Abstract
Infertility is a complex condition influenced by genetic and biological factors. In men, it is often caused by abnormalities in chromosome number or structure, alterations in DNA structure, specific gene mutations, or missing segments of the Y chromosome. Recent studies have also highlighted the impact of epigenetic mechanisms—such as DNA methylation, histone modifications, and noncoding RNAs—on gene function and fertility. Abnormal methylation patterns, particularly in genes such as DNMT3A/B/L, H19, MEST, SOX30, and those involved in the piRNA pathway, have been linked to poor sperm production and reduced fertility. These epigenetic changes can lower sperm quality, impair embryo health, and decrease the chances of success with fertility treatments. The role of biological and epigenetic factors in female reproduction is more limited. This narrative review aims to examine well-established genetic and epigenetic mechanisms associated with infertility, with a focus on key molecular pathways and regulatory processes. A literature review was conducted to summarize the most relevant and recent publications addressing genetic and epigenetic mechanisms as contributing factors to human infertility.
Plain language summary
Infertility is a complex health issue that can be caused by both genetic and biological factors. In men, common causes include problems with the number or structure of chromosomes, changes in DNA, specific gene mutations, or missing pieces of the Y chromosome. Research also shows that epigenetic mechanisms—changes that affect how genes work without altering the DNA sequence—play a role in fertility. These include DNA methylation, histone modifications, and non-coding RNAs (ncRNAs). Abnormal methylation in certain genes, such as DNMT3A/B/L H19, MEST, SOX30, and genes involved in the piRNA pathway, has been linked to poor sperm production and reduced fertility. Such changes can lower sperm quality, harm embryo health, and reduce the success rates of fertility treatments. The influence of genetic and epigenetic factors in female reproduction was also reviewed, although there is less data available. This article summarizes current knowledge on the most important genetic and epigenetic mechanisms involved in infertility, focusing on key molecular pathways and regulatory processes. It is based on a review of recent and relevant scientific publications on this topic.
Introduction
Infertility is a highly heterogeneous disorder in which genetic, epigenetic, and environmental factors play a role in its etiology, resulting from a wide variety of causes affecting both germ and somatic cells, including single-gene mutations, incomplete development of the testis, hormonal imbalance, immaturity of somatic cells, and an increase in apoptosis and inflammation in men. 1 Nevertheless, not all the causes of the decrease in spermatogenesis have been clarified.
The ASRM and AUA recommend considering karyotype analysis and Y chromosome microdeletion testing for men with non-obstructive azoospermia (NOA) or severe oligospermia (<5 × 106/mL). Genetic abnormalities are identified in approximately 2%–7% of infertile men and in up to 35% of those with NOA or severe oligozoospermia. Currently, there is no clinically available NOA gene panel for diagnosing patients with NOA or other types of male infertility. 2 Therefore, a considerable proportion of male infertility cases remains classified as idiopathic, with the precise molecular mechanisms largely unknown. Recently, the association between infertility and aberrant DNA methylation of the whole genome, specific genes, or male or female imprinted genes in spermatozoa was shown in many studies. In addition, many studies have indicated a role for different histone tail modifications and short noncoding RNAs (ncRNAs) in infertility.
A literature review was conducted to summarize the most relevant and recent publications addressing genetic and epigenetic mechanisms as contributing factors to human infertility. PubMed was searched with terms genetic, epigenetic, single gene, structural and numerical aberrations, sperm DNA methylation, genomic imprinting, X chromosome inactivation (XCI), male infertility, and female infertility. In this narrative review, we focus on the recent findings and summarize the current data on genetic and epigenetic mechanisms, such as aneuploidies, such as Klinefelter and Turner, Y chromosome microdeletion, and gene mutations, DNA methylation, and different metabolites produced during methylation and demethylation in women and men. In addition, the molecular mechanisms of genomic imprint formation and XCI, and their relationship with male and female infertility.
Genetic mechanisms of infertility
Infertility is a highly heterogeneous disorder in which both genetic and environmental factors play a role in its etiology. Approximately 15% of the cases are estimated to result from genetic causes. 3 These genetic causes include numerical and structural chromosomal aneuploidies, structural aberrations, autosomal gene mutations, and Y chromosome microdeletions in men, as well as structural chromosomal aneuploidies, structural aberrations, and single-gene mutations in women (Figure 1).
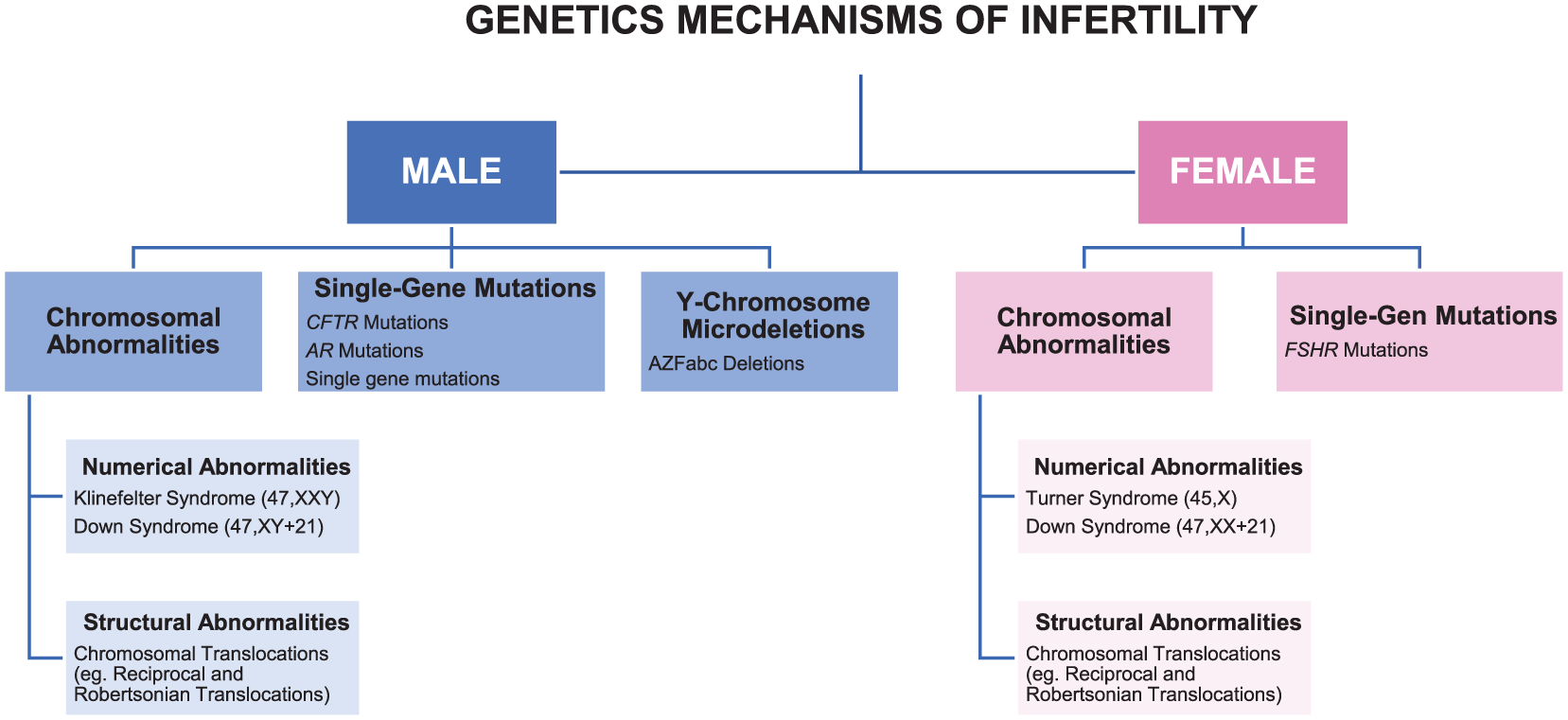
Genetic abnormalities causing infertility in men and women.
Chromosomal abnormalities
Chromosomal abnormalities refer to numerical or structural deviations from the normal chromosomal composition and represent a significant risk factor for fertility, including distinct alterations of sperm count and quality. 3 These anomalies can be observed in autosomal chromosomes as well as in sex chromosomes.
Numerical abnormalities
Klinefelter syndrome (47,XXY)
Klinefelter syndrome (KS) is the most prevalent chromosomal abnormality related to male infertility and is present in approximately 1 in 500–1000 male newborns. 4 This rate increases to 1%–2% among infertile males and up to 11% among azoospermic males. 5 The syndrome is characterized by an extra X chromosome in males, and these individuals are karyotyped as 47,XXY. The occurrence of this karyotype results from nondisjunction during gametogenesis in meiosis I or II in one of the parents. 6 Affected males present with gynecomastia, low testosterone levels, and small and firm testes, accompanied by hyalinization and fibrosis of the seminiferous tubules. 3 Along with these histological changes, extensive apoptosis of spermatogonial germ cells occurs, and testosterone secretion from Leydig cells is reduced. The progressive loss of germ cells leads to impaired spermatogenesis and, hence, azoospermia, while decreased androgen secretion results in hypogonadism. 7 These conditions, in turn, contribute to infertility. Although 91%–99% of males with the syndrome are estimated to be azoospermic, not all patients exhibit azoospermia. 8 The remaining portion includes males with ejaculated sperm, 8.4% of whom are classified as oligospermia. 8 An important factor influencing the presence of sperm is mosaicism, as the existence of germ cells with a 46,XY karyotype alongside 46,XXY results in a milder phenotype and enables spermatogenesis. 9 Although it was initially believed that only 46,XY cells could successfully complete meiosis, subsequent findings have demonstrated the presence of sperm in the ejaculate of non-mosaic patients. 9 These observations suggest that 46,XXY cells may also have the potential to complete meiosis and contribute to spermatogenesis. 10
Patients who desire fatherhood are advised to undergo microsurgical testicular sperm extraction (microTESE). The sperm retrieved through this method can be used in assisted reproductive techniques (ART) such as in vitro fertilization (IVF) or intracytoplasmic sperm injection (ICSI), enabling them to achieve parenthood. Age is a crucial factor in the success of this procedure. It has been observed that the previously mentioned histological changes and the progressive loss of germ cells begin at an early stage, with sperm production gradually declining with advancing age. 7 Due to the higher sperm retrieval rates in younger patients, early fertility preservation is recommended. 11 Although the optimal age for cryopreservation is a subject of debate, it is important to consider it between the ages of 15 and 35. 8 A recent systematic review and meta-analysis (SRMA) showed that men with non-mosaic KS have similar sperm retrieval success rates whether the procedure is done in adolescence or during reproductive adulthood, though success may decline after age 40. 12 Despite modest overall live birth rates due to challenges with sperm retrieval and assisted reproduction, patients can be reassured that waiting until mid-adulthood for sperm retrieval will not significantly impact their chances of achieving biological fatherhood. The main question is whether 47,XXY spermatogonia can complete meiosis or lose the supernumerary X chromosome and revert to a normal 46,XY karyotype. Therefore, genetic counseling is essential to determine whether PGT or prenatal genetic testing should be offered. 13
Turner syndrome (45,X)
Turner syndrome (TS), the most common sex chromosome anomaly in females, affects approximately 1 in 2000–2500 live-born girls. 14 While the condition is primarily characterized by the partial or complete loss of one X chromosome, resulting in a typical 45,X karyotype, some cases exhibit mosaicism, X isochromosome formation, or the presence of a partial or complete Y chromosome. 15 The integrity of the X chromosome is crucial for female fertility; therefore, deficiencies in this chromosome are closely associated with infertility. 15 Gonadal dysgenesis, a hallmark of Turner syndrome and a major cause of infertility, underlies the accelerated depletion of ovarian germ cells. 16 In Turner syndrome, there is a remarkably rapid atresia of the ovarian germ cell pool. 17 Initiated during the prenatal period and continuing in the early postnatal stage, accelerated oocyte apoptosis and disrupted folliculogenesis lead to premature ovarian failure, causing most patients to experience menopause during childhood or early adulthood.17,18 Moreover, the karyotype plays a significant role in the progression of the disease. Mosaicism alleviates the symptoms, increasing the spontaneous menarche rate from 4% to 12% in monosomic cases to 28%–45%. 18
Various candidate genes located on the X chromosome have been associated with ovarian function and gonadal insufficiency in Turner syndrome. Among the genes identified with expression and methylation differences in Turner syndrome patients are BMP15, KDM6A, ZFX, and USP9X.16,18 BMP15, which is expressed in oocytes, encodes bone morphogenetic protein 15, a member of a protein family that regulates ovarian function, and stimulates anti-Müllerian hormone (AMH) production and folliculogenesis. USP9X, a ubiquitin-specific protease, is thought to play a role in oogenesis, whereas KDM6A, a histone demethylase, and ZFX, a DNA-binding protein functioning as a transcription factor, regulate the expression of genes involved in germ cell development. 17
Spontaneous pregnancy occurs in approximately 10% of Turner syndrome cases. 19 Given the risk of premature ovarian insufficiency (POI) due to the rapid depletion of ovarian follicles, fertility preservation efforts in Turner syndrome patients should be initiated at an early age. The preservation options include oocyte cryopreservation for those who experience spontaneous puberty and ovarian tissue cryopreservation for others, though success rates remain largely unknown. Predictive markers of fertility potential include mosaic karyotype, spontaneous puberty, and biomarkers such as AMH and follicle-stimulating hormone (FSH). 19 For those unable to conceive with their own gametes, alternative options such as oocyte donation (OD) or embryo donation and gestational carriers exist but require careful cardiological and reproductive management to ensure safety. A recent SRMA reported between 17% and 40% live birth rate per embryo transfer using OD in TS patients, indicating that OD represents a reasonably effective assisted reproductive technology option for women with TS who wish to experience pregnancy and motherhood. 20 However, research has shown that when TS patients use OD for fertility treatment, they face an increased risk of pregnancy complications, including gestational hypertension, preterm birth, fetal death, and miscarriage. 21 Pregnancies in individuals with TS carry higher risks of miscarriage, preterm birth, chromosomal abnormalities, and cardiovascular complications, necessitating specialized medical care. 22
Testicular disorder of sex development (46,XX)
Testicular disorder of sex development with 46,XX karyotype (46,XX TDSD) is an uncommon and diverse condition defined by the development of testicular tissue with 46,XX karyotype. Patients with sex-determining region Y (SRY)-positive 46,XX TDSD are usually asymptomatic males diagnosed in puberty or adulthood because of hypergonadotropic hypogonadism, microorchidism, and infertility due to azoospermia and Sertoli cells only. SRY-positive 46,XX TDSD results from an aberrant Y to X translocation during the paternal meiosis. Approximately 80% of the cases are SRY-positive with complete deletions of azoospermia factor (AZF) regions on Y chromosome, while the remaining cases are SRY-negative. 23 Patients with 46,XX TDSD present with varying clinical manifestations from adult males seeking fertility treatment with normal male external and internal genitalia to children brought to pediatric urology or general pediatric clinics due to ambiguous genitalia or micropenis. Testicular sperm extraction is not recommended in these patients, and adoption or IVF with a sperm donor should be considered. 24
Trisomy 21 (Down syndrome)
Down syndrome is the most common viable autosomal aneuploidy and is characterized by trisomy of chromosome 21. 22 In addition to being associated with various health problems, Down syndrome is frequently linked to impaired fertility in both sexes.25,26
The literature includes only a very limited number of cases in which men with Down syndrome have fathered children spontaneously.27,28 While a significant contributing factor is the lack of sexual activity and desire for parenthood due to cognitive impairment, partial gonadal dysfunction may also play a role. 29 Although males with Down syndrome typically enter puberty at the expected age and exhibit normal pubertal development, progressive gonadal impairment becomes evident over time. Post-pubertally, a decline in testicular volume is commonly observed alongside a compensatory rise in gonadotropin levels, including luteinizing hormone (LH) and FSH, indicating disrupted testicular function. 30 This dysfunction is further supported by reduced levels of AMH, collectively pointing to compromised activity of both Sertoli and Leydig cells. 31 These findings suggest that gonadal deterioration in men with DS advances with age despite initially normal pubertal onset.
Studies reporting childbirth in women with Down syndrome indicate that these women are indeed fertile. 32 As in males, both the onset and progression of puberty appear to be normal in females with Down syndrome. Although there are not many recent studies on the subject, the literature suggests that individuals with Down syndrome commonly exhibit ovarian abnormalities, including reduced follicle numbers and impaired follicular development. 33 Histological studies show predominantly quiescent ovaries with limited or absent folliculogenesis, suggesting that the condition disrupts normal ovarian maturation from early stages. It has also been reported that AMH levels, a reliable marker of the ovarian reserve, are significantly lower in women with Down syndrome, which may help explain the earlier onset of menopause observed in this population. 34
Infertility in men with Down syndrome is often attributed to defective spermatogenesis, although a lack of knowledge about the sexual act may also contribute. 27
Structural abnormalities
Translocations are classified among the balanced chromosomal abnormalities that do not alter the overall DNA content. These, of course, depend on whether the chromosomal breakpoints disrupt dominant genes or alter the expression of adjacent genes due to segment repositioning. 35 Balanced translocations that are compatible with life include reciprocal translocations and Robertsonian translocations. These translocation carriers typically exhibit normal behavior but are at increased risk of producing abnormal gametes, which can result in fertility issues. 36 During gametogenesis, translocated chromosomes form a quadrivalent structure in meiosis I, and while alternate segregation produces balanced gametes, other segregation patterns—adjacent-1, adjacent-2, 3:1, and 4:0—lead to unbalanced gametes. 35 The fusion of an unbalanced gamete with a normal one results in an unbalanced zygote, which typically leads to embryonic lethality in cattle. 37 This, in turn, contributes to pregnancy loss. Cytogenetic studies have revealed that balanced translocations are the most common cause of recurrent miscarriages.38,39
The impact of translocation is influenced by several factors, including the structure and size of the involved chromosomes, their gene content, the size of the translocated segments, the location of the breakpoints, and the sex of the carrier.35,36 According to Wu et al., translocations involving chromosomes 1–5, which are considered long chromosomes, may pose a risk to blastocyst formation. 36 In another study, translocations involving chromosome 2 were commonly associated with male infertility and recurrent pregnancy loss. 40 In addition, studies in the literature have also linked translocations involving chromosome 5, chromosome 10, and chromosome 17 with sperm parameters and male reproductive failure.40,41
The impact of sex on fertilization outcomes in translocation carriers remains controversial. While some studies suggest that being male may pose a risk factor, others report lower fertilization rates in female carriers, and additional studies indicate no significant influence of sex at all. 36 Nevertheless, the majority of studies in the literature have predominantly focused on the association between chromosomal translocations and male infertility. This is partly attributed to the increased susceptibility of sperm cells to chromosomal abnormalities due to inherent differences in gametogenesis. The higher number of mitotic and meiotic divisions during spermatogenesis raises the likelihood of chromosomal missegregations or structural rearrangements, including translocations. In individuals with chromosomal translocations, abnormal configurations during meiosis, such as improper chromosomal alignment and tension defects, can activate meiotic checkpoints, interrupting the normal progression of meiosis. 42 As a result, affected cells may undergo apoptosis, leading to spermatogenic failure and reduced sperm output. The effect of balanced translocations on sperm parameters is highly variable, depending potentially on the specific chromosomes involved, carriers may present with a wide range of clinical conditions, from normozoospermia to complete azoospermia. 43 Translocations involving the X and Y chromosomes have been especially associated with azoospermia. 44 Another study reported that asthenozoospermia and teratozoospermia were more frequently observed in carriers of autosomal reciprocal translocations. 45 Similarly, Robertsonian translocations have been shown to lead to male infertility by causing oligozoospermia or azoospermia. 46 Moreover, in their retrospective study, Chen et al. demonstrated an association between Robertsonian and autosomal reciprocal translocations and reduced sperm concentration and motility. 42 Dong et al., in their study involving 1056 infertile men, reported that chromosomal translocations may be associated with reduced testicular volume and decreased testosterone levels, which can impair spermatogenesis and lead to conditions such as azoospermia or oligozoospermia. They also suggested that Robertsonian translocations were seven times, and reciprocal translocations were fifteen times more frequent in infertile men compared to the general population. 47
Complex chromosomal rearrangements
Complex chromosomal rearrangements (CCRs) are inherited structural changes in chromosomes that affect at least three chromosomes or involve multiple breakpoints. These conditions are undetected in the general population. Men with CCRs are diagnosed primarily due to infertility; however, women with CCRs are generally diagnosed with a history of repeated pregnancy loss or with an affected child with multiple congenital anomalies. CCRs develop through several different cellular processes, including DNA strand breakage, errors during DNA replication, DNA repair, or genetic recombination. These findings demonstrate the need for making cytogenetic analysis in azoospermic and severely oligozoospermic men, as well as women with recurrent pregnancy loss or those who have had a child with multiple congenital anomalies. 48
Single-gene mutations
Numerous genes play a role in the processes of spermatogenesis and oogenesis. Therefore, it is challenging to predict the number of single-gene mutations that may lead to infertility and address all of them comprehensively. In addition, during the normal germ cell development, multiple stages of mitosis take place, and in each of these mitoses, one mutation takes place. Therefore, single-gene mutations may be higher spermatozoa and may increase the gene mutations in men. 49
Next-generation sequencing (NGS) studies have uncovered many potential gene–disease relationships (GDRs). A recent review classifies GDRs underlying monogenic forms of human male infertility and their clinical validity. Most well-established GDRs were associated with isolated infertility phenotypes and endocrine disorders or reproductive system syndromes, whereas a smaller number were related to syndromic infertility. 50
Genes associated with reproductive system syndrome infertility include AR (Partial androgen insensitivity syndrome); AMH and AMHR2 (Persistent Müllerian duct syndrome); ANOS1, CCDC141, FGF1, FGF8, FGFR1, HS6ST1, KISS1R, PLXNA1, PROK2, PROKR2, SEMA3A, SOX10, TACR3, and WDR11 (Kallmann syndrome); BMP4, BMP, and BNC2 (Hypospadias); ANOS1, CHD7, GNRH1, SOX2, GNRHR, KISS1R, and LHB (Isolated hypogonadotropic hypogonadism); CYP17A1, DHX37, SOX9, SRD5A2, SRY, MAMLD1, and MYRF (46,XY Disorders of sexual development); WT1 (46,XY Disorders of sexual development (Prader scale 4; 5 or 6) without Wilm’s tumor); SOX3 (46,XX Disorders of sexual development), CYP19A1 (46,XX Disorders of sexual development (Prader scale 4; 5 or 6) due to aromatase deficiency); CYP21A2 (Classic congenital adrenal hyperplasia or non-classic adrenal hyperplasia). 50
ADGRG2 and CFTR (congenital bilateral absence of the vas deferens); AR, DMRT1, M1AP, MEI1, STAG3, SYCP3, TEX11, TEX14, TEX15, and XRCC2 (NOA); AURKC (macrozoospermia); CFAP91 and DPY19L2 (globozoospermia), FANCM and KLHL10 (oligozoospermia); ARMC2, CFAP251, CFAP43, CFAP44, CFAP65, CFAP69, CFAP91, DNAH1, DNAH17, FSIP2, QRICH2, SEPTIN12, SPEF2, and TTC29 (confidently linked to MMAF); PLCZ1 (fertilization failure); PMFBP1, SUN5, and TSGA10 (acephalic spermatozoa); SYCP2 (severe oligozoospermia); USP26 (azoospermia or oligozoospermia) are genes represent the best-characterized monogenic causes of isolated male infertility to date. 50
Mutations in several genes contribute to syndromic phenotypes in which infertility presents as part of a broader clinical spectrum, such as APOA1 (testicular amyloidosis); CATSPER2 (deafness infertility syndrome); CCDC39, CCDC40, DNAAF2, DNAAF4, DNAAF6, LRRC6, and RSPH3 (primary ciliary dyskinesia); CDC14A (oligoasthenoteratozoospermia); CEP290 (Leber congenital amaurosis); FANCA (occult fanconi anemia), MNS1 (asthenoteratozoospermia), NLRP3 (Muckle-Wells syndrome), PKD1 (polycystic kidney disease and asthenozoospermia); SPEF2 (primary ciliary dyskinesia with multiple morphological abnormalities of the sperm flagellum); TRIM37 (Mulibrey nanism). 50
Collectively, these findings underscore the genetic heterogeneity of male infertility, encompassing both isolated defects of spermatogenesis and syndromic disorders where infertility arises as part of a broader clinical phenotype.
Cystic fibrosis (CFTR gene mutations)
The cystic fibrosis transmembrane conductance regulator (CFTR) is a cAMP-activated anion channel primarily involved in the epithelial transport of chloride and bicarbonate ions, playing a critical role in maintaining hydration and viscosity of secretions across various organ systems, including the male reproductive tract. 51 Mutations in the CFTR gene, which span over 2000 identified variants, can impair protein synthesis, processing, gating, or stability, leading to a spectrum of conditions ranging from classic cystic fibrosis (CF) to milder CFTR-related disorders (CFTR-RDs). 52
Nearly all male patients with CF are azoospermic due to congenital bilateral absence of the vas deferens (CBAVD), clearly demonstrating the impact of the CFTR gene in reproduction. 53 The detection of CFTR mutations in cases of absence of the vas deferens suggests that an adequate amount of functional CFTR protein is essential for the proper development of the vas deferens. 54 Beyond CBAVD-associated obstructive azoospermia (OA), CFTR mutations have also been proposed to contribute to non-obstructive male infertility by affecting processes such as spermatogenesis, sperm capacitation, maintenance of the blood-testis barrier integrity, and the regulation of various signaling pathways involved in reproductive processes. 55
The role of CFTR mutations in male infertility has been well-studied; however, their impact on female reproductive health has not been sufficiently elucidated. In females, CFTR dysfunction can contribute to infertility through disrupted chloride and bicarbonate transport, which alters cervical and endometrial mucus hydration and consistency. 56 These changes may hinder sperm migration through the reproductive tract, interfere with fertilization, and impair endometrial receptivity. In addition, abnormal CFTR expression may affect uterine fluid dynamics and hormonal regulation, further compromising the environment needed for successful conception and implantation.
Androgen insensitivity syndrome (AR gene mutations)
The androgen receptor (AR) plays a pivotal role in regulating spermatogenesis by mediating the effects of testosterone and dihydrotestosterone within the testis. AR is expressed in Sertoli cells, peritubular myoid cells, and Leydig cells, where it modulates the expression of genes critical for germ cell development and testicular function. Disruption or dysfunction of AR signaling impairs spermatogenesis, leading to infertility. 56 Mutations or altered expression of the AR gene have been associated with various forms of male infertility, including idiopathic azoospermia and androgen insensitivity syndromes (AISs), which are classified as complete (CAIS), partial (PAIS), or mild (MAIS), highlighting its essential role in male reproductive health.57,58
Since the AR gene is located on the X chromosome, AIS is classified among X-linked disorders. Although considered a rare disease, it represents one of the most common XY disorders of sex development. 59 The phenotypic variability observed in AIS is determined by the degree of residual AR function and ranges from an utterly female phenotype to a male phenotype characterized by infertility and/or incomplete virilization. 60 CAIS results in a female phenotype with external genitalia and absent Müllerian structures, while testes are typically undescended. PAIS, which represents an intermediate level of AR activity, leads to varying degrees of hypospadias and ambiguous genitalia. MAIS, the mildest form, presents with normal male external genitalia but under-virilization, such as reduced body hair and infertility. 61 While infertility is inevitable in CAIS and PAIS, it has been suggested that fertility potential may be preserved in MAIS. 59
Follicle-stimulating hormone receptor (FSHR) mutations
FSH is essential for gonadal development, sexual maturation during puberty, and the production of gametes. It exerts its effects by binding to the FSH receptor (FSHR) located on the plasma membrane of granulosa cells in the ovaries and Sertoli cells in the testes. 62 Proper functioning of this receptor is vital for follicular development and estradiol production in females, and for regulating Sertoli cell activity and spermatogenesis in males. 63 The gene encoding the FSHR is located on the short arm of chromosome 2 (2p), and numerous mutations and polymorphisms have been identified within this gene. 64 Given the receptor’s central role in reproductive function, such genetic variations are not unexpected in individuals with infertility.
Mutations in the FSHR gene may affect the receptor’s structure and ligand-binding capacity, alter its substrate specificity, and interfere with multiple signaling pathways involving FSH, including those responsible for second messenger production and downstream cellular responses. FSHR mutations are primarily classified as either activating or inactivating, depending on their impact on receptor function. Inactivating mutations exert effects ranging from impaired receptor function to a complete functional blockade, resulting in disrupted receptor-ligand complex formation and, consequently, impaired FSH signal transduction. 64 The clinical manifestations of these mutations in females include primary or secondary amenorrhea, ovarian dysgenesis, polycystic ovary syndrome (PCOS), and infertility.63–65 In males, they are associated with small testicular volume and impaired spermatogenesis; however, studies have indicated that fertility is not directly affected. 63 On the other hand, activating mutations in the FSHR gene lead to constitutive receptor activity and a loss of ligand specificity, rendering the receptor abnormally responsive to human chorionic gonadotropin (hCG) and thyroid stimulating hormone (TSH). Stimulation of mutant FSHRs by elevated levels of these hormones, particularly hCG, has been implicated in the development of ovarian hyperstimulation syndrome (OHSS). To date, all activating FSHR mutations identified in women have been associated with spontaneous OHSS. 63
Y chromosomal microdeletions
The Y chromosome contains vital genes involved in male sex determination, testicular development, and spermatogenesis (Figure 2). 66 In particular, the AZF region located on its long arm is critically important for sperm production. Microdeletions of the 16 genes identified within this region have been associated with male infertility. 67 The long arm of the Y chromosome contains numerous ampliconic and palindromic sequences, which render this region particularly susceptible to deletions arising from recombination errors during spermatogenesis. 66 Yq deletions represent the most common known genetic cause of spermatogenic failure, and such microdeletions are frequently identified in cases of NOA and severe oligospermia. 68 Accordingly, Y chromosome microdeletion screening is now routinely performed in the diagnosis and evaluation of male infertility. 68

AZF regions on Y chromosome.
The AZF region, located on Yq11, is divided into three subregions: AZFa, AZFb, and AZFc, with AZFc microdeletions. 69 The most commonly observed complete deletion is in the AZFc region (70%–80%), followed by deletions in AZFa (0.5%–9%), AZFb (1%–7%), and the combined AZFbc region (1%–20%). 70 Deletions may involve a single subregion or multiple subregions of the AZF locus, and the resulting phenotypic manifestations vary depending on the specific region(s) affected. While AZFa and AZFb deletions are typically associated with azoospermia, AZFc deletions do not invariably lead to azoospermia. A full deletion of the AZFa region completely halts spermatogenesis in the seminiferous tubules, leading to Sertoli cell-only syndrome (SCOS) and resulting in azoospermia. AZFb deletions disrupt spermatogenesis at the primary spermatocyte stage, where mature sperm are absent, although earlier germ cells may still be present in testicular biopsies. 71 In contrast, AZFc deletions exhibit a wider phenotypic spectrum, ranging from azoospermia to oligozoospermia. Therefore, unlike carriers of AZFa or AZFb deletions, individuals with AZFc deletions may still have the potential to father children through assisted reproductive technologies such as ICSI. 72
Epigenetic mechanisms of infertility
DNA methylation
DNA methylation and demethylation are the main epigenetic rearrangements of the human genome. Fertilization and early embryo development require normal sperm DNA methylation, affecting also the health of offspring in mammals. 73 DNA methylation is the transfer of a methyl to the fifth carbon atom of the cytosine by DNA methyltransferases (DNMTs), therefore resulting in the formation of 5-methylcytosine (5mC).74,75 DNA methylation is necessary for the regulation of tissue-specific gene expression in germ cell development, silencing of retroviral elements, XCI, and genomic imprinting.74–76 DNA methylation is a reversible chemical modification, and its removal is regulated by an active erasure mechanism primarily mediated by Ten-Eleven Translocation (TET) enzymes, including TET1, TET2, and TET. These enzymes catalyze the oxidation of 5mC to 5-hydroxymethylcytosine (5hmC), initiating DNA demethylation through either passive or active pathways. In passive demethylation, 5hmC is not recognized by DNMT1 during replication, resulting in progressive loss of demethylation in proliferating cells. 77 In the active demethylation, TET enzymes further convert 5hmC to 5-formylcytosine and then to 5-carboxylcytosine, which are subsequently excised by the base-excision repair machinery and replaced with unmodified cytosine. 74
Disruptions in this essential process contribute to the pathogenesis of many common diseases, affect reproductive potential in both men and women, and may also result in abnormal post-fertilization events and impaired embryo development. Appropriate DNA methylation is critical for proper chromatin condensation within the sperm head, a process that supports sperm maturation and optimizes its ability to fertilize the oocyte and participate in subsequent post-fertilization events. Methylation and demethylation of DNA have critical importance for the development of male germ cells.78–81 DNMT family catalyzes and regulates DNA methylation. DNMT1, maintenance DNMT, binds immediately to newly synthesized DNA following the replication and copies the previous methylation marks to the new sequence. 74 It can also repair DNA methylation aberrations and preserve the original methylation marks. 76 The other two DNMTs, DNMT3A and DNMT3B, de novo methyltransferases, can add methyl groups to previously unmethylated. 81 Despite their structural and functional similarities, DNMT3A is expressed in almost all tissues, while DNMT3B is more abundant in specific tissues, such as the testis.76,82 DNMT3L is the last member of the DNMT family and plays a role in methylation-mediated silencing of retrotransposons with PIWI-loaded piRNAs in the male germ cell line. 81 Studies have revealed that errors in silencing of retrotransposons, which constitute the vast majority of transposable elements, may cause absence of spermatozoa in the ejaculate.78,83,84 DNMT3L is required for spermatogenesis, and animal studies have shown that Dnmt3L null males cannot progress beyond the spermatocyte stage; therefore, they are entirely azoospermic. 85 Similarly, earlier research has shown that aberrant DNA methylation may play a key role in the onset of disorders such as PCOS, endometriosis, and POI. Such methylation defects can disrupt the regulation of genes involved in immune function and hormone production. 86
Methylation mechanism of imprinted genes
Genomic imprinting mechanism
Genomic imprinting is a nonrandom monoallelic expression of a gene depending on its parental origin. Imprinted genes are crucial for fetal and placental growth and development. This expression pattern results from epigenetic marking during the differentiation of male and female germ lines (Figure 3).87,88 Therefore, imprinted genes are found under strict parental (maternal or paternal allele expression) control.87,89 Recognizing these facts demonstrated that, despite the lack of genetic justification, chromosomes from both parents are required for normal development, highlighting genomic imprinting as a fundamental driving force in normal development. 89 The first gene associated with genomic imprinting was the maternally expressed and paternally repressed IGF2 gene, discovered in 1991. 90 In the same year, the paternally expressed IGF2 and the maternally expressed H19 genes were also identified.91,92 Allelic epigenetic marks are inherited from the parental germ line and regulate expression through DNA methylation, suppressing gene activity.92,93 This natural imprinting pattern has been identified in mammals and is highly conserved across imprinted gene clusters. Approximately 6%–10% of the genome is monoallelically expressed, or one allele is excluded from expression. Normal (canonical) imprinting is well-characterized in mammals and is highly conserved across imprinted gene clusters. This imprinting process is mediated by allelic DNA methylation.
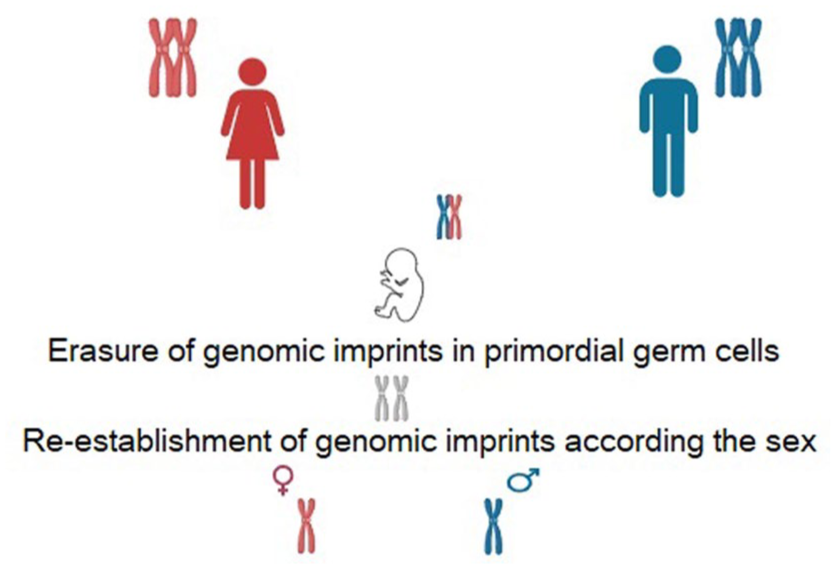

Maternal and paternal genomic imprinting.
Maternal and paternal origins of genomic imprinting
The parental origin of an imprinted allele plays a role in phenotype determination. In familial cases, pathogenic variants in imprinted genes are inherited in an autosomal dominant manner, but their penetrance depends on the contributing gender of the parent. In contrast, the sex of the affected individual does not influence phenotypic expression, and the prevalence of the phenotype is similar in both males and females. 89
It is well-established that DNA methylation is the primary driving force behind genomic imprinting. Imprinting control regions regulate the function of one or more imprinted genes during imprinting in oocytes and sperm. 94 In germ cells, de novo methylation is carried out by the methyltransferase DNMT3A and its cofactor DNMT3L. 95 In oocytes, DNA methylation is almost exclusively limited to transcribed genes. 96 In spermatozoa, however, the majority of the genome is highly methylated by DNMT3A and DNMT3L. 97 Unlike in oocytes, where methylation is linked to H3K36me3, spermatogenesis is associated with H3K36me2. 98
There is no de novo mechanism in the germline that specifically targets imprinted loci for methylation. However, in both oocytes and sperm, de novo DNMTs are dependent on H3K36 methylation. 99 Different DNA methylation characteristics exist between oocytes and sperm. For instance, oocytes contain approximately 2000 highly methylated CpG islands, whereas sperm lack these regions. 96 The methylation patterns transmitted through embryonic gametes are critical for defining and maintaining the number of permanent gametic differentially methylated regions (DMRs) and imprinted loci. Two proteins from the Krüppel-associated box-containing zinc finger protein family, ZFP57 100 and ZNF445, 101 have been identified as key imprinting protection factors. All methylation marks in primordial germ cells are erased through passive or active demethylation before being re-established according to parental origin during germ line development.74,102,103 (Figure 3). Passive demethylation occurs during DNA replication when newly synthesized DNA strands are not properly methylated due to DNMT1. Active demethylation, in contrast, does not require DNA replication and is mediated by one or more enzymatic factors. 104 The process continues through successive germ cell stages, including spermatocytes and spermatogonia, leading to the transmission of paternal methylation patterns to offspring via spermatozoa. Specific genes such as Igf2/H19, Rasgrf1, Dlk-Gtl2, and Zdbf2 are typically highly methylated and paternally imprinted. The H19 gene, located at 11p15.5, consists of five exons and four introns, encoding a 2.7-kb transcript involved in RNA processing and protein transport. H19 DMR is unmethylated on the maternal allele, allowing H19 expression while preventing the action of the IGF2 gene enhancer. On the paternal allele, H19 DMR methylation permits IGF2 gene expression. Maternally imprinted genes include MEST, ZAC1, PEG3, and SNRPN, which are typically unmethylated in spermatozoa and methylated in oocytes, leading to paternal allele expression. 105 The MEST gene, localized at 7q32, encodes an alpha/beta hydrolase family member and is paternally expressed during fetal development. It plays a crucial role in protein folding and fetal mesoderm development. A meta-analysis evaluating the methylation levels of MEST and SNRPN DMRs found that these genes were more highly methylated in infertile men compared to fertile men. 106 Methylation alterations in either maternal or paternal germ lines have been strongly linked to male infertility. In addition, such sperm DNA methylation changes may negatively impact sperm quality and success rates in ART. 107 Studies on human placentas have shown that H19 expression is higher in ICSI and IVF compared to natural conception.108,109 Furthermore, research suggests an increased incidence of Angelman syndrome in children born through ART.108,110
Non-imprinted genes
DNA methylation abnormalities in specific genes and genome-wide have been linked to male infertility.79,111–113 One key gene, MTHFR, involved in folate metabolism,114–116 shows varying methylation patterns in testicular tissues of infertile patients, suggesting a role in impaired spermatogenesis. Silencing of the MTHFR gene via the CpG promoter methylation could affect the function of sperm and also pregnancy outcome and embryo development. Despite the importance of folate in spermatogenesis, results across studies are inconsistent, and comprehensive and well-designed studies are necessary in certain phenotypic subtypes of infertility. 81
Other genes, such as DDR1, 117 SPATA16, 118 MAEL, 78 and genes related to the PIWI-interacting RNA (piRNA) pathway (e.g., PIWIL1, PIWIL2, and BOLL), also show promoter hypermethylation and altered expression in male infertility, indicating their involvement in germ cell development and transposon silencing. 119 SOX30, critical for sperm development, was found to be hypermethylated in NOA, with knockout studies in mice confirming its essential role. 120 Similarly, ZCCHC13 shows a negative correlation between promoter methylation and gene expression, suggesting epigenetic regulation in spermatogenic failure. 121 In addition, the genes involved in piRNA pathways, methylation changes in the maelstrom spermatogenic transposon silencer (MAEL) gene, which expresses a protein known to have a function in suppressing transposable elements, have also been reported in NOA.78,122 In NOA patients with hypospermatogenesis (HS), including idiopathic cases, it was found that CpGs in the promoter regions of boule homolog, RNA binding protein (BOLL), DEAD-box helicase 4 (DDX4), HORMA domain containing 1 (HORMAD1), and MAEL genes showed increased methylation and decreased mRNA expressions in the testis. 123 Mouse studies showed that MAEL was expressed in spermatocytes and elongated spermatids, and decreased piRNA levels and impaired silencing of LINE-1 in the testes of MAEL null animals. 122 In another study involving 26 patients with NOA and HS, a specific site known to affect the expression of the MAEL promoter was found to be hypermethylated in the testes of patients with HS compared to controls with normal spermatogenesis. 78 However, LINE-1 transcripts were also observed to increase in cell culture models obtained by enriching the germ cells of HS patients. 78 In addition, a deficiency in the expression of SOX30 from the SOX family, which is known to be necessary for male fertility, has been reported to reduce the expression of genes that provide hyperactivation and motility of sperm. 111 Testicular specimens from 58 patients with NOA were compared with those of patients with OA and more than 5800 genome-wide DMRs were analyzed. 124 This study showed that 25 CpGs in the SOX30 promoter were hypermethylated in patients with NOA and SOX30 methylation was strongly associated with testicular impairment in NOA. 124 Sox30 null mice showed complete arrest at the beginning of spermiogenesis, without spermatid or sperm production. 120 A recent study revealed that germ cell differentiation in the testes of Sox30 null male mice was halted at the zygotene stage of meiosis, and reexpression of Sox30 also reversed testicular damage. 124 In another study analyzing both gene expression and DNA methylation in testicular samples from patients with NOA and OA, the zinc finger CCHC-type containing 13 (ZCCHC13) gene exhibited a strong negative correlation between methylation and expression. 79 Immunohistochemistry results revealed that the level of ZCCHC13 protein decreased in NOA samples compared to OA. Using a mouse cell line, the researchers also reported that ZCCHC13 expression was higher in germ cells compared to Sertoli cells, and demethylation in the ZCCHC13 promoter increased its expression level. 79 Subsequent studies have shown that ZCCHC13 acts as a cell cycle regulatory signaling molecule in AKT and ERK pathways. 125 As a noninvasive method, it has been reported that the level of cell-free DNA promoter methylation may be associated with idiopathic NOA. 126 Testis-specific promoter methylations were correlated in both the seminal cell-free DNA and in testicular biopsy samples from patients with NOA and SCOS, MA, and HS phenotypes. 126 However, methylation variations were detected among patients with NOA, which may be due to epigenetic differences during gametogenesis. 126 Among testis-specific genes, especially cyclin A1 (CCNA1) and doublesex and mab-3 related transcription factor 1 (DMRT1), methylation patterns in cell-free DNA samples were different in HS compared to other NOA groups. 126
Until now, limited studies have performed global DNA methylation analyses from the tissues of infertile men. Although genome-wide methylation studies are important to identify candidate genes responsible for the DNA methylation changes observed in infertile men, the presence of somatic DNA is a condition that should be taken into account, especially in the epigenetic evaluation of testicular tissue. 119 High-throughput studies in determining the DNA methylation profile allow screening of both gene bodies and intergenic regions.79,127 Thus, genes potentially involved in spermatogenic failure, along with DNA methylation changes in ncRNAs, may be identified. 128
Epigenetic regulation can be shaped not only by internal mechanisms but also by external factors, such as environmental exposures, diet, stress, and lifestyle habits. Excessive production of reactive oxygen species induces oxidative stress, which disrupts spermatogenesis and leads to impaired sperm function and reduced fertility.129,130 Moreover, oxidative stress has been shown to cause epigenetic alterations, including aberrant sperm DNA methylation, altered histone modification patterns, and dysregulated ncRNA expression. 131 Notably, oxidative stress-induced epigenetic changes can be transmitted across generations, potentially compromising the health and fertility of offspring.132,133
Inactivation of X chromosome
In female mammals, one of the two X chromosomes is randomly inactivated early in development to ensure gene dosage compensation. 87 XCI occurs in females and, in rare cases, in individuals with X chromosome numerical anomalies (typically XX, but also XXY or XXXY). It serves to compensate for differences in X chromosome copy number between males and females (typically XY). One of the X chromosomes is randomly selected for silencing and remains silenced throughout subsequent mitotic divisions, but is reactivated during meiosis. The X-inactivation center (Xic) contains genes encoding ncRNAs. The best-characterized genes in this region include X inactive-specific transcript (Xist) and its antisense RNA Tsix. Tsix, an ncRNA gene, binds to X Xic contains genes encoding ncRNAs during XCI (Figure 4). The name “Tsix” is derived from the reverse spelling of “Xist.” Xist, expressed only from the inactive X chromosome, coats the X chromosome and silences it. Tsix suppresses Xist and is a known regulator of its expression. Other genes in the Xic region also regulate Xist expression. 134

Mechanism of X chromosome inactivation.
This step determines whether there is more than one X chromosome in the diploid genome of a cell. While males typically have one X chromosome and females have two, meiotic errors can lead to individuals with karyotypes such as XXX or XXY. Nevertheless, it has long been known that only one X chromosome remains active in such individuals. 135 Sex chromosome abnormalities (Klinefelter and Turner syndromes) are the largest chromosome abnormality class and the common genetic cause of infertility in men and women; therefore, understanding the function and mechanism of XCI is essential. 136
Noncoding RNAs
MicroRNAs (miRNAs) and small interfering RNAs (siRNAs) are both small ncRNAs that regulate gene expression post-transcriptionally, and they play significant roles in male and female fertility at the molecular level.
MiRNAs and male fertility
miRNAs are crucial in every stage of spermatogenesis, including the proliferation and differentiation of spermatogonia, meiosis of spermatocytes, and maturation of spermatids. Therefore, spermatogenic cells at all stages, including differentiating and undifferentiated spermatogonia that form reserve stem cells, Sertoli cells, harbor different miRNA expressions. 137 In addition, studies suggest that a small but important portion of spermatozoal miRNAs is acquired for embryo development during the epididymal passage of spermatozoa. 138
Many studies investigate the effects of miRNAs on the formation of male germ cells, and some miRNAs stand out in this regard. miRNAs may promote or inhibit apoptosis in germ cells. Dysregulation may lead to excess germ cell death or survival of defective cells, affecting fertility. miR-17-92 cluster supports germ cell survival by targeting pro-apoptotic genes. 139 It was observed that testicular atrophy developed and sperm production was absent due to increased germ cell apoptosis in the testes of adult mice in which miR-17-92 was deleted. 139 Also, the level of pro-apoptotic proteins such as Bim, Stat3, c-Kit, and Socs3 increased in the infertile animals. 139 Expression analyses in spermatogenic cells of model animals revealed that miR-34c was highly expressed in these cells.140–142 MiR-34c, which is known to be associated with these tumor suppressor proteins in somatic cells, was shown to be able to play a role in the differentiation of sperm cells in the male germ cell line independently of this pathway. 141 It has also been revealed that mir-34c is one of the miRNAs carried to the zygote from sperm and is necessary for the first cell division of the zygote by affecting Bcl-2 expression. 142 Subsequent studies have shown that miR-34c targets Smad7, which is effective in modulating the TGF-β pathway involved in the regulation of the blood-testis barrier formed by Sertoli cells. 140 It has been reported that transfection of mir-34c or silencing of its target Smad7 reduces the rate of apoptosis in adult mouse Sertoli cells. 140 Another miRNA found to alter the expression of TGF-β1 in the male germ cell line, miR-106b, was also shown to increase the proliferation of SSCs by inducing differentiation inhibitors in SSCs. 143 Two other miRNAs that have been studied to play a role in apoptosis in seminiferous tubules are miR-21 and miR-615. Analysis of RNA-seq data from cell lines showed that increased expression of miR-21 can inhibit proliferation and increase apoptosis in Leydig cells, while miR-615 reduced the apoptosis rate. 144 miR-449 family has been shown to have roles in regulating sperm flagella and motility. The lack of expression of the miR-449 family in double knockout mice caused the mice to be infertile along with neurological and respiratory problems. 145 This effect could be due to the deterioration in the testis and the lack of motile cilia in the efferent ducts, preventing sperm cells from passing to the epididymis. 145 Another ncRNA, miR-23a, which was associated with the deterioration of semen parameters due to its effect on sperm motility, has been shown to reduce sperm motility by suppressing both HMMR and PFKFB4. 146 HMMR plays a role in critical processes for fertilization, such as sperm capacitation, acrosome reaction, and regulation of sperm motility, while PFKFB4 is an enzyme involved in glucose metabolism and is found in critical areas for movement in sperm cells, especially the acrosomal region and flagella. It is thought that increased expression of miR-23a in oligoasthenozoospermic infertile men disrupts both the acrosome reaction and energy metabolism by decreasing the expression of HMMR and PFKB4. 146
In conclusion, miRNAs play multifaceted and indispensable roles in regulating spermatogenesis, from the maintenance and differentiation of spermatogonia to the maturation and functional optimization of spermatozoa. By modulating key signaling pathways, apoptosis, and cellular differentiation processes, specific miRNAs such as miR-17-92, miR-34c, miR-106b, miR-21, miR-615, the miR-449 family, and miR-23a exert a critical influence on germ cell survival, Sertoli cell function, sperm motility, and fertilization potential.
miRNAs and female fertility
miRNAs play critical roles in regulating female fertility by affecting folliculogenesis, oocyte maturation, ovulation, embryo development, and hormonal regulation. miRNAs regulate granulosa cell proliferation, differentiation, and survival, all of which are vital for follicle development. Increased expression of miR-145 in mouse granulosa cells decreased the expression of activin receptor type I B (AcvrIB) at both mRNA and protein levels, affecting Smad2 phosphorylation, which is the continuation of the intracellular signaling of activin receptor. 147 Thus, it was shown that increased expression of miR-145 inhibited granulosa cell proliferation. Later, another study conducted with patient samples and in vitro cell lines also addressed the effects of miR-145 on granulosa cells. 148 In follicular fluid samples from patients with PCOS, miR-145 expression was decreased, while SE Translocation (SET) expression, another gene reported to be its direct target, was increased. 148 Previous studies have shown that the SET protein promotes cell proliferation and reduces apoptosis via AKT and ERK signaling pathways. 148 These studies suggest that disruptions in the expression of miR-145, together with its target genes, may contribute to follicular developmental disorders.
miR-21, on the other hand, as a miRNA induced by LH surge in mouse granulosa cells, was shown to protect granulosa cells from apoptosis.149,150 Six hours after the LH surge, miR-21 expression was detected at the highest level in granulosa cells, and at the same time, the level of cleaved caspase 3 was reduced. 149 It was reported that apoptosis in mouse granulosa cells increased when miR-21 expression was reduced by exogenous means compared to controls. 149 Later studies showed that this antiapoptotic effect of miR-21 was mediated through its targets PDCD4 and PTEN, and that the toxic effect of bisphenol A may also be mediated by miR-21. 150 Another supporting study also showed that one of the experimental targets of miR-21 is SNHG7, a member of the SMAD protein family that plays a role in growth signaling. 151 The study showed that miR-21 expression was reduced in PCOS patient samples and cell lines, underlining its contribution to the regulation of granulosa cell proliferation. 151
miRNAs also control oocyte gene expression during the transition from germinal vesicle to metaphase II stages. 152 Studies have shown that miR-503 represses Cdc25A and cyclin Ds,152–154 which are important for the cell cycle, especially in the transition of G1/S, and may impact oocyte maturation. Studies with samples from patients undergoing oophorectomy revealed that cell proliferation in human endometriotic cyst stromal cells was reduced due to gene hypermethylation of expression of miR-503, which has roles in suppressing cell cycle progression. 152 Similarly, miR-125b was shown to affect oocyte survival.155,156 Animal studies have shown that miR-125b, abundant in the ovary, reduces the expression of proliferation and apoptosis markers by targeting bone morphogenetic protein receptor type 1B (BMPR1B) and may play a regulatory role.155,156
Thus, various miRNAs, including miR-145, miR-21, miR-503, and miR-125b, are essential regulators of female fertility, regulating key processes from folliculogenesis to embryo development through precise modulation of granulosa cell function, oocyte maturation, and hormonal signaling pathways.
SiRNAs in female and male reproductive systems
While siRNAs are less naturally abundant than miRNAs in mammalian systems, they are important in germline gene silencing and genome integrity. 157 Studies with male germline-specific Dicer and Drosha knockout mice were used to assess whether siRNAs are necessary for fertilization and preimplantation development.158–160 It was determined that zygotes formed from normal eggs fertilized with sperm with altered siRNA profiles had reduced development. It was emphasized that paternal siRNAs and paternal miRNAs are important in post-fertilization zygotes and two-cell embryos. 158 siRNAs also participate in post-transcriptional gene silencing by degrading maternal mRNAs during oocyte maturation and after fertilization. Oocytes from Dicer knockout mice failed to progress through meiosis with defects in spindle formation, spindle checkpoints, and chromosomal segregation because they could not complete maternal transcripts essential for oocyte maturation.161,162 siRNAs, particularly in early germ cells, help silence transposable elements, protecting genome integrity during gametogenesis.157,159,163 Endogenous siRNAs originate from double-stranded RNAs formed by bidirectional transcription of transposable elements or repetitive sequences. After being processed by Dicer into 21–23 nucleotide mature siRNAs, they are loaded onto the RNA-induced silencing complex, which then binds to complementary transposon mRNA transcripts and cleaves them, preventing their translation and activity. 163 This function overlaps with piRNAs, but especially endogenous siRNAs also contribute. 159 In addition, spermatogenic defects were detected in male Dicer knockout animals due to errors in silencing of repeats located in centromeric regions, again demonstrating the importance of siRNAs in contributing to genome integrity.159,160 Although siRNAs are less prevalent than miRNAs in mammalian systems, they play indispensable roles in germline development, fertilization, and the preservation of genome stability. Their functions extend from regulating maternal mRNA turnover during oocyte maturation to safeguarding the genome against transposable elements in early germ cells.
Epigenetic regulation is fundamental to spermatogenesis, with DNA methylation dynamics, histone modifications, and ncRNAs intricately controlling the development of mature spermatozoa. Oxidative stress disrupts these epigenetic mechanisms by inducing DNA damage, altering histone modification patterns, and modulating ncRNA expression, leading to impaired sperm function and male infertility. Environmental and lifestyle factors, including aging, exposure to pollutants, unhealthy habits, and radiation, exacerbate oxidative stress, contributing to epigenetic alterations that not only affect individual fertility but may also have transgenerational implications. While animal studies have demonstrated the potential for epigenetic modifications to impact offspring health and fertility, human studies are limited and often present contradictory findings. 132
Conclusion
This narrative review underscores well-recognized genetic causes of infertility in men with azoospermia or severe oligozoospermia include chromosomal abnormalities, microdeletions in the AZF regions, and mutations in the CFTR gene 31650616. 164 In addition, numerous autosomal and sex chromosome-linked genes have been associated with spermatogenic failure and male infertility 32314821. 75 Identifying potential genetic etiologies in infertile men is challenging due to the subtle phenotypic differences between infertile and fertile men, the limited availability of informative pedigrees, and the involvement of over 2000 genes in spermatogenesis. 165 In this context, NGS is a high-throughput technology capable of rapidly sequencing DNA to detect genetic variations linked to disease. It enables the simultaneous sequencing of billions of DNA bases and allows multiple patient samples to be analyzed in a single run within hours. NGS approaches are generally categorized into whole exome sequencing (WES), whole genome sequencing (WGS), and targeted sequencing, all of which share similar laboratory workflows and offer high analytical accuracy. WES focuses on all protein-coding regions of the genome (~1%–2%) along with selected flanking regulatory or splice sites, making it a widely adopted and relatively cost-efficient tool in clinical genetics. Given that approximately 85% of known pathogenic mutations occur in exons, WES holds substantial clinical relevance. WGS sequences the entire genome, providing the most comprehensive genomic data, albeit at a higher cost. TS, in contrast, is a hypothesis-driven approach targeting predefined genes or genomic regions, and is frequently applied in conditions such as CBAVD caused by CFTR mutations, or in gene panels for disorders of the hypothalamic–pituitary axis. 166
Epigenetic changes influence not just fertility potential in men and women, but also have a crucial impact on inheritance patterns across generations, and the health of children conceived through ART. Indeed, previous research has identified abnormal DNA methylation in both imprinting genes and non-imprinting genes, which have been linked to different forms of male and female infertility.74,167
