Abstract
Background:
Acute upper gastrointestinal bleeding (AUGIB) is a major cause of morbidity and mortality. This presentation however is not universally high risk as only 20–30% of bleeds require urgent haemostatic therapy. Nevertheless, the current standard of care is for all patients admitted to an inpatient bed to undergo endoscopy within 24 h for risk stratification which is invasive, costly and difficult to achieve in routine clinical practice.
Objectives:
To develop novel non-endoscopic machine learning models for AUGIB to predict the need for haemostatic therapy by endoscopic, radiological or surgical intervention.
Design:
A retrospective cohort study
Method:
We analysed data from patients admitted with AUGIB to hospitals from 2015 to 2020 (n = 970). Machine learning models were internally validated to predict the need for haemostatic therapy. The performance of the models was compared to the Glasgow-Blatchford score (GBS) using the area under receiver operating characteristic (AUROC) curves.
Results:
The random forest classifier [AUROC 0.84 (0.80–0.87)] had the best performance and was superior to the GBS [AUROC 0.75 (0.72–0.78), p < 0.001] in predicting the need for haemostatic therapy in patients with AUGIB. A GBS cut-off of ⩾12 was associated with an accuracy, sensitivity, specificity, positive predictive value (PPV) and negative predictive value (NPV) of 0.74, 0.49, 0.81, 0.41 and 0.85, respectively. The Random Forrest model performed better with an accuracy, sensitivity, specificity, PPV and NPV of 0.82, 0.54, 0.90, 0.60 and 0.88, respectively.
Conclusion:
We developed and validated a machine learning algorithm with high accuracy and specificity in predicting the need for haemostatic therapy in AUGIB. This could be used to risk stratify high-risk patients to urgent endoscopy.
Introduction
Acute upper gastrointestinal bleeding (AUGIB) is one of the most common gastrointestinal emergencies and a major cause of morbidity and mortality in Western populations. There are over 50,000 admissions annually to UK hospitals, with an average mortality rate of 7%.1,2 The impact of AUGIB on utilizing hospital resources is considerable, creating a burden on the National Health Service, with 26% of the mean in-hospital cost per patient being attributable to endoscopy. 3
Among patients who have underlying high-risk stigmata causing AUGIB, such as bleeding or visible vessels, timely therapy to provide haemostasis and prevent re-bleeding is necessary. 4 As a result, national and international guidelines recommend endoscopy in all patients presenting with an AUGIB and admission to an inpatient bed within 24 h.5–7 In practice, this may not be possible, especially given it is reported in a nationwide UK audit that only 52% of hospitals had a consultant-led out-of-hours rota, 8 and Canadian and UK audits showing that endoscopy within 24 h for AUGIB only occurs in 50–65% of cases.8,9 Importantly, however, endoscopy may not be necessary for all patients in this time frame, as 70–80% of patients have low-risk lesions, such as clean-based ulcers or a normal examination, which do not require therapy. 8 Therefore, accurate non-endoscopic risk scoring systems are needed to identify high-risk patients requiring haemostatic therapy and low-risk patients who do not.
Several scoring systems exist for AUGIB, which can be calculated on presentation to hospital but few have found their way into routine clinical practice.10–12 The Glasgow-Blatchford score (GBS) has been recommended by UK and international guidelines to identify patients at low risk of requiring blood transfusion, and invasive therapy and at low risk of death, who can therefore be managed as outpatients without the need for urgent endoscopy.6,13,14 The score is, however, not accurate at identifying high-risk patients requiring haemostatic therapy with endoscopic, radiological or surgical intervention. 15
Machine learning (ML) presents a promising advancement in the field of risk stratification in patients with an AUGIB. ML models can extract patterns from raw data that are available through patient health records and can analyse large, complex datasets creating more unique risk scores for individual patients. The algorithms can simultaneously analyse multiple variables and have been shown to outperform standard risk scoring systems for AUGIB in predicting low-risk cases compared to a GBS of zero. 16 Given the current lack of accurate risk stratification tools for predicting invasive management in patients presenting with AUGIB, this study aimed to develop and validate novel ML risk models for the prediction of the need for haemostatic therapy, including endoscopic treatment, interventional radiology and surgery.
Methods
This manuscript was reported in accordance with the Transparent Reporting of a multivariable prediction model for Individual Prognosis or Diagnosis (TRIPOD) statement. 17
Study design and data collection
This was a retrospective cohort study. This study was performed by analysing an existing anonymized database of patients presenting with AUGIB collated for audit to three large acute hospitals; St Marys, Charing Cross and Hammersmith Hospitals, London, UK (November 2015–February 2020). The study was approved by the Joint Research Compliance office at Imperial College Healthcare NHS Trust (ref 125HH25060). The office confirmed that no formal ethical review or informed consent was required as the study involved existing anonymized routinely collected data, no new data were being collected and there was no clinical intervention.
Inclusion criteria were patients over the age of 18 years presenting to the emergency departments or ambulatory care with a suspected diagnosis of AUGIB based on a history of haematemesis or melaena, or those who developed an AUGIB whilst an inpatient. Patients with missing data on their records were excluded.
Data were collected on an Excel spreadsheet and included patient demographics (age, sex), observations (heart rate, blood pressure), laboratory results [haemoglobin, urea, creatinine, C-reactive protein (CRP)], medication (proton pump inhibitor use), past medical history [hepatic disease, cardiac failure, respiratory disease, hypertension (HTN), stroke], findings on endoscopy, and treatments (none, blood transfusion, endoscopy, interventional radiology, surgery).
General treatment of patients
Management of patients with AUGIB followed UK NICE guidelines. 7
Follow-up
Patients in this cohort were followed up for 30 days using electronic health records and telephone calls to patients and primary care providers.
Outcomes
The predetermined outcome of this study was the need for haemostatic therapy. Patients were determined as needing haemostasis, and therefore high risk, if they received appropriate endoscopic therapy, interventional radiology or surgery to achieve haemostasis during the time period of the study. As the delivery of haemostasis can be subjective, we examined the appropriateness of therapy in patients who underwent endoscopy which was the presence of high-risk stigmata that require therapy. 18 This was assessed independently by blinding to final risk scores. Low-risk patients were defined as those patients who had not received any of these interventions.
Development and validation of ML models
Novel handcrafted features were designed and extracted from the datasets for the training of the classification models. To design an effective prediction model, we incorporated the importance of accurate prediction together with the development of an interpretable model, that is, one that used key features from the physiological parameters to determine the estimation. For this reason, we used Mean Decrease Accuracy, also known as permutation importance, 19 to validate the performance of the selected features.
Permutation importance is a model inspection method, especially useful for non-linear estimators, in which the concept is to break the relationship between the target and each particular feature by random shuffling across different subjects. In this way, the drop in estimation result is indicative of how much the model depends on this particular feature. The clinical features used to build the ML model are shown in Table 1. Before feeding the extracted features to the networks, feature scaling was implemented using the sklearn library (StandardScaler) to ensure that the model would not be affected by one single feature with a large magnitude.
Clinical features used to build machine learning models.

The data within the database were separated into training and test sets in an 80:20 split, and five-fold cross-validation was performed. The training data were used to develop the ML model, which was then validated using the test set. The ground truth for the ML model was based on each patient’s endoscopic findings and management. High-risk lesions were defined as those requiring either endoscopic treatment, interventional radiology or a surgical procedure.
Following feature selection and data balance, analysis of the datasets was performed using various supervised ML classifiers, such as random forest, extra tree, gradient boost, multi-layer perceptron (MLP), K-nearest neighbours (KNN), decision tree and ensemble learning. Given the limited training data and sparsity of the data, supervised ML classifiers were chosen instead of conventional deep learning methods.
Comparison of proposed models with GBS
A direct comparison of the validation results of the ML model with the GBS scoring system was carried out. We compared the ML models with a GBS of ⩾12, which has been suggested as the most accurate cut-off to identify patients needing endoscopic therapy. 20
Statistical analysis
ML classifiers were evaluated in terms of accuracy, sensitivity, specificity, positive predictive value (PPV) and negative predictive value (NPV). These outputs were also calculated for GBS. In addition, the areas under the receiver operating characteristics (AUROC) curves and corresponding 99% confidence intervals for the ML models were calculated. AUROC curves of 0.90–1.00 were considered excellent, 0.80–0.89 good and 0.70–0.79 moderate. Excellent and good accuracies were considered useful for clinical decision-making. Sensitivity and Specificity were also calculated for all models. A model with a sensitivity of ⩾90% was considered excellent and would be useful in clinical practice to rule out the need for haemostatic therapy and therefore an urgent endoscopy (useful in identifying low-risk patients). A model with a specificity ⩾90% was considered excellent and would be useful to rule in need for haemostatic therapy and therefore an urgent endoscopy (useful in identifying high-risk patients).
The best-performing prediction model was evaluated in terms of calibration (a plot of predicted versus observed risk). A decision curve analysis was performed to determine the clinical utility of the prediction model.
Statistical analysis was carried out using Python 3.6 (Python Software Foundation, Wilmington, NC, USA). Patient characteristics were analysed using Python (with the library numpy and scipy.stats). The discriminative ability of the prediction of the need for haemostasis by the two scoring systems, as measured by AUROCs, was assessed using the method by Delong et al. 21
Results
Patient characteristics
The database contained 977 patients of which a total of 970 patients were included in this study. Due to missing information (observations at presentation not noted, missing laboratory results) and inability to calculate the GBS, a total of seven patients were excluded. The patient characteristics are shown in Table 2. The median age of all the patients was 61 years old with the majority of patients being male (62.9%). Approximately 22% of patients required haemostatic therapy (endotherapy, radiological embolization or surgery).
Patient characteristics, endoscopic findings and haemostatic intervention.

CRP, C-reactive protein; IQR, interquartile range.
Training versus validation
Patients were randomly allocated to training and validation sets, in an 80:20 split, as part of a five-fold cross-validation method. A total of 776 patients’ data was used to train the ML algorithms, and 194 patients’ data were used to validate the classifiers. The study design flowchart 22 is shown in Figure 1.

Study design flowchart.
Performance of ML classifiers in comparison to GBS
Results of the AUROC for the ML classifiers and GBS to detect high-risk patients are shown in Table 3 and Figure 2. After feature selection and data balancing, the random forest classifier [AUROC 0.84 (0.80–0.87)], ensemble learning [AUROC 0.83 (0.80–0.87)], extra tree [AUROC 0.81 (0.79–0.84)] and gradient boost [AUROC 0.83 (0.80–0.87)] performed better than GBS [AUROC 0.75 (0.72–0.78)] in predicting the need for endoscopic, surgical or radiological haemostatic therapy in patients presenting with an AUGIB. The network architecture of the random forest and ensemble learning models is illustrated in Figure 3.
AUROC curve values of the ML classifiers and GBS.

AUROC, area under the receiver operating characteristics; GBS, Glasgow-Blatchford score; ML, machine learning; MLP, multi-layer perceptron.
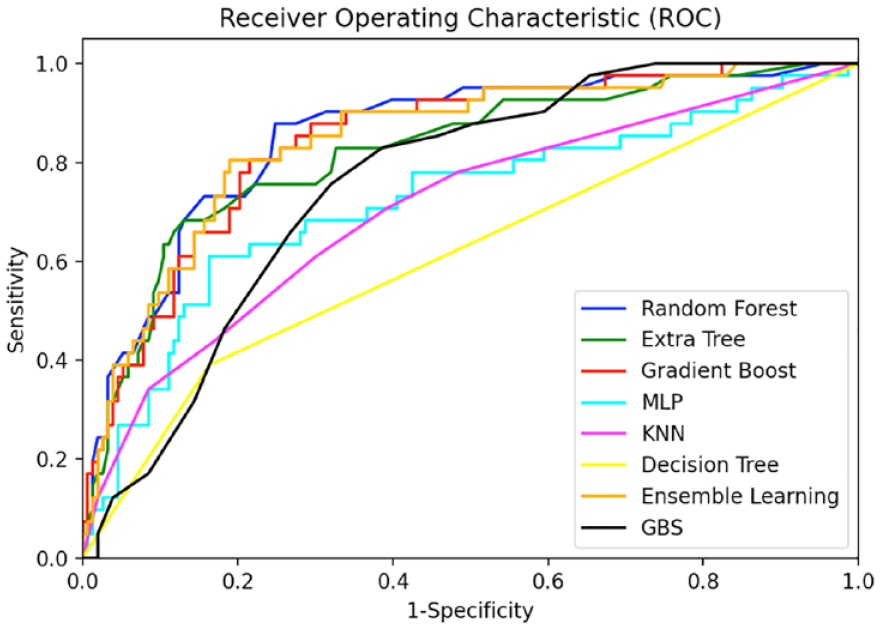
AUROC curve values of the ML classifiers and GBS.
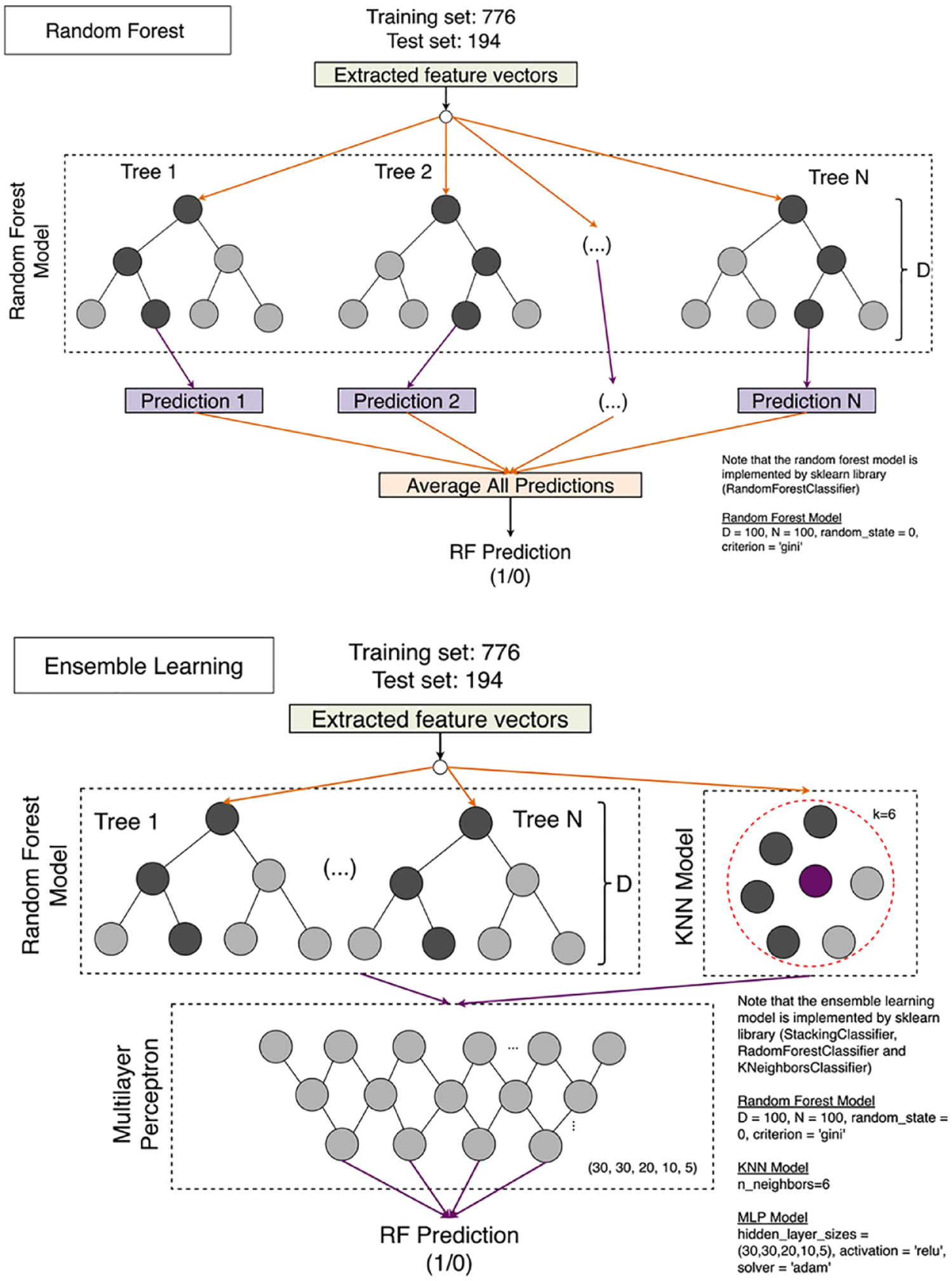
The network architecture of the random forest and ensemble learning models.
A GBS cut-off of ⩾ 12 was associated with an accuracy of 0.74, a sensitivity of 0.49, a specificity of 0.81, a PPV of 0.41 and an NPV of 0.85. The most accurate ML classifier was the random forest with the following metrics: accuracy 0.82, sensitivity 0.54, specificity 0.90, PPV 0.60 and NPV 0.88. The performance metrics of the other ML classifiers can be accessed in Supplemental Table 1.
The calibration plot predicting the need for haemostatictherapy in AUGIB highlighted that the random forest model had good calibration on the validation set, with predicted risk close to observed risk. The calibration plot is illustrated in Figure 4. The decision curve analysis showed that the random forest model had the highest net benefit across the whole range of possible threshold probabilities. The decision curve analysis is shown in Figure 5.

Calibration plot for the random forest model.
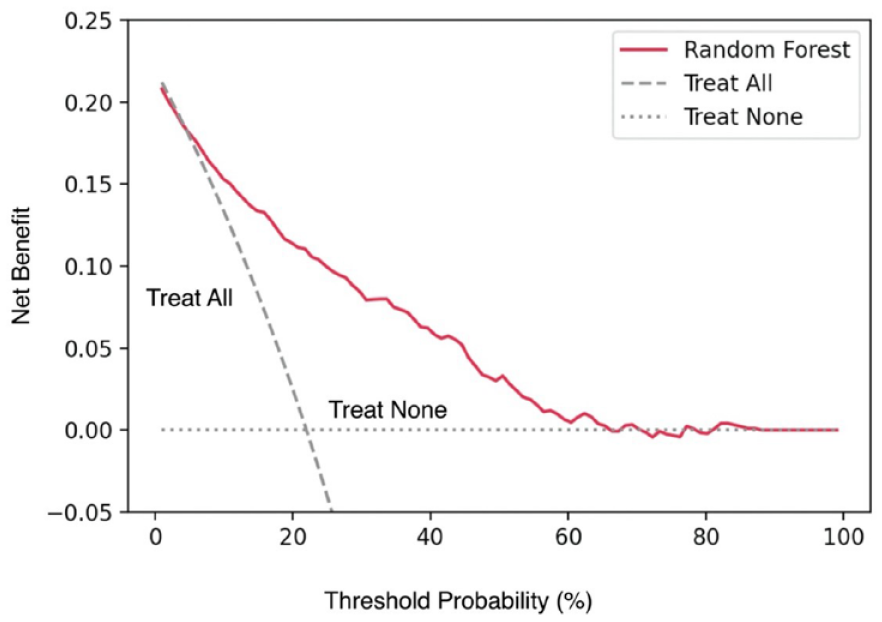
Decision curve analysis for random forest classifier.
Discussion
In this study, we developed and validated ML algorithms (random forest, ensemble learning, extra tree and gradient boost) with high accuracy when compared to a conventional risk scoring system (GBS), in predicting the need for endoscopic, surgical or radiological haemostatic therapy in patients presenting with an AUGIB. The random forest classifier demonstrated an excellent AUROC score of 0.84, compared to 0.75 for GBS (p < 0.001).
Other studies have used a type of ML algorithm (artificial neural networks) to develop predictive tools for patients presenting with AUGIB. However, the studies by Rotondano et al. 23 and Grossi et al., 24 who achieved an AUROC of 0.95 and 0.87, respectively, focus on predicting mortality rather than haemostatic therapy. Shung et al. 16 used ML (Gradient Boost) to develop a model to predict a composite outcome of blood transfusion, hospital-based intervention and death which identified very low-risk patients rather than the need for haemostatic intervention alone. In addition, a large systematic review analysing ML to predict outcomes in patients presenting with all acute gastrointestinal bleeding highlighted the lack of studies evaluating the use of ML specifically for upper GI bleeding and for predicting the need for therapy. When assessing any GI bleeding, the median intervention AUROC was 0.84 (0.40–0.95) when using ML models. 25 Das et al. 26 did investigate a neural network to predict the need for endoscopic therapy which achieved an AUROC of 0.89 for internal validation and 0.78 for external validation. The study cohort was, however, limited to those with non-variceal bleeding, unlike our study which included patients presenting with haematemesis or melaena which may be more clinically relevant, as clinicians who are deciding to endoscope often do not know the final diagnosis.
A GBS of 12 has been suggested as the most accurate cut-off for predicting the need for endoscopic therapy, that is, patients with scores ⩾12 require endoscopic therapy and those with scores <12 do not.20,26 In our study, the accuracy of this cut-off was moderate at 0.74, limiting clinical utility. By contrast, the Random Forest algorithm had a good accuracy of 0.82 and an excellent specificity of 0.90. In clinical practice, this would mean that our model would be able to identify patients that would likely require therapy, with high certainty, and that a positive test result would likely mean that endoscopic treatment was needed. In this way, patients could be prioritized for urgent endoscopy and higher levels of care if appropriate. This is invaluable in a healthcare system in which resources are limited. Despite the high specificity, however, the sensitivity of the random forest model was not high so could not be used to rule out patients requiring haemostatic therapy.
Strengths of this study include the multicentre design, large cohort and validation of results. Most importantly, the ML algorithms could predict the outcome accurately which supports the clinician to make a different clinical decision than is currently routine, that is, the option to not do an urgent endoscopy. This could allow for delayed inpatient endoscopy after treatment of concurrent illnesses such as sepsis or outpatient endoscopy.
This study has limitations in that while the results were validated, they would also need to be verified with a different dataset (external cohort). While our study benefits from a robust internal validation through a five-fold cross-validation method, we acknowledge the importance of an independent external test set to further validate the robustness and generalisability of our ML models across diverse clinical contexts. The next step in our research aims to enhance the external validity of the models and ensure their applicability in varied healthcare settings. Nevertheless, the internal validation employed in our study contributes to reducing the chance of bias and provides a foundation for the robustness of our findings. We are actively pursuing opportunities to obtain an external dataset for further validation and refinement of our predictive models. Secondly, all the patients in this study did not undergo endoscopy which is in line with similar studies and clinical practice.8,15 This is mainly because the hospitals have a guideline to discharge patients from the emergency department with a GBS of ⩽1 and many then do not attend a scheduled outpatient endoscopy appointment. 27 We do not believe this impacted the outcome of the study as patients were followed up for 30 days after AUGIB for rebleeding and mortality. Moreover, a risk model that required endoscopy would be of limited value as the clinical need is to avoid or delay endoscopy. The best ML algorithm (random forest) did not have a specificity of 100% (but was close to this) which means that some patients who have underlying high-risk stigmata may not undergo urgent endoscopy within 24 h. However, the adoption of this model would be an improvement on the scenario in routine clinical practice where about 40–50% of patients would not get an endoscopy within 24 h.8,28 Moreover, patients are already in the hospital so can be triaged to more urgent endoscopy if deterioration occurs. In addition, for acute lower gastrointestinal bleeding, an Oakland score of ⩽8 predicts a 95% chance of safe discharge 29 and has been recommended for use in routine clinical practice, 30 demonstrating that clinicians recognize that 100% certainty from risk scores or clinical impression are not available nor necessary.
Furthermore, although the ML model has more variables than the GBS, all features included in the model would be information gathered on a patient’s presentation to the hospital and therefore easily accessible, so we do not foresee this to significantly prolong the time taken to enter data and achieve a result. To allow the adoption of the model into clinical practice, an application providing an easy-to-use graphical user interface would be integrated into the medical records workflow, for use by clinicians.
Conclusion
Current risk scoring tools for AUGIB cannot accurately discriminate high-risk from low-risk patients requiring haemostatic intervention. We have developed an ML model that can risk stratify these patients with high accuracy and identify a group of high-risk patients likely to require haemostatic therapy.
Supplemental Material
sj-docx-1-cmg-10.1177_26317745241246899 – Supplemental material for Development and validation of machine learning models to predict the need for haemostatic therapy in acute upper gastrointestinal bleeding
Supplemental material, sj-docx-1-cmg-10.1177_26317745241246899 for Development and validation of machine learning models to predict the need for haemostatic therapy in acute upper gastrointestinal bleeding by Scarlet Nazarian, Frank Po Wen Lo, Jianing Qiu, Nisha Patel, Benny Lo and Lakshmana Ayaru in Therapeutic Advances in Gastrointestinal Endoscopy
Supplemental Material
sj-docx-2-cmg-10.1177_26317745241246899 – Supplemental material for Development and validation of machine learning models to predict the need for haemostatic therapy in acute upper gastrointestinal bleeding
Supplemental material, sj-docx-2-cmg-10.1177_26317745241246899 for Development and validation of machine learning models to predict the need for haemostatic therapy in acute upper gastrointestinal bleeding by Scarlet Nazarian, Frank Po Wen Lo, Jianing Qiu, Nisha Patel, Benny Lo and Lakshmana Ayaru in Therapeutic Advances in Gastrointestinal Endoscopy
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
