Abstract
Background
With multimorbidity becoming the norm rather than the exception, the management of multiple chronic diseases is a major challenge facing healthcare systems worldwide.
Methods
Using a large, nationally representative database of electronic medical records from the United Kingdom spanning the years 2005–2016 and consisting over 4.5 million patients, we apply statistical methods and network analysis to identify comorbid pairs and triads of diseases and identify clusters of chronic conditions across different demographic groups. Unlike many previous studies, which generally adopt cross-sectional designs based on single snapshots of closed cohorts, we adopt a longitudinal approach to examine temporal changes in the patterns of multimorbidity. In addition, we perform survival analysis to examine the impact of multimorbidity on mortality.
Results
The proportion of the population with multimorbidity has increased by approximately 2.5 percentage points over the last decade, with more than 17% having at least two chronic morbidities. We find that the prevalence and the severity of multimorbidity, as quantified by the number of co-occurring chronic conditions, increase progressively with age. Stratifying by socioeconomic status, we find that people living in more deprived areas are more likely to be multimorbid compared to those living in more affluent areas at all ages. The same trend holds consistently for all years in our data. In general, hypertension, diabetes, and respiratory-related diseases demonstrate high in-degree centrality and eigencentrality, while cardiac disorders show high out-degree centrality.
Conclusions
We use data-driven methods to characterize multimorbidity patterns in different demographic groups and their evolution over the past decade. In addition to a number of strongly associated comorbid pairs (e.g., cardiac-vascular and cardiac-metabolic disorders), we identify three principal clusters: a respiratory cluster, a cardiovascular cluster, and a mixed cardiovascular-renal-metabolic cluster. These are supported by established pathophysiological mechanisms and shared risk factors, and largely confirm and expand on the results of existing studies in the medical literature. Our findings contribute to a more quantitative understanding of the epidemiology of multimorbidity, an important pre-requisite for developing more effective medical care and policy for multimorbid patients.
Introduction
Multimorbidity, defined as the coexistence of two or more chronic medical conditions in an individual patient, 1 is a growing public health concern for healthcare systems worldwide. It has been found to be associated with adverse health outcomes, including a higher risk of mortality, a lower quality of life, increased utilization of health care, and correspondingly higher healthcare costs.2–14 It is most prevalent in the elderly population, as organs gradually lose full function with the aging process.8,15–17 With an increasing life expectancy and an aging population, the number of people with multiple health conditions is set to rise, as is public expenditure on long-term medical care. Unfortunately, current healthcare systems are largely designed to treat single diseases, resulting in the need to use multiple services to manage multimorbidity.11,18–20 Due to poor coordination and integration in medical care, causing a lack of continuity in treatment, disorders not designated as the primary condition are often undertreated. 21
In order to align medical care more closely to the needs of patients with multiple health conditions, a better understanding of the epidemiology of multimorbidity in the general population is necessary. Studies have shown that multimorbidity can be present in all age groups, including the pediatric population. 22 In particular, significant attention has been paid to multimorbidity in the elderly, due to its high prevalence in that population. Data sources used range from structured databases (e.g., electronic health records and insurance billing data) to self-administered questionnaires and research interviews. While the former tend to be more reliable, the latter are typically subject to self-reporting bias. Some analyses are based on small sample sizes from selected populations, which likely do not generalize well. Lastly, many studies employ only a narrow range of methods to study multimorbidity patterns (e.g., identifying the most prevalent pairs and triads, or calculating the odds ratio) although some have explored more novel clustering approaches as well (e.g., matrix factorization, association rules, and undirected network analysis).2,19,20,23–31
In this paper, we aim to characterize multimorbidity patterns not only in older patients, but also across groups with different demographic and socioeconomic statuses, using a large, nationally representative primary care electronic medical records database. Unlike many previous studies, which generally adopt cross-sectional designs based on single snapshots of closed cohorts, we examine temporal changes in the patterns of multimorbidity across a decade of open-cohort patient data. Among previous longitudinal studies, few have examined disease trajectories. 32 Here, we apply various statistical methods to identify common comorbid pairs and triads of diseases, and use directed and undirected network analysis algorithms to measure temporal multimorbidity progression and identify clusters of chronic conditions. In addition, we analyze the impact of multimorbidity on mortality using survival analysis models.
Methods
Data
We use anonymized electronic medical records from The Health Improvement Network (THIN) 33 database for our analysis. The database contains longitudinal patient data collected at primary care clinics throughout the UK, covering approximately 6% of the UK population. The average length of follow-up in the THIN database is around 9 years. We extract demographic information (e.g., date of birth, sex, geographical location, and socioeconomic group), baseline vitals (e.g., smoking and alcohol status), and medical history (e.g., medical condition and date of diagnosis) from patient records between 2005 and 2016. To capture temporal trends in the population, we perform our analysis sequentially on each year of data in the sample period (i.e., one set of results for each year). We categorize the subjects into seven mutually exclusive age groups based on Medical Subject Headings (MeSH) definitions (see Supplementary Material A). 34 In contrast with studies that use static baseline demographics collected at the beginning of follow-up, we use the point-in-time patient age for our analyses. For example, a patient that is 16 years old in 2005 will be classified as an Adolescent for analyses between 2005 and 2007, and subsequently reclassified as an Adult from 2008 onwards.
Diagnoses are recorded in the THIN database using Read Codes, a coded thesaurus of clinical terms used by the UK National Health Service since 1985.
35
There is no standard method for the selection and definition of morbidities in the literature. After consulting with medical officers and Life & Health (L&H) actuaries at Swiss Re, we identify chronic conditions in the records, that is, diseases that are either permanent, caused by nonreversible pathological alterations, or require long periods of rehabilitation and care,19,36 and map them to a list of 46 higher level morbidities. Furthermore, we classify the morbidities into 14 System Organ Classes (SOCs) as defined in the Medical Dictionary for Regulatory Activities (MedDRA). (See Figures 1 and 2 and Supplementary Material A for lists of morbidities and classifications.) As in similar studies, we define multimorbidity as the presence of at least 2 of the 46 morbidities in a patient. Mapping between index and chronic conditions. CAD: Coronary Artery Disease; HVD: Heart Valve Disorder; MI: Myocardial Infarction; COPD: Chronic Obstructive Pulmonary Disease; PAD: Peripheral Artery Disease; TIA: Transient Ischemic Attack. Abbreviations for MedDRA SOCs used in figures.

Statistical analysis
We examine the distribution of multimorbidity in relation to age and socioeconomic status, as done in Barnett et al. 11 However, we use the Index of Multiple Deprivation (IMD) as a proxy for socioeconomic status. The IMD is a widely used measure of relative deprivation or poverty of wards and districts in the UK. It is computed using census data as a weighted index of deprivation in seven domains, including income, employment, education, health, crime, barriers to housing and services, and living environment. 37 (IMD data was available only for a subset of the patients. See Supplementary Material B for the sample sizes used in this analysis.) We note that the same approach, defining socioeconomic status by the area of residence, has been used in previous studies.11,38
For each age group, we also compute the observed prevalence for all individual, pairs, and triplets of morbidities. By the assumptions of probability theory, we expect diseases that are independent to co-occur at a rate close to the product of the observed prevalence of each individual constituent disease (i.e., the expected prevalence). Therefore, by comparing the ratio of the observed prevalence versus the expected prevalence (i.e., the lift), we can identify pairs and triads of diseases that occur together more frequently than expected by chance, possibly driven by an underlying pathophysiological mechanism. As a second metric, we estimate the odds ratio using logistic regression models to determine the association between each pair of diseases, both without adjustment and adjusted by age, sex, and all other diseases.
Next, we construct multimorbidity networks to study the natural clustering of diseases in the dataset. We consider diseases as nodes with sizes proportional to their observed prevalence. For each pair of diseases, we connect their nodes with an undirected edge weighted by the estimated lift, a measure of the strength of the association between the comorbid pair. This creates a dense network where each node is linked to almost every other node. This density, however, makes visualization and inference difficult. As a pre-processing step for subsequent analysis, we extract the main graph structure by removing edges from the adjacency matrix that are peripheral and relatively unimportant. We prune the edges between nodes that have joint prevalence below the 90th percentile, and keep only the edges that have a lift above 2.0, that is, those edges between pairs that co-occur two times more frequently than expected by chance. Similar thresholds have been used in related studies.2,39–42
We compute measures of centrality to identify the most important vertices in the multimorbidity network. In particular, for each node, we compute the degree centrality, which is defined as the number of links incident on a node, a direct measure of the connectivity of a node. In this context, a disease with high degree centrality is important because it often co-occurs with a large number of pathologies. We also estimate the eigenvector centrality, a measure of the transitive influence of nodes. To calculate the eigencentrality, each node is assigned a score that is proportional to the sum of the scores of all of its neighbors. Nodes with high eigencentrality either have many connections, or are connected to important neighbors. In addition, we compute the graph clustering coefficient (also known as the transitivity) as a quantitative measure of the network’s tendency to aggregate in smaller subgroups. To identify any clusters embedded in the multimorbidity networks, we apply a community detection algorithm based on modularity maximization43–45 to partition nodes into groups that have dense intra-group connections and sparse inter-group connections. Communities identified in this manner can be interpreted as clusters of diseases that tend to co-occur together.
To gain insight into temporal disease associations, we construct directed multimorbidity networks. We extract from each patient’s medical history a sequence of diseases ordered by the time of diagnosis. Using these trajectories, we can derive the probability of any given disease conditional on some prior diagnosis, that is, Prob(Disease B given Disease A). We use these probabilities as weights of the directed edges in the network. As before, we prune the network based on node prevalence and edge weights. Since these connections are directed, we can compute the in-degree and out-degree centralities, defined as the number of edges directed to the node, and the number of edges directed from the node to others, respectively. A node with a high in-degree centrality is often diagnosed following other diseases; a node with a high out-degree centrality often leads to subsequent diagnoses in other diseases. These metrics are useful for understanding disease progression, and any causal or contributory relationships between diseases.
Finally, we examine the association between multimorbidity and mortality by performing predictive survival analysis on the dataset. We use five-year overall survival as the primary outcome variable, and consider in our models a range of features, including demographic group, baseline vitals, baseline medical history, the severity of multimorbidity as quantified by the number of co-occurring chronic conditions, and the presence of any of the top ten most prevalent pairs and triplets of morbidities as observed in the Aged and Elderly age groups. We exclude those subjects aged 65 or less from this part of the analysis, as younger age groups have five-year overall mortality rates close to zero.
We explore three standard methods used in survival modeling—the Cox proportional hazards model, 46 the regularized Cox model, and the accelerated failure time model—and additionally, we apply a nonlinear and non-parametric neural network survival model. 47 For model estimation and validation, we randomly split the original dataset into two disjoint sets, a training set that comprises 70% of the data, and a testing set that comprises the remaining 30%. We use the training set to estimate our models, and keep the testing set as an out-of-sample dataset for performance validation. We use the concordance index (C-index) as the metric for model performance. This metric is commonly used in survival analysis to evaluate its predictive power. 48 It is a measure of the concordance between orderings of observed survival times and the predicted times or risks. (A C-index of 0.5 corresponds to a random model, while a value of 1.0 corresponds to a perfect model.) We use cross-validation to tune the hyperparameters of the models.
In addition to discriminative power, we assess the calibration of our models by comparing the actual and the predicted survival probabilities at 36, 48, and 60 months of overall survival. For each time cutoff, we divide the test set into quintiles based on the predicted risk scores. We then compute the average predicted score and the true survival probability observed in each of the quintiles. Last, we create calibration plots by plotting the observed probabilities against the predicted probabilities. In the ideal case, the points should lie as close as possible to the diagonal line, which represents perfect calibration.
Results
Summary statistics
Demographics of the dataset between 2005 and 2016.

The proportion of the population with multimorbidity has increased by approximately 2.5 percentage points over the last decade, with more than 17% of all patients having at least two chronic morbidities in 2016. We find that the prevalence and the severity of multimorbidity increase progressively with age (see Figure 3). By age 60, approximately half the population has been diagnosed with at least one chronic condition, after which we observe a steep rise in multimorbidity, with close to 1 in 3 patients having at least two morbidities by age 70. Stratifying the prevalence of multimorbidity by IMD, we find that people living in more deprived areas are more likely to be multimorbid compared to those living in more affluent areas at all ages. The same trend holds for all years in our data. (See Supplementary Material C.) Single-disease prevalence by age group in 2016. See Figures 1 and 2 for disorder to index mapping and MedDRA SOC abbreviations. See Supplementary Material C for breakout by age group and by year.
Individual, pairs, and triplets
Lift and odds ratio of the top 10 most prevalent multimorbidity pairs in 2016. See Supplementary Material C for other years. We include only the top four for the Infant age group due to the small sample size.

Hypertension is most associated with a second condition in the older age groups, although most pairs do not necessarily occur more frequently than by chance. The combination of hypertension and diabetes stands out with a relatively high lift and an odds ratio that is greater than 2.0. Angina and coronary artery disease (CAD) also demonstrate a strong association in the Aged and Elderly age groups with unusually high lift and odds ratio.
To better visualize the data, we plot the lift of all combinations of disease pairs in heat maps, stratified by MedDRA system organ classes. (See Figure 4 and Supplementary Material C for other age groups and years.) The co-occurrence of cardiac-cardiac and cardiac-respiratory disorders is a major risk across all age groups. We observe significant coupling between cardiac and hepatobiliary disorders in the Adolescent and Child (2–13 years old) age groups. On the other hand, combinations of cardiac-vascular and cardiac-metabolic disorders are the most dominant in the Middle-Aged and older age groups. We observe the same general patterns across time. Heat map of lift of multimorbidity pairs in the Aged subgroup in 2016. See Figures 1 and 2 for disorder to index mapping and MedDRA SOC abbreviations. See Supplementary Material C for other age groups and years.
Lift of the top 10 most prevalent multimorbidity triplets in 2016. See Supplementary Material C for other years. We exclude the Infant age group and include only the top five for the Child subgroup due to the small sample size.

Multimorbidity networks
In Figures 5 and 6, we plot the undirected and directed multimorbidity networks observed in the Aged age group in 2016. (See Supplementary Material C for other age groups and years.) Instead of a force-directed layout, we place the nodes in fixed positions around a circle to allow easy visualization of temporal changes in connections and clusters when comparing plots from different years. The edge thickness is proportional to the lift between each disease pair. Apart from single-node clusters, the communities detected using modularity maximization are given different colors. Undirected multimorbidity network in the Aged subgroup in 2016. Edge thickness is proportional to the lift between each disease pair. Intra-group edges and inter-group edges are represented by solid lines and dashed lines, respectively. Only communities with more than one node are colored. See Figures 1 and 2 for mapping of disorder to index and MedDRA SOC abbreviations. See Supplementary Material C for other age groups and years. Directed multimorbidity network in the Aged subgroup in 2016. Edge thickness is proportional to the lift between each disease pair. Intra-group edges and inter-group edges are represented by solid lines and dashed lines, respectively. Only communities with more than one node are colored. See Figures 1 and 2 for mapping of disorder to index and MedDRA SOC abbreviations. See Supplementary Material C for other age groups and years.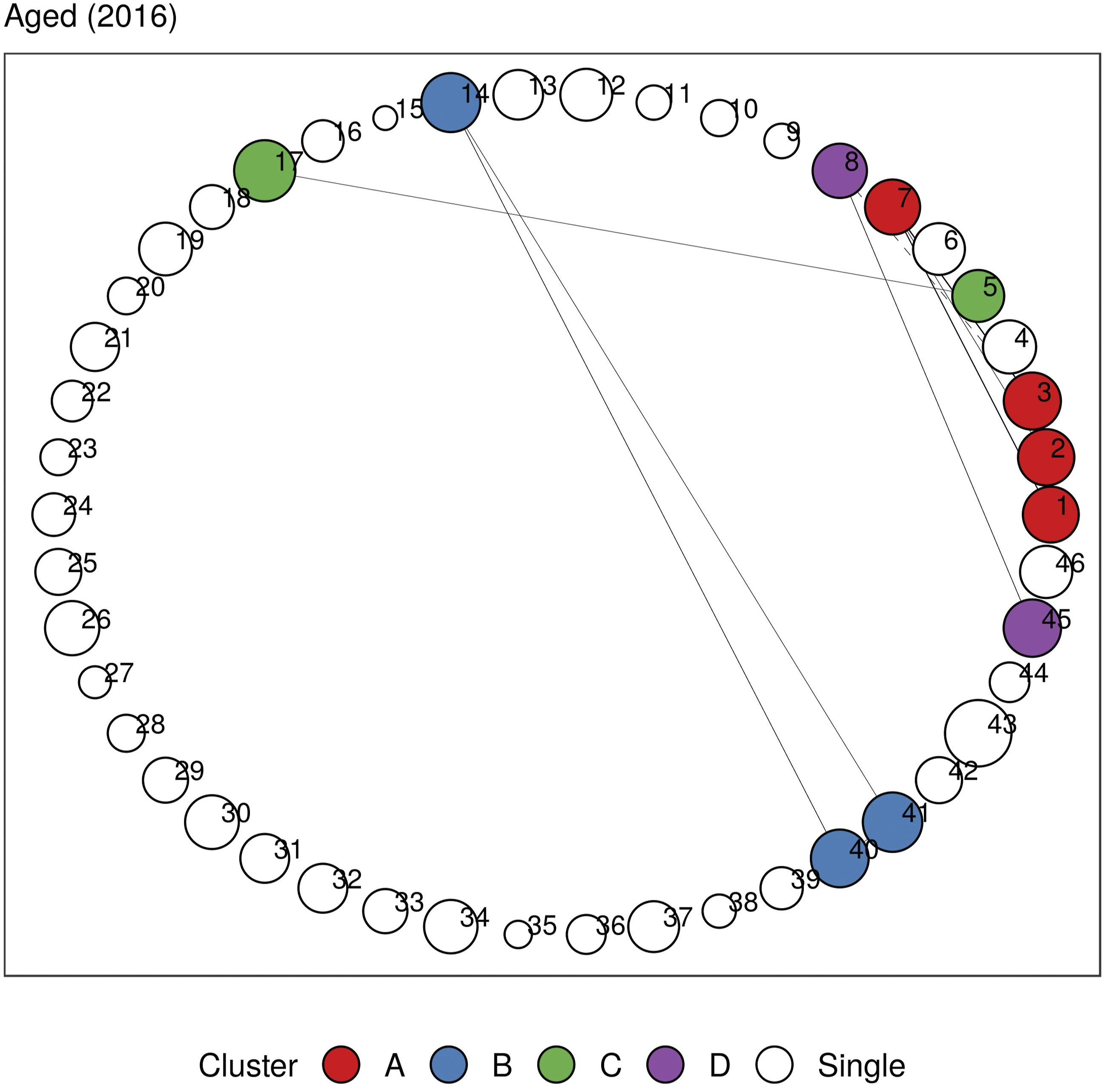

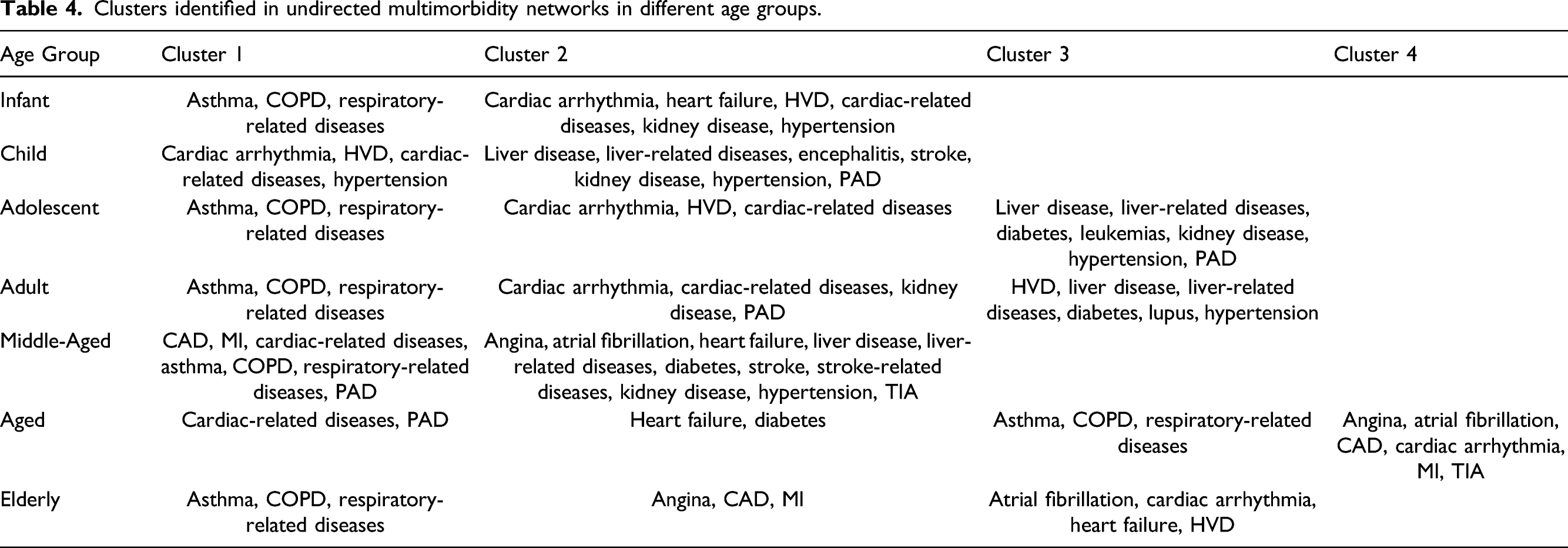
Clusters identified in undirected multimorbidity networks in different age groups.
Clusters identified in directed multimorbidity networks in different age groups.

A respiratory cluster of asthma, chronic obstructive pulmonary disease (COPD), and respiratory-related diseases appears to be present in all age groups in both undirected and directed graphs. Similarly, a vascular-metabolic-hepatobiliary-renal cluster that is characterized by hypertension, diabetes, liver diseases, and kidney diseases, with the occasional appearance of cardiac disorders, is also present in almost all cohorts. As observed in previous analyses, we also find several clusters dominated by cardiovascular disorders such as angina, CAD, myocardial infarction (MI), atrial fibrillation, cardiac arrhythmia, heart failure, heart valve disorder (HVD), stroke, peripheral artery disease (PAD), and transient ischemic attack (TIA).
Centrality measures for top five diseases in undirected multimorbidity networks with mean computed over time.

Centrality measures for top five diseases in directed multimorbidity networks with mean computed over time. We exclude eigencentralities that are close to zero.

We observe similar results in directed networks. In general, hypertension, diabetes, and respiratory-related diseases demonstrate high in-degree centrality and eigencentrality, while cardiac disorders show high out-degree centrality. In the Middle-Aged and younger age groups, asthma emerges as a new central node with high in-degree centrality, while the top five diseases for the Aged and Elderly age groups remain dominated by cardiovascular diseases.
Survival analysis
We summarize the dataset used for survival analysis in Supplementary Material D. The sample consists of approximately 390,000 patients in the Aged and Elderly age groups for each year between 2010 and 2012. More than 50% of the patients are multimorbid. In terms of predicting five-year overall survival, we find the performance of the linear and nonlinear survival models explored to be very similar. We focus on the Cox model here due to its ease of interpretability. The model achieves a promising C-index of 0.81 (95% CI 0.80–0.81) on out-of-sample data in 2012. In addition, its calibration curves lay close to the ideal diagonal, indicating that the model is well calibrated, that is, the model does not systematically overestimate or underestimate survival rates in any of the quintiles. (See Supplementary Material C for plots.)
We extract the top ten coefficients in the Cox model to identify specific risk factors (see Supplementary Material D). To correct for multiple testing, we perform the Benjamini–Hochberg adjustment with a 5% false discovery rate for identifying significant factors. Apart from cancers, we find the presence of multimorbidity to be a strong adverse risk factor, that is, the higher the number of co-occurring chronic conditions, the greater the mortality risk. For example, the hazard ratio of having four or more chronic conditions is 2.44 (95% CI 2.22–2.69). We also find a high IMD, corresponding to a lower socioeconomic status, to be significantly associated with increased risk, although this factor is not in the top ten coefficients.
Discussion
With multimorbidity becoming the norm rather than the exception,2,12,17,25,49,50 the management of multiple chronic diseases in older adults is a major challenge facing healthcare systems worldwide. It is clear that a better understanding of the epidemiology of multimorbidity is required to develop more effective preventive interventions and better primary medical care for multimorbid patients. In this paper, we use data-driven methods to characterize multimorbidity patterns in different demographic groups and their evolution over the past decade, using a large, representative electronic medical records database consisting of over 4.5 million patients.
Consistent with other studies, we find that the prevalence and severity of multimorbidity increase substantially with age. In addition, we observe social inequalities in multimorbidity, with patients in socioeconomically deprived areas more likely to be multimorbid.11,12,38,49,51–53 Our findings also support the role of hypertension as an important risk factor in older adults, as reported in the literature.2,40,54,55 Hypertension is one of the most prevalent and most central chronic conditions in our dataset, and one that serves as an important bridge between many diseases in our networks. Other trends identified in our analysis, such as the falling prevalence of angina56–59 and the growing prevalence of diabetes, 60 are also well documented in previously published population studies.
In our pairwise analysis, we find strong association between multiple pairs of chronic conditions, including between asthma and respiratory-related diseases61,62 in the Adolescent age group, between hypertension and diabetes28,63–65 and between CAD and angina 66 among older patients, and between cardiovascular and respiratory disorders in all age groups. 2 Triplets involving cardiovascular and metabolic disorders, such as CAD, hypertension, and diabetes, also occurred more frequently than expected by chance.2,25,28,67,68
Our network analysis further identified several meaningful communities that are common across all demographics, including a respiratory cluster (e.g., asthma and COPD), 69 a cardiovascular cluster,19,70,71 and a mixed cardiovascular-renal-metabolic cluster,39,72–74 all of which are supported by either established pathophysiological mechanisms or shared risk factors. For example, it is well known that cardiovascular diseases are one of the most common complications of diabetes. While we do not find any particular multimorbidity pattern to have a significant effect on mortality, our models do indeed verify the substantial burden of multimorbidity (as quantified by the number of co-occurring chronic conditions) on overall survival in older patients.7,12,26,75,76
However, we must emphasize that our results do not necessarily imply any causal link between diseases identified to be in the same cluster. The association might be attributable to shared risk factors (e.g., smoking) or other adverse events, and any temporal relationships to be inferred from the multimorbidity directed networks might be administrative in nature (e.g., incomplete medical records that are rectified in subsequent visits) or biased by delayed diagnosis.
In general, the lack of an accepted standard for defining multimorbidity makes it difficult for any meaningful comparison of results across different studies.77,78 Moreover, because results can be highly dependent on the study population, the disease ontology used, and the number of chronic conditions considered, it is not uncommon for studies to report seemingly conflicting findings. In this paper, we consider a wide range of demographic groups and a total of 46 morbidities, which is more than most similar studies, 11 and well above the minimum of 11–12 as recommended by systematic reviews in this field of research.78,79 In addition, our findings are largely consistent with existing studies in the medical literature.
Lastly, we note that cancer appears to be under represented in the THIN database. This is because many cancer patients are treated separately in cancer centers under the care of specialized clinical teams. Unfortunately, data on such patients rarely make their way back to the primary care clinics where the THIN data is collected, leading to a gap in this area.
Conclusions
Current healthcare systems are largely centered on single-disease approaches to treatment, resulting in the fragmentation of care and a lack of continuity in the management of multiple diseases. Even most clinical trials exclude multimorbid patients. Because multimorbidity is more common in disadvantaged groups, the current structure exacerbates health inequalities in society.
In this paper, we apply statistical methods and network analysis to characterize multimorbidity associations in the general UK population using a large electronic medical records database spanning the years 2005–2016. We find that the proportion of the population with multimorbidity has increased over the last decade, and the prevalence and severity of multimorbidity increase substantially with age. We identify strongly associated comorbid pairs of cardiac-vascular and cardiac-metabolic disorders. In addition, our clustering algorithm reveals three principal clusters: a respiratory cluster, a cardiovascular cluster, and a mixed cardiovascular-renal-metabolic cluster. In our directed network analysis, hypertension, diabetes, and respiratory-related diseases demonstrate high in-degree centrality, while cardiac disorders show high out-degree centrality. Our findings largely confirm and expand on the results of existing studies in the literature. We believe that our results contribute to a better understanding of multimorbidity that may be useful for the early detection and prevention of comorbidities, for example, prescribing lifestyle interventions (i.e., adopting healthy dietary and exercise regimens) to hypertension patients as a preventive measure for diabetes. 80
There is a pressing need for a universal framework that standardizes the way that multimorbidity is assessed (e.g., the appropriate number of diseases and the choice of chronic conditions to include) in order to facilitate comparisons between studies and populations. With the “Omics” revolution, the combination of phenotypic, genomic, and epigenomic data has the potential to provide deeper insights into the underlying pathophysiological associations between comorbid diseases. Unfortunately, the availability of such linked datasets remains very limited. Further research is also needed to better understand the impact of multimorbidity on different health outcomes, such as quality of life and healthcare costs, in order to align the healthcare system more closely to the needs to multimorbid patients.
Supplemental Material
Supplemental Material—Multimorbidity and mortality: A data science perspective
Supplemental Material for Multimorbidity and mortality: A data science perspective by Kien Wei Siah, Chi Heem Wong, Jerry Gupta, and Andrew W Lo in Journal of Multimorbidity and Comorbidity
Footnotes
Acknowledgments
We thank Christoph Nabholz for supporting the project and Jayna Cummings for editorial support. Research support from the MIT Laboratory for Financial Engineering is gratefully acknowledged. The views and opinions expressed in this article are those of the authors only, and do not necessarily represent the views and opinions of any institution or agency, any of their affiliates or employees, or any of the individuals acknowledged above.
Author contributions
Conceptualization, K.W.S., C.H.W., J.G., and A.W.L.; resources, J.G., and A.W.L.; methodology, K.W.S., C.H.W., J.G., and A.W.L.; software, K.W.S.; formal analysis, K.W.S.; writing—original draft, K.W.S. and A.W.L.; writing—review and editing, K.W.S., C.H.W., J.G., and A.W.L.; supervision, A.W.L.; project administration, J.G. and A.W.L.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: K.W.S. and C.H.W. declare no competing interests. J.G. is an employee of Swiss Re and declares no competing interests. A.W.L. reports personal investments in private biotech companies, biotech venture capital funds, and mutual funds. A.W.L. is a co-founder and partner of QLS Advisors, a healthcare analytics and consulting company; an advisor to Apricity Health, Aracari Bio, BrightEdge Ventures, Enable Medicine, FINRA, Lazard, NIH/NCATS, Quantile Health, SalioGen Therapeutics, Swiss Finance Institute, and Thalēs; and a director of AbCellera, Annual Reviews, Atomwise, BridgeBio Pharma, and Roivant Sciences. During the most recent six-year period, A.W.L. has received speaking/consulting fees, honoraria, or other forms of compensation from: AbCellera, AlphaSimplex Group, Annual Reviews, Apricity Health, Aracari Bio, Atomwise, Bernstein Fabozzi Jacobs Levy Award, BridgeBio Pharma, Cambridge Associates, Chicago Mercantile Exchange, Enable Medicine, Financial Times, Harvard University, IMF, Journal of Investment Management, Lazard, National Bank of Belgium, New Frontier Advisors/Markowitz Award, Oppenheimer, Princeton University Press, Q Group, QLS Advisors, Quantile Health, Research Affiliates, Roivant Sciences, SalioGen, Swiss Finance Institute, and WW Norton.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: No direct funding was received for this study; general research support was provided by the MIT Laboratory for Financial Engineering and its sponsors. The authors were personally salaried by their institutions during the period of writing (though no specific salary was set aside or given for the writing of this paper).
Data availability
The data that support the findings of this study are available from The Health Improvement Network (THIN; ![]() ) but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are however available from the authors upon reasonable request and with permission of THIN.
) but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are however available from the authors upon reasonable request and with permission of THIN.
Supplemental material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
