Abstract
The case presented here illustrates that interdisciplinary teamwork can be essential for the understanding of the COVID-19 disease presentation and enlightening of the pathophysiology. A 60-year-old woman without any comorbidities, apart from overweight, was found dead in her apartment after 14 days of home isolation due to suspicion of COVID-19. A forensic autopsy was performed. This revealed severely condensed, almost airless, firm lungs, and the cause of death was severe acute respiratory distress syndrome-associated with COVID-19 (SARS-CoV-2). In addition, SARS-CoV-2 was detected with reverse transcription polymerase chain reaction (RT-PCR) in cerebrospinal fluid, lung tissue, and tracheal sample and specific antibodies for SARS-CoV-2 were detected in cerebrospinal fluid and serum. Subsequent sequencing of the SARS-CoV-2 virus showed variation in nucleotides at 3 sites between SARS-CoV-2 isolates recovered from the tracheal sample, cerebrospinal fluid, and tissues from both lungs, and phylogenetic analysis revealed that the spinal fluid sample differed the most from the other 3 samples. This case supports the hypothesis that SARS-CoV-2 may be neuroinvasive and cause central nervous system infection.
Background
Coronavirus disease 2019 (COVID-19) is an infection caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which arose in Wuhan Hubei Province, China in late 2019.1,2 SARS-CoV-2 is a relatively large single-stranded RNA virus with clinical similarities including neurological manifestations to closely related severe acute respiratory syndrome coronavirus-1 (SARS-CoV-1).1,3-5 Severe acute respiratory syndrome coronavirus-2 has since spread and caused more than 1.5 million deaths worldwide. 6 In Denmark (total population 5.8 million), by the middle of December 2020, a total of 137,000 patients have tested positive for SARS-CoV-2 and 1053 (0.8%) have died. 7
The primary cause of death associated with the SARS-CoV-2 is acute respiratory failure with exudative diffuse alveolar damage 8 followed by bronchopneumonia, massive pulmonary embolism, and alveolar haemorrhage.8,9 However, symptoms such as anosmia, ageusia, as well as ataxia, seizures have led to the hypothesis that SARS-CoV-2 may also have neuroinvasive capacities and initiate neuroinflammatory events. The route of central nervous system (CNS) infection is thought to be hematogenous or by an axonal transport through ACE2-receptors or directly by use of the olfactory nerve.5,10 Furthermore, neuroinvasion of SARS-CoV-2 has been suggested to play a role in the respiratory failure of COVID-19 patients. 11
Here we describe a patient with possible CNS involvement of COVID-19, discovered through a forensic autopsy and microbiological examinations. The autopsy was performed due to the Danish legal act on sudden death and according to ISO 17020 standards. It included a whole-body postmortem computer tomography (PMCT) before the external and internal examination, followed by thorough bio sampling, and additional supplementary toxicological and histopathological tests, as well as extensive microbiological analyses including full genome sequencing.
Case Presentation
A 60-year-old woman of European origin with body mass index (BMI) 28.5 kg/m2, with no significant past medical history, was found dead in her bed by her partner. The circumstantial information in the police report revealed that she had been in home isolation for 14 days due to suspicion of COVID-19 infection. In the days preceding her death, she reported symptoms of tachycardia (around 100 beats per minute), fever, and increasing respiratory difficulty according to her family. Clinical records were checked for pre-existing medical conditions and medications and none were reported.
Diagnostic assessment
Postmortem computer tomography revealed uniformly consolidated lungs bilaterally with air entrapment (Figure 1A). Postmortem computer tomography of the brain was unremarkable. Macroscopically, the lungs were heavily consolidated and firm, almost airless with a total weight of 1460 g. Pleura was without fibrin deposits. The tracheobronchial mucosa was hyperaemic, without purulent changes. Tracheobronchial lymph nodes were enlarged. The brain was oedematous weighing 1512 g (normal weight 1250 g ± 20 g), 12 with slight flattening of the gyri, unremarkable arachnoid, grey and white matter, and without signs of inflammation or incarceration. The kidneys showed signs of acute shock with dark coloration of the renal marrow opposed to a pale cortex.
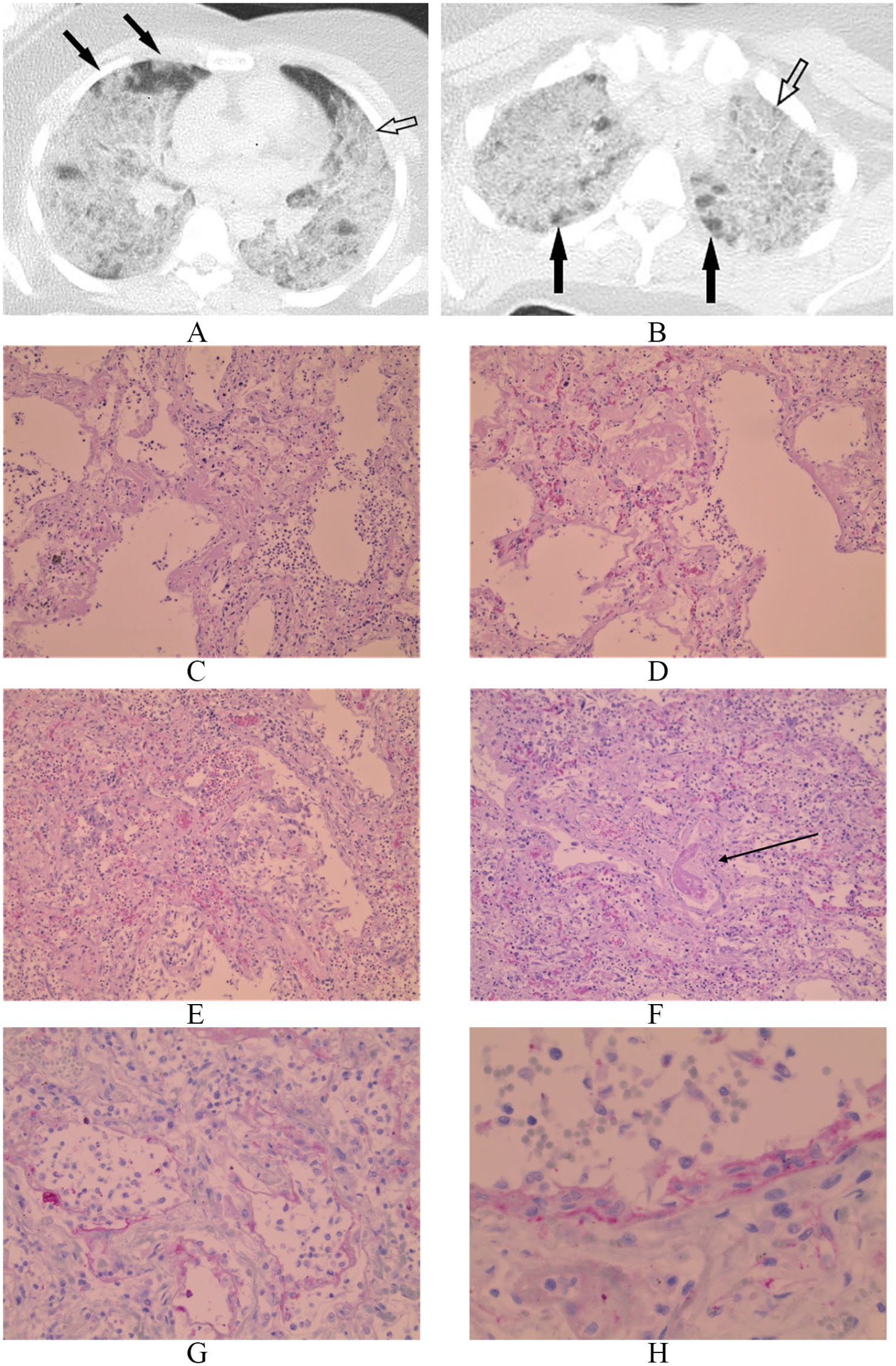
(A) Axial postmortem computer tomography scan of the consolidated lungs with peripheral ground glass opacities (solid arrows) and ‘crazy paving’ indicating oedema of the interstium (open arrow). (B) Axial postmortem computer tomography scan of the consolidated lungs with air entrapment (solid arrows) and “crazy paving,” indicating oedema of the interstitium (open arrow). Microscopic pulmonary findings of diffuse alveolar damage showing a spectrum of acute exudative lesions (C-D) to a more consolidated phase with proliferation and organization (E), including a thrombus in the vasculature (F, arrow) (Hematoxylin and eosin staining, 200×). Microscopic immune-histochemical findings showing the deposition of C4d complement fraction in the alveolar structures (G) and microvasculature (H) (IHC for C4d stain; 200×).
The histological examination was performed according to standard protocols13,14 and supplementary immunohistochemistry was performed for structural cell types comprising cytokeratin 7 (CK7), thyroid transcription factor 1 (TTF-1), platelet/endothelial cell adhesion molecule-1 (CD31), smooth muscle actin (SMA), and inflammatory markers of T-lymphocytes (CD3, CD4, CD8, FOX-P3), B-lymphocytes (CD20), plasma cells (CD79a, CD138, IgG, IgM), epithelial cells (HLA-DR), and complement fractions (C4d and C3d). The histopathology of the lung tissue revealed universally severe diffuse alveolar damage ranging from characteristic acute changes to more chronic changes (Figure 1C-F). There were no viral inclusions. The endothelium in the vasculature was severely hypoplastic with necrotizing, thrombotic microangiopathy. The complement fraction C4d (Figure 1G and H) was expressed in the endothelium. In addition, the cellular immunohistochemistry showed a dominance of CD3-positive T-lymphocytes and only a few CD20-positive B-lymphocytes. The T-lymphocytes were predominantly CD4 positive, with fewer CD8 and only very few FOX P3 cells. The abundant CD68-positive macrophages and plasma cells were with severe deposition of IgM, IgG as well as HLA-DR in the epithelial, endothelial, and inflammatory cells.
Tissue from the CNS comprised cerebrum with the motor cortex, basal ganglia, hippocampus, pons, and cerebellum. The meninges were sparsely represented. The tissue showed sparse nonspecific reactive minimal focal perivascular inflammation consisting of macrophages. There was no oedematous change, vasculitis, microthrombi, and no deposition of complement fractions. Myocardial tissue was with nonspecific subtle oedema.
Toxicologic analyses of blood and vitreous fluid for medication, illicit drugs, alcohol, diabetes-related analyses, and sodium as well as potassium concentration was performed. The results showed a treatment-related concentration of paracetamol. Illicit drugs or alcohol were not detected. The diabetes-related analyses results were normal. The postmortem electrolyte concentrations derived from the vitreous humour showed hyponatremia (103 mmol/L, defined < 134 mmol/L) 15 while the potassium concentration was high (21 mmol/L), which is as expected in postmortem material in which the sodium levels are quite constant, while the potassium level increases rapidly postmortem.
Microscopy of the cerebrospinal fluid (CSF) was performed but interpreted as inconclusive due to blood contamination. Further examination of the CSF such as cell count, differential and gramme-strain was not completed. Culturing of the CSF was with growth of Staphylococcus capitis. Growth of Bacillus cereus was identified in tissue from the right lung. Both findings were interpreted as either postmortem bacterial migration or contamination.
Polymerase chain reaction
RNA extraction was performed in the ISO-certified routine using automated nucleic acid extraction (MagNA Pure 96, Roche) according to manufacture recommendations. The detection of SARS-CoV-2 was performed by real-time reverse transcription polymerase chain reaction (RT-PCR)-based assay as described previously. 16 Severe acute respiratory syndrome coronavirus-2 was identified in CSF with cycle threshold (Ct) at 38.23, lungs tissue from both left and right lungs with Ct-values at 28.18 and 25.10, respectively, and in the tracheobronchial specimen with Ct 26.76. The blood specimen and the heart tissue specimen were negative for SARS-CoV-2.
Multiplex RT-PCR-based assay for 17 different respiratory tract virus (influenza type A and B, respiratory syncytial virus type A and B, metapneumovirus, parainfluenzavirus type 1, 2, 3 and 4, adenovirus, rhinovirus, enterovirus, parechovirus and coronavirus OC43, 229E, NL63, HKU1) was performed. Tissue samples from the heart and both lungs, as well as tracheobronchial swab, blood, and CSF, were analysed by this assay. None of the viruses was detected in any of the samples.
SARS-CoV-2-specific antibodies testing
The qualitative enzyme-linked immunosorbent assay (ELISA), Wantai SARS-CoV-2 Total Antibody ELISA (Beijing Wantai Biological Pharmacy Enterprise, Beijing, China) was used for measuring total developed SARS-CoV-2 antibodies. The protocol for performance of the Wantai SARS-CoV-2 Total Antibody ELISA assay has been described earlier. 17 The assay uses a double-antigen sandwich principle and detects total antibodies binding SARS-CoV-2 spike protein receptor binding domain in human serum and plasma. So far, the assay has not been validated for the use on CSF. However, the assay was used, according to the manufacturer’s instructions on both serum and CSF. Both samples were positive with absorbance values at 1.102 in blood and 3.092 in CSF at 450 nm after blanking.
Sequencing and genotyping of SARS-CoV-2
Total RNA was converted to cDNA with Superscript IV First-Strand Synthesis Kit (Thermo) using 11 μL of extracted RNA and random hexamers according to the manufacturer’s specifications. Polymerase chain reaction-based amplification of the viral genome was performed according to the ARTIC protocol using 2 separate multiplex primer pools reactions of the version 3 release of the primer set (https://artic.network/ncov-2019). Nanopore Sequencing libraries were prepared according to the Artic protocol and loaded onto an FLO-MIN106D Type R9.4.1 flow cell and sequenced on the MinION device (Oxford Nanopore, UK). Illumina libraries were prepared using the Nextera XT DNA Library Prep Kit (Illumina, CA). Paired-end sequencing was performed on the MiSeq platform (Illumina, CA).
Sequence analysis was performed in CLC genomics workbench 20.0.4 with the Long-read support (Beta) plug-in installed for the import and mapping of Oxford Nanopore reads. Demultiplexed Reads were imported as Oxford Nanopore reads and paired Illumina reads, respectively. Reads were trimmed against the V3 primer set (n = 218) to remove primer sequences from subsequent analysis. Oxford Nanopore reads and Illumina reads were mapped individually to the MT135044 SARS-CoV-2 reference sequence at CLCs default settings. To enhance the fraction of the genome sequenced and read-coverage (depth), Nanopore and Illumina mappings were merged into one for sequences from each of the 4 samples. This mapping had the following number of mapped reads, fraction (%) of the genome sequenced and average read coverages: spinal: 101 181, 98%, 294.12; tracheal: 346 914, 99%, 1041.55; left lung: 5 044 442, 99.9%, 14 041.95; and right lung: 4 431 907, 100%, 12 191.83. Consensus sequences were generated from the merged read mappings both at any coverage and at a minimum coverage of 5 reads and inserting N as a symbol at sequence positions not covered sufficiently. Sequence reads from the 4 different samples were individually aligned with the reference sequence and manually inspected and corrected in BioEdit 7.2.5. 18 Only inter-sample variants observed in the MiSeq mapping covered by ⩾5 reads and also observed in the Nanopore mapping were included in the consensus sequences. Nonresolved variant sites were marked with an N in the consensus sequence and excluded from further analysis. After this curation step, the fraction of the genome sequenced (in %) were: spinal: 92.9%; tracheal: 98.9%; left lung: 99.9%, and right lung: 100%.
The clade designation system 19 was used by downloading the Nextstrain Metadata set on the May 28, 2020. From this Metadata list, 27 sequences, representing all major SARS-CoV-2 clades (A1a, A2, A2a, A3, A6, B, B1, B2, and B4; only clades represented by at least 10 sequences were included) were selected along with the Wuhan-Hu-1 sequence (NC_045512). These 28 reference sequences were aligned with 4 patient sequences (named after their anatomical origin: left lung, right lung, tracheal, and spinal) in Mafft. 20 Phylogenetic analysis was performed in Mega 1021-23 using the Maximum Likelihood statistical method and the Hasegawa–Kishino–Yano model with a Gamma parameter of 2 and 100 bootstrap replicates and complete deletion of any gaps or missing data. The phylogenetic analysis (Figure 2) showed that all 4 virus samples belonged to the A2a clade (corresponding to clade 20B by the classification system implemented in Nextclade: https://clades.nextstrain.org/results).

Phylogenetic analysis of patient sequences. Phylogenetic tree rooted with the Wuhan-Hu-1 sequence (NC_045512) and GISAID sequences, shown for example EPI ISL 420347 (GISAID identifier) A2 (Nextstrain clade) and the 4 patient sequences (spinal, tracheal, right lung, and left lung).
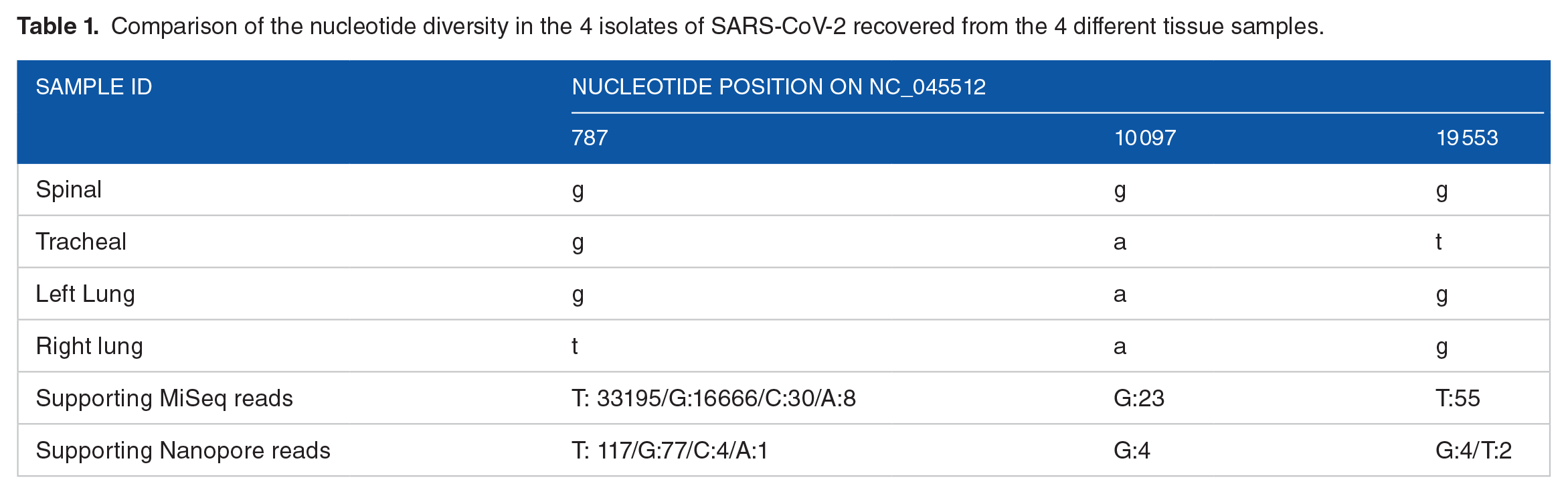
Sequences from 3 of the samples (spinal, tracheal, and right lung) had a dominant single nucleotide polymorphism (SNP) at 3 sites in the ORF1ab gene (Table 1), 2 of which (right lung and tracheal) were both mixed mutations. In sequences from the right lung, the SNP (T) was present in 66% of the reads while the G shared with the sequences from other samples was present in 33% of reads. Two minor variants, C and A constituted 0.01% and 0.06% of the reads and probably is most likely background seen at the high coverage observed at this site. The SNPs in the spinal and the tracheal samples resulted in changes in the encoded amino acid (Table 2). The majority of viral sequences in the spinal sample encoded a Glycine (G) instead of a Serine (S) in the 3C proteinase gene, and the majority of viral sequences in the tracheal sample encoded a Valine (V) instead of a Glycine (G) in the NSP11 encoding gene.
Comparison of the nucleotide diversity in the 4 isolates of SARS-CoV-2 recovered from the 4 different tissue samples.

Variations of the amino acids in the 4 isolates of SARS-CoV-2 recovered from the 4 different tissue samples.

Discussion and Conclusions
In this case, thorough examination revealed that the cause of death was severe acute respiratory distress syndrome-associated with COVID-19 with both severe acute and subacute to chronic diffuse alveolar damage, including necrosis of the endothelium and vascular changes with microvascular thrombosis. These pulmonary findings were accompanied by expression of the complement fraction C4d in endothelium, predominance of CD4- and CD3-positive T-lymphocytes, CD68-positive macrophages, and plasma cells with severe deposition of IgM, IgG, and HLA-DR in the epithelial, endothelial, and inflammatory cells consistent with systemic activation of complement pathways. The demonstration of intrathecal presence of SARS-CoV-2-specific antibodies and SARS-CoV-2-RNA detected in CSF suggest possible CNS involvement.
A previous report found that 36.4% of COVID-19 patients had neurological manifestations and that the rate of neurological symptoms was higher in patients with severe infection. 24 Also, in a recent report from Japan, a 24-year-old man was diagnosed with COVID-19-related meningitis/encephalitis, where specific SARS-CoV-2 RNA was detected in CSF, while it was not detected in the nasopharyngeal swabs. 25 In our case, specific SARS-CoV-2 RNA was detected in the tracheal sample, CSF, and tissue from both lungs. Central nervous system infection in relation to COVID-19 is further supported, by a report where the presence of the virus in neural and capillary endothelial cells, in the frontal lobe of a patient with COVID-19 infection and neurological symptoms, were documented by electron microscopy. 26 The microscopic examination of the brain revealed no signs of encephalitis, microthrombi, or inflammation, which is in keeping with Menter et al, 8 who, apart from nonspecific microscopic findings, registered a low viral load in neurological tissue.
Phylogenetic analysis showed that all 4 samples belonged to the same A2a clade. Most European SARS-CoV-2-sequences on Nextstrain as per May 28, 2020, belong to this clade (3177 of 4082; 77.8%). In Denmark, this clade is the most dominant per May 28, 2020 (206 of 208; 99.03%) (https://nextstrain.org/ncov/europe). The phylogenetic analysis showed that the spinal fluid sample differed the most from the other 3 samples. As only a partial sequence was obtained from this sample and differed at one nucleotide position, it is not clear if it differed more or if this has any functional significance.
For sequencing/PCR-results, we always consider the possible error sources. In addition, for the spinal sample, 7.9% of the genome was not covered or the quality was too low for analysis, which might have led to some mutations not being detected. However, since this is postmortem material, which cannot be recovered once it is used, and since we used state-of-the-art methods, this limitation cannot be further addressed with currently available methods. Another limitation to this case presentation, especially regarding the possible CNS involvement is the lack of clinical information, for example, if the patient had symptoms corresponding to meningitis/encephalitis before her death.
In the above-described case, the patient had been ill for 14 days before her death. This timeline is supported by the finding of SARS-CoV-2-specific antibodies. 27 Also the microscopic changes in the lung tissue with representation of severe diffuse alveolar damage in the early phase and an organizing and proliferative intermediary phase, indicating a disease process lasting around 1 to 2 weeks with a continuing severe acute component affecting both the pulmonary parenchyma, respiratory epithelium, and vasculature.8,27,28 The microscopic changes in the brain tissue were sparse and unspecific and therefore not suggestive of a primary CNS infection.
Biochemical analysis showed severe hyponatremia, which we suspect to be the result of several components: kidney involvement and possibly increased water intake during the disease; however, due to the fact that the deceased had been in isolation in the weeks before her death, information regarding food and water intake in her last days is not available. We conclude that hyponatraemia is a secondary result of a prolonged disease course. A well-known complication to hyponatraemia is cerebral oedema.29,30 At the autopsy, cerebral oedema was noted, but there were no signs of incarceration, and we therefore believe that severe pulmonary disease is the most likely cause of death.
Several publications reported association between SARS-CoV-2 and CNS infection documented by RT-PCR techniques,5,25,31,32 while only a few studies have confirmed the presence of SARS-CoV-2 in CNS by deep sequencing of CSF samples. 33 To the best of our knowledge, this is the first report that confirms CNS involvement of SARS-CoV-2 documented by RT-PCR followed by deep sequencing of the virus and supplemented with antibody analysis.
Histopathological and observational clinical studies suggest that the virus can create multiple microscopic ischemic infarcts in the subcortical white matter.5,10 In addition, an experimental study has shown that ACE2 expression is functionally required for SARS-CoV-2 infection in human brain organoids, by demonstrating that blocking ACE2 with antibodies or by administering CSF from a COVID-19 patient prevented neuronal infection. 6
The likely mechanisms of SARS-CoV-2-associated neuroinfection are thoughts to be through either axonal transport or a hematogenous route.34,35 During the axonal transport, SARS-CoV-2 enters the CNS through the olfactory and trigeminal nerve endings and infects neurons via ACE2-receptors on the surface of these cells. Through the hematogenous route SARS-CoV-2 enters the blood vessels through mucosa and infected lungs, disrupting the endothelial cells of the CNS and the virus can pass the blood-brain barrier resulting in a cerebral oedema. 34 COVID-19-associated autoimmune encephalitis has also been described by endothelial cells being attacked by autoantibodies produced after SARS-CoV-2 infection. 35
To our knowledge, this is the first report were both specific antibodies for SARS-CoV-2 and SARS-CoV-2-specific RNA were detected in CSF. Detection of intrathecal specific antibodies is diagnostic for a number of encephalo-meningeal virus infections, for example, herpes simplex virus and Varicella Zoster virus. 36 The lack of microscopic changes in the brain tissue might suggest a reduced immune response in the brain tissue. In addition, the detection of specific antibodies for SARS-CoV-2 in CSF is not validated, and the precise clinical significance of the findings is therefore unclear.
To conclude, this case supports the hypotheses that SARS-CoV-2 may cause CNS infection. The pathogenesis, incidence, and clinical impact, as well as the detection of specific antibodies for SARS-CoV-2 in CSF, need to be investigated further.
Footnotes
Acknowledgements
The authors acknowledge the staff of the Laboratory for Serology, Virus and Microbiological Special Diagnostics and Infectious Disease Preparedness, Statens Serum Institute, and Section of Forensic Pathology, Department of Forensic Medicine, University of Copenhagen for excellent laboratory support. Finally, they thank the next-of-kin for giving their consent for publication of this case report.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: All authors were funded by their departments. J.B. is involved with a project supported by the Lundbeck foundation. The funds had no role in study design, data collection, and analysis, decision to publish, or preparation of the manuscript.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
V.V.S.J., L.H.M., and J.B. designed the study. J.B., C.B., and C.J. contributed to the collection of the samples. C.J., J.F., C.B., and M.R. carried out laboratory analysis. L.H.M., C.J., J.F., C.B., M.R., A.F., J.B., and V.V.S.J. carried out data analysis. All authors wrote the manuscript and approved its publication.
Ethics Approval and Consent to Participate
The written consent from the next-of-kin, as well as consent from the police authorities, was obtained.
Availability of Data and Materials
The data and materials are available on request from the corresponding author (Veronika Vorobieva Solholm Jensen, Dept. of Virus and Microbiological Special Diagnostics, Statens Serum Institute; Artillerivej 5, DK-2300 Copenhagen S, Denmark. Email:
