Abstract
Oncogenic neurotrophic tropomyosin receptor kinase gene fusions occur in less than 1% of common cancers. These mutations have emerged as new biomarkers in cancer genomic profiling with the approval of selective drugs against tropomyosin receptor kinase fusion proteins. Nevertheless, the optimal pathways and diagnostic platforms for this biomarker’s screening and genomic profiling have not been defined and remain a subject of debate. A panel of national experts in molecular cancer diagnosis and treatment was convened by videoconference and suggested topics to be addressed in the literature review. The authors proposed a testing algorithm for oncogenic neurotrophic tropomyosin receptor kinase gene fusion screening and diagnosis for the Brazilian health system. This review aims to discuss the latest literature evidence and international consensus on neurotrophic tropomyosin receptor kinase gene fusion diagnosis to devise clinical guidelines for testing this biomarker. We propose an algorithm in which testing for this biomarker should be requested to diagnose advanced metastatic tumors without known driver mutations. In this strategy, Immunohistochemistry should be used as a screening test followed by confirmatory next-generation sequencing in immunohistochemistry-positive cases.
Background
Malignant neoplasms have been diagnosed and treated based on topography and histological type. Recently, the FDA (Food and Drug Administration) has approved the first classes of drugs for tissue-agnostic indications, based on a common biomarker as opposed to the primary site of origin: neurotrophic tyrosine kinase receptor (NTRK) inhibitors; pembrolizumab for microsatellite instability-high (MSI-H) or defective mismatch repair (dMMR) solid tumors and for high tumor mutational burden cancer (TMB-H). Even more recently, 2 additional agnostic indications have been approved by the FDA: the combination of dabrafenib and trametinib for metastatic solid tumors with BRAFV600E mutations, and selpercatinib for advanced or metastatic tumors with RET fusion.1 -3
NTRK gene fusions generally occur in less than 1% of most common cancers such as lung, pancreas, breast, colorectal, kidney, and melanomas. However, the incidence of NTRK gene fusions is higher in some specific rare tumors. Infantile fibrosarcoma, breast, thyroid, and salivary gland secretory carcinomas, gastrointestinal stromal tumors (pan-negative), and high-grade glioma have a high pre-test probability of harboring NTRK gene fusions. 4
Selective inhibition of tropomyosin receptor kinase (TRK) fusion proteins represents an advance in precision medicine to treat different solid tumors. 5 Entrectinib and larotrectinib were approved by the FDA for the treatment of tumors harboring NTRK oncogenic gene fusions in 2017 and 2018, respectively. The NTRK gene family comprises 3 genes: NTRK1, NTRK2, and NTRK3, which encode 3 tyrosine kinase receptors: TRK A, B, and C, respectively. NTRK1/2/3 gene fusions have emerged as novel targets for cancer therapy as they can be successfully inhibited by targeted kinase inhibitors.5,6
With the growing use of target therapies for TRK fusion proteins, NTRK1/2/3 gene fusion has become a new biomarker in cancer genomic profiling. Despite these advances, the optimal pathways and diagnostic platforms for screening and identifying these fusions remain a subject of debate. Next-generation sequencing techniques (NGS) are evolving rapidly, with lower costs and shorter turnaround times each year. 7 However, sequencing large numbers of tumor samples is still expensive and challenging. 8 Thus, it is important to establish clear criteria for deciding whether a sample should be sequenced or not.
The latest European Society for Medical Oncology (ESMO), the Canadian Consensus for Biomarker Testing and Treatment of TRK Fusion Cancer in Adults, and an international expert consensus of several oncology societies have recommended that NTRK immunohistochemistry IHC testing may be performed in advanced solid tumors without other actionable and driver mutations or fusions, but tumor types with a high probability of harboring NTRK fusions should be sequenced straightforward.6,9,10 As part of this testing strategy, IHC has been proposed as a screening test, with positive results requiring subsequent confirmation by NGS. 10
The purpose of this article is to critically appraise the latest international consensus, discuss the role of current diagnostic platforms available for the screening and clinical diagnosis of NTRK fusion genes, as well as to make evidence-based recommendations for pathologists and laboratories on how to implement and optimize these tests in routine clinical practice, correctly identify NTRK fusions, and propose a diagnostic pathway for tumors with distinct probabilities of harboring actionable alterations in these oncogenes.
Methods
After the approval of the first agnostic drug in Brazil, larotrectinib, there is now an urgent need to discuss and devise clinical guidelines to guide oncologists and pathologists regarding the testing flow of NTRK1/2/3 genes, according to the specificities of the healthcare system. A panel of 10 national experts with an active interest in molecular cancer diagnosis and treatment was convened via videoconference on February 12, 2021. Panel members had either clinical or pathology experience with tumors harboring NTRK gene fusions.
The Chairpersons (FS and KL) pre-defined practical questions on handling the diagnostic management of NTRK in clinical practice and suggested topics to be addressed. A literature review was performed for selected topics before the online meeting and the experts led active discussions during the virtual meeting involving a multidisciplinary team comprising pathologists and clinical oncologists. Recent publications of international societies, such as the European Society of Medical Oncology (ESMO), the American Society of Clinical Oncology (ASCO), and the Japanese Society of Medical Oncology (JSMO), were reviewed to discuss the methodology and consensus writing, and the panelists sought to adapt these recommendations to the context of the Brazilian health system.6,9 -15 Whenever possible, a consensus was sought among the experts.
The meeting served to discuss the most relevant issues among the practical questions. The Panel addressed 2 major areas concerning NTRK1/2/3: (1) the biology of fusions; and (2) diagnostic strategies for detecting NTRK fusions. These algorithms were then used as the basis for discussion and further refinement. Consensus on these algorithms was reached through a series of teleconferences and emails. The algorithms and draft text were subsequently revised and recirculated through an iterative process until all authors agreed on the final content. This article describes the current landscape for NTRK tests in an agnostic cancer setting in Brazil.
Discussion
NTRK fusion biology
Reciprocal and non-reciprocal fusions
Oncogenic TRK activation can occur through mutation, overexpression, and splice variants. However, the most well-described oncogenic mechanism of this activation is fusions involving NTRK1, NTRK2, or NTRK3 gene activation.4,16
The fusions are usually intrachromosomal or interchromosomal rearrangements of hybrid genes in which 3′ sequences of NTRK1, NTRK2, or NTRK3 harboring the kinase domain coding region are juxtaposed to 5′ sequences of a different gene, called the partner gene or fusion partner. 4 In nearly all cases, the fusion eliminates the ligand-binding site, resulting in ligand-independent dimerization and phosphorylation. 16 In order to be oncogenic, the TRK fusion protein needs to have an intact tyrosine kinase domain.
The product of the fusions between the NTRK (1, 2, or 3) and partner gene is a chimeric oncoprotein characterized by ligand-independent constitutive activation of the TRK kinase and its downstream signaling cascades. 4
Novel NTRK fusion partners
Tumor genomic complexity may lead to clinical findings of new unpublished NTRK fusions of uncertain clinical significance, also known as non-published partners. 1 In some cases, it may be challenging to properly understand and classify a novel fusion even with RNA sequencing data. 17
In clinical practice, physicians must weigh the risk-benefit of using an unpublished functional NTRK-expressed fusion and indicate NTRK inhibitors on a case-by-case basis.
Diagnostic strategies and practical recommendations for NTRK fusion detection
What tests for NTRK fusion are recommended?
IHC is recommended for NTRK fusion screening, and NGS for NTRK fusion diagnostic confirmation (Figure 1). Under some circumstances, fluorescence in-situ hybridization (FISH) and reverse-transcriptase real-time PCR (RT-qPCR) can be acceptable for diagnosis.
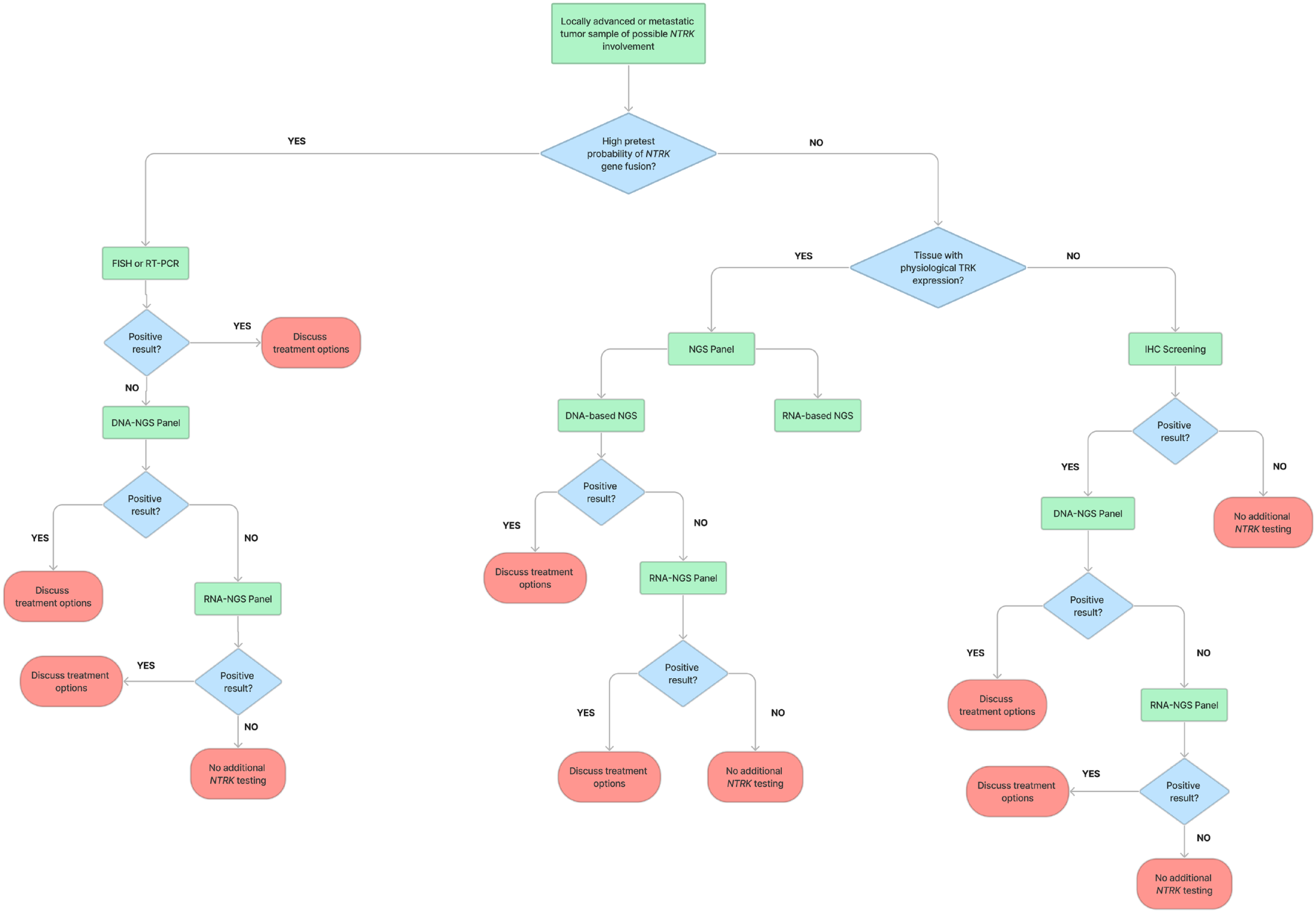
Proposed NTRK Testing algorithm for a newly diagnosed advanced or metastatic tumor sample.
Immunohistochemistry
IHC is a common technique that is widely available across pathology labs. IHC is a useful and cost-effective test for detecting the presence of TRK protein that offers rapid turnaround time, low cost, high sensitivity, high negative predictive value, and requires less amount of tissue compared to other techniques such as NGS.7,9 TRK proteins are physiologically expressed in neural and/or muscle cells of the brain, adrenal glands, testis, ovary, pancreas, kidney, gallbladder, stomach, colon, heart muscle, smooth muscle, and skeletal muscle. 18 Currently, IHC is recommended by the European Society for Medical Oncology (ESMO) Translational Research and Precision Medicine Working Group as a screening method for NTRK fusion detection, to be followed by a confirmatory molecular-based methodology, such as NGS, FISH, or RT-qPCR. 6 Regarding cost, the benefit of using IHC as a screening method is that it may be reimbursed or covered by insurance in many countries, such as Brazil, when performed as part of an IHC panel.
A large study by Solomon et al. tested 38 095 samples and reported that the sensitivity for detection of NTRK 1/2/3 fusions by IHC was 96.2%, 100%, and 79.4%, respectively. It was hypothesized that, for some tumor types, the reduced sensitivity of NTRK3 fusion could be due to a fainter staining and nuclear pattern, resulting in a higher number of false negatives. The overall specificity of the technique was 81.1%, with lower specificity in tumors derived from tissues that physiologically express TRK proteins, such as gliomas, neuroblastomas, and some subtypes of sarcomas, limiting the applicability of IHC in these specific cases.16,19
The most widely used monoclonal antibodies for TRK IHC detect an epitope that is common to the 3 protein isoforms and does not distinguish between wild type and fused TRK. However, IHC yields high sensitivity and negative predictive values, conferring greater clinical value as a screening test when compared to NGS, given the low pretest probability of NTRK fusions in most cancers, high costs of NGS, and the still limited availability of molecular testing and sequencing for large numbers of patients in most countries.20,21 In the next sections of the article, we will discuss and recommend evidence-based actions for adoption by pathology laboratories to optimize the TRK IHC testing routine.
Monoclonal antibodies
Several antibody options for evaluating TRK protein expression in formalin-fixed paraffin-embedded (FFPE) tumor tissue sections are commercially available. 6 Some antibodies target specific TRK isoforms (eg, TRKA—rabbit monoclonal antibody clones EP1058Y and Sab76291; and TRKB—rabbit monoclonal antibody clone 80G2). However, these antibodies are mostly used in academic settings or specific case investigations.22,23 There is no added clinical value in routinely identifying the specific TRK protein in the screening IHC step, as the driver NTRK gene fusion will be further identified by molecular methods. Additionally, the use of specific clones aiming just one of the isoforms means that more than 1 IHC protocol needs to be performed for each sample analyzed for TRK expression, and so adding unnecessary work to the laboratory routine.
The most frequently used, validated, and well-characterized clone is the pan-TRK—rabbit recombinant monoclonal antibody, clone EPR17341, which identifies the expression of all 3 TRK proteins.16,19,24 The A7H6R pan-TRK—rabbit recombinant monoclonal antibody is an alternative option that has yielded similar results in a recent interlaboratory comparison. 25 Thus, EPR17341 is the current first-choice antibody option for pathology laboratories.
Regardless of the chosen antibody, proper validation is necessary to ensure the efficiency and reliability of the methodology. The College of American Pathologists and Laboratory Quality Center Guideline contains all recommendations to allow proper pre-analytical and analytical validation and consequently ensure accuracy and reproducibility of the IHC test. 26
Tumor sample quality assurance
The pre-analytical phase is the main source of interference and altered results in clinical laboratories. Although some variables are usually not the responsibility of the pathology laboratory such as tissue ischemia, there are many factors that determine the quality of preservation of macromolecules in tissue and influence the quality of IHC results: (1) tissue sampling and viability, time, and quality of fixation, paraffin embedding, and sectioning, which determine the quality of the tissue to be labeled and stained; and (2) storage time, quality of the staining platform, antigen retrieval method, washing, blockage, antibody labeling, incubation, and contra staining. These are all factors that have an important influence on the final quality of the slides. Alterations in any of these factors can lead to increased rates of false-negative or false-positive results.27,28
Biopsy sampling
When testing for TRK expression, careful consideration should be given to the selection of the biopsy site in patients with multiple metastatic lesions. Factors such as lesion accessibility, safety, size, and location should be taken into account. The biopsy should be ideally obtained from the primary lesion, as it is usually the most clinically significant site. However, in cases where a biopsy from metastatic sites is required, it is recommended to target lesions that are of sufficient size and quality, easily accessible through minimally invasive procedures, and pose minimal risk to the patient. 5
Positive control
An important factor in ensuring the quality of reactions, optimizing sensitivity, and minimizing the risk of false-negative results, is the presence of positive control for each IHC reaction. The control may be internal, represented by some component already present in the tissue analyzed, or external, manually placed on a slide. 20 TRK proteins are physiologically expressed in neural structures and so tissues rich in ganglion cells, such as the appendix, testis, and colonic wall, have been widely used as internal positive controls.19,20 For external controls, options include cell lines such as KM12 (TPM3::NTRK1), MO-91 (ETV6::NTRK3), and CUTO-3.29 (MPRIP::NTRK1), although these tend to be more expensive. Formalin-fixed paraffin-embedded cell pallets can also be used as external positive controls as well as samples of tumors that previously tested positive for NTRK fusion. 6
Because the presence of internal positive controls such as neural cells in the tissue examined cannot always be guaranteed, an external positive control should be included on all slides to ensure the quality and effectiveness of the immunohistochemical reactions for TRKs. 20 Thus, the aforementioned TRK-expressing tissues (especially the appendix) are considered the most accessible external positive control options for inclusion in the pan-TRK testing routine.
TRK immunohistochemical analysis and interpretation
Although there is no interpretation guide for any of the available IHC clones, in the recently published largest series of tumors with NTRK fusions, pan-TRK labeling of ⩾1% cells above background has been defined as a positive result (Table 1).16,30 Pan-TRK IHC for NTRK3 has demonstrated lower sensitivity of 79.4%, compared to 96.2% and 100% for NTRK1 and NTRK2, respectively. 16 This disparity is believed to occur because NTRK3 fusion-positive tumors exhibit a more perinuclear, focal, and weak staining pattern compared to the other NTRK fusion genes. 7 Secretory carcinomas and infantile fibrosarcoma have higher rates of NTRK3 fusion genes, and this should be taken into account in the evaluation of these tumors when NTRK fusion involvement is suspected. 8
Albeit cytoplasmic staining is the most frequently observed pattern in most cancers with NTRK fusions, it is also the pattern of physiological TRK expression and, consequently, of the majority of false-positive cases 20 (Figure 2A). The threshold of ⩾1% cells above background is applied to pan-TRK expression identified in any subcellular localization, as the staining pattern varies depending on the subcellular localization associated with the fusion partner gene. Cytoplasmic, nuclear, perinuclear, and membranous staining have all been observed, requiring pathologists to be aware of the variable staining patterns in order to improve the accuracy of NTRK fusion testing, 30 especially regarding the possible presence of weak nuclear staining attributed to the reduced sensitivity of pan-TRK IHC for NTRK37 (Figure 2B-E). Table 1 describes the staining patterns of some known NTRK fusions. As outlined above, gliomas, neuroblastomas, and sarcomas may have physiological TRK expression, hampering the interpretation of results (Figure 2A).

Different TRK IHC staining patterns (examples): (A) physiological cytoplasmic expression (false positive), (B) membranous expression, (C) cytoplasmic and membranous expression, (D) perinuclear expression, (e) weak cytoplasmic + nuclear expression (common ETV6-NTRK3 pattern), and (F) mammary analog secretory carcinoma of salivary.
A meta-analysis of 224 solid tumors with NTRK rearrangements from 13 studies showed that pan-TRK IHC had an overall sensitivity of 82%, with a false-negative rate of 18% compared to molecular diagnosis. The false-negative rate was significantly higher in cases with NTRK3 fusions (27%) compared to NTRK1 (6%) and NTRK2 (21%) fusions (Table 2). It is important to note that a significantly higher number of NTRK3 fusions exhibited nuclear staining in IHC compared to NTRK1 and NTRK2 fusions (45, 8, and 1 case, respectively). 31 Additionally, another retrospective study of 87 patients with NTRK fusions also revealed that pan-TRK-IHC had a lower sensitivity for detecting NTRK3 fusions (79%) compared to NTRK1 (96%) and NTRK2 (100%) fusions. Hence, this evidence highlights the potential problems related to the presence of weak nuclear staining in the interpretation of pan-TRK IHC.
Sensitivity and false-negative rates of pan-TRK IHC in solid tumors across different NTRK genes. 31

Staining in ⩾1% tumor cells was considered a positive result.
This is important for tumors that have a high occurrence of NTRK3 fusions, such as secretory carcinomas and infantile fibrosarcoma. However, pan-TRK IHC demonstrated a sensitivity of 96% and 92% for detecting ETV6::NTRK3 fusions in 2 case series comprising 27 and 25 salivary gland secretory carcinomas, respectively.32,33 In the latter study only nuclear staining, which was exhibited in 20 out of 25 cases, was considered in the analysis. 33 Thus, IHC can be considered a useful and sensitive screening tool for secretory carcinomas. Nevertheless, these tumors may show strong nuclear staining and are well-known for their high frequency of NTRK3 fusions, which could contribute to the increased sensitivity observed in these case series of secretory carcinomas, as well as other cancers with a high pre-test probability of NTRK3 fusions.
While nuclear, perinuclear, and membranous staining is highly suggestive of the presence of NTRK fusions, they account for 8% to 10% of false-positive cases, 20 the purpose of IHC screening is to rapidly distinguish between patient specimens that are pan-negative for NRTKs and those showing some level of staining, which may indicate harboring of gene fusions. 6 Molecular biology methods such as NGS must be performed to confirm the results detected in pan-TRK IHC-positive samples.
Moderate-to-strong diffuse cytoplasmic or membranous pan-TRK IHC staining can be considered a surrogate of the presence of NTRK1/NTRK2 fusions (in the absence of tumor muscle or neural differentiation), while nuclear pan-TRK IHC can be considered a surrogate of NTRK3 fusions until the results of additional tests become available (Figure 2B-F). NTRK fusion should be mandatorily confirmed by other molecular methods to ensure the presence of fusions in those patients who are considered candidates for target therapies. 6
Limitations of immunohistochemistry
Besides the known issues related to the pre-analytical phase of IHC, the diagnosis of NTRK-fusion expression in tumors may pose a number of challenges, even for experienced pathologists, namely (1): difficult interpretation in tissues where TRKs are physiologically expressed or in tumors with neural or smooth muscle differentiation; and (2) large variability of subcellular staining patterns and degrees of expression. Because these gene fusions are relatively rare events, this may make it even more difficult for pathologists to acquire the necessary expertise and standardize the interpretation of results among different professionals.6,7
Additionally, the clones used in IHC detect the presence of a TRK protein c-terminus epitope, present in both wildtype and mutant proteins. Thus, the presence of NTRK-fusions cannot be confirmed by IHC, but its results can be strongly suggestive in many cases. 8
Next-generation sequencing
NGS, also known as high-throughput sequencing, is a modern sequencing approach comprising different commercial platforms that can generate large amounts of sequenced data. The most used platforms are based on sequencing by synthesis: Illumina (Illumina, San Diego, CA, USA) and ThermoFisher sequencers (ThermoFisher Scientific, Waltham, MA, USA). The platforms differ regarding their throughput, cost, error profile, and read structure. 34
Briefly, the step-by-step workflow of an NGS sequencing assay includes sample preparation, library preparation, sequencing, and data analysis. Library preparation involves steps such as DNA or cDNA fragmentation and adapters ligation, allowing the samples to proceed to sequencing. 35 For sequencing, the Illumina platform uses the principle of fluorophore excitation involving the incorporation of fluorescently labeled deoxyribonucleotide triphosphates (dNTPs) into a DNA template during the cycles of DNA synthesis across millions of fragments in a massively parallel fashion. The ion torrent technology used in ThermoFisher sequencers uses the principle of H+ release and a pH change in each nucleotide incorporation that is detected by a semiconductor chip. 34
An oncogenic chimeric NTRK1/2/3 rearrangement transcript is characterized by the formation of an in-frame molecule harboring the 3′ portion of the NTRK1/2/3 gene with an intact kinase domain and the 5′ of its fusion partner. The resultant protein contains the C-terminus of the TRK protein joined to the N-terminus of the fusion partner. The actionable NTRK fusions are those harboring an in-frame and intact kinase domain.8,36 A number of different gene partners have been reported as partners with NTRK1/2/3, and these may represent intrachromosomal (when both genes involved are from the same chromosome) or interchromosomal (when genes involved are from different chromosomes) rearrangements. The position in the gene where the fusion with the other gene occurs is called a breakpoint. In general, the majority of the genomic breakpoints producing fusion genes are located in intronic, untranscribed regions. 36
NGS for the detection of gene fusions, such as NTRK fusions, has the advantage of, besides providing potentially highly sensitive results, also performing a concomitant evaluation of many other actionable oncogenic drivers, optimizing patient care and tumor tissue utilization. NGS for gene fusion detection may be performed using either the tumor DNA or RNA as a starter material and be based on different enrichment protocols such as those employing hybridization capture or amplicon. Awareness of limitations regarding the use of DNA or RNA and the different enrichment protocols is crucial to understanding the coverage of each type of assay for fusion detection.35,37 In the present report, we examine NGS approaches in the context of NTRK fusion diagnosis and recommend evidence-based actions for adoption by pathology laboratories to improve the identification of these gene fusions using NGS.
Library assembly
There are 2 main approaches to preparing DNA or cDNA libraries for targeted NGS: amplicon-based enrichment; and hybridization capture. The amplicon-based enrichment of target sequences may be subdivided into those based on classical multiplex PCR and those based on anchored multiplex PCR (AMP). This first approach consists of a classic multiplex PCR reaction with forward and reverse primers designed to flank the regions of interest in the genome. In the context of cancer fusion detection, this approach requires that the sequence of both the target gene and its fusion partner be known because both primers are gene-specific. 37
AMP is similar to classical PCR reactions. However, an NGS adapter sequence is ligated to the DNA or cDNA fragments, and the PCR is performed with 1 primer designed to pair with the target gene and the other designed to pair with the adapter sequence. With this approach, only one of the fusion genes needs to be known, and novel fusion partners can be amplified.37,38
The hybrid capture approach uses a probe with a specific nucleotide sequence, complementary to regions of interest in the genome, to capture the intended regions of the designed panel. One of the main differences is that the capture probe is larger than the primers used in the classical and anchored amplicon approaches, allowing the presence of some mismatches in the probe site without losing the target region.
Although the hybridization-capture approach is theoretically able to detect more variants because its probe allows pairing with some mismatches, it also requires a greater amount of starter genetic material that is not always available from FFPE tumor samples and requires a longer workflow. Thus, the AMP approach may represent a very feasible library preparation technique for cancer clinical diagnosis, as it minimizes time and sample requirements compared to the hybridization-capture technique while allowing the detection of novel fusion genes. AMP provides greater sensitivity than classical PCR amplicon panels when detecting fusions through NGS, as previously unknown NTRK fusions are regularly being reported. 39 Figure 3 illustrates the different approaches used to build a DNA or cDNA library for NGS.

cDNA library assembly approaches comparing classical amplicon, hybrid capture and anchored amplicon based PCR methods.
Regarding the respective performance of classical amplicon multiplex PCR versus anchored amplicon multiplex PCR, a comparison of a small series of 37 FFPE NSCLC samples analyzed on an Ion Torrent platform showed that ArcherDX AMP delivered better accuracy. 40 However, further data is necessary to draw any definitive conclusions related to the analytical superiority of one method over the other.
DNA versus RNA sequencing
DNA-based NGS assays to detect fusions can be based on either a classical amplicon approach or a hybridization approach. The classical amplicon approach is designed to detect only known fusion partners and, out of all the different combinations, is the least sensitive for fusion detection because of the limitations inherent to this method of library preparation.8,39 However, improvements in amplicon libraries have been made by manufacturers, such as the Ion Torrent Genexus System (ThermoFisher Scientific, Waltham, MA, USA), which integrates and automates sequencing preparation and analysis. 41 Hybridization enrichment, when properly designed, can target a variety of intronic regions and detect rearrangements with an unknown partner. DNA NGS hybrid capture assay sensitivity is dictated by the assay design. Two different assays claiming to cover NTRK1/2/3 fusions at the DNA level may have different sensitivity in detecting these alterations, depending on their pool of designed primers and covered introns of these genes.39,42 Amplicon-based DNA NGS assays may have limitations when handling genes with the large and/or repetitive intronic regions often involved in rearrangements which hamper primer design and the optimization of multiplex reactions and sequencing efficiencies, as observed in NTRK2/3 genes, and exemplified in Figure 4. Thus, sensitivity for detecting fusions in the NTRK3 gene, other than the well-known ETV6, is lower than for NTRK1 genes.37,42,43

Structural differences between DNA and messenger RNA (mRNA). Intron exclusion in mRNA (cDNA) facilitates primer/probe library design allowing more sensitivity in identifying fusions in cancer cells.
The RNA-based NGS assay for NTRK1/2/3 fusion has greater sensitivity than the DNA assay. The assay is based on RNA extracted from the tumor tissue, and thus represents the processed gene-coding molecule after intronic exclusion by splicing, eliminating the technical challenge of intronic coverage involved in the DNA assay (Figure 4). 37 After extraction, the RNA is converted to cDNA (complementary DNA) (Figure 2) RNA sequencing protocols may also be based on amplicon, AMP, or hybridization-based enrichment panels. Classical amplicon panels based on RNA may not be able to characterize unknown fusion partners, but a 3′/5′ ratio can be calculated for the genes included in the panel to assess the possibility of the presence of novel gene fusions. 44 The hybridization capture and anchored-multiplex PCR (AMP) approaches allow for the detection of genes with novel fusion partners.
RNA sequencing allows the detection of alternatively spliced transcripts, novel gene fusions, mutations/single nucleotide polymorphisms (SNPs), and changes in gene expression. DNA sequencing also allows the detection of novel gene fusions and single-nucleotide polymorphisms. However, this sequencing does not provide information related to alternatively spliced transcript nor, most importantly, on quantitative changes in gene expression. Thus, RNA sequencing has the advantage of providing more accurate information on the expression of the gene fusion of interest in tumor tissues. 37
In spite of these benefits, RNA-based assays may face challenges with regard to obtaining good quality RNA from clinical FFPE samples and having enough samples for both DNA and RNA analyses, considering the potential need for a comprehensive analysis of the DNA alterations, as well as of the tumor sample in other protocols, such as IHC and FISH.30,37,39
Fusions detected by DNA sequencing, besides all the shortcomings described above, might also detect sequences that do not code a functional, translated protein. Meanwhile, RNA sequencing detects the mRNA translated into the oncogenic protein, which is the functional product driving cancer cell growth. Therefore, RNA sequencing is the recommended test for detecting NTRK1/2/3 fusions.
Assessment of nucleic acid quality for NGS
One of the key issues related to sequencing FFPE tissues is DNA and RNA quality, as the fixation, tissue processing, and storage methods may cause nucleic acid degradation, characterized by fragmentation of DNA and RNA. Cold ischemia, volume, fixative quality, and storage temperature are important factors influencing DNA and RNA quality. Rapid time between tumor resection and fixation, adequate temperature, and usage of 10% neutral phosphate-buffered formalin are some of many good practices to be incorporated by the laboratories to ensure good quality genetic material for molecular pathology tests.45,46
The minimum amounts of DNA or RNA from FFPE samples required for NGS vary widely across different library preparation kits available in the market, ranging from 10 to 200 ng per sample. 37 The same kit may require different amounts of nucleic acids, depending on the quality of the extracted material, given that sequencing low-quality samples requires larger amounts of DNA or RNA. 47 The Agilent RNA Integrity Number (RIN), which is a measure of the relative proportion of ribosomal 28S and 18S RNA, has been widely used to determine the RNA quality of FFPE samples. However, there have been suggestions that this method has low analytic sensitivity, especially for smaller fragments. DV200, which denotes the percentage of RNA fragments containing over 200 nucleotides, was developed as a more accurate metric for assessing RNA quality in FFPE samples and is currently recommended by library preparation kit developers, such as Illumina®.47,48 Determinations of both these measures can be provided by bioanalyzer systems.
Nonetheless, each library preparation kit developer has its own recommendations and measurements for RNA quality assessment, which must be observed in order to correctly perform the assays.
Turnaround time
NGS is a complex testing methodology that depends on many factors, including analytical variables related to the NGS workflow, which differ among sequencing platforms, as well as pre-analytical variables related to the distance between the patient treatment center and the molecular laboratory. There is scant data on mean NGS turnaround times in the literature, with periods ranging from 8 to 37 days and large variability reported within studies.49,50 Patients that have access to hospitals with their own molecular pathology laboratories receive results earlier than individuals treated at smaller or distant centers, who can face sample transportation and logistics delays. Additionally, both laboratory demand and sequencing platforms impact the turnaround time of cancer genetic profiling. General pathology laboratories may improve TAT by speeding up the release of biopsy samples upon requests for molecular tests.
FISH
FISH is a method based on the labeling of samples with fluorescent probes that are interpreted under a fluorescence microscope. In the context of NTRK gene fusion, both fusion probes or break-apart probes targeting specific DNA regions may be used. There are established commercial fusion probes for the detection of NTRK fusions, such as the ETV6::NTRK3, offering good sensitivity and specificity. The ETV6::NTRK3 assay is very effective for identifying this specific gene fusion in tumors where it is highly prevalent, such as infantile fibrosarcoma (IF), secretory breast carcinoma (SBC), and secretory carcinoma of the salivary gland (MASC).6,20 Another option is using break-apart probes for NTRK1/2/3, which detect the existence of breakpoints and structural variants of the genes but cannot identify whether the gene itself is being transcribed, or its fusion partner.7,8 FISH has a rapid TAT, but tests demand 1 slide per probe. Although it can detect alterations at the gene level, it cannot confirm the presence of in-frame functional fusions. Another disadvantage of FISH is the occurrence of short split lengths that can result in false-negative results, particularly for NTRK1 fusions, which often occur from intrachromosomal events involving chromosome 1. 36 Therefore, RT-qPCR or NGS should be employed to mitigate the effects of short split length in FISH.
RT-qPCR
RT-qPCR is a relatively low-cost methodology with a rapid turnaround time compared to sequencing approaches. However, the PCR primers design requires that both fused genes and their exon breakpoints be known, increasing the false-negative rates of the method. It may be a suitable methodology to confirm the presence of specific fusions (such as ETV6::NTRK3) with a high probability in cancers such as IF, SBC, and MASC. RT-qPCR is unable to detect mutations/single nucleotide polymorphisms, alternatively spliced genes or differences in gene breakpoints. Although RT-qPCR was important for gene-fusion profiling in the past, it is becoming increasingly less useful with the advancement of more sensitive and sophisticated methods, such as RNA sequencing. 8
We provide a NTRK methodology guidance overview in Table 3.
NTRK methodology guidance.
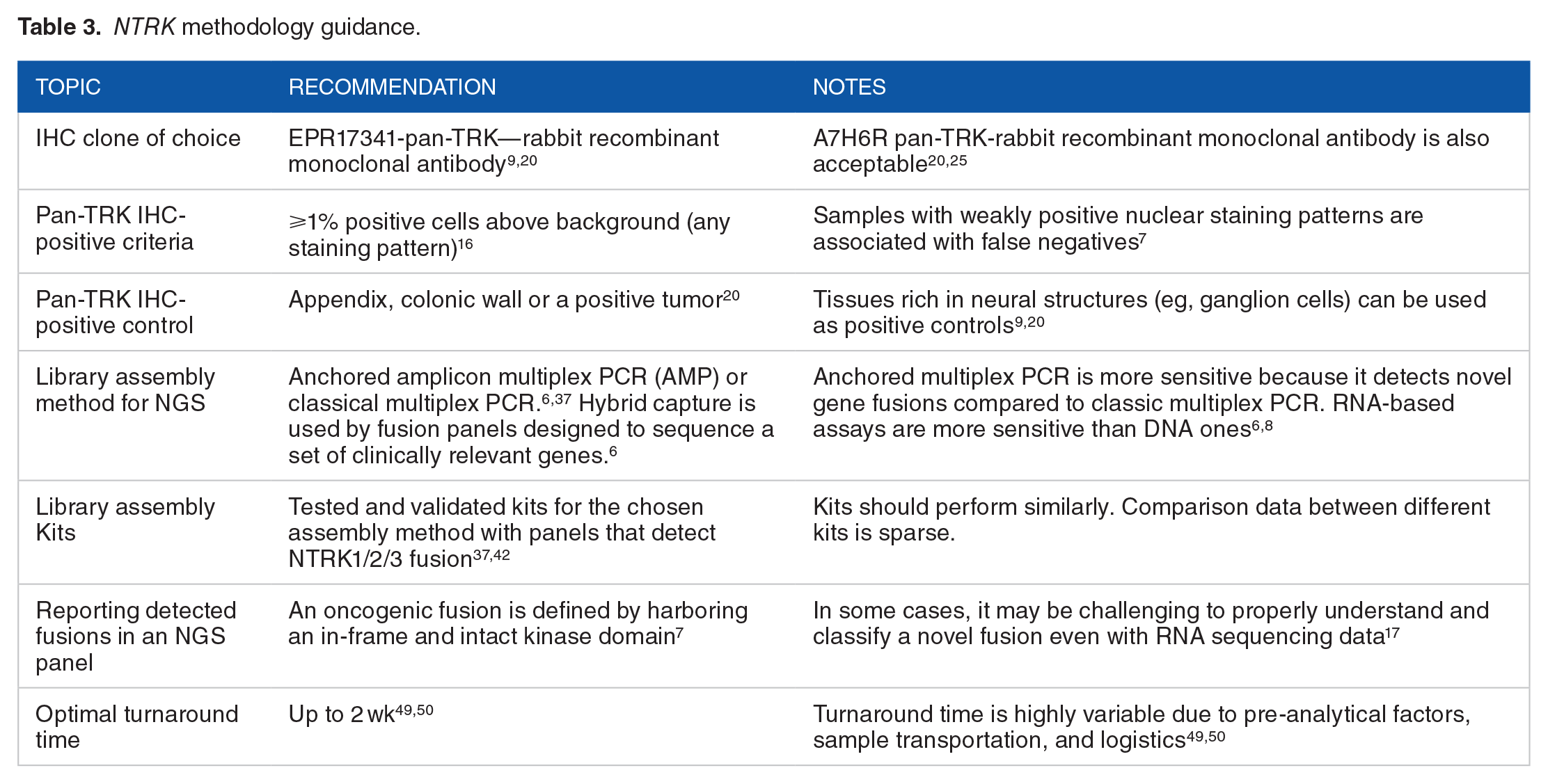

Expert recommendations
Who should be eligible for NTRK fusion testing?
All patients with locally advanced or metastatic solid tumors. We present the rate of NTRK fusion across different solid tumors in Table 4.
Prevalence of NTRK fusion for different solid tumors.

When should NTRK fusion testing be ordered?
NTRK fusion testing should be ordered for the diagnosis of advanced or metastatic tumors without known driver mutations
Comprehensive genomic panels in Brazil
Comprehensive genomic panels are the most straightforward and in-depth method to identify genomic modifications related to tumor oncogenesis. However, high costs are the main access barrier to these tests. The panels should be considered if they are covered by the patient’s health insurance plan or if the patient is able to cover the cost of testing out of pocket.
Screening strategies (IHC) followed by confirmatory testing (NGS)
Screening with pan-TRK IHC followed by a confirmatory test (NGS) in IHC-positive cases is recommended for locally advanced or metastatic solid tumors with a low incidence of NTRK fusions.
Direct confirmatory testing
NTRK gene fusion testing is required for reliable diagnosis of tumor types with a very high prevalence of NTRK gene fusions (>80%) (eg, secretory breast cancer and infantile fibrosarcoma). In this context, we recommend using confirmatory testing methods directly (Figure 1) instead of initial IHC screening due to the high pre-test probability of these tumors containing NTRK gene fusions. FISH can be used to detect specific gene fusions that are highly frequent in these cancers, such as ETV6::NTRK3, and should be employed if NGS is not accessible or financially viable. However, RNA based NGS panels should be the preferred choice whenever possible.
What is the order of preference of sample type for performing NTRK fusion testing?
(1) Surgical specimens; (2) Biopsy samples; (3) Cytopathology samples; (4) Bone tissue samples (only if no other available specimen and after adequate fixation and EDTA decalcification); and (5) Liquid biopsy (lower sensitivity than tissue).
How should samples be handled?
Ten percent buffered formalin is the ideal fixating solution for prompt sample handling. The pathology laboratory must fix biopsy samples for a minimum of 6 hours up to a maximum of 48 hours, and surgical samples for a minimum of 12 hours up to a maximum of 72 hours. Reagents must be subject to rigorous quality control, including formalin, alcohol, xylene, and paraffin. Paraffin should be of high quality with a fusion point not exceeding 68°C. The other procedures must observe national and international quality standards. Bone samples should be handled adequately, with the sectioning of larger specimens for appropriate fixation, followed by decalcification with EDTA for a minimum of 24 hours up to a maximum of 48 hours 54 to allow adequate decalcification while avoiding nucleic acid degeneration. The morphological features (H&E) should be analyzed by a pathologist in order to establish a tumor diagnosis and ensure that sections containing at least 20% tumor cells are selected for further processing in the subsequent tests. Complementary tests, such as IHC and molecular diagnostic assays, should be performed rationally in order to conserve sample material and resources. FFPE samples must be stored separately according to recommended conditions.
Optimizing the patient journey
When a molecular test is requested by the oncologist, patients usually need to request the pathology laboratory to retrieve the biopsy sample from its archives and send it to the molecular biology laboratory. However, this process usually takes up to 2 weeks, which is valuable time for patients with advanced cancer. In this regard, general pathology laboratories may improve the patient journey by establishing protocols to speed up sample retrieval and shipping to the molecular biology laboratory when a molecular test is requested for patients with locally advanced or metastatic cancer.
Treatment with TRK inhibitors
The presence of TRK fusion is a new predictive biomarker for solid tumors for TRK inhibitors. We recommend that adult and pediatric patients with locally advanced or metastatic solid tumors with TRK fusion-positive cancers be offered screening strategies to identify those who might benefit from treatment with a TRK inhibitor.
Conclusions
Although oncogenic NTRK fusions are rare events in common cancer types (less than 1% of cases), taking all cancer types together, they represent a significant number of patients. Targeted TRK inhibition is the treatment of choice for TRK fusion-positive cancers and may confer a significant survival benefit and meaningful anti-tumor response in these patients. Thus, optimal diagnosis of the presence of NTRK fusions is pivotal to correctly identifying patients eligible for this potentially life-saving intervention.
We have proposed a testing algorithm based on the latest ESMO consensus for adoption by pathology laboratories. For common tumors with a low pretest probability of harboring NTRK fusions, we propose pan-TRK IHC as a screening test, followed by confirmation of positive results by NGS. However, when tumor histology is derived from tissues that physiologically express TRK proteins, we recommend direct NGS profiling due to the high rates of IHC false-positive results reported in these tissues. Finally, for rare tumors with a high incidence of harboring NTRK fusions, such as infantile fibrosarcoma and secretory breast carcinomas, we recommend FISH testing or real-time PCR for the classical ETV6::NTRK3 fusion, followed by NGS investigation of negative cases or direct NGS tumor profiling.
In this consensus, we have provided evidence-based recommendations on how to implement, optimize and analyze the results of the methodologies described, serving as practical help for pathologists and laboratories during their testing routines.
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This paper was supported by an unconditional grant from Bayer for editorial assistance provided by corebox (São Paulo, Brazil). The sponsor had no role in manuscript drafting or in the decision to submit it.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MPM, ECTN, FS and KMRL were responsible for conceiving the study, participating in the study design, contributed to writing and critically revising the manuscript. All other authors contributed to writing and critically revising the manuscript.

