tests, single-cell expression analysis, and protein–protein interaction networks. Both CFB and KRT18 demonstrated evidence of shared causal variants with PE through colocalization analysis. Following thorough validation, proteins were classified into three tiers: KRT18 (Tier 1) successfully passed all analytical evaluations; KDELC2 (Tier 3) did not meet multiple testing criteria, while the remaining five proteins (ARG2, RANBP1, FYN, AMIGO1, and CFB) were designated as Tier 2. Furthermore, single-cell expression analysis revealed increased expression of KRT18, ARG2, RANBP1, and FYN specifically in trophoblast cells, and phenome-wide MR analysis revealed associations between CFB and various digestive diseases. Collectively, these findings support the biological plausibility and therapeutic promise of the identified proteins, particularly KRT18, in the context of PE pathogenesis.
Despite the development of new drugs for many years, the current therapeutic options for PE remain unsatisfactory. Considering the pathogenesis of PE, which is characterized by compromised placental vasodilation and increased maternal blood pressure, resulting in a reduced blood supply to the fetus, 23 we explored the causal proteins for PE and their distribution in placenta-associated cells. Notably, in our study, we found that the identified circulating plasma proteins were mainly distributed trophoblast cells. 24 We also conducted bidirectional MR analysis among the seven identified proteins and failed to observe any significant associations. The presence of the placental barrier may explain the absence of any significant bidirectional correlations. Although the evidence remains scarce, the current study suggested that plasma might be a valuable resource for identifying proteins associated with PE and that the proteins circulating in the plasma might be promising drug targets for PE treatment.
Among the five potential proteins identified in this study, the roles of ARG2 and KRT18 have been relatively explored in PE in previous studies.25–28 Unlike our study, these studies only reported the levels of these proteins in the plasma of patients with PE. Owing to the importance of identifying protein drug targets for the success of precision or personalized medicine approaches, our study provides new insights into the development of therapeutic strategies for PE.
KRT18, also known as cytokeratin 18, is an intermediate filament protein that acts as proinflammatory cytokine in serum and apoptotic marker The correlation between circulating KRT18 levels and PE risk is positive, as previous studies on plasma KRT18 levels have reported significantly increased KRT18 expression in the plasma of patients with PE. Moreover, the genes encoding the protein KRT18 were primarily distributed in VCT and EVT cells. Despite the difference in tissue-specific expression of KRT18, it was the only protein that was validated as Tier 1, indicating a greater probability of KRT18 being a causal protein for PE.
KRT18, commonly referred to as cytokeratin 18, is classified as an intermediate filament protein that serves as an indicator of apoptosis and inflammation in serum samples.29,30 Our research highlights a significant association between the levels of circulating KRT18 and the risk of developing PE, corroborating previous studies that have documented increased KRT18 levels in patients diagnosed with PE. 28 Moreover, KRT18 is expressed predominantly in VCT and EVT cells, and it has been recognized as a Tier 1 causal protein in the context of PE, providing compelling evidence for its role in the pathogenesis of this disease. Nevertheless, owing to the critical biological functions of KRT18, its ability to act as a direct therapeutic target necessitates careful consideration, despite its notable correlation with PE. As a fundamental component of epithelial intermediate filaments, KRT18 plays a vital role in maintaining cytoskeletal stability, facilitating cell–cell adhesion, responding to stress, and preserving tissue barriers, with pronounced expression observed in placental, hepatic, and gastrointestinal epithelial tissues. Direct inhibition of KRT18 may result in detrimental consequences, including disruption of epithelial structure, which can lead to apoptosis or necrosis; dysfunction of the placental barrier, affecting maternal–fetal exchange, and damage to hepatocytes. This assertion is further supported by its recognized role as a diagnostic biomarker in liver diseases. In light of these potential risks, KRT18 may be more appropriately regarded as a circulating biomarker or as a reflection of upstream regulatory mechanisms, such as those related to inflammation or oxidative stress, rather than a direct target for therapeutic intervention. Subsequent research should aim to identify specific modulators that can influence KRT18 expression or activity, facilitating safer, indirect therapeutic strategies. In summary, while KRT18 demonstrates a strong causal relationship with PE, its structural significance within epithelial tissues emphasizes its potential utility as a monitoring tool for disease rather than as a direct target for pharmacological treatment.
Human ARG2 is a mammalian arginase isoform encoded by ARG2, which is located on chromosome 14q2427. 31 As a key hydrolase in the urea cycle, ARG2, which contains 354 amino acids, is abundantly expressed in mitochondria and is preferentially expressed in the kidney, lactating mammary gland and even macrophages. 32 Consistent with our results, previous studies have indicated that a low concentration of ARG2 is associated with an increased risk of PE. 33 In addition, we found a relationship between ARG2 and VCT cells in PE tissues. Given the lack of colocalization of ARG2 and the specific distribution of ARG2 in VCT cells revealed by proteome-wide MR analysis, we speculated that the effect of ARG2 in PE might be blocked by the placental barrier. In other words, ARG2 might be a promising druggable target in the circulation of the placenta. In addition, our PPI analysis revealed that ARG2, APOD, and LILRB4 were significantly enriched in the negative regulation of chemokine production. We noticed that a reduction in plasma ARG2 levels was reported to improve the responsiveness to antihypertensive treatment in PE patients, which might suggest that A may have therapeutic value in PE and deserves further study. RANBP1 belongs to the RAS superfamily of small GTPases that participate in the internuclear transport of proteins, nucleic acids, and microRNAs and contribute to the cellular epigenomic signature. 34 According to previous studies, RANBP1 is not associated with the pathogenesis of PE. However, MR analysis revealed that RANBP1 is a potential target for PE medications, and RANBP1 was confirmed to be primarily distributed in VCT cells in this study. Because RANBP1 is an intracellular protein with high extracellular and intracellular expression, we hypothesized that it might be a drug target for PE.
FYN and AMIGO1 were protective proteins against PE in our study. FYN is a member of the src family of protein kinases that regulates multiple cellular processes, including cell adhesion, invasion, proliferation, survival, apoptosis, and angiogenesis. 35 FYN interacts with and phosphorylates a wide variety of proteins, such as RS1, SOCS3, and PLAUR, suggesting that FYN might act by reducing the stability of the amniotic membrane, promoting the development of PE. Notably, FYN was also confirmed to be enriched in T/NK cells, which are immune cells that play a major causative role in the pathology of PE. 36 Maternal immune tolerance is a special contributor to pregnancy. Insufficient T cells or inadequate functional competence are implicated in PE, which stems from placental insufficiency. 37 Taken together, these findings indicate that FYN might be a potential therapeutic target for PE and may be involved in immune responses. AMIGO1 is an LRR-domain cell adhesion molecule preferentially expressed on nerve cells that mediates the fasciculation and myelination of developing axons. 38 In contrast to the other identified PE targets in this study, AMIGO1 has been shown to affect neuronal genes.39,40
In a recent study examining the pathogenesis of PE, Xu et al. performed MR and colocalization analyses involving 734 plasma proteins within the FinnGen cohort. 41 Their research revealed several potential candidate proteins, such as CXCL10, PZP, AHSG, and UROS, indicating the involvement of immune and inflammatory pathways in the progression of this disease. Building upon the findings of Xu et al., the current investigation broadens the proteomic analysis by incorporating cis-pQTLs data from five extensive studies, which encompass a total of 2,968 plasma proteins.
From a methodological standpoint, we employed a two-stage design that included both discovery and replication phases within the FinnGen and UKBB cohorts. This approach was further enhanced by meta-analysis and validation techniques, including Bayesian colocalization, SMR/HEIDI tests, single-cell expression profiling, and the exploration of protein–protein interaction networks. Our analysis revealed a unique set of proteins—KRT18, ARG2, FYN, RANBP1, AMIGO1, and CFB—that are linked to cytoskeletal integrity, metabolic regulation, and cellular adhesion, thereby contrasting with the immune-inflammatory targets identified by Xu et al. Rather than presenting conflicting outcomes, these disparate findings underscore the multifaceted nature of PE and illustrate how diverse methodological frameworks can uncover complementary pathological mechanisms. These results imply that the pathogenesis of PE involves simultaneous dysregulation across immune, structural, and metabolic pathways. Future research should integrate multiomics data from a variety of populations to clarify the interactions among these mechanisms and facilitate the advancement of targeted therapeutic interventions.
Inevitably, there are several limitations in our study. First, we screened prioritized proteins, and the sensitivity analysis was adjusted in the analysis; therefore, potential bias could not be excluded. Moreover, most participants in the GWAS datasets were of European descent, and the results of this study might not be entirely applicable to subjects of non-European descent, which means that we need to be cautious when applying our results to other populations. In addition, some factors associated with PE, such as parity and maternal age, were not included. Due to the limited availability of clinical data from Asian groups, the causal correlation between these potential proteins and PE was not validated. Furthermore, importantly, the odd ratios of the analyzed proteins correlated with PE were not substantial, warranting clarification. Constrained by existing data, the observed effects of these proteins on PE were modest, whether positively or negatively correlated. However, significant p values and previous studies suggest their potential value as targets for PE. Finally, the examination of five proteins (KRT18, AMIGO1, ARG2, FYN, and RANBP1) was limited by the availability of SNPs, which impedes the practicality of the findings, particularly concerning heterogeneity and pleiotropy. Because a large amount of data is required, analyses of proteins other than CFB may not be entirely reliable, and the results of these analyses should be interpreted with caution. Therefore, future work should focus on the proteins related to PE, and additional experimental validation is required to substantiate these findings.
Conclusions
In general, our MR analysis suggested that the plasma levels of the identified proteins (KRT18, FYN, RANBP1, AMIGO1, and ARG2) are causally associated with PE risk. The identified proteins may be potential biomarkers or druggable targets for PE, especially circulating KRT18. Based on our results, we hypothesized that the protein KRT18 might play a role in the development of PE, and future clinical studies should be conducted to verify the effects of this protein on PE. Further work is needed to evaluate the credibility of these candidate proteins in PE treatment.
Author Disclosure Statement
No competing financial interests exist.
Footnotes
Supplemental Material
Supplemental Material
Supplemental Material
Supplemental Material
Abbreviations Used
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
